Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Cosmos Ifeanyi Onyiba | -- | 5409 | 2022-10-25 11:01:44 | | | |
| 2 | Amina Yu | + 3 word(s) | 5412 | 2022-11-03 07:58:20 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Onyiba, C.I.; Scarlett, C.J.; Weidenhofer, J. Sirtuins in Breast and Prostate Cancer. Encyclopedia. Available online: https://encyclopedia.pub/entry/32589 (accessed on 15 May 2026).
Onyiba CI, Scarlett CJ, Weidenhofer J. Sirtuins in Breast and Prostate Cancer. Encyclopedia. Available at: https://encyclopedia.pub/entry/32589. Accessed May 15, 2026.
Onyiba, Cosmos Ifeanyi, Christopher J. Scarlett, Judith Weidenhofer. "Sirtuins in Breast and Prostate Cancer" Encyclopedia, https://encyclopedia.pub/entry/32589 (accessed May 15, 2026).
Onyiba, C.I., Scarlett, C.J., & Weidenhofer, J. (2022, November 03). Sirtuins in Breast and Prostate Cancer. In Encyclopedia. https://encyclopedia.pub/entry/32589
Onyiba, Cosmos Ifeanyi, et al. "Sirtuins in Breast and Prostate Cancer." Encyclopedia. Web. 03 November, 2022.
Copy Citation
In mammals, seven sirtuins (SIRT1–7) have been identified, which primarily function as NAD-dependent deacetylases (SIRT1–3 and SIRT5–7) and ADP-ribosyl transferases (SIRT4 and 6). Additionally, sirtuins have been reported to function as demyristoylases (SIRT1–3 and 6), lipoamidases (SIRT4), and desuccinylases/demalonylases/deglutarylases (SIRT5). The forcus herein is the information on the tumor promotion or suppression roles of SIRT1–7 in breast and prostate cancers. Precisely, it is highlighted that sirtuins regulate various proteins implicated in proliferation, apoptosis, autophagy, chemoresistance, invasion, migration, and metastasis of both breast and prostate cancer.
sirtuins
breast
prostate
cancer
tumor
miRNA
suppression
promotion
1. Introduction
The silent information regulator 2 (sir2) family of proteins, simply known as sirtuins (sir-two-ins), was first discovered in yeast, where its upregulation increased lifespan[1]. Consistent with this finding, subsequent studies revealed that sirtuins promote longevity in Caenorhabditis elegans and Drosophila melanogaster[2]. As members of the highly conserved class III histone deacetylase, sirtuins, in addition to their initial description as mono-adenosine diphosphate (ADP) transferases, were later found to be a nicotinamide (NAM) adenine dinucleotide (NAD)-dependent group of enzymes[3]. Sirtuins are reported to play a dual role in the pathogenesis of breast and prostate cancer via the targeting of different proteins, including transcription factors, signaling molecules, and catalytic proteins. Consequently, sirtuins have exhibited both tumor-suppressing and tumor-promoting effects in breast and prostate cancer cells (Figure 1). In this section, the mechanistic roles of nuclear (SIRT1, 6, and 7), cytoplasmic (SIRT2), and mitochondrial sirtuins (SIRT3, 4, and 5) in breast and prostate cancer will be discussed.
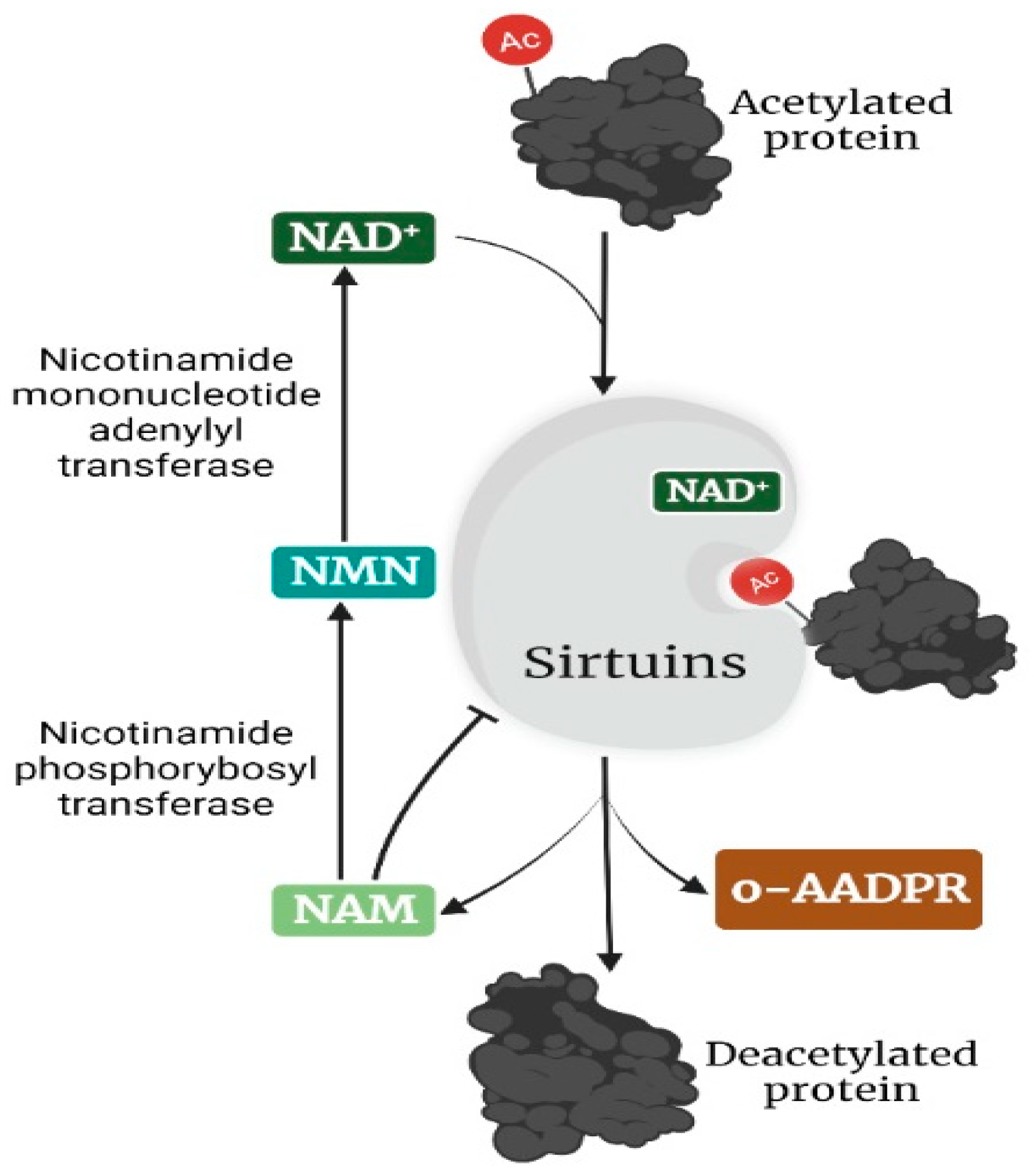
Figure 1. Outcomes of sirtuins modulation of various targets in breast (A) and prostate (B) cancers. SIRT1–7 regulate the expression and activities of various targets in breast (A) and prostate (B) cancer, resulting in either tumor promotion or suppression. ?—Not Known. The image was created with BioRender.com.
2. Nuclear Sirtuins in Breast Cancer
2.1. SIRT1
Recent studies have revealed that SIRT1 is constitutively upregulated in breast cancer cells [4][5][6]. Additionally, SIRT1 has been shown to modulate different targets in breast cancer, including transcription factors, signaling molecules, and certain enzymes, showing opposing effects by its exhibition of either tumor promotion or tumor suppression. The most prominent transcription factors targeted by SIRT1 in breast cancer include forkhead box O3 (FOXO3), paired related homeobox 1 (PRRX1), and tumor suppressor proteins (p53 and p21). For instance, Mahmud et al. demonstrated that the downstream anti-proliferative activity of acetylated FOXO3, a tumor suppressor whose activity is vulnerable to posttranslational modifications, is activated upon SIRT1 gene silencing in BT474 breast cancer cells, suggesting that SIRT1 deacetylates and inhibits FOXO3 activity to promote breast cancer [7]. They also demonstrated that SIRT1 gene silencing increases BT474 cells’ sensitivity to lapatinib treatment, further suggesting that SIRT1 promotes BT474 cells’ resistance to lapatinib treatment. The protein expression of PRRX1, a transcription factor that induces epithelial-to-mesenchymal transition (EMT) and inhibits cancer stemness, was shown to be downregulated by SIRT1 gene silencing in mouse 4T1 breast cancer cells [8]. Interestingly, SIRT1 gene silencing upregulated the expression of the EMT-related protein E-cadherin and cancer stemness markers kruppel-like factor 4 (KLF4) and aldehyde dehydrogenase 1 family member A1 (ALDH1A1) [8]. Based on their findings, it was concluded that SIRT1 deacetylates and stabilizes PRRX1 to inhibit breast cancer stemness and metastasis. Importantly, inhibition of SIRT1 by sirtinol, a classic inhibitor of sirtuins, resulted in the activation of p53 and consequently increased p21 protein levels in SK-BR-3 breast cancer cells, thus suggesting a tumor-promoting role of SIRT1 in breast cancer [9]. Moreover, SIRT1 expression has been previously shown to downregulate tumor suppressor genes, such as cyclin G2 and p53, in estrogen-dependent breast cancer cells [10].
SIRT1 has been demonstrated to modulate several intracellular signaling protein molecules, including protein kinase B (PKB), B-cell lymphoma 2 (Bcl-2), metadherin (MTDH), Frizzled 7, EMT-related proteins, and the cluster of differentiation 36 (CD36), in breast cancer cells. In a study by Jin et al., SIRT1 was shown to directly interact with and phosphorylate PKB to promote subsequent downstream proliferative effects in MDA-MB-231 and BT-549 breast cancer cells [5]. To substantiate this finding, the authors further investigated Akt activities in SIRT1-knockdown breast cancer cells and found that proliferation was inhibited both in vitro and in vivo. Conversely, a previous study reported that SIRT1 represses estrogen receptor activation by deactivating Akt activity to suppress the growth of MCF-7 breast cancer cells [11]. These contradictory results may be due to genetic differences between the breast cancer cells used in the studies. However, further investigations are required to illuminate the possible reasons for these contradictions. The expression of Bcl-2, an anti-apoptotic or pro-survival protein, was markedly decreased in MCF-7 and MDA-MB-231 breast cancer cells following the inhibition of SIRT1 with sirtinol [12], which indicates that SIRT1 promotes the growth of breast cancer through the upregulation of the Bcl-2 protein.
Inhibition of SIRT1 with either EX-527 or SIRT1 siRNA resulted in the upregulation of c-Myc (a pro-oncogenic protein) and MTDH (an oncogenic protein) in MDA-MB-231 and BT-549 breast cancer cells. Further, in the presence of c-Myc siRNA, EX-527 could not upregulate MTDH in breast cancer cells [13]. As a mechanism, it was suggested that the SIRT1-mediated inhibition of the proliferation of the breast cancer cells was due to the downregulation of c-Myc, which resulted in the downregulation of MTDH. Simmons et al. demonstrated that the inhibition of SIRT1 by both inhibitor VII and SIRT1 siRNA markedly downregulated the expression of Frizzled 7, a chronically activated oncogenic receptor in cancers [14]. This finding supports the tumor-promoting role of SIRT1 in breast cancer. Furthermore, another research group investigated the effect of SIRT1 inhibition by SIRT1 siRNA on EMT-related proteins (vimentin, snail-1, and E-cadherin) in MDA-MB-231, MDA-MB-436, and MDA-MB-468 breast cancer cell lines and found that the siRNA inhibition of SIRT1 downregulated the expression of vimentin (a protein required for cell migration, motility, and adhesion) and snail-1 (a repressor of E-cadherin) but upregulated the expression of E-cadherin (a cell-cell adhesion molecule), indicating that SIRT1 induces the invasion of breast cancer cells by modulating the EMT pathway [15]. On the contrary, SIRT1 overexpression by resveratrol (a classical activator of sirtuins) was reported to increase the expression of CD36 (a transmembrane receptor that regulates apoptosis and angiogenesis) to cytotoxic levels, thus inhibiting the proliferation of MCF-7 breast cancer cells [6]. Moreover, there is evidence that resveratrol induces toxicity in breast cancer cells through a SIRT1-dependent mechanism [16][17].
Interestingly, SIRT1 has also been demonstrated to regulate the expression of certain catalytic proteins in breast cancer. For instance, the transcription of CYP19A1 (a gene encoding the aromatase enzyme) was downregulated in MDA-MB-231 breast cancer cells expressing aromatase following the inhibition of SIRT1 by treatment with both cambinol and inhibitor VII [18]. This finding indicates that SIRT1 promotes the growth of estrogen-dependent breast cancer through the upregulation of CYP19A1 expression. Additionally, another study demonstrated that the knock-out of SIRT1 by shRNA reduced the proliferation, migration, and invasion of MCF-7 breast cancer cells [4]. The shRNA-mediated silencing of SIRT1 downregulated the expression of human DNA polymerase delta 1 (POLD1), a gene that encodes p125 (the catalytic domain of DNA polymerase δ) and regulates cell proliferation and the cell cycle. SIRT1 upregulated the activities of POLD1/p125 to aid the proliferation, migration, and invasion of MCF-7 breast cancer cells. Recently, a study demonstrated that CRISPR-Cas9-mediated deletion of SIRT1 promotes the release of exosomes (containing unique cargoes and hydrolases that degrade the extracellular matrix), impairs lysosomal function, and inhibits lysosomal acidification in MDA-MB-231 breast cancer cells [19]. It was showed that the deletion of SIRT1 reduces the expression of the vacuolar-type H+ ATPase (V-ATPase) subunit responsible for proper lysosomal acidification and protein degradation. This indicates that SIRT1 upregulated V-ATPase activities to increase lysosomal function, which halted the secretion of exosomes containing hydrolases that degrade the extracellular matrix, thereby ultimately inhibiting the aggressiveness of MDA-MB-231 breast cancer cells.
2.2. SIRT6
Although previous studies have demonstrated that SIRT6 expression is downregulated in several human cancers [20], including invasive breast tumors [21], and that SIRT6 expression positively correlates with the survival of breast cancer patients [22], there are few studies demonstrating the mechanistic effects of SIRT6 in breast cancer. Ioris et al. demonstrated that the overexpression of SIRT6 downregulates the transcription of PI3K-controlled genes, independent of its deacetylase activity, to suppress the progression and stem cell-like capacity of PyMT-induced breast cancer in mice, thus indicating a tumor suppressor role of SIRT6 in breast cancer [23]. Conversely, a study recently demonstrated that SIRT6 acts as a tumor promoter, given that its expression promoted MCF-7 cell growth as well as resistance to oxidative stress [24]. These effects were driven by SIRT6 deacetylation and activation of nicotinamide phosphoribosyl transferase and glucose-6-phosphate dehydrogenase, which increase NADH and NADPH levels, respectively, to protect against oxidative stress. Corroborating the tumor-promoting activities of SIRT6, it was recently demonstrated that SIRT6 deletion downregulated the expression and activity of pyruvate dehydrogenase and oxidative phosphorylation-related genes, suppressed the activity of respiratory complexes, and reduced the ATP/AMP ratio in MDA-MB-231 and MCF7 breast cancer cells [25]. The above findings reveal a vital role of SIRT6 in remodeling the metabolic profile to consistently provide energy and nutrients to drive the growth of breast cancer. Moreover, low SIRT6 expression was recently found to be associated with better overall survival of patients with breast cancer [26]. Notwithstanding, more studies are required to investigate the expression level of SIRT6 and its effect on the metabolic profile of breast cancer at various stages of carcinogenesis.
2.3. SIRT7
The mRNA expression level of SIRT7 was found to be higher in patients with metastatic breast cancer [27]. More recently, high expression of SIRT7 was associated with poor prognosis of breast cancer [28][29] and positively correlated with increased M1 macrophage infiltration and exhaustion of T-cells in luminal breast cancer [29]. There are conflicting reports on the function of SIRT7 in breast cancer. In BT-549 and MDA-MB-231 breast cancer cells, SIRT7 deacetylated and destabilized SMAD4, resulting in reduced formation of the SMAD2/SMAD3-SMAD4 complex. On translocation to the nucleus, this complex activates EMT-related transcription factors (such as Snail1/2, Twist1/2, and Zeb1/2), thus facilitating the migration and invasion of cancer cells [30]. It was also demonstrated that the deacetylation and destabilization of SMAD4 promote its degradation by β-TrCP1, a substrate recognition subunit of E3 ubiquitin ligase. Additionally, it was corroborated their findings with knockout experiments, in which SIRT7 knockout with either shSIRT7 or siSIRT7 activated TGF-β signaling to promote EMT-related gene transcription and lung metastasis of breast cancer. Taken together, it was demonstrated a tumor-suppressing role of SIRT7 in breast cancer. Furthermore, a recent study demonstrated that SIRT7 antagonizes TGF-β signaling and inhibits metastasis of breast cancer [28]. On the contrary, a more recent study revealed that SIRT7 promotes Adriamycin-induced metastasis in breast cancer by interacting with TIE2 [31], a tyrosine kinase receptor that induced the dormancy of breast cancer cells, resulting in increased resistance to chemotherapy [32]. Given the limited studies conducted to date and contradictory findings, more studies are required to provide insight into the modulation of other signaling proteins (including transcription factors) by SIRT7 causing either tumor-suppressing or tumor-promoting effects in breast cancer.
3. Nuclear Sirtuins in Prostate Cancer
3.1. SIRT1
Several studies have shown that SIRT1 is highly expressed in prostate cancer [33][34][35][36], suggesting an important role for SIRT1 in its progression. Regardless of its predominant expression in prostate cancer cells, SIRT1 has been demonstrated to either promote or suppress prostate cancer via different mechanisms. Although the mechanisms were not fully elucidated, studies reported that SIRT1 promotes cell growth and chemoresistance in PC3 and DU145 prostate cancer cells [36] as well as the migration and invasion of DU145 prostate cancer cells [37]. A mechanistic study, using PC3 prostate cancer cells with active p53, or PC3 prostate cancer cells regardless of p53 status, demonstrated that SIRT1 inhibition by SIRT1 shRNA or sirtinol downregulates the deacetylation of p53 and FOXO1, thereby activating antiproliferative responses, such as apoptosis and senescence [33]. This suggests that SIRT1 expression may promote the development of prostate cancer by deacetylating and deactivating p53 and FOXO1. Moreover, using DU145 prostate cancer cells, it was previously demonstrated that SIRT1 deacetylates and deactivates FOXO1 transcription activities [38]. To further corroborate these findings, later studies demonstrated that SIRT1 deacetylates FOXO3, a tumor suppressor transcription factor that has been shown to inhibit cell proliferation and promote apoptosis [39], to promote its poly-ubiquitination and subsequent proteasomal degradation [40]. These findings suggest that SIRT1 may mediate its tumor-promoting activities in prostate cancer by preventing the tumor-suppressing function of FOXOs. Nonetheless, animal studies are required to investigate how the tumor microenvironment of prostate cancer influences the outcome of SIRT1 activities in this regard.
The tumor-promoting effects of SIRT1 in prostate cancer have been demonstrated through the regulation of EMT-related protein expression. A 2012 study revealed that SIRT1 promotes the migration and metastasis of DU145 and PC3 prostate cancer cells by downregulating the expression of the epithelial marker E-cadherin and upregulating the expression of mesenchymal markers (N-cadherin and fibronectin) and the EMT-inducing transcription factor ZEB1 [41]. Similarly, Cui et al. [42] showed that siRNA-mediated silencing of SIRT1 expression suppressed the migration and invasion of PC3 prostate cancer cells by upregulating E-cadherin and downregulating N-cadherin and vimentin expression, thus suggesting that SIRT1 activates EMT to promote the migration and metastasis of prostate cancer. Additionally, SIRT1 can promote the migration and invasion of PC3 prostate cancer cells by antagonizing p300/CBP-associated protein (PCAF)-catalyzed MPP8-K439 acetylation, thereby preventing ubiquitin-proteasome-mediated proteolysis of M-phase phosphoprotein 8 [43], a methyl-H3K9 binding protein that promotes E-cadherin gene silencing and EMT of tumor cells.
It is well-established that androgen receptors (AR) play a pivotal role in prostate carcinogenesis [44]. In this regard, studies have investigated the effect of SIRT1 expression on AR, as a mechanism of SIRT1 in the development and progression of prostate cancer. For instance, SIRT1 was demonstrated to deacetylate AR and suppress coactivator-induced interactions between AR amino and carboxyl termini to inhibit ligand-induced (dihydrotestosterone) AR transcription activity, thereby inhibiting the growth of AR-expressing LNCaP (lymph node carcinoma of the prostate) cells [45]. Similarly, it was also demonstrated that SIRT1 deacetylates AR and histone H3 at the promoter region of the prostate-specific antigen gene to suppress AR-mediated gene transcription, thereby inhibiting the proliferation of androgen-responsive LNCaP prostate cancer cells [46]. These findings suggest that low expression of SIRT1 is particularly beneficial for the growth of androgen-responsive prostate cancer. Conversely, androgen-independent prostate cancer cells, such as PC3 and DU145, have been shown to express higher levels of SIRT1 than androgen-dependent LNCaP prostate cancer cells [34]. This may suggest a differential role of SIRT1 in various sub-types of prostate cancer. In contrast with the above-mentioned studies, a more recent study revealed that SIRT1 promotes the progression of LNCaP prostate cancer cells by upregulating AR signaling [47]. Although this contradiction may be influenced by the cancer stage or context [48], it provides opportunities for future studies to ascertain the molecules or intermediates in AR signaling that are modulated by SIRT1 in favor of the progression of prostate cancer.
SIRT1 has been shown to modulate catalytic protein activity as a part of its mechanisms of action in prostate cancer. Notably, it has been revealed that SIRT1 deacetylates and upregulates the expression of matrix metalloproteinase-2 (MMP2) [49], a zinc-dependent endopeptidase that degrades the extracellular matrix to promote the invasion of cancer cells [50]. To corroborate their finding, it was further demonstrated that siRNA-mediated knockdown of SIRT1 reduced MMP2 protein stability and zymographic activity and consequently inhibited the invasion of PC3 prostate cancer cells. This suggests that SIRT1 upregulates MMP2 activity to promote the progression of prostate cancer. Additionally, shRNA-mediated inhibition of SIRT1 expression induces proliferation and inhibits autophagy of DU145 and PC3 prostate cancer cells by repressing the phosphorylation of p70 ribosomal S6 kinases (S6K) and 4E-binding protein 1 [34], which are downstream effectors of the mammalian target of rapamycin 1 (mTORC1)-mediated protein synthesis regulation [51]. It was further suggested that SIRT1 promotes the tumorigenesis of prostate cancer via SIRT1/S6K-mediated inhibition of autophagy. In other words, they speculated that the deacetylation of S6K by SIRT1 may trigger its dephosphorylation, leading to a halt in the synthesis of apoptotic or autophagic proteins. To elucidate this mechanism, further studies are required to clarify/verify their claims and ascertain the specific apoptotic or autophagic proteins regulated through this axis.
3.2. SIRT6
Unlike in other types of cancer, including breast cancer, SIRT6 is overexpressed in prostate tumor tissues and cells (PC3, DU145, 22RV1, and LNCaP), with high SIRT6 expression correlating with poor overall survival of prostate cancer patients [52][53][54][55]. This indicates an important role of SIRT6 in prostate cancer, which needs to be fully explored. Liu et al. demonstrated that siRNA-mediated inhibition of SIRT6 induces cell cycle arrest at the sub-G1 phase, decreases Bcl-2 expression, induces apoptosis, increases DNA damage, and enhances chemotherapeutic sensitivity of PC3 and DU145 prostate cancer cells [53]. This indicates that SIRT6 promotes the survival and proliferation of prostate cancer by increasing Bcl-2 expression and preventing cell cycle arrest. To corroborate these findings, Xie et al. demonstrated that siRNA-mediated inhibition of SIRT6 reduced the EMT-protein N-cadherin and anti-EMT protein E-cadherin to promote the migration and invasion of PC3 and DU145 prostate cancer cells. Furthermore, a recent study reported that SIRT6 promotes the progression of prostate cancer by abrogating necroptosis-facilitated innate immunity [52]. Another recent study also demonstrated that SIRT6 promotes the progression of prostate cancer by negatively regulating the Wnt/β-Catenin signaling pathway, as shown by a decrease in the expression levels of c-Myc, cyclin D1, and β-Catenin [55]. To date, all studies unequivocally portray SIRT6 as a promoter of prostate cancer.
3.3. SIRT7
SIRT7 is overexpressed in various human cancers [56], including prostate cancer [57][58], suggesting a pivotal role of SIRT7 in the progression of prostate cancer. Malik et al. observed an inverse relationship between SIRT7 and the expression of E-cadherin in highly aggressive PC3 prostate cancer cells; that is, there was a low expression of E-cadherin and a high expression of SIRT7 [59]. To verify their findings, using SIRT7-deficient PC3 prostate cancer cells, they demonstrated that the E-cadherin protein level was upregulated, while the mesenchymal marker vimentin and EMT-inducing transcription factor slug were both downregulated [59]. Taken together, these findings indicate that SIRT7 promotes the migration and metastasis of PC3 prostate cancer cells by upregulating EMT. Similarly, corroborating the above-mentioned findings, Haider et al. demonstrated that SIRT7-knockdown reduced the migration of DU145 prostate cancer cells and that SIRT7 overexpression promoted the aggressiveness and docetaxel resistance of LNCaP prostate cancer cells by triggering EMT as demonstrated by an increase in fibronectin [57]. Overall, these findings indicate that SIRT7 overexpression induces EMT to promote the aggressiveness of both androgen-dependent (LNCaP) and androgen-independent (DU145) prostate cancer cells.
More recently, Ding et al. demonstrated that SIRT7 depletion in LNCaP and 22RV1 prostate cancer cells significantly reduces the conversion of LC3B-I to LC3B-II, a vital process that results in autophagy, and that SIRT7 knockdown downregulates vimentin, slug, MMP2, and MMP9 in 22Rv1 prostate cancer cells [58]. They further demonstrated that SIRT7 expression is positively correlated with AR signaling in both prostate cancer tissues and cells (LNCaP and 22Rv1) and that SIRT7 knockdown upregulates SMAD4, a tumor suppressor protein [60], which interacts with AR to control its signaling in prostate cancer [61]. Taken together, these findings indicate that SIRT7 expression promotes the proliferation, metastasis, and androgen-induced autophagy of prostate cancer via induction of AR signaling. Moreover, it has been previously demonstrated that androgens promote the growth of prostate cancer cells via the induction of autophagy [62], thus indicating a favorable effect of autophagy in androgen-responsive prostate cancer cells.
4. Cytoplasmic Sirtuins in Breast Cancer
SIRT2
Studies have shown that SIRT2 expression is downregulated in breast cancer tissues compared to normal breast tissues [63][64][65]. Furthermore, SIRT2 was shown to be predominantly involved in the tumorigenesis and prognosis of breast cancer and that, depending on the tumor grade, SIRT2 acts as either a tumor suppressor or tumor promoter in breast tumors [64]. In this regard, studies have investigated the tumor-suppressing or tumor-promoting mechanisms of SIRT2 in breast cancer. Given that SIRT2 translocates to the nucleus during mitotic cell division, Kim et al. demonstrated that SIRT2 deacetylates CDH1 and CDC20 to positively regulate the activity of the anaphase-promoting complex/cyclosome (APC/C) [63], a multi-subunit protein that mediates ubiquitination of distinctly functional substrates, including Aurora-A and -B, survivin Plk1, Nek2A, securin, and cyclins-A and -B [66]. However, some of these APC/C substrates, such as Aurora A, cyclins, and Plk1, are overexpressed in human cancers and promote tumorigenesis [67][68][69]. Hypothetically, the overexpression of the APC/C substrates in human cancers may be due to the downregulated expression of SIRT2, which may result in aberrant mitotic cell division. To verify this hypothesis, Kim et al. demonstrated that SIRT2 deficiency impairs the activity of APC/C, leading to the accumulation of mitotic regulatory proteins (including Aurora A and Plk1), mitotic catastrophe, genetic instability, and, consequently, tumorigenesis [63]. Overall, their findings indicate a tumor-suppressing role of SIRT2 in breast cancer through the promotion of normal mitosis. To further support the tumor-suppressing role of SIRT2 in breast cancer, Fiskus et al. demonstrated that SIRT2 deacetylates and inhibits the peroxidase activity of peroxiredoxin-1 (an antioxidant), thereby sensitizing MCF-7 and MDA-MB-231 breast cancer cells to the cytotoxic effects of increased reactive oxygen species and DNA damage [70].
Conversely, Park et al. demonstrated that SIRT2 deacetylates the lysine 305 (K305) residue of the pyruvate kinase M2 (PKM2) isoform to direct glycolysis and promote tumor growth [71]. To corroborate their findings, they demonstrated that SIRT2 knockout by shSIRT2 RNA altered the PKM2 activity and glycolytic metabolism, thus indicating that SIRT2 ensures the growth and survival of breast cancer through the regulation of glycolysis and PKM2 activity. Additionally, using MCF10A breast cancer cells, Zhou et al. demonstrated that SIRT2 promotes the growth and aggressiveness of basal-like breast cancer by deacetylating the K116 domain of slug [72], a tumor promoter protein that, upon stabilization, facilitates tumor progression and metastasis through EMT, leading to loss of cell adhesion, as well as the enhancement of migratory and invasive properties of tumors [73]. As a mechanism, it was demonstrated that the protein slug is stabilized by the deacetylation of its K116 domain. Moreover, the stabilization of slug has been shown to prevent its proteasomal degradation, facilitate its transcriptional repression of E-cadherin, and potentiate its anti-apoptotic and cell-invasive functions [74][75]. To corroborate their findings, Zhou et al. further demonstrated that silencing of SIRT2 with shSIRT2 causes the loss of aggressive basal-like breast cancer features in SUM149 and SUM1315 cancer cells [72]. Furthermore, a recent study demonstrated that SIRT2 deacetylase activity promotes the heterodimerization, nuclear retention, and stability of the breast cancer type I susceptibility protein (BRCA1) and BRCA1-associated RING domain protein I [76], a tumor suppressor protein involved in DNA repair and genome stability [77]. Another recent study revealed that SIRT2 positively regulates T cell differentiation, specifically the CD8+ T cells, to facilitate tumor immune response during breast cancer progression [78]. In summary, SIRT2 is highly involved in the initiation and progression of breast cancer, playing either a tumor-promoting or tumor-suppressing role. This opens further opportunities to explore the distinct roles of SIRT2 in breast cancer, particularly in the initiation processes of breast cancer, such as DNA damage, genetic instability, and mitotic catastrophe, as this could provide insights into the early detection and treatment of breast cancers.
5. Cytoplasmic Sirtuins in Prostate Cancer
SIRT2
Unlike in breast cancer, SIRT2 was previously shown to be highly expressed in prostate cancer cells compared with normal prostate cells [33]. However, in a recent study, SIRT2 expression was shown to decline from benign to malignant and then metastatic prostate cancer [79]. To corroborate this finding, Lee et al. showed that SIRT2 protein levels are reduced in castrate-resistant prostate cancer. These findings suggest a tumor stage-dependent expression of SIRT2 in prostate cancer, such that the SIRT2 protein level reduces as the prostate tumor progresses. Unfortunately, to date, there are no studies that have investigated the mechanistic role of SIRT2 in prostate cancer. In this regard, future studies are required to determine the oncogenic or tumor-suppressing effect of SIRT2 in prostate cancer.
6. Mitochondrial Sirtuins in Breast Cancer
6.1. SIRT3
The primary mitochondrial-localized deacetylase SIRT3 is known to play a vital role in mitochondrial metabolism [80]. Loss of SIRT3 culminates in hyperacetylation of mitochondrial proteins, which subsequently leads to the reduction of mitochondrial ability to produce ATP and induction of oxidative stress [81]. By deacetylating several enzymes involved in mitochondrial metabolism, such as isocitrate dehydrogenase and manganese superoxide dismutase, SIRT3 protects against pathological conditions, including aging-associated pathophysiologies [82][83]. Furthermore, the dual roles of SIRT3 in various types of cancers have been extensively reviewed [84], and their role in breast cancer is discussed below.
The expression of SIRT3 is decreased in human breast cancers [85][86]. To substantiate the decreased expression of SIRT3 in breast cancer, further investigations revealed that the SIRT3 gene is deleted in approximately 20 percent of human cancers and 40 percent of breast cancers [86]. Corroborating this, the SIRT3 gene is deleted in breast cancer more than in other cancers [85]. Armed with these findings, several researchers have investigated the tumor-suppressive role of SIRT3 in breast cancer. Finley et al. demonstrated that SIRT3 suppresses the Warburg effect and proliferation of human breast cancers [86]. Mechanistically, it was demonstrated that SIRT3 opposes the Warburg effect by destabilizing hypoxia-inducible factor 1α (HIF1α), the main facilitator of increased glycolysis and lactate production during hypoxic conditions [86]. It was also revealed that SIRT3 overexpression suppresses the proliferation of CAMA1 breast cancer cells in the presence of high glucose [86]. To corroborate their findings, it was also demonstrated that the knockout of SIRT3 promoted the Warburg effect in human breast cancer cells, thus promoting the survival of the breast cancer cells [86].
Additionally, Zou et al. demonstrated that the loss of SIRT3 (following treatment with SIRT3 shRNA) in MCF-7 breast cancer cells resulted in increased acetylation at lysine 413 of isocitrate dehydrogenase 2 (IDH2), a key enzyme in the Krebs cycle that oxidizes and decarboxylates isocitrate into α-ketoglutarate, and that SIRT3 loss decreases the level of IDH2 dimerization [87]. It was further demonstrated that SIRT3 loss reduced IDH2 activities by decreasing IDH2 dimerization and that IDH2 acetylation at lysine 413 impairs mitochondrial respiration and detoxification, increases ROS production, and correlates with Luminal B breast cancer risk [87]. These findings suggest that SIRT3 expression may reverse the tumorigenic phenotype in breast cancer cells. Pinteric et al. demonstrated that de novo overexpression of SIRT3 downregulates the expression of vegfr1 (a proangiogenic protein involved in cell proliferation), EMT markers (vimentin and slug), lactate dehydrogenase A (LDHA; a glycolytic marker), peroxisome proliferator-activated receptor gamma coactivator 1-alpha (PGC1α), SIRT1, superoxide dismutase 2 (SOD2), and catalase (CAT) in MCF-7 breast cancer cells, thus indicating a tumor suppressor effect of SIRT3 in breast cancer [88]. Supporting this finding, a recent study demonstrated that the overexpression of SIRT3 promoted apoptosis and inhibited the proliferation of MCF-7 and MDA-MB-231 breast cancer cells [89].
Intriguingly, another recent study showed that while SIRT3 expression improved mitochondrial mass and potential metabolism (Warburg effect) and increased SOD2 and CAT expression, it also increased mitochondrial ROS, DNA damage, and expression of tissue inhibitor of metalloproteinase 1 (a major inhibitor of MMP9), which have been shown to promote malignant progression and metastasis [90]. Further, SIRT3 expression also induced the formation of multinucleated cells and apoptosis and inhibited the proliferation of MDA-MB-231 breast cancer cells [91].
Further supporting the tumor suppressor role of SIRT3 in breast cancer, SIRT3 suppresses estrogen-induced cytosolic and mitochondrial ROS production, diminishes estrogen-induced DNA synthesis, and upregulates p53 expression in MCF-7 breast cancer cells [92], despite its increasing antioxidant activities and cytosolic ROS. From the above-mentioned studies, it is evident that there is a well-established tumor suppressor role of SIRT3 in breast cancer.
6.2. SIRT4
Like other mitochondrial sirtuins, SIRT4 is a mitochondrial deacetylase involved in energy metabolism [93]. However, in addition to its deacetylase activity, SIRT4 possesses ADP-ribosyl transferase activity, which catalyzes the transfer of ADP-ribosyl units to targets, such as glutamate dehydrogenase [94]. SIRT4 has been reported to be a therapeutic target that has both oncogenic and tumor suppressor effects in cancers [95]; however, the underlying mechanism is still a matter of debate. So far, most studies have demonstrated that SIRT4 regulates glutamine metabolism [96]. Notably, it is through the inhibition of glutamine metabolism that SIRT4 exhibits tumor suppressor effects in B Cell Lymphoma [97], 2014), colorectal cancer [98], and thyroid cancer [99].
In breast cancer, the expression pattern of SIRT4 is controversial. While some studies report that SIRT4 expression is upregulated in breast cancer [100][101], others report downregulation [60][96][102][103]. The differences in the expression patterns of SIRT4 in breast cancer cells may be attributed to differences in the breast cancer cell types and quantitation methods for SIRT4 used in these studies. Given its differential pattern of expression, it can be speculated that SIRT4 exhibits both tumor-suppressing and tumor-promoting effects. For instance, although the underlying mechanism was not investigated, a 2017 study demonstrated that SIRT4 upregulation promotes the proliferation, migration, and invasion of MDA-MB-435S breast cancer cells [101], although it should be noted that there is also controversy over whether this cell line is of breast origin [104]. Additionally, a recent study demonstrated that SIRT4 deacetylates methylenetetrahydrofolate dehydrogenase/methylenetetrahydrofolate cyclohydrolase 2 (MTHFD2), a vital bifunctional enzyme in folate metabolism, to remodel folate metabolism against breast cancer cells. As a mechanism, it was revealed that SIRT4 deacetylation of MTHFD2 destabilizes MTHFD2 and causes its proteasomal degradation, leading to NADPH reduction and intracellular ROS accumulation in the breast tumors which, in turn, inhibits the proliferation of the breast cancer cells [105].
On the contrary, another study demonstrated that SIRT4 overexpression downregulates IL-6 expression and STAT3 Y705 phosphorylation, as well as the transcription and translation of STAT3 target genes (MYC and CNDD1), to enhance the sensitivity of MCF7 and T47D breast cancer cells to tamoxifen, thus potentiating the effect of this chemotherapy against breast cancer cells [106].
6.3. SIRT5
In addition to its deacetylase activity, SIRT5 has been shown to exhibit demalonylase, deglutarylase, and succinylase activities [107][108]. As a member of the mitochondrial sirtuins, SIRT5, through its deacetylase activities, participates in several biological processes, such as metabolic regulation, aging, and oxidative stress [109]. SIRT5 expression has been evaluated in various kinds of cancer, particularly in head and neck squamous cell carcinoma [110], endometrial carcinoma [111], basal carcinoma [112], and hepatocellular carcinoma [113]. Although SIRT5 expression was found to be upregulated in breast cancer [103][114], which may be indicative of its role in breast cancer progression, only a few studies have revealed the mechanisms underlying the role of SIRT5 in breast cancer.
7. Mitochondrial Sirtuins in Prostate Cancer
7.1. SIRT3
The expression of SIRT3 in prostate cancer remains controversial. SIRT3 expression has been reported to be decreased in prostate cancer by two studies: Quan et al. [115] found that SIRT3 is moderately downregulated in prostate carcinoma tissues and Li et al. [116] detected a decrease in metastatic tissues compared to prostate tumor tissues. Singh et al. [117] found that SIRT3 was upregulated in cancerous prostatic tissues and cell lines compared to normal tissues and prostatic epithelia cells, respectively. These conflicting findings may suggest a differential pattern of SIRT3 expression in prostate cancer tissues and cells, which may depend on the cell type, tumor stage or grade, and genetic differences in the prostate cancer tissues or cells. In this regard, further studies are needed to clarify the expression pattern of SIRT3 in prostate cancer tissues and cells. Similarly, mechanistic studies have been conducted in prostate cancer, which indicated that SIRT3 has a dual role in prostate carcinogenesis.
7.2. SIRT4
Unlike in other human cancers, including breast cancer, the expression pattern of SIRT4 and its mechanistic roles in prostate cancer have been sparsely investigated. Nevertheless, a study found that SIRT4 suppresses the proliferation of DU145 prostate cancer cells by inhibiting the uptake of glutamine [102], a metabolite vital for proliferating and cancerous cells [118]. Furthermore, Li et al. demonstrated that SIRT4 promotes mitochondrial-mediated apoptosis in PC3 prostate cancer cells by deacetylating and promoting the ubiquitination and degradation of adenine nucleotide translocase-2 [119], a mitochondrial inner membrane protein that is highly expressed in cancer cells [120]. Notwithstanding, further studies are required to clarify the expression pattern of SIRT4 and its mechanistic roles in prostate cancer.
7.3. SIRT5
As in breast cancer, SIRT5 was found to be highly expressed in prostate tumor tissues compared to normal prostate tissues [121]. However, in contrast, recent studies have revealed that SIRT5 expression is downregulated in advanced prostate cancer [122][123], suggesting that SIRT5 downregulation is dependent on the progression or stage of the prostate cancer cells. These findings may indicate a tumor-stage-dependent role of SIRT5 in prostate carcinogenesis. There are few studies on the mechanistic roles of SIRT5 in prostate cancer. Recently, it was demonstrated that SIRT5 promotes the proliferation and migration of LNCaP and PC3 prostate cancer cells [121]. This was shown to occur through increased expression of SIRT5, which resulted in the upregulation of cyclin D1, MMP9, and MAPK signaling-related proteins. It was further demonstrated that SIRT5 decreased the protein levels of acetyl-CoA acetyltransferase 1 (ACAT1), a promoter of prostate cancer [124]. Since ACAT1 negatively regulates the MAPK signaling pathway [121], Peck and Shulze [124] concluded that SIRT5 promotes the proliferation, invasion, and migration of prostate cancer via the downregulation of ACAT1 protein levels.
References
- Matt Kaeberlein; Mitch McVey; Leonard Guarente; The SIR2/3/4 complex and SIR2 alone promote longevity in Saccharomyces cerevisiae by two different mechanisms. Genes & Development 1999, 13, 2570-2580, 10.1101/gad.13.19.2570.
- Johannes H. Bauer; Siti Nur Sarah Morris; Chengyi Chang; Thomas Flatt; Jason G. Wood; Stephen L. Helfand; dSir2 and Dmp53 interact to mediate aspects of CR-dependent life span extension in D. melanogaster. Aging 2008, 1, 38-48, 10.18632/aging.100001.
- Shin-Ichiro Imai; Christopher M. Armstrong; Matt Kaeberlein; Leonard Guarente; Transcriptional silencing and longevity protein Sir2 is an NAD-dependent histone deacetylase. Nature 2000, 403, 795-800, 10.1038/35001622.
- Xu, Y.; Qin, Q.; Chen, R.; Wei, C.; Mo, Q. SIRT1 promotes proliferation, migration, and invasion of breast cancer cell line MCF-7 by upregulating DNA polymerase delta1 (POLD1). Biochem. Biophys. Res. Commun. 2018, 502, 351–357.
- Jin, X.; Wei, Y.; Xu, F.; Zhao, M.; Dai, K.; Shen, R.; Yang, S.; Zhang, N. SIRT1 promotes formation of breast cancer through modulating Akt activity. J. Cancer 2018, 9, 2012–2023.
- Yao, Y.; Liu, T.; Wang, X.; Zhang, D. The Contrary Effects of Sirt1 on MCF7 Cells Depend on CD36 Expression Level. J. Surg. Res. 2019, 238, 248–254.
- Mahmud, Z.; Gomes, A.R.; Lee, H.J.; Aimjongjun, S.; Jiramongkol, Y.; Yao, S.; Zona, S.; Alasiri, G.; Gong, G.; Yagüe, E.; et al. EP300 and SIRT1/6 Co-Regulate Lapatinib Sensitivity Via Modulating FOXO3-Acetylation and Activity in Breast Cancer. Cancers 2019, 11, 1067.
- Shi, L.; Tang, X.; Qian, M.; Liu, Z.; Meng, F.; Fu, L.; Wang, Z.; Zhu, W.-G.; Huang, J.-D.; Zhou, Z.; et al. A SIRT1-centered circuitry regulates breast cancer stemness and metastasis. Oncogene 2018, 37, 6299–6315.
- Santolla, M.F.; Avino, S.; A Pellegrino, M.; De Francesco, E.M.; De Marco, P.; Lappano, R.; Vivacqua, A.; Cirillo, F.; Rigiracciolo, D.C.; Scarpelli, A.; et al. SIRT1 is involved in oncogenic signaling mediated by GPER in breast cancer. Cell Death Dis. 2015, 6, e1834.
- Elangovan, S.; Ramachandran, S.; Venkatesan, N.; Ananth, S.; Gnana-Prakasam, J.P.; Martin, P.M.; Browning, D.D.; Schoenlein, P.V.; Prasad, P.D.; Ganapathy, V.; et al. SIRT1 is essential for oncogenic signaling by estrogen/estrogen receptor α in breast cancer. Cancer Res. 2011, 71, 6654–6664.
- Moore, R.L.; Faller, D.V. SIRT1 represses estrogen-signaling, ligand-independent ERα-mediated transcription, and cell proliferation in estrogen-responsive breast cells. J. Endocrinol. 2013, 216, 273–285.
- Kuo, S.J.; Lin, H.-Y.; Chien, S.-Y.; Chen, D.-R. SIRT1 suppresses breast cancer growth through downregulation of the Bcl-2 protein. Oncol. Rep. 2013, 30, 125–130.
- Gollavilli, N.; Kanugula, A.K.; Koyyada, R.; Karnewar, S.; Neeli, P.K.; Kotamraju, S. AMPK inhibits MTDH expression via GSK3β and SIRT1 activation: Potential role in triple negative breast cancer cell proliferation. FEBS J. 2015, 282, 3971–3985.
- Simmons, G.E.J.; Pandey, S.; Nedeljkovic-Kurepa, A.; Saxena, M.; Wang, A.; Pruitt, K. Frizzled 7 expression is positively regulated by SIRT1 and β-catenin in breast cancer cells. PLoS ONE 2014, 9, e98861.
- Jin, M.-S.; Hyun, C.L.; Park, I.A.; Kim, J.Y.; Chung, Y.R.; Im, S.-A.; Lee, K.-H.; Moon, H.-G.; Ryu, H.S. SIRT1 induces tumor invasion by targeting epithelial mesenchymal transition-related pathway and is a prognostic marker in triple negative breast cancer. Tumour Biol. 2016, 37, 4743–4753.
- Deus, C.M.; Serafim, T.L.; Magalhães-Novais, S.; Vilaça, A.; Moreira, A.C.; Sardão, V.A.; Cardoso, S.M.; Oliveira, P.J. Sirtuin 1-dependent resveratrol cytotoxicity and pro-differentiation activity on breast cancer cells. Arch. Toxicol. 2017, 91, 1261–1278.
- Kala, R.; Shah, H.N.; Martin, S.L.; Tollefsbol, T.O. Epigenetic-based combinatorial resveratrol and pterostilbene alters DNA damage response by affecting SIRT1 and DNMT enzyme expression, including SIRT1-dependent γ-H2AX and telomerase regulation in triple-negative breast cancer. BMC Cancer 2015, 15, 672.
- Holloway, K.R.; Barbieri, A.; Malyarchuk, S.; Saxena, M.; Nedeljkovic-Kurepa, A.; Mehl, M.C.; Wang, A.; Gu, X.; Pruitt, K. SIRT1 positively regulates breast cancer associated human aromatase (CYP19A1) expression. Mol. Endocrinol. 2013, 27, 480–490.
- Latifkar, A.; Ling, L.; Hingorani, A.; Johansen, E.; Clement, A.; Zhang, X.; Hartman, J.; Fischbach, C.; Lin, H.; Cerione, R.A.; et al. Loss of Sirtuin 1 Alters the Secretome of Breast Cancer Cells by Impairing Lysosomal Integrity. Dev. Cell 2019, 49, 393–408.e7.
- Kugel, S.; Feldman, J.L.; Klein, M.A.; Silberman, D.M.; Sebastian, C.; Mermel, C.; Dobersch, S.; Clark, A.R.; Getz, G.; Denu, J.M.; et al. Identification of and Molecular Basis for SIRT6 Loss-of-Function Point Mutations in Cancer. Cell Rep. 2015, 13, 479–488.
- Wang, D.; Li, C.; Zhang, X. The promoter methylation status and mRNA expression levels of CTCF and SIRT6 in sporadic breast cancer. DNA Cell Biol. 2014, 33, 581–590.
- Thirumurthi, U.; Shen, J.; Xia, W.; LaBaff, A.M.; Wei, Y.; Li, C.-W.; Chang, W.-C.; Chen, C.-H.; Lin, H.-K.; Yu, D.; et al. MDM2-mediated degradation of SIRT6 phosphorylated by AKT1 promotes tumorigenesis and trastuzumab resistance in breast cancer. Sci. Signal 2014, 7, ra71.
- Ioris, R.M.; Galié, M.; Ramadori, G.; Anderson, J.G.; Charollais, A.; Konstantinidou, G.; Brenachot, X.; Aras, E.; Goga, A.; Ceglia, N.; et al. SIRT6 Suppresses Cancer Stem-like Capacity in Tumors with PI3K Activation Independently of Its Deacetylase Activity. Cell Rep. 2017, 18, 1858–1868.
- Sociali, G.; Grozio, A.; Caffa, I.; Schuster, S.; Becherini, P.; Damonte, P.; Sturla, L.; Fresia, C.; Passalacqua, M.; Mazzola, F.; et al. SIRT6 deacetylase activity regulates NAMPT activity and NAD(P)(H) pools in cancer cells. FASEB J. 2019, 33, 3704–3717.
- Becherini, P.; Caffa, I.; Piacente, F.; Damonte, P.; Vellone, V.G.; Passalacqua, M.; Benzi, A.; Bonfiglio, T.; Reverberi, D.; Khalifa, A.; et al. SIRT6 enhances oxidative phosphorylation in breast cancer and promotes mammary tumorigenesis in mice. Cancer Metab. 2021, 9, 6.
- Wu, X.; Wang, S.; Zhao, X.; Lai, S.; Yuan, Z.; Zhan, Y.; Ni, K.; Liu, Z.; Liu, L.; Xin, R.; et al. Clinicopathological and prognostic value of SIRT6 in patients with solid tumors: A meta-analysis and TCGA data review. Cancer Cell Int. 2022, 22, 84.
- Aljada, A.; Saleh, A.M.; Alkathiri, M.; Shamsa, H.B.; Al-Bawab, A.; Nasr, A. Altered Sirtuin 7 Expression is Associated with Early Stage Breast Cancer. Breast Cancer 2015, 9, 3–8.
- Tang, X.; Shi, L.; Xie, N.; Liu, Z.; Qian, M.; Meng, F.; Xu, Q.; Zhou, M.; Cao, X.; Zhu, W.-G.; et al. SIRT7 antagonizes TGF-β signaling and inhibits breast cancer metastasis. Nat. Commun. 2017, 8, 318.
- Huo, Q.; Li, Z.; Cheng, L.; Yang, F.; Xie, N. SIRT7 Is a Prognostic Biomarker Associated with Immune Infiltration in Luminal Breast Cancer. Front. Oncol. 2020, 10, 621.
- Thiery, J. Epithelial—Mesenchymal transitions in tumour progression. Nat. Rev. Cancer 2002, 2, 442–454.
- Yang, F.; Hu, Y.; Shao, L.; Zhuang, J.; Huo, Q.; He, S.; Chen, S.; Wang, J.; Xie, N. SIRT7 interacts with TEK (TIE2) to promote adriamycin induced metastasis in breast cancer. Cell Oncol. 2021, 44, 1405–1424.
- Drescher, F.; Juárez, P.; Arellano, D.L.; Serafín-Higuera, N.; Olvera-Rodriguez, F.; Jiménez, S.; Licea-Navarro, A.F.; Fournier, P.G. TIE2 Induces Breast Cancer Cell Dormancy and Inhibits the Development of Osteolytic Bone Metastases. Cancers 2020, 12, 868.
- Jung-Hynes, B.; Nihal, M.; Zhong, W.; Ahmad, N. Role of sirtuin histone deacetylase SIRT1 in prostate cancer. A target for prostate cancer management via its inhibition? J. Biol. Chem. 2009, 284, 3823–3832.
- Li, G.; Rivas, P.; Bedolla, R.; Thapa, D.; Reddick, R.L.; Ghosh, R.; Kumar, A.P. Dietary resveratrol prevents development of high-grade prostatic intraepithelial neoplastic lesions: Involvement of SIRT1/S6K axis. Cancer Prev. Res. 2013, 6, 27–39.
- Hoffmann, M.J.; Engers, R.; Florl, A.R.; Otte, A.P.; Muller, M.; Schulz, W.A. Expression changes in EZH2, but not in BMI-1, SIRT1, DNMT1 or DNMT3B are associated with DNA methylation changes in prostate cancer. Cancer Biol. Ther. 2007, 6, 1403–1412.
- Kojima, K.; Ohhashi, R.; Fujita, Y.; Hamada, N.; Akao, Y.; Nozawa, Y.; Deguchi, T.; Ito, M. A role for SIRT1 in cell growth and chemoresistance in prostate cancer PC3 and DU145 cells. Biochem. Biophys. Res. Commun. 2008, 373, 423–428.
- Nakane, K.; Fujita, Y.; Terazawa, R.; Atsumi, Y.; Kato, T.; Nozawa, Y.; Deguchi, T.; Ito, M. Inhibition of cortactin and SIRT1 expression attenuates migration and invasion of prostate cancer DU145 cells. Int. J. Urol. 2012, 19, 71–79.
- Yang, Y.; Hou, H.; Haller, E.M.; Nicosia, S.V.; Bai, W. Suppression of FOXO1 activity by FHL2 through SIRT1-mediated deacetylation. EMBO J. 2005, 24, 1021–1032.
- Paik, J.H.; Kollipara, R.; Chu, G.; Ji, H.; Xiao, Y.; Ding, Z.; Miao, L.; Tothova, Z.; Horner, J.W.; Carrasco, D.R.; et al. FoxOs are lineage-restricted redundant tumor suppressors and regulate endothelial cell homeostasis. Cell 2007, 128, 309–323.
- Wang, F.; Chan, C.-H.; Chen, K.; Guan, X.; Lin, H.-K.; Tong, Q. Deacetylation of FOXO3 by SIRT1 or SIRT2 leads to Skp2-mediated FOXO3 ubiquitination and degradation. Oncogene 2012, 31, 1546–1557.
- Byles, V.; Zhu, L.; Lovaas, J.D.; Chmilewski, L.K.; Wang, J.; Faller, D.V.; Dai, Y. SIRT1 induces EMT by cooperating with EMT transcription factors and enhances prostate cancer cell migration and metastasis. Oncogene 2012, 31, 4619–4629.
- Cui, Y.; Li, J.; Zheng, F.; Ouyang, Y.; Chen, X.; Zhang, L.; Chen, Y.; Wang, L.; Mu, S.; Zhang, H. Effect of SIRT1 Gene on Epithelial-Mesenchymal Transition of Human Prostate Cancer PC-3 Cells. Med. Sci. Monit. 2016, 22, 380–386.
- Sun, L.; Kokura, K.; Izumi, V.; Koomen, J.M.; Seto, E.; Chen, J.; Fang, J. MPP8 and SIRT1 crosstalk in E-cadherin gene silencing and epithelial-mesenchymal transition. EMBO Rep. 2015, 16, 689–699.
- Fujita, K.; Nonomura, N. Role of Androgen Receptor in Prostate Cancer: A Review. World J. Mens Health 2019, 37, 288–295.
- Fu, M.; Liu, M.; Sauve, A.A.; Jiao, X.; Zhang, X.; Wu, X.; Powell, M.J.; Yang, T.; Gu, W.; Avantaggiati, M.L.; et al. Hormonal control of androgen receptor function through SIRT1. Mol. Cell Biol. 2006, 26, 8122–8135.
- Dai, Y.; Ngo, D.; Forman, L.W.; Qin, D.C.; Jacob, J.; Faller, D.V. Sirtuin 1 is required for antagonist-induced transcriptional repression of androgen-responsive genes by the androgen receptor. Mol. Endocrinol. 2007, 21, 1807–1821.
- Huang, S.B.; Thapa, D.; Munoz, A.R.; Hussain, S.S.; Yang, X.; Bedolla, R.G.; Osmulski, P.; Gaczynska, M.E.; Lai, Z.; Chiu, Y.-C.; et al. Androgen deprivation-induced elevated nuclear SIRT1 promotes prostate tumor cell survival by reactivation of AR signaling. Cancer Lett. 2021, 505, 24–36.
- Huang, S.B.; Rivas, P.; Yang, X.; Lai, Z.; Chen, Y.; Schadler, K.L.; Hu, M.; Reddick, R.L.; Ghosh, R.; Kumar, A.P. SIRT1 inhibition-induced senescence as a strategy to prevent prostate cancer progression. Mol. Carcinog. 2022, 61, 702–716.
- Lovaas, J.D.; Zhu, L.; Chiao, C.Y.; Byles, V.; Faller, D.V.; Dai, Y. SIRT1 enhances matrix metalloproteinase-2 expression and tumor cell invasion in prostate cancer cells. Prostate 2013, 73, 522–530.
- Noël, A.; Jost, M.; Maquoi, E. Matrix metalloproteinases at cancer tumor-host interface. Semin. Cell Dev. Biol. 2008, 19, 52–60.
- Hay, N.; Sonenberg, N. Upstream and downstream of mTOR. Genes Dev. 2004, 18, 1926–1945.
- Fu, W.; Li, H.; Fu, H.; Zhao, S.; Shi, W.; Sun, M.; Li, Y. The SIRT3 and SIRT6 Promote Prostate Cancer Progression by Inhibiting Necroptosis-Mediated Innate Immune Response. J. Immunol. Res. 2020, 2020, 8820355.
- Liu, Y.; Xie, Q.R.; Wang, B.; Shao, J.; Zhang, T.; Liu, T.; Huang, G.; Xia, W. Inhibition of SIRT6 in prostate cancer reduces cell viability and increases sensitivity to chemotherapeutics. Protein. Cell 2013, 4, 702–710.
- Xie, Q.; Wong, A.S.; Xia, W. Abstract 1151: SIRT6 induces EMT and promotes cancer cell invasion and migration in prostate cancer. Cancer Res. 2014, 74, 1151.
- Zhang, X.; Chen, R.; Song, L.-D.; Zhu, L.-F.; Zhan, J.-F. SIRT6 Promotes the Progression of Prostate Cancer via Regulating the Wnt/β-Catenin Signaling Pathway. J. Oncol. 2022, 2022, 2174758.
- Barber, M.F.; Michishita-Kioi, E.; Xi, Y.; Tasselli, L.; Kioi, M.; Moqtaderi, Z.; Tennen, R.I.; Paredes, S.; Young, N.L.; Chen, K.; et al. SIRT7 links H3K18 deacetylation to maintenance of oncogenic transformation. Nature 2012, 487, 114–118.
- Haider, R.; Massa, F.; Kaminski, L.; Clavel, S.; Djabari, Z.; Robert, G.; Laurent, K.; Michiels, J.-F.; Durand, M.; Ricci, J.-E.; et al. Sirtuin 7: A new marker of aggressiveness in prostate cancer. Oncotarget 2017, 8, 77309–77316.
- Ding, M.; Jiang, C.-Y.; Zhang, Y.; Zhao, J.; Han, B.-M.; Xia, S.-J. SIRT7 depletion inhibits cell proliferation and androgen-induced autophagy by suppressing the AR signaling in prostate cancer. J. Exp. Clin. Cancer Res. 2020, 39, 1–18.
- Malik, S.; Villanova, L.; Tanaka, S.; Aonuma, M.; Roy, N.; Berber, E.; Pollack, J.R.; Michishita-Kioi, E.; Chua, K.F. SIRT7 inactivation reverses metastatic phenotypes in epithelial and mesenchymal tumors. Sci. Rep. 2015, 5, 9841.
- Du, X.; Li, Q.; Yang, L.; Liu, L.; Cao, Q.; Li, Q. SMAD4 activates Wnt signaling pathway to inhibit granulosa cell apoptosis. Cell Death Dis. 2020, 11, 373.
- Zhu, M.L.; Partin, J.V.; Bruckheimer, E.M.; Strup, S.E.; Kyprianou, N. TGF-beta signaling and androgen receptor status determine apoptotic cross-talk in human prostate cancer cells. Prostate 2008, 68, 287–295.
- Shi, Y.; Han, J.J.; Tennakoon, J.B.; Mehta, F.F.; Merchant, F.; Burns, A.R.; Howe, M.K.; McDonnell, D.P.; Frigo, D.E. Androgens promote prostate cancer cell growth through induction of autophagy. Mol. Endocrinol. 2013, 27, 280–295.
- Kim, H.S.; Vassilopoulos, A.; Wang, R.-H.; Lahusen, T.; Xiao, Z.; Xu, X.; Li, C.; Veenstra, T.D.; Li, B.; Yu, H.; et al. SIRT2 maintains genome integrity and suppresses tumorigenesis through regulating APC/C activity. Cancer Cell 2011, 20, 487–499.
- McGlynn, L.M.; Zino, S.; MacDonald, A.I.; Curle, J.; Reilly, J.E.; Mohammed, Z.M.A.; McMillan, D.C.; Mallon, E.; Payne, A.P.; Edwards, J.; et al. SIRT2: Tumour suppressor or tumour promoter in operable breast cancer? Eur. J. Cancer 2014, 50, 290–301.
- Shi, P.; Zhou, M.; Yang, Y. Upregulated tumor sirtuin 2 expression correlates with reduced TNM stage and better overall survival in surgical breast cancer patients. Ir. J. Med. Sci. 2020, 189, 83–89.
- Li, M.; Zhang, P. The function of APC/CCdh1 in cell cycle and beyond. Cell Div. 2009, 4, 2.
- Casimiro, M.C.; Casimiro, M.C.; Crosariol, M.; Loro, E.; Li, Z.; Pestell, R.G. Cyclins and cell cycle control in cancer and disease. Genes Cancer 2012, 3, 649–657.
- Saeki, T.; Ouchi, M.; Ouchi, T. Physiological and oncogenic Aurora-A pathway. Int. J. Biol. Sci. 2009, 5, 758–762.
- Schmit, T.L.; Zhong, W.; Nihal, M.; Ahmad, N. Polo-like kinase 1 (Plk1) in non-melanoma skin cancers. Cell Cycle 2009, 8, 2697–2702.
- Fiskus, W.; Coothankandaswamy, V.; Chen, J.; Ma, H.; Ha, K.; Saenz, D.T.; Krieger, S.S.; Mill, C.P.; Sun, B.; Huang, P.; et al. SIRT2 Deacetylates and Inhibits the Peroxidase Activity of Peroxiredoxin-1 to Sensitize Breast Cancer Cells to Oxidant Stress-Inducing Agents. Cancer Res. 2016, 76, 5467–5478.
- Park, S.H.; Ozden, O.; Liu, G.; Song, H.Y.; Zhu, Y.; Yan, Y.; Zou, X.; Kang, H.-J.; Jiang, H.; Principe, D.R.; et al. SIRT2-Mediated Deacetylation and Tetramerization of Pyruvate Kinase Directs Glycolysis and Tumor Growth. Cancer Res. 2016, 76, 3802–3812.
- Zhou, W.; Ni, T.K.; Wronski, A.; Glass, B.; Skibinski, A.; Beck, A.; Kuperwasser, C. The SIRT2 Deacetylase Stabilizes Slug to Control Malignancy of Basal-like Breast Cancer. Cell Rep. 2016, 17, 1302–1317.
- Polyak, K.; Weinberg, R.A. Transitions between epithelial and mesenchymal states: Acquisition of malignant and stem cell traits. Nat. Rev. Cancer 2009, 9, 265–273.
- Wang, S.; Wang, W.-L.; Chang, Y.-L.; Wu, C.-T.; Chao, Y.-C.; Kao, S.-H.; Yuan, A.; Lin, C.-W.; Yang, S.-C.; Chan, W.-K.; et al. p53 controls cancer cell invasion by inducing the MDM2-mediated degradation of Slug. Nat. Cell Biol. 2009, 11, 694–704.
- Wu, W.S.; Heinrichs, S.; Xu, D.; Garrison, S.P.; Zambetti, G.P.; Adams, J.M.; Look, A.T. Slug antagonizes p53-mediated apoptosis of hematopoietic progenitors by repressing puma. Cell 2005, 123, 641–653.
- Minten, E.V.; Kapoor-Vazirani, P.; Li, C.; Zhang, H.; Balakrishnan, K.; Yu, D.S. SIRT2 promotes BRCA1-BARD1 heterodimerization through deacetylation. Cell Rep. 2021, 34, 108921.
- Tarsounas, M.; Sung, P. The antitumorigenic roles of BRCA1-BARD1 in DNA repair and replication. Nat. Rev. Mol. Cell Biol. 2020, 21, 284–299.
- Jiang, C.; Liu, J.; Guo, M.; Gao, X.; Wu, X.; Bai, N.; Guo, W.; Li, N.; Yi, F.; Cheng, R.; et al. The NAD-dependent deacetylase SIRT2 regulates T cell differentiation involved in tumor immune response. Int. J. Biol. Sci. 2020, 16, 3075–3084.
- Damodaran, S.; Damaschke, N.; Gawdzik, J.; Yang, B.; Shi, C.; Allen, G.O.; Huang, W.; Denu, J.; Jarrard, D. Dysregulation of Sirtuin 2 (SIRT2) and histone H3K18 acetylation pathways associates with adverse prostate cancer outcomes. BMC Cancer 2017, 17, 874.
- Weir, H.J.; Lane, J.D.; Balthasar, N. SIRT3: A Central Regulator of Mitochondrial Adaptation in Health and Disease. Genes Cancer 2013, 4, 118–124.
- Marcus, J.M.; Andrabi, S.A. SIRT3 Regulation Under Cellular Stress: Making Sense of the Ups and Downs. Front. Neurosci. 2018, 12, 799.
- Brown, K.; Xie, S.; Qiu, X.; Mohrin, M.; Shin, J.; Liu, Y.; Zhang, D.; Scadden, D.T.; Chen, D. SIRT3 reverses aging-associated degeneration. Cell Rep. 2013, 3, 319–327.
- Sidorova-Darmos, E.; Sommer, R.; Eubanks, J.H. The Role of SIRT3 in the Brain Under Physiological and Pathological Conditions. Front. Cell Neurosci. 2018, 12, 196.
- Torrens-Mas, M.; Oliver, J.; Roca, P.; Sastre-Serra, J. SIRT3: Oncogene and Tumor Suppressor in Cancer. Cancers 2017, 9, 90.
- Kim, H.S.; Patel, K.; Muldoon-Jacobs, K.; Bisht, K.S.; Aykin-Burns, N.; Pennington, J.D.; van der Meer, R.; Nguyen, P.; Savage, J.; Owens, K.M.; et al. SIRT3 is a mitochondria-localized tumor suppressor required for maintenance of mitochondrial integrity and metabolism during stress. Cancer Cell 2010, 17, 41–52.
- Finley, L.W.; Carracedo, A.; Lee, J.; Souza, A.; Egia, A.; Zhang, J.; Teruya-Feldstein, J.; Moreira, P.; Cardoso, S.M.; Clish, C.; et al. SIRT3 opposes reprogramming of cancer cell metabolism through HIF1α destabilization. Cancer Cell 2011, 19, 416–428.
- Zou, X.; Zhu, Y.; Park, S.-H.; Liu, G.; O’Brien, J.; Jiang, H.; Gius, D. SIRT3-Mediated Dimerization of IDH2 Directs Cancer Cell Metabolism and Tumor Growth. Cancer Res. 2017, 77, 3990–3999.
- Pinterić, M.; Podgorski, I.I.; Sobočanec, S.; Hadžija, M.P.; Paradžik, M.; Dekanić, A.; Marinović, M.; Halasz, M.; Belužić, R.; Davidović, G.; et al. De novo expression of transfected sirtuin 3 enhances susceptibility of human MCF-7 breast cancer cells to hyperoxia treatment. Free Radic Res. 2018, 52, 672–684.
- Zu, Y.; Chen, X.-F.; Li, Q.; Zhang, S.-T.; Si, L.-N. PGC-1α activates SIRT3 to modulate cell proliferation and glycolytic metabolism in breast cancer. Neoplasma 2021, 68, 352–361.
- Mehner, C.; Hockla, A.; Miller, E.; Ran, S.; Radisky, D.C.; Radisky, E.S. Tumor cell-produced matrix metalloproteinase 9 (MMP-9) drives malignant progression and metastasis of basal-like triple negative breast cancer. Oncotarget 2014, 5, 2736–2749.
- Podgorski, I.I.; Pinterić, M.; Marčinko, D.; Hadžija, M.P.; Filić, V.; Ciganek, I.; Pleše, D.; Balog, T.; Sobočanec, S. Combination of sirtuin 3 and hyperoxia diminishes tumorigenic properties of MDA-MB-231 cells. Life Sci. 2020, 254, 117812.
- Pinterić, M.; Podgorski, I.I.; Hadžija, M.P.; Filić, V.; Paradžik, M.; Proust, B.L.J.; Dekanić, A.; Ciganek, I.; Pleše, D.; Marčinko, D.; et al. Sirt3 Exerts Its Tumor-Suppressive Role by Increasing p53 and Attenuating Response to Estrogen in MCF-7 Cells. Antioxidants 2020, 9, 294.
- Nasrin, N.; Wu, X.; Fortier, E.; Feng, Y.; Bare, O.C.; Chen, S.; Ren, X.; Wu, Z.; Streeper, R.S.; Bordone, L. SIRT4 regulates fatty acid oxidation and mitochondrial gene expression in liver and muscle cells. J. Biol. Chem. 2010, 285, 31995–32002.
- Haigis, M.C.; Mostoslavsky, R.; Haigis, K.M.; Fahie, K.; Christodoulou, D.C.; Murphy, A.J.; Valenzuela, D.M.; Yancopoulos, G.D.; Karow, M.; Blander, G.; et al. SIRT4 inhibits glutamate dehydrogenase and opposes the effects of calorie restriction in pancreatic beta cells. Cell 2006, 126, 941–954.
- Huang, G.; Zhu, G. Sirtuin-4 (SIRT4), a therapeutic target with oncogenic and tumor-suppressive activity in cancer. Onco Targets Ther. 2018, 11, 3395–3400.
- Jeong, S.M.; Xiao, C.; Finley, L.W.; Lahusen, T.; Souza, A.L.; Pierce, K.; Li, Y.-H.; Wang, X.; Laurent, G.; German, N.J.; et al. SIRT4 has tumor-suppressive activity and regulates the cellular metabolic response to DNA damage by inhibiting mitochondrial glutamine metabolism. Cancer Cell 2013, 23, 450–463.
- Jeong, S.M.; Lee, A.; Lee, J.; Haigis, M.C. SIRT4 protein suppresses tumor formation in genetic models of Myc-induced B cell lymphoma. J. Biol. Chem. 2014, 289, 4135–4144.
- Miyo, M.; Yamamoto, H.; Konno, M.; Colvin, H.; Nishida, N.; Koseki, J.; Kawamoto, K.; Ogawa, H.; Hamabe, A.; Uemura, M.; et al. Tumour-suppressive function of SIRT4 in human colorectal cancer. Br. J. Cancer 2015, 113, 492–499.
- Chen, Z.; Lin, J.; Feng, S.; Chen, X.; Huang, H.; Wang, C.; Yu, Y.; He, Y.; Han, S.; Zheng, L. SIRT4 inhibits the proliferation, migration, and invasion abilities of thyroid cancer cells by inhibiting glutamine metabolism. Onco Targets Ther. 2019, 12, 2397–2408.
- Shi, Q.; Liu, T.; Zhang, X.; Geng, J.; He, X.; Nu, M.; Pang, D. Decreased sirtuin 4 expression is associated with poor prognosis in patients with invasive breast cancer. Oncol. Lett. 2016, 12, 2606–2612.
- Huang, G.; Lin, Y.; Zhu, G. SIRT4 is upregulated in breast cancer and promotes the proliferation, migration and invasion of breast cancer cells. Int. J. Clin. Exp. Pathol. 2017, 10, 11849–11856.
- Csibi, A.; Fendt, S.-M.; Li, C.; Poulogiannis, G.; Choo, A.Y.; Chapski, D.J.; Jeong, S.M.; Dempsey, J.M.; Parkhitko, A.; Morrison, T.; et al. The mTORC1 pathway stimulates glutamine metabolism and cell proliferation by repressing SIRT4. Cell 2013, 153, 840–854.
- Igci, M.; Kalender, M.E.; Borazan, E.; Bozgeyik, I.; Bayraktar, R.; Bozgeyik, E.; Camci, C.; Arslan, A. High-throughput screening of Sirtuin family of genes in breast cancer. Gene 2016, 586, 123–128.
- Prasad, V.V.; Gopalan, R.O. Continued use of MDA-MB-435, a melanoma cell line, as a model for human breast cancer, even in year, 2014. NPJ Breast Cancer 2015, 1, 15002.
- Zhang, F.; Wang, D.; Li, J.; Su, Y.; Liu, S.; Lei, Q.-Y.; Yin, M. Deacetylation of MTHFD2 by SIRT4 senses stress signal to inhibit cancer cell growth by remodeling folate metabolism. J. Mol. Cell Biol. 2022, 14, mjac020.
- Xing, J.; Li, J.; Fu, L.; Gai, J.; Guan, J.; Li, Q. SIRT4 enhances the sensitivity of ER-positive breast cancer to tamoxifen by inhibiting the IL-6/STAT3 signal pathway. Cancer Med. 2019, 8, 7086–7097.
- Du, J.; Zhou, Y.; Su, X.; Yu, J.J.; Khan, S.; Jiang, H.; Kim, J.; Woo, J.; Kim, J.H.; Choi, B.H.; et al. Sirt5 is a NAD-dependent protein lysine demalonylase and desuccinylase. Science 2011, 334, 806–809.
- Tan, M.; Peng, C.; Anderson, K.A.; Chhoy, P.; Xie, Z.; Dai, L.; Park, J.; Chen, Y.; Huang, H.; Zhang, Y.; et al. Lysine glutarylation is a protein posttranslational modification regulated by SIRT5. Cell Metab. 2014, 19, 605–617.
- Yang, L.; Ma, X.; He, Y.; Yuan, C.; Chen, Q.; Li, G.; Chen, X. Sirtuin 5: A review of structure, known inhibitors and clues for developing new inhibitors. Sci. China Life Sci. 2017, 60, 249–256.
- Lai, C.C.; Lin, P.-M.; Lin, S.-F.; Hsu, C.-H.; Lin, H.-C.; Hu, M.-L.; Yang, M.-Y. Altered expression of SIRT gene family in head and neck squamous cell carcinoma. Tumour. Biol. 2013, 34, 1847–1854.
- Bartosch, C.; Monteiro-Reis, S.; Almeida-Rios, D.; Vieira, R.; Castro, A.; Moutinho, M.; Rodrigues, M.; Graça, I.; Lopes, J.M.; Jerónimo, C. Assessing sirtuin expression in endometrial carcinoma and non-neoplastic endometrium. Oncotarget 2016, 7, 1144–1154.
- Temel, M.; Koç, M.N.; Ulutaş, S.; Göğebakan, B. The expression levels of the sirtuins in patients with BCC. Tumour Biol. 2016, 37, 6429–6435.
- Zhang, R.; Wang, C.; Tian, Y.; Yao, Y.; Mao, J.; Wang, H.; Li, Z.; Xu, Y.; Ye, M.; Wang, L. SIRT5 Promotes Hepatocellular Carcinoma Progression by Regulating Mitochondrial Apoptosis. J. Cancer 2019, 10, 3871–3882.
- Greene, K.S.; Lukey, M.J.; Wang, X.; Blank, B.; Druso, J.E.; Lin, M.-C.J.; Stalnecker, C.A.; Zhang, C.; Abril, Y.N.; Erickson, J.W.; et al. SIRT5 stabilizes mitochondrial glutaminase and supports breast cancer tumorigenesis. Proc. Natl. Acad. Sci. USA 2019, 116, 26625–26632.
- Quan, Y.; Wang, N.; Chen, Q.; Xu, J.; Cheng, W.; Di, M.; Xia, W.; Gao, W.-Q. SIRT3 inhibits prostate cancer by destabilizing oncoprotein c-MYC through regulation of the PI3K/Akt pathway. Oncotarget 2015, 6, 26494–26507.
- Li, R.; Quan, Y.; Xia, W. SIRT3 inhibits prostate cancer metastasis through regulation of FOXO3A by suppressing Wnt/β-catenin pathway. Exp. Cell Res. 2018, 364, 143–151.
- Singh, C.; Singh, C.K.; Chhabra, G.; Nihal, M.; Iczkowski, K.A.; Ahmad, N. Abstract 539: Pro-proliferative function of the histone deacetylase SIRT3 in prostate cancer. Cancer Res. 2018, 78, 539.
- DeBerardinis, R.J.; Mancuso, A.; Daikhin, E.; Nissim, I.; Yudkoff, M.; Wehrli, S.; Thompson, C.B. Beyond aerobic glycolysis: Transformed cells can engage in glutamine metabolism that exceeds the requirement for protein and nucleotide synthesis. Proc. Natl. Acad. Sci. USA 2007, 104, 19345–19350.
- Li, T.; Li, Y.; Liu, T.; Hu, B.; Li, J.; Liu, C.; Liu, T.; Li, F. Mitochondrial PAK6 inhibits prostate cancer cell apoptosis via the PAK6-SIRT4-ANT2 complex. Theranostics 2020, 10, 2571–2586.
- Chevrollier, A.; Loiseau, D.; Chabi, B.; Renier, G.; Douay, O.; Malthièry, Y.; Stepien, G. ANT2 Isoform Required for Cancer Cell Glycolysis. J. Bioenerg. Biomembr. 2005, 37, 307–317.
- Guan, J.; Jiang, X.; Gai, J.; Sun, X.; Zhao, J.; Li, J.; Li, Y.; Cheng, M.; Du, T.; Fu, L.; et al. Sirtuin 5 regulates the proliferation, invasion and migration of prostate cancer cells through acetyl-CoA acetyltransferase 1. J. Cell. Mol. Med. 2020, 24, 14039–14049.
- Choi, S.Y.; Jeon, J.M.; Na, A.Y.; Kwon, O.K.; Bang, I.H.; Ha, Y.-S.; Bae, E.J.; Park, B.-H.; Lee, E.H.; Kwon, T.G.; et al. SIRT5 Directly Inhibits the PI3K/AKT Pathway in Prostate Cancer Cell Lines. Cancer Genom. Proteom. 2022, 19, 50–59.
- Kwon, O.K.; Bang, I.H.; Choi, S.Y.; Jeon, J.M.; Na, A.-Y.; Gao, Y.; Cho, S.S.; Ki, S.H.; Choe, Y.; Lee, J.N.; et al. SIRT5 Is the desuccinylase of LDHA as novel cancer metastatic stimulator in aggressive prostate cancer. Genom. Proteom. Bioinform. 2022.
- Peck, B.; Schulze, A. Cholesteryl Esters: Fueling the Fury of Prostate Cancer. Cell Metabolism 2014, 19, 350–352.
More
Information
Subjects:
Biochemistry & Molecular Biology; Cell Biology; Oncology
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
972
Revisions:
2 times
(View History)
Update Date:
03 Nov 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





