Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Youhang Li | -- | 2070 | 2022-11-08 09:09:22 | | | |
| 2 | Conner Chen | Meta information modification | 2070 | 2022-11-09 08:34:07 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Zhao, Y.; Hao, S.; Wu, W.; Li, Y.; Hou, K.; Liu, Y.; Cui, W.; Xu, X.; Wang, H. Lysine Crotonylation in DNA Damage Response. Encyclopedia. Available online: https://encyclopedia.pub/entry/33513 (accessed on 01 June 2026).
Zhao Y, Hao S, Wu W, Li Y, Hou K, Liu Y, et al. Lysine Crotonylation in DNA Damage Response. Encyclopedia. Available at: https://encyclopedia.pub/entry/33513. Accessed June 01, 2026.
Zhao, Yuqin, Shuailin Hao, Wenchi Wu, Youhang Li, Kaiping Hou, Yu Liu, Wei Cui, Xingzhi Xu, Hailong Wang. "Lysine Crotonylation in DNA Damage Response" Encyclopedia, https://encyclopedia.pub/entry/33513 (accessed June 01, 2026).
Zhao, Y., Hao, S., Wu, W., Li, Y., Hou, K., Liu, Y., Cui, W., Xu, X., & Wang, H. (2022, November 08). Lysine Crotonylation in DNA Damage Response. In Encyclopedia. https://encyclopedia.pub/entry/33513
Zhao, Yuqin, et al. "Lysine Crotonylation in DNA Damage Response." Encyclopedia. Web. 08 November, 2022.
Copy Citation
The DNA damage response (DDR) system plays an important role in maintaining genome stability and preventing related diseases. The DDR network comprises many proteins and posttranslational modifications (PTMs) to proteins, which work in a coordinated manner to counteract various genotoxic stresses. Lysine crotonylation (Kcr) is a newly identified PTM occurring in both core histone and non-histone proteins in various organisms. This novel PTM is classified as a reversible acylation modification, which is regulated by a variety of acylases and deacylases and the intracellular crotonyl-CoA substrate concentration. Kcr links cellular metabolism with gene regulation and is involved in numerous cellular processes.
lysine crotonylation
DNA damage response (DDR)
DNA double-strand break (DSB)
1. Introduction
DNA in cells is under constant threat from various genotoxic agents derived from endogenous cellular metabolism and the environment. These agents attack DNA to cause DNA damage, leading to the accumulation of lesions in each cell. These DNA lesions must be removed to ensure that cells function properly because they interfere with crucial functions of DNA, including replication and transcription. Failure to properly repair these DNA lesions can affect the stability of the genome, which is associated with cancer, developmental defects, infertility, immune deficiency, neurodegenerative diseases, and premature aging [1][2][3]. Mammalian cells have evolved a complex and versatile signaling network called DNA damage response (DDR) to mitigate the toxic effects of DNA lesions [4][5]. DDR is a concerted process involving the detection of DNA lesions, cell cycle checkpoint activation, DDR factor recruitment, chromatin reorganization, and DNA processing and repair. Initially, different DNA lesions are recognized by specific proteins that trigger and coordinate the recruitment of other DDR factors to DNA damage sites and initiate DDR [5][6][7]. With the recruitment of DDR factors, cascades of posttranslational modifications (PTMs), including phosphorylation, ubiquitylation, sumoylation, acetylation, and poly-ADP-ribosylation, are induced, which are associated with the modulation of related cellular processes such as the cell cycle, replication, transcription, and DNA repair [8][9][10][11][12][13].
2. Kcr Is an Evolutionary Conserved and Abundant PTM
The roles of reversible acetylation of the ɛ-amino group of histone and non-histone lysines in DDR have been studied extensively [14][15][16][17][18][19]. In recent years, with the help of high-sensitivity mass spectrometry (MS), some short-chain acylation modifications, which also occur to the ɛ-group of lysine, including propionylation, butyrylation, 2-hydroxyisobutyrylation, succinylation, malonylation, glutarylation, crotonylation, β-hydroxybutyrylation, and lactylation, have been discovered successively. These acylation modifications are distinct in hydrocarbon chain length, hydrophobicity and charge but are structurally similar to the well-studied acetylation of lysine. These modifications also act like acetylation to neutralize the positive charge of the lysine side chain or to provide a larger lysine side chain for protein recognition and binding, thereby affecting the properties and functions of modified proteins [20][21][22][23][24][25][26][27]. Currently, the functional significance of these newly discovered PTMs, especially in DDR, remains largely unknown.
Kcr was first discovered as a new PTM of histone about ten years ago using an MS-based proteomics approach by Tan et al. [26]. In this study, Kcr was identified as a new type of PTM, 28 Kcr sites on core histones were discovered, and it was demonstrated that histone Kcr marks either active promoters or potential enhancers in both human somatic and mouse male germ cells [26]. This study opened a new direction in the field of PTMs and has attracted significant attention. Subsequently, histone Kcr was found to be associated with many diseases such as acute renal injury, depression, HIV latency, and the process of cancer. Kcr is found in different species, such as mouse, Saccharomyces cerevisiae, Caenorhabditis elegans, Drosophila melanogaster, and plants. Non-histone proteins have also been found to contain Kcr modifications. Numerous reports have shown that Kcr is an evolutionarily conserved and abundant PTM [28][29][30][31][32][33][34][35][36][37][38][39][40][41].
3. Enzymes Responsible for Reversible Kcr Regulation
Histone acetylation is one of the most widely studied epigenetic modifications related to chromatin remodeling, gene transcriptional regulation, and other key biological processes [42][43][44][45][46][47]. Protein lysine acetylations are regulated by the concerted actions of lysine acetyltransferases (KATs) (Kac “writer”) and lysine deacetylases (KDACs) (Kac “eraser”) that function by adding and removing acetyl groups from lysine residues, respectively. Because histones were the first identified Kac substrates, KATs and KDACs are often referred to as histone acetyltransferases (HATs) and histone deacetylases (HDACs), respectively. HATs/HDACs catalyze reversible acetylation of histone and non-histone proteins via transferring the acetyl group from acetyl-CoA to the ɛ-group of lysine or removing the acetyl group from a modified lysine [48][49][50][51][52][53][54][55][56]. Currently, no crotonyl-specific “writer” or “eraser” has been identified. Enzymes that regulate lysine acetylation also appear to regulate lysine crotonylation, although the two acyl groups are not identical in size and structure. The well-characterized HAT CBP (CREB-binding protein) and P300 have been shown to possess histone crotonyltransferase (HCT) activity and are possibly major HCTs in mammalian cells [57][58][59]. In addition, GNAT (GCN5-related N-acetyltransferase) family PCAF (P300/CBP-associated factor), MYST (Moz, Ybf2, Sas2, and Tip60) family hMOF (human males absent on the first), and HBO1 (histone acetyltransferase binding to ORC) were also revealed to possess HCT activity [60][61]. In mammals, Zn2+-dependent HDACs (HDAC 1–11, grouped into classes I, II, and IV) and NAD+ dependent sirtuins (SIRT 1–7, class III HDAcs) are responsible for removing acetyl groups from lysine residues in histone and non-histone substrates [62][63]. It is now clear that lysine crotonylation is also as dynamic as lysine acetylation. Class I HDACs (HDAC1/2/3) and the sirtuin family of HDACs (SIRT1/2/3/6) harbor histone decrotonyltransferase (HDCR) or lysine decrotonyltransferase (KDCR) activity, and class I HDACs are probably the major HDCR in mammalian cells [64][65][66][67][68]. Noteworthily, Wong’s lab generated CBP/p300 mutants with HCT but impaired HAT activity and HDAC1/HDAC3 mutants with DHCT but impaired DHAT, which may help to specifically study the function of histone Kcr and the molecular mechanism of its regulation [59][64]. Further studies are needed to determine whether these mutants apply to all histone Kcr sites and whether they also apply to non-histone proteins.
4. Regulation of Kcr by the Cellular Concentrations of Crotonyl-CoA
In vitro, acetyl-CoA and crotonyl-CoA can promote Kac and Kcr, respectively, without support from enzymes [69][70][71]. In cells, acetyl-CoA and crotonyl-CoA are necessary substrates for HATs to catalyze Kac or HCTs to catalyze Kcr. Acetyl-CoA is the most abundant CoA species measured, and crotonyl-CoA is approximately 1000-fold less abundant than acetyl-CoA in cells [58]. The relatively low intracellular abundance of crotonyl-CoA makes it easier to change the concentration of crotonyl-CoA than that of acetyl-CoA in cells and thus more likely to affect intracellular Kcr levels experimentally. Consistent with this, supplementation with crotonate significantly increased the cellular crotonyl-CoA and Kcr levels [26]. In contrast, depletion of acyl-CoA synthetase short-chain family member 2 (ACSS2), which converts crotonate into crotonyl-CoA, reduces the cellular crotonyl-CoA and Kcr levels [58]. Moreover, crotonyl-CoA is an endogenous intermediate metabolite during fatty acid oxidation and lysine/tryptophan metabolism. The key enzymes in these metabolic pathways that control crotonyl-CoA production can also affect the crotonyl-CoA and Kcr levels in cells. Depletion of mitochondrial short-chain acyl-CoA dehydrogenase (ACADS) or peroxisomal acyl-CoA oxidase (ACOX3), two key enzymes that catalyze the conversion of butyryl-CoA to crotonyl-CoA during fatty acid oxidation, also decrease the Kcr level by specifically reducing the cellular crotonyl-CoA level [72]. In addition to these enzymes capable of catalyzing crotonyl-CoA production, intracellular crotonyl-CoA can also be negatively regulated by crotonyl-CoA hydratase. Liu et al. reported that the chromodomain Y-like (CDYL) protein acts as a crotonyl-CoA hydratase to negatively regulate Kcr by converting crotonyl-CoA to β-hydroxybutyryl-CoA. CDYL transgenic mice showed dysregulation of Kcr and related phenotypes [71].
5. Recognition of Kcr by Chromatin-Associated Proteins
Increasing evidence suggests that crotonylation of histones is functionally different from histone acetylation and may regulate various cellular processes [73]. The identification of candidate proteins or structural modules that can “read” Kcr and translate Kcr of histones into diverse functional outcomes in cells has attracted much attention in the field. Chromatin-associated proteins are often involved in epigenetic regulation by recognizing PTMs of histones, such as phosphorylation and acetylation of histones, and thus were first investigated as Kcr reader candidates. Currently, no protein or structural module that specifically recognizes Kcr has been identified. Classical Kac “reader” proteins that contain YEATS (Yaf9, ENL, AF9, Taf14, and Sas5) and double plant homeodomain finger (DPF) domains preferentially bind Kcr when compared with Kac [74][75][76].
The YEATS domain, named for its five founding domain-containing proteins (Yaf9, ENL, AF9, Taf14, and Sas5), is evolutionarily conserved from yeast to human [77]. Originally, Li et al. reported that the YEATS domain of human AF9 strongly binds histone H3 acetylation at K9 (H3K9ac) and constitutes a novel family of histone Kac readers. Li et al. solved the crystal structure of the human AF9 YEATS domain bound to H3K9ac and found that the human YEATS domain organizes a unique aromatic “sandwich” pocket using highly conserved residues for Kac readout. The reader pocket of YEATS is characteristic of an “end-open” feature, indicating that it may accommodate the longer and more rigid Kcr better than Kac. This observation was confirmed by subsequent studies showing that Kcr enhances its binding to different YEATS domains by 2–5-fold when compared with Kac [74][78]. Structural analysis showed that AF9 YEATS uses the same Kac-binding aromatic sandwich pocket for Kcr recognition and the extended side chain of Kcr fits perfectly into the “end-open” pocket. The planar crotonylamide group is sandwiched by two aromatic residues that enable “aromatic-π-aromatic” stacking, which is optimal for Kcr recognition. The aromatic-π-stacking mechanism for Kcr recognition is observed consistently in the crystal structure of AF9, YEATS2, and Taf14 in complex with Kcr, and these YEATS domain proteins represent the first class of selective Kcr readers [74].
Although the “aromatic-π-stacking” mechanism perfectly accounts for the high affinity of the YEATS domain toward Kcr, it does not seem to be the recognition mechanism between all “readers” and Kcr. Recently, the DPF domains of the MYST family member monocytic leukemic zinc-finger (MOZ) and DPF2 were also found to recognize Kcr selectively. Crystal structures of the DPF/MOZ domain in complex with H3K14cr revealed that the DPF domain generates an intimate hydrophobic pocket without the participation of aromatic sandwiching residues to house the Kcr [79]. Recently, Klein et al. revealed another case of Kcr recognition by a DPF domain. The DPD domain of monocytic leukemic zinc-finger related factor (MORF) was also shown to bind H3K14cr over H3K14ac preferentially. The structure of the DPF/MORF-H3K14cr complex is very similar to the previously reported structure of the DPF/MOZ-H3K14cr complex, indicating that DPF/MORF and DPF/MOZ recognize Kcr by a conserved mechanism [80]. The DPF domains in MOZ and MORF represent the second class of selective Kcr readers.
6. The Emerging Role of Kcr in DSB Repair
Is non-histone Kcr also involved in DDR? The answer is yes. The deletion of CDYL1 attenuates crotonyl-CoA hydrolysis, increases the global intracellular Kcr level, and then improves the detection of Kcr. Using quantitative MS to detect Kcr in CDYL1 deficient HeLa cells, Yu et al. identified a large crotonylome dataset containing 14,311 Kcr sites in 3734 proteins [81]. Using bioinformatics analysis, they found that Kcr levels of a series of protein factors related to DNA repair, including RPA1, POLD1, APEX1, XRCC5, and KDM1A, were increased significantly in CDYL1-deficient HeLa cells. They focused on RPA1 and found that CDYL1 negatively regulates K88cr, K379cr, and K595cr of RPA1. Further analysis showed that the Kcr level of RPA1 increases after hydroxyurea (HU), ultraviolet, ionizing radiation, VP16 or camptothecin insult, suggesting the level of RPA1 Kcr is upregulated by DNA damage and differs from the aforementioned histone Kcr. Through mutation analysis, Yu et al. hypothesized that Kcr of RPA1 enhances its interaction with single-strand DNA (ssDNA) (Figure 1) and major HR factors, such as MRE11 and BLM, and then promotes the recruitment of RAD51 recombinase and HR-mediated DSB repair [81]. These findings revealed for the first time the important roles of non-histone Kcr in DNA repair. According to Enas R. Abu-Zhayia et al., CDYL1 does not affect HR-mediated DSB repair through its crotonyl-CoA hydratase activity. In other words, CDYL1 does not regulate HR by influencing Kcr of RPA1 through modulating intracellular crotonly-CoA [82]. Other regulatory mechanisms, such as HCT/HDCR enzymes mediating crotonylation/decrotonylation, must be involved in DSBs damage-induced Kcr of RPA1 and subsequent regulated DSB repair, which need to be revealed in future efforts.
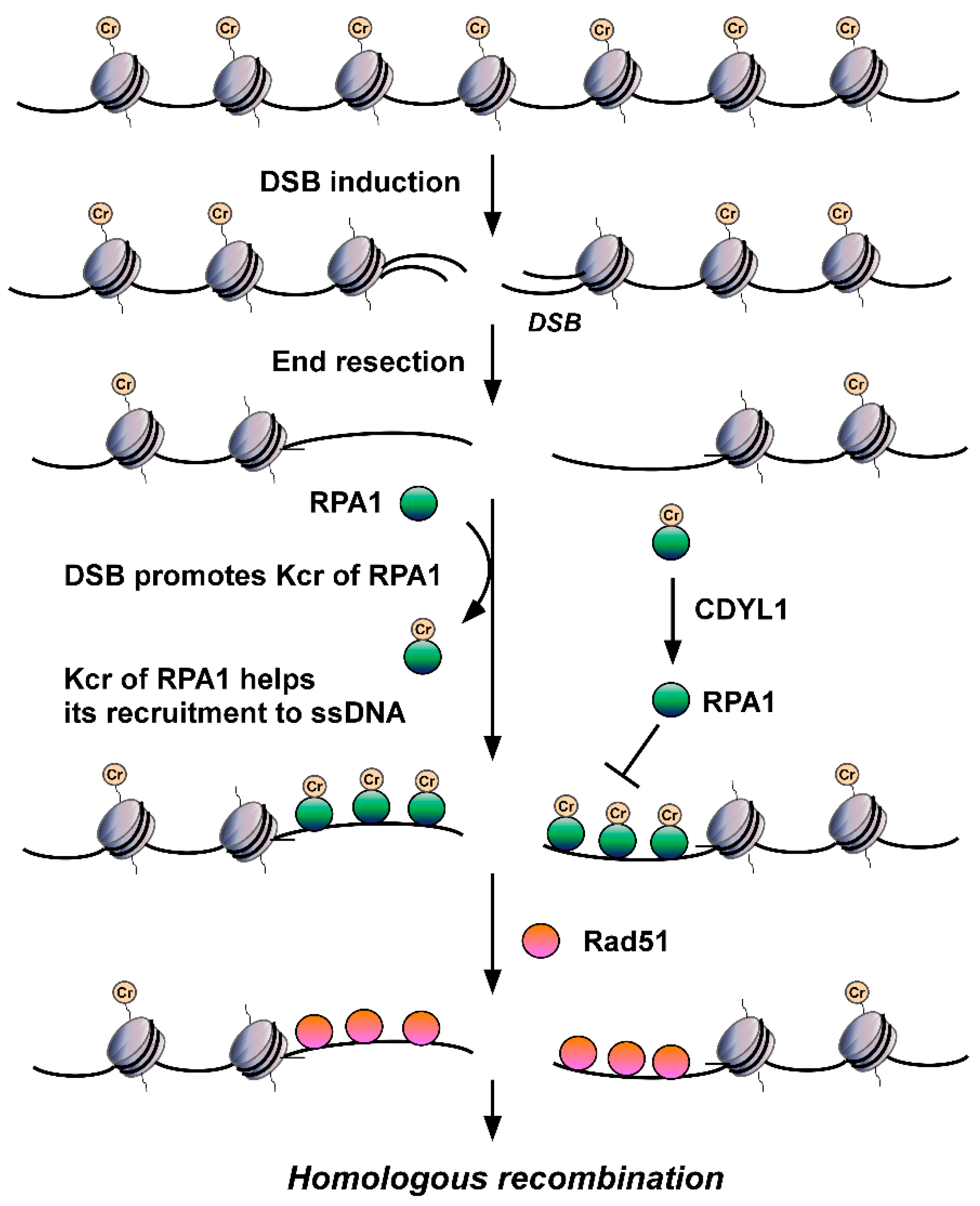
Figure 1. The roles of RPA Kcr in DSB repair. DSB injury promotes Kcr of RPA1. Kcr can increase the affinity of RPA for ssDNA, facilitate its recruitment to ssDNA generated by DSB end resection, enhance its interactions with major HR factors, such as RAD51, and promote HR-mediated DSB repair.
References
- Hoeijmakers, J.H. DNA damage, aging, and cancer. N. Engl. J. Med. 2009, 361, 1475–1485.
- Varon, R.; Vissinga, C.; Platzer, M.; Cerosaletti, K.M.; Chrzanowska, K.H.; Saar, K.; Beckmann, G.; Seemanova, E.; Cooper, P.R.; Nowak, N.J.; et al. Nibrin, a novel DNA double-strand break repair protein, is mutated in Nijmegen breakage syndrome. Cell 1998, 93, 467–476.
- Savitsky, K.; Bar-Shira, A.; Gilad, S.; Rotman, G.; Ziv, Y.; Vanagaite, L.; Tagle, D.A.; Smith, S.; Uziel, T.; Sfez, S.; et al. A single ataxia telangiectasia gene with a product similar to PI-3 kinase. Science 1995, 268, 1749–1753.
- Ciccia, A.; Elledge, S.J. The DNA damage response: Making it safe to play with knives. Mol. Cell 2010, 40, 179–204.
- Jackson, S.P.; Bartek, J. The DNA-damage response in human biology and disease. Nature 2009, 461, 1071–1078.
- Wang, W. Emergence of a DNA-damage response network consisting of Fanconi anaemia and BRCA proteins. Nat. Rev. Genet. 2007, 8, 735–748.
- Harper, J.W.; Elledge, S.J. The DNA damage response: Ten years after. Mol. Cell 2007, 28, 739–745.
- Huen, M.S.; Chen, J. The DNA damage response pathways: At the crossroad of protein modifications. Cell Res. 2008, 18, 8–16.
- Gong, F.; Miller, K.M. Histone methylation and the DNA damage response. Mutat. Res. Rev. Mutat. Res. 2019, 780, 37–47.
- Wang, Z.; Zhu, W.G.; Xu, X. Ubiquitin-like modifications in the DNA damage response. Mutat. Res. 2017, 803–805, 56–75.
- Brown, J.S.; Jackson, S.P. Ubiquitylation, neddylation and the DNA damage response. Open Biol. 2015, 5, 150018.
- Hou, W.H.; Chen, S.H.; Yu, X. Poly-ADP ribosylation in DNA damage response and cancer therapy. Mutat. Res. Rev. Mutat. Res. 2019, 780, 82–91.
- Kolobynina, K.G.; Rapp, A.; Cardoso, M.C. Chromatin Ubiquitination Guides DNA Double Strand Break Signaling and Repair. Front Cell Dev. Biol. 2022, 10, 928113.
- Kim, J.J.; Lee, S.Y.; Miller, K.M. Preserving genome integrity and function: The DNA damage response and histone modifications. Crit. Rev. Biochem. Mol. Biol. 2019, 54, 208–241.
- Tang, J.; Cho, N.W.; Cui, G.; Manion, E.M.; Shanbhag, N.M.; Botuyan, M.V.; Mer, G.; Greenberg, R.A. Acetylation limits 53BP1 association with damaged chromatin to promote homologous recombination. Nat. Struct. Mol. Biol. 2013, 20, 317–325.
- Jacquet, K.; Fradet-Turcotte, A.; Avvakumov, N.; Lambert, J.P.; Roques, C.; Pandita, R.K.; Paquet, E.; Herst, P.; Gingras, A.C.; Pandita, T.K.; et al. The TIP60 Complex Regulates Bivalent Chromatin Recognition by 53BP1 through Direct H4K20me Binding and H2AK15 Acetylation. Mol. Cell 2016, 62, 409–421.
- Miller, K.M.; Tjeertes, J.V.; Coates, J.; Legube, G.; Polo, S.E.; Britton, S.; Jackson, S.P. Human HDAC1 and HDAC2 function in the DNA-damage response to promote DNA nonhomologous end-joining. Nat. Struct. Mol. Biol. 2010, 17, 1144–1151.
- Elia, A.E.; Boardman, A.P.; Wang, D.C.; Huttlin, E.L.; Everley, R.A.; Dephoure, N.; Zhou, C.; Koren, I.; Gygi, S.P.; Elledge, S.J. Quantitative Proteomic Atlas of Ubiquitination and Acetylation in the DNA Damage Response. Mol. Cell 2015, 59, 867–881.
- Gong, F.; Chiu, L.Y.; Miller, K.M. Acetylation Reader Proteins: Linking Acetylation Signaling to Genome Maintenance and Cancer. PLoS Genet 2016, 12, e1006272.
- Sabari, B.R.; Zhang, D.; Allis, C.D.; Zhao, Y. Metabolic regulation of gene expression through histone acylations. Nat. Rev. Mol. Cell Biol. 2017, 18, 90–101.
- Huang, H.; Sabari, B.R.; Garcia, B.A.; Allis, C.D.; Zhao, Y. SnapShot: Histone modifications. Cell 2014, 159, 458.
- Chen, Y.; Sprung, R.; Tang, Y.; Ball, H.; Sangras, B.; Kim, S.C.; Falck, J.R.; Peng, J.; Gu, W.; Zhao, Y. Lysine propionylation and butyrylation are novel post-translational modifications in histones. Mol. Cell Proteomics 2007, 6, 812–819.
- Dai, L.; Peng, C.; Montellier, E.; Lu, Z.; Chen, Y.; Ishii, H.; Debernardi, A.; Buchou, T.; Rousseaux, S.; Jin, F.; et al. Lysine 2-hydroxyisobutyrylation is a widely distributed active histone mark. Nat. Chem. Biol. 2014, 10, 365–370.
- Xie, Z.; Dai, J.; Dai, L.; Tan, M.; Cheng, Z.; Wu, Y.; Boeke, J.D.; Zhao, Y. Lysine succinylation and lysine malonylation in histones. Mol. Cell Proteomics 2012, 11, 100–107.
- Tan, M.; Peng, C.; Anderson, K.A.; Chhoy, P.; Xie, Z.; Dai, L.; Park, J.; Chen, Y.; Huang, H.; Zhang, Y.; et al. Lysine glutarylation is a protein posttranslational modification regulated by SIRT5. Cell Metab. 2014, 19, 605–617.
- Tan, M.; Luo, H.; Lee, S.; Jin, F.; Yang, J.S.; Montellier, E.; Buchou, T.; Cheng, Z.; Rousseaux, S.; Rajagopal, N.; et al. Identification of 67 histone marks and histone lysine crotonylation as a new type of histone modification. Cell 2011, 146, 1016–1028.
- Xie, Z.; Zhang, D.; Chung, D.; Tang, Z.; Huang, H.; Dai, L.; Qi, S.; Li, J.; Colak, G.; Chen, Y.; et al. Metabolic Regulation of Gene Expression by Histone Lysine beta-Hydroxybutyrylation. Mol. Cell 2016, 62, 194–206.
- Jiang, G.; Nguyen, D.; Archin, N.M.; Yukl, S.A.; Mendez-Lagares, G.; Tang, Y.; Elsheikh, M.M.; Thompson, G.R., 3rd; Hartigan-O’Connor, D.J.; Margolis, D.M.; et al. HIV latency is reversed by ACSS2-driven histone crotonylation. J. Clin. Investig. 2018, 128, 1190–1198.
- Ntorla, A.; Burgoyne, J.R. The Regulation and Function of Histone Crotonylation. Front. Cell Dev. Biol. 2021, 9, 624914.
- Wan, J.; Liu, H.; Chu, J.; Zhang, H. Functions and mechanisms of lysine crotonylation. J. Cell Mol. Med. 2019, 23, 7163–7169.
- Ruiz-Andres, O.; Sanchez-Nino, M.D.; Cannata-Ortiz, P.; Ruiz-Ortega, M.; Egido, J.; Ortiz, A.; Sanz, A.B. Histone lysine crotonylation during acute kidney injury in mice. Dis. Model Mech. 2016, 9, 633–645.
- Li, K.; Wang, Z. Histone crotonylation-centric gene regulation. Epigenetics Chromatin 2021, 14, 10.
- Hou, J.Y.; Zhou, L.; Li, J.L.; Wang, D.P.; Cao, J.M. Emerging roles of non-histone protein crotonylation in biomedicine. Cell Biosci. 2021, 11, 101.
- Xu, W.; Wan, J.; Zhan, J.; Li, X.; He, H.; Shi, Z.; Zhang, H. Global profiling of crotonylation on non-histone proteins. Cell Res. 2017, 27, 946–949.
- Kwon, O.K.; Kim, S.J.; Lee, S. First profiling of lysine crotonylation of myofilament proteins and ribosomal proteins in zebrafish embryos. Sci. Rep. 2018, 8, 3652.
- Chen, W.; Tang, D.; Xu, Y.; Zou, Y.; Sui, W.; Dai, Y.; Diao, H. Comprehensive analysis of lysine crotonylation in proteome of maintenance hemodialysis patients. Medicine 2018, 97, e12035.
- Sun, J.; Qiu, C.; Qian, W.; Wang, Y.; Sun, L.; Li, Y.; Ding, Z. Ammonium triggered the response mechanism of lysine crotonylome in tea plants. BMC Genomics 2019, 20, 340.
- Lin, H.; Tang, D.; Xu, Y.; Zhang, R.; Ou, M.; Zheng, F.; Chen, J.; Zhang, Y.; Zou, G.; Xue, W.; et al. Quantitative analysis of protein crotonylation identifies its association with immunoglobulin A nephropathy. Mol. Med. Rep. 2020, 21, 1242–1250.
- Liu, K.; Yuan, C.; Li, H.; Chen, K.; Lu, L.; Shen, C.; Zheng, X. A qualitative proteome-wide lysine crotonylation profiling of papaya (Carica papaya L.). Sci. Rep. 2018, 8, 8230.
- Liu, S.; Xue, C.; Fang, Y.; Chen, G.; Peng, X.; Zhou, Y.; Chen, C.; Liu, G.; Gu, M.; Wang, K.; et al. Global Involvement of Lysine Crotonylation in Protein Modification and Transcription Regulation in Rice. Mol. Cell Proteomics 2018, 17, 1922–1936.
- Yin, D.; Jiang, N.; Zhang, Y.; Wang, D.; Sang, X.; Feng, Y.; Chen, R.; Wang, X.; Yang, N.; Chen, Q. Global Lysine Crotonylation and 2-Hydroxyisobutyrylation in Phenotypically Different Toxoplasma gondii Parasites. Mol. Cell Proteomics 2019, 18, 2207–2224.
- Grunstein, M. Histone acetylation in chromatin structure and transcription. Nature 1997, 389, 349–352.
- Bonnaud, E.M.; Suberbielle, E.; Malnou, C.E. Histone acetylation in neuronal (dys)function. Biomol. Concepts 2016, 7, 103–116.
- Koprinarova, M.; Schnekenburger, M.; Diederich, M. Role of Histone Acetylation in Cell Cycle Regulation. Curr. Top Med. Chem. 2016, 16, 732–744.
- Choi, J.K.; Howe, L.J. Histone acetylation: Truth of consequences? Biochem. Cell Biol. 2009, 87, 139–150.
- Wade, P.A.; Pruss, D.; Wolffe, A.P. Histone acetylation: Chromatin in action. Trends Biochem. Sci. 1997, 22, 128–132.
- Gong, F.; Miller, K.M. Mammalian DNA repair: HATs and HDACs make their mark through histone acetylation. Mutat. Res. 2013, 750, 23–30.
- Roth, S.Y.; Denu, J.M.; Allis, C.D. Histone acetyltransferases. Annu. Rev. Biochem. 2001, 70, 81–120.
- Ogryzko, V.V. Mammalian histone acetyltransferases and their complexes. Cell Mol. Life Sci. 2001, 58, 683–692.
- Marmorstein, R.; Roth, S.Y. Histone acetyltransferases: Function, structure, and catalysis. Curr. Opin. Genet. Dev. 2001, 11, 155–161.
- Marmorstein, R. Structure and function of histone acetyltransferases. Cell Mol Life Sci 2001, 58, 693–703.
- Aka, J.A.; Kim, G.W.; Yang, X.J. K-acetylation and its enzymes: Overview and new developments. Handb. Exp. Pharmacol. 2011, 206, 1–12.
- Chen, H.P.; Zhao, Y.T.; Zhao, T.C. Histone deacetylases and mechanisms of regulation of gene expression. Crit. Rev. Oncog. 2015, 20, 35–47.
- De Ruijter, A.J.; van Gennip, A.H.; Caron, H.N.; Kemp, S.; van Kuilenburg, A.B. Histone deacetylases (HDACs): Characterization of the classical HDAC family. Biochem. J. 2003, 370, 737–749.
- Seto, E.; Yoshida, M. Erasers of histone acetylation: The histone deacetylase enzymes. Cold Spring Harb. Perspect. Biol. 2014, 6, a018713.
- Shvedunova, M.; Akhtar, A. Modulation of cellular processes by histone and non-histone protein acetylation. Nat. Rev. Mol. Cell Biol. 2022, 23, 329–349.
- Zhao, S.; Zhang, X.; Li, H. Beyond histone acetylation-writing and erasing histone acylations. Curr. Opin. Struct. Biol. 2018, 53, 169–177.
- Sabari, B.R.; Tang, Z.; Huang, H.; Yong-Gonzalez, V.; Molina, H.; Kong, H.E.; Dai, L.; Shimada, M.; Cross, J.R.; Zhao, Y.; et al. Intracellular crotonyl-CoA stimulates transcription through p300-catalyzed histone crotonylation. Mol. Cell 2015, 58, 203–215.
- Liu, X.; Wei, W.; Liu, Y.; Yang, X.; Wu, J.; Zhang, Y.; Zhang, Q.; Shi, T.; Du, J.X.; Zhao, Y.; et al. MOF as an evolutionarily conserved histone crotonyltransferase and transcriptional activation by histone acetyltransferase-deficient and crotonyltransferase-competent CBP/p300. Cell Discov. 2017, 3, 17016.
- Kollenstart, L.; de Groot, A.J.L.; Janssen, G.M.C.; Cheng, X.; Vreeken, K.; Martino, F.; Cote, J.; van Veelen, P.A.; van Attikum, H. Gcn5 and Esa1 function as histone crotonyltransferases to regulate crotonylation-dependent transcription. J. Biol. Chem. 2019, 294, 20122–20134.
- Xiao, Y.; Li, W.; Yang, H.; Pan, L.; Zhang, L.; Lu, L.; Chen, J.; Wei, W.; Ye, J.; Li, J.; et al. HBO1 is a versatile histone acyltransferase critical for promoter histone acylations. Nucleic Acids Res. 2021, 49, 8037–8059.
- Yang, X.J.; Seto, E. The Rpd3/Hda1 family of lysine deacetylases: From bacteria and yeast to mice and men. Nat. Rev. Mol. Cell Biol. 2008, 9, 206–218.
- Houtkooper, R.H.; Pirinen, E.; Auwerx, J. Sirtuins as regulators of metabolism and healthspan. Nat. Rev. Mol. Cell Biol. 2012, 13, 225–238.
- Wei, W.; Liu, X.; Chen, J.; Gao, S.; Lu, L.; Zhang, H.; Ding, G.; Wang, Z.; Chen, Z.; Shi, T.; et al. Class I histone deacetylases are major histone decrotonylases: Evidence for critical and broad function of histone crotonylation in transcription. Cell Res. 2017, 27, 898–915.
- Kelly, R.D.W.; Chandru, A.; Watson, P.J.; Song, Y.; Blades, M.; Robertson, N.S.; Jamieson, A.G.; Schwabe, J.W.R.; Cowley, S.M. Histone deacetylase (HDAC) 1 and 2 complexes regulate both histone acetylation and crotonylation in vivo. Sci. Rep. 2018, 8, 14690.
- Hao, S.; Wang, Y.; Zhao, Y.; Gao, W.; Cui, W.; Li, Y.; Cui, J.; Liu, Y.; Lin, L.; Xu, X.; et al. Dynamic switching of crotonylation to ubiquitination of H2A at lysine 119 attenuates transcription-replication conflicts caused by replication stress. Nucleic Acids Res. 2022, 50, 9873–9892.
- Bao, X.; Wang, Y.; Li, X.; Li, X.M.; Liu, Z.; Yang, T.; Wong, C.F.; Zhang, J.; Hao, Q.; Li, X.D. Identification of ’erasers’ for lysine crotonylated histone marks using a chemical proteomics approach. Elife 2014, 3, e02999.
- Fellows, R.; Denizot, J.; Stellato, C.; Cuomo, A.; Jain, P.; Stoyanova, E.; Balazsi, S.; Hajnady, Z.; Liebert, A.; Kazakevych, J.; et al. Microbiota derived short chain fatty acids promote histone crotonylation in the colon through histone deacetylases. Nat. Commun. 2018, 9, 105.
- Wagner, G.R.; Hirschey, M.D. Nonenzymatic protein acylation as a carbon stress regulated by sirtuin deacylases. Mol. Cell 2014, 54, 5–16.
- Baddiley, J.; Kekwick, R.A.; Thain, E.M. A new method for acetylating proteins. Nature 1952, 170, 968–970.
- Liu, S.; Yu, H.; Liu, Y.; Liu, X.; Zhang, Y.; Bu, C.; Yuan, S.; Chen, Z.; Xie, G.; Li, W.; et al. Chromodomain Protein CDYL Acts as a Crotonyl-CoA Hydratase to Regulate Histone Crotonylation and Spermatogenesis. Mol. Cell 2017, 67, 853–866.e5.
- Fang, Y.; Xu, X.; Ding, J.; Yang, L.; Doan, M.T.; Karmaus, P.W.F.; Snyder, N.W.; Zhao, Y.; Li, J.L.; Li, X. Histone crotonylation promotes mesoendodermal commitment of human embryonic stem cells. Cell Stem Cell 2021, 28, 748–763.e7.
- Baumann, K. Post-translational modifications: Crotonylation versus acetylation. Nat. Rev. Mol. Cell Biol. 2015, 16, 265.
- Li, Y.; Sabari, B.R.; Panchenko, T.; Wen, H.; Zhao, D.; Guan, H.; Wan, L.; Huang, H.; Tang, Z.; Zhao, Y.; et al. Molecular Coupling of Histone Crotonylation and Active Transcription by AF9 YEATS Domain. Mol. Cell 2016, 62, 181–193.
- Andrews, F.H.; Shinsky, S.A.; Shanle, E.K.; Bridgers, J.B.; Gest, A.; Tsun, I.K.; Krajewski, K.; Shi, X.; Strahl, B.D.; Kutateladze, T.G. The Taf14 YEATS domain is a reader of histone crotonylation. Nat. Chem. Biol. 2016, 12, 396–398.
- Zhao, D.; Guan, H.; Zhao, S.; Mi, W.; Wen, H.; Li, Y.; Zhao, Y.; Allis, C.D.; Shi, X.; Li, H. YEATS2 is a selective histone crotonylation reader. Cell Res. 2016, 26, 629–632.
- Schulze, J.M.; Wang, A.Y.; Kobor, M.S. YEATS domain proteins: A diverse family with many links to chromatin modification and transcription. Biochem. Cell Biol. 2009, 87, 65–75.
- Li, Y.; Wen, H.; Xi, Y.; Tanaka, K.; Wang, H.; Peng, D.; Ren, Y.; Jin, Q.; Dent, S.Y.; Li, W.; et al. AF9 YEATS domain links histone acetylation to DOT1L-mediated H3K79 methylation. Cell 2014, 159, 558–571.
- Xiong, X.; Panchenko, T.; Yang, S.; Zhao, S.; Yan, P.; Zhang, W.; Xie, W.; Li, Y.; Zhao, Y.; Allis, C.D.; et al. Selective recognition of histone crotonylation by double PHD fingers of MOZ and DPF2. Nat. Chem. Biol. 2016, 12, 1111–1118.
- Klein, B.J.; Jang, S.M.; Lachance, C.; Mi, W.; Lyu, J.; Sakuraba, S.; Krajewski, K.; Wang, W.W.; Sidoli, S.; Liu, J.; et al. Histone H3K23-specific acetylation by MORF is coupled to H3K14 acylation. Nat. Commun. 2019, 10, 4724.
- Yu, H.; Bu, C.; Liu, Y.; Gong, T.; Liu, X.; Liu, S.; Peng, X.; Zhang, W.; Peng, Y.; Yang, J.; et al. Global crotonylome reveals CDYL-regulated RPA1 crotonylation in homologous recombination-mediated DNA repair. Sci. Adv. 2020, 6, eaay4697.
- Abu-Zhayia, E.R.; Bishara, L.A.; Machour, F.E.; Barisaac, A.S.; Ben-Oz, B.M.; Ayoub, N. CDYL1-dependent decrease in lysine crotonylation at DNA double-strand break sites functionally uncouples transcriptional silencing and repair. Mol. Cell 2022, 82, 1940–1955.e7.
More
Information
Subjects:
Biochemistry & Molecular Biology
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
828
Revisions:
2 times
(View History)
Update Date:
09 Nov 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





