
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Jason Zhu | -- | 3716 | 2022-10-28 01:38:14 | | | |
| 2 | Jason Zhu | Meta information modification | 3716 | 2022-10-28 11:16:35 | | | | |
| 3 | Jason Zhu | Meta information modification | 3716 | 2022-10-28 11:16:57 | | | | |
| 4 | Jason Zhu | Meta information modification | 3716 | 2022-10-28 11:17:23 | | |
Video Upload Options
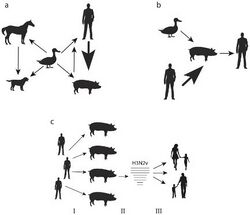
Reverse zoonosis, also known as zooanthroponosis, and sometimes anthroponosis (Greek zoon "animal", anthropos "man", nosos "disease"), refers to pathogens reservoired in humans that are capable of being transmitted to other non-human animals.
1. Terminology
Anthroponosis refers to pathogens sourced from humans and can include human to non-human animal transmission but also human to human transmission. The term zoonosis technically refers to disease transferred between any animal and another animal, human or non-human, without discretion, and also been defined as disease transmitted from animals to humans and vice-versa.[1] Yet because of human-centered medical biases, zoonosis tends to be used in the same manner as anthropozoonosis which specifically refers to pathogens reservoired in non-human animals that are transmissible to humans.[1]
Additional confusion due to frequency of scientists using "anthropozoonosis" and "zooanthroponosis" interchangeably was resolved during a 1967 Joint Food and Agriculture and World Health Organization committee meeting that recommended the use of "zoonosis" to describe the bidirectional interchange of infectious pathogens between animals and humans.[1][2]
Furthermore, because humans are rarely in direct contact with wild animals and introduce pathogens through "soft contact", the term "sapronotic agents" must be introduced. Sapronoses (Greek sapros "decaying") refers to human diseases that harbor the capacity to grow and replicate (not just survive or contaminate) in abiotic environments such as soil, water, decaying plants, animal corpses, excreta, and other substrata.[1] Additionally, sapro-zoonoses can be characterized as having both a live host and a non-animal developmental site of organic matter, soil, or plants.[1] It must be noted that obligate intracellular parasites that cannot replicate outside of cells and are entirely reproductively reliant on entering the cell to use intracellular resources such as viruses, rickettsiae, chlamydiae, and Cryptosporidium parvum cannot be sapronotic agents.[1]
2. Etymological Pitfalls
Categorizing of disease into epidemiologic classes by the infection’s supposed source or the direction of transmission raises a number of contradictions that could be resolved by the use of cyclical models as demonstrated by the following scenarios:
2.1. Zoonosis vs Reverse Zoonosis vs Anthroponosis
In the case of diseases transferred from arthropod vectors such as urban yellow fever, dengue, epidemic typhus, tickborne relapsing fever, zika fever, and malaria,[1] the differentiation between terms becomes ever more hazy. For example, a human infected with malaria is bitten by a mosquito that is subsequently infected as well. This is a case of reverse zoonosis (human to animal). However, the newly infected mosquito then infects another human. This could be a case of zoonosis (animal to human) if the mosquito is considered the original source, or anthroponosis (human to human) if the human is considered the original source. If this infected mosquito instead infected a non-human primate, it could be considered a case of reverse zoonosis/zooanthroponosis (human to animal) if the human is considered the primary source, or simply zoonosis (animal to animal) if the mosquito is considered the primary source.
2.2. Zoonosis vs Anthroponosis
Similarly, HIV originating in simians (crossover due to humans consuming wild chimpanzee bushmeat) and influenza A viruses originating in avians (crossover due to an antigenic shift) could have initially been considered a zoonotic transference as the infections first came from vertebrate animals, but could currently be regarded as an anthroponosis because of its potential to transfer between human to human.
2.3. Sapronosis vs Sapro-Zoonosis
Typical examples of sapronotic agents are fungal such as coccidioidomycosis, histoplasmosis, aspergillosis, cryptococcosis, Microsporum gypseum. Some can be bacterial from the sporulating clostridium and bacillus to Rhodococcus equi, Burkholderia pseudomallei, Listeria, Erysipelothrix, Yersinia pseudotuberculosis, legionellosis, Pontiac fever, and nontuberculous mycobacterioses. Other sapronotic agents are amebic as in primary amebic meningoencephalitis. Yet again, difficulties in classification arise in the case of sporulating bacteria whose infectious spores are only produced after a significant period of inactive vegetative growth within an abiotic environment, yet this is still considered a case of sapronoses.[1] However, cases of zoo-sapronoses involving Listeria, Erysipelothrix, Yersinia pseudotuberculosis, Burkholderia pseudomallei, and Rhodococcus equi can be transferred by an animal or an abiotic substrate but usually occur via a fecal-oral route between humans and other animals. [3]
3. Cases with Modes of Transmission
3.1. Arthropod Vectors
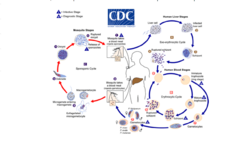
Malaria
Malaria involves the cyclical infection of animals (human and non-human) and mosquitoes from the genus Anopheles with a number of Plasmodium species. The Plasmodium parasite is transferred to the mosquito as it feeds on the blood of the infected animal whereupon it begins a sporogenic cycle in the gut of the mosquito that will infect another animal at the next blood meal. There does not seem to be any deleterious effects to the mosquito as a result of the parasitic infection.[4] The Plasmodium brasilianum parasite normally found in primates is morphologically similar to the malarial inducing Plasmodium malariae that is more commonly found in humans and it is contested as to whether the two are actually different species.[5] Nevertheless, 12 reports of malaria in the remotely located indigenous Yanomami communities of the Venezuelan Amazon arose where it was surprisingly found to be caused by a strain of P. brasilianum with 100% identical to sequences found in Alouatta seniculus monkies.[6] This suggests a definite zoonosis and high possibility of spillback back into non-human primate bands as reverse zoonoses.
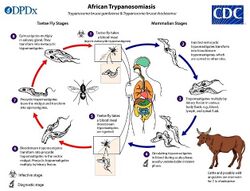
African Sleeping Sickness
Trypanosoma brucei gambiense (T. b. gambiense) is a species of African trypanosomes which are protozoan hemoflagellates responsible for trypanosomiasis (more commonly known as African sleeping sickness) in humans and other animals. The protozoa are transferred via Tsetse flies where they multiply and can be transferred to yet another animal host during the fly’s blood meal feeding.[7] Outbreaks of sleeping sickness in certain human communities have been eliminated but only temporarily as constant re-introduction from unknown sources statistically suggests the presence of a non-human reservoir where spillback of the pathogen is maintained in a sylvatic cycle and re-introduced into the urban cycle.[8] The presence of T. b. gambiense has been found separately in humans and livestock. This spurred a molecular study comparing serum reactivity of pigs, goats, and cows to human serum where notable similarities in all samples but especially in pig samples.[9] Combined, these findings implicate a reverse zoonotic human to animal transmission.
Arboviruses
Yellow fever viruses, Dengue fever viruses, and Zika viruses are of the Flavivirus genera and Chikungunya virus is of the Alphavirus genera. All of them are considered arboviruses denoting their ability to be transmitted through arthropod vectors.[10][11] Sylvatic transmission cycles for arboviruses within non-human primate communities have the potential to spillover into an urban cycle within humans where humans could be dead-end hosts in scenarios where further intermingling is eliminated but much more probable is a reemergence of these viruses into either cycle due to spillback.[12] Apparently the maintenance of an arboviral urban cycle between humans requires a rare or understudied conjunction of factors to occur. One of the following situations occurs:
- An infected human in an urban environment feeds a sylvatic (typically remotely located) mosquito such as Haemogogus (which has a relatively long lifespan compared to other mosquitoes and can transmit the virus for extended periods) that infects another human or non-human animal that will serve as a reservoir.
- An urban Aedes (more commonly found in urban areas [13] feeds and transmits the virus to another human or non-human animal that will serve as a reservoir.
- Sufficient numbers of sylvatic vector mosquito and the animal reservoir inhabit the same ecologic niche in close contact to promote and sustain the zoonotic cycle of the virus.
- The animal reservoir of the virus maintains a suitable virus level in the blood to allow the infection of a vector mosquito.
- A bridge-vector mosquito such as Aedes albopictus, which can survive in an urban area and spread to rural, semi-rural, and forest areas could carry the virus to a sylvatic environment.[14]Confronting data sparsity to identify potential sources of Zika virus spillover infection among primates
- Zika fever: The Zika virus is caused by the single stranded RNA Flavivirus that uses the Aedes mosquito as a vector to infect other human and animal hosts.[15] A 2015 zika virus strain isolated from a human in Brazil was used to infect pregnant rhesus macaques intravenously and intraamniotically. Both the dams and the placentas were infected with Zika positive tissue samples being recorded for up to 105 days. This confirms a reverse zoonotic transference potential between humans and non-human primates. [16]
- Yellow fever: Yellow fever virus also transmitted by the bite of an infected Aedes or Haemagogus species of mosquitoes that feed off an infected animal. The historical course of the American slave trade is a prime example of introduction of a pathogen to create a completely new sylvatic cycle. Previous hypotheses of a "New World YFV" were laid to rest in a 2007 study that examined rates of nucleotide substitution and divergence to determine that yellow fever was introduced into the Americas approximately 400 years ago from West Africa. It was also around the 17th century that yellow fever was documented by Europeans complicit in slave trafficking. The actual mode of introduction could have played out in a number of scenarios whether a viremic Old World human, infected Old World mosquito, eggs laid by an Old World infected mosquito, or all three were transported to the Americas seeing that yellow fever transmission was not uncommon on sailing vessels.[17] Amidst more recent yellow fever outbreaks in southeastern Brazil, the spillback potential was highly indicated.[18] Molecular comparisons of non-human primate outbreak strains proved to be more closely related to human strains than strains derived from other non-human primates thus suggesting a continuing reverse zoonosis.[19]
- Chikungunya: The Chikungunya virus is a single stranded RNA alphavirus typically transmitted by the Aedes mosquitoes to another animal host. There is no evidence to suggest a barrier to Chikungunya switching hosts between humans and non-human primates because it has no preferences in any given primate species. It has a high potential to spill-over or spill-back into sylvatic cycles as was the case with the similar arbovirus that was imported to the Americas during the slave trade.[20] Studies have proven chikungunya’s potential to orally infect sylvatic types of mosquitoes including Haemagogus leucocelaenus and Aedes terrens. Moreover, in a serologic survey carried out in non-human primates of urban and peri-urban areas of Bahia State, 11 animals showed chikungunya neutralizing antibodies.[12]
- Dengue fever: The Dengue virus is a flavivirus also transmissible by Aedes mosquito vectors to other animal hosts. Dengue was also introduced to the Americas by the slave trade along with Aedes aegypti.[21] A 2009 study in French Guiana found that infections of dengue viruses types 1 through 4 were present in various different types of neotropical forest mammals other than primates such as rodents, marsupials, and bats. After sequence analyses, it was revealed that the 4 non-human mammalian strains had an 89% to 99% similarity index to human strains circulating at the same time. This confirms that other mammals in the vicinity have the potential to be infected by human sources and indicates presence of an urban cycle.[22][23] A case to prove the arthropod vectors are capable of being infected comes from Brazil where Aedes albopictus (which frequents the backyards of human houses but easily spreads into rural, semi-rural, and wild environments) was found infected with dengue virus 3 in São Paulo State. Meanwhile, in the State of Bahia, the sylvatic vector Haemogogus leucocelaenus was found to be infected with dengue virus 1.[24] In another study carried out in the Atlantic Forest of Bahia, primates (Leontopithecus chrysomelas and Sapajus xanthosternos) were found with antibodies dengue viruses 1 and 2 while sloths (Bradypus torquatus) had antibodies for dengue virus 3 therefore suggesting the possible presence of an established sylvatic cycle.
3.2. Wild Animals
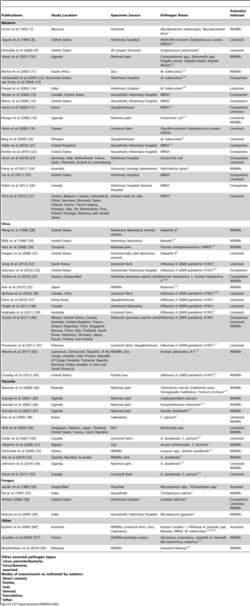
A large number of wild animals with habitats that have yet to be encroached upon by humans are still affected by sapronotic agents through contaminated water.
Giardia
- Beavers: Giardia was introduced to beavers through runoff of human sewage upstream of a beaver colony.[25]
Influenza A virus subtype H1N1
- Seals: In 1999, wild seals were admitted into a Dutch seal rehabilitation center with flu-like symptoms and it was found that they were in fact infected with a human influenza B like virus that had circulated in humans in 1995 and had undergone an antigenic shift since adaptation to its new seal host.[26]
Tuberculosis
- Red deer, wild boar: In areas of intensive game management that included big game fencing, supplementary feeding locations, and grazing livestock, cases of tuberculosis lesions in wild red deer and wild boars appeared. Some boars and deer shared the same strains of tuberculosis which were similar to those found in livestock and humans suggesting a possible sapronotic or sapro-zoonotic contamination of shared water sources, supplemental feed, direct contact with humans or livestock, or their excretions.[27]
3.3. Domesticated Companionship Animals
E. coli
- Dogs, horses: Evidence of infection by human E. coli strains in several dogs and horses across Europe was found, thus implicating the possibility of zoonotic inter-special transmission of multiresistant strains from humans to companion animals and vice-versa.[28]
Tuberculosis
- Dog: A Yorkshire terrier was admitted into a veterinary clinic with a chronic cough, poor weight retention, and vomiting being reported for months where it was revealed that the owner had recovered from tuberculosis, however the dog initially tested negative for tuberculosis in 2 different molecular assays and was discharged. 8 days later the dog was euthanized because of a urethral obstruction. A necroscopy was performed where liver and tracheobronchial lymph node samples in fact tested positive for the exact same strain of tuberculosis the owner had previously. This is a very clear case of reverse zoonosis.[29]
Influenza A virus subtype H1N1
- Ferrets: Ferrets are often used in human clinical studies thus the potential for human influenza to infect them was previously confirmed. However confirmation of natural transference of a human H1N1 strain from the 2009 outbreak in household pet ferrets further implicates human to animal transference.[30]
Amidst the 2020 global pandemic of COVID-19, Susceptibility of cats, ferrets, dogs, chickens, pigs, and ducks to the SARS-CoV-2 coronavirus was examined and it was found that it can be replicated in cats and ferrets with lethal results.
- Cats: The virus can be transmitted in the air between cats. Viral RNA was detected in feces within 3–5 days of infection and pathological studies detected viral RNA in the soft palate, tonsils, and trachea. Kittens acquired massive lesions in the lungs, nasal and tracheal mucosa epitheliums. Surveillance for SARS-CoV-2 in cats should be considered as an adjunct to elimination of COVID-19 in humans.[31]
- Ferrets: Ferrets were inoculated with viral strains from the environment of the Huanan Seafood Market in Wuhan, China as well as human isolates from Wuhan. It was found that with both isolates, that the virus can replicate in the upper respiratory tract of ferrets for up to 8 days without causing disease or death and viral RNA was detected in rectal swabs. Pathological studies performed after 13 days of infection revealed mild peribronchitis in the lungs, severe lymphoplasmacytic perivasculitis and vasculitis amongst other ailments with antibody production against SARS-CoV-2 detected in all ferrets. The fact that SARS-CoV-2 replicates efficiently in the upper respiratory tract of ferrets makes them a candidate animal model for evaluating antiviral drugs or vaccine candidates against COVID-19.[31]
- Dogs: Of the Beagle dogs tested, viral RNA was detected in fecal matter and 50% of the Beagles that were inoculated seroconverted after 14 days while the other 50% remained seronegative demonstrating a low susceptibility to SARS-CoV-2 in dogs.[31]
- Chicken, duck, pig: There was no evidence of susceptibility in chickens, ducks, or pigs with all viral RNA swabs returning negative results and seronegative after 14 days post inoculation.[31]
3.4. Domesticated Livestock Animals
Influenza A virus subtype H1N1
- Turkeys: A Norwegian turkey breeder’s flock exhibited a decrease in egg production with no other clinical signs after a farm hand reported having H1N1. A study revealed that the turkeys also had H1N1 and were seropositive to its antigens. Maternally derived H1N1 antibodies were detected in egg yolks and further genetic analyses revealed an identical H1N1 strain in the turkeys as the farm worker who likely infected the turkeys during artificial insemination.[32]
- Pigs: Human to pig H1N1 transmission was reported in Canada,[33] Korea,[34] and eventually came to include every continent save Antarctica during the 2009 outbreak.[35] It has also been known to spread during seasonal epidemics in France between humans and pigs.[36]
Methicillin-resistant staphylococcus aureus
- Horses: 11 equine patients admitted into a veterinary hospital for various reasons from different farms over the span of approximately one year exhibited MRSA infections later. Considering that MRSA isolates are extremely rare in horses, it was suggested that the MRSA outbreak was due to nosocomial infection derived from a human during the horse’s stay at the hospital.[37]
- Cows, turkeys, pigs: A case of reverse zoonosis was proposed to explain how a particular human Methicillin Sensitive Streptococcus Aureus strain was found in livestock (pigs, turkeys, cows) with not only a loss of human virulence genes (which could decrease zoonotic potential for human colonization) but also the addition of methicillin resistance and a tetracycline (which will increase occurrence of MRSA infections). The concern here being that excessive antibiotic usage in livestock productions exacerbate the creation of novel antibiotic resistant zoonotic pathogens.[38]
3.5. Wild Animals in Captivity
Tuberculosis
- Elephants: In 1996, The Hawthorne Circus Corporation reported 4 of their elephants and 11 of their keepers harboring M. tuberculosis infections. Unfortunately, these elephants had been sub-leased out to different circus acts and zoological gardens all over America. This spurred a nation-wide epidemic, but because tuberculosis isn’t a disease that’s typically transmitted from animals to humans, it was suggested that the epidemic was because of transference from a human handler to a captive elephant.[39]
- Alpacas: A 2007 outbreak of alpaca coronavirus because of the intermingling happening at a national alpaca exhibition led to a comparison between human and alpaca coronaviruses in an attempt to deduce the source of the outbreak. It was found that the alpaca coronavirus is most evolutionarily similar to a human coronavirus strain that was isolated in the 1960s suggesting that an alpaca coronavirus could have very well been circulating for decades causing respiratory illness in herds undetected for lack of diagnostic capabilities. It also suggests a human to alpaca mode of transmission.[40]
Helminths and protozoans
- Primates: At the University of Ibadan Zoological Garden in Nigeria, fecal samples of zookeepers tested positive for helminths (Ancylostoma duodenale, Ascaris lumbricoides and Trichuris trichiura) and protozoan cysts (Giardia lamblia and Entamoeba histolytica). The exact same strain of helminths and protozoan cysts were found in the zoo’s non-human primates thus implicating a zoonosis between the two.[41]
Measles
- Non-human primates: In 1996, a measles outbreak occurred in a sanctuary in 94 non-human primates. Although the source of the outbreak was never determined, serum and urine testing proved that the virus was definitely associated with recent human cases of measles in the U.S.[42]
Helicobacter pylori
- Marsupials: The stripe-face dunnart is an Australia n marsupial that has faced multiple outbreaks of Helicobacter pylori in captivity. Stomach sampling from the marsupial revealed that the H. pylori strain responsible for the outbreaks aligned 100% with a strain originating from the human intestinal tract. Thus, it can be assumed that the outbreak was caused by the handlers.[43]
3.6. Wild Animals in Conservation Areas
- Chimpanzees: The transmission of the human coronavirus (HCoV) OC43 to wild chimpanzees (Pan troglodytes verus) living in the Taï National Park, Côte d´Ivoire was reported in 2016 to 2017. These chimpanzees were accustomed to human presence that had been studying these particular communities since the 1980s [44] The HCoV-OC43, belonging to the species Betacoronavirus 1 (BetaCoV1), normally causes episodes of common cold in humans (this excludes SARS and MERS) human coronaviruses [45] but has also been detected in ungulates, carnivores, and lagomorphs.[46] Therefore it is completely plausible that researchers or poachers could have inadvertently spread the virus to the chimpanzees thus revealing yet another interface in coronavirus host switching.[47]
Rhinovirus C
- Chimpanzees: Though previously considered a uniquely human pathogen, human Rhinovirus C was determined to be the cause of a 2013 outbreak of respiratory infections in chimpanzees in Uganda. Examination of chimpanzees from all over Africa found that they show a universal homozygosity for the 3 CDHR3-Y529 allele (cadherin related family member) which is a receptor that drastically increases susceptibility to rhinovirus C infection and asthma in humans. If respiratory viruses of human origin are capable of maintaining circulation in non-human primates, this would prove to be harmful should the infection spillback into human communities.[48]
Tuberculosis
- Elephants: A necroscopy of a free-ranging African elephant (Loxodonta africana) in Kruger National Park in South Africa found significant lung damage due to a human strain of M. tuberculosis. Elephants explore their environment with their trunks therefore it was very likely that aerosolized pathogens from domestic waste, contaminated water from a human community upstream, human excrement, or contaminated food from tourists was the source of the infection.[49]
Pneumoviruses
- Chimpanzees: In Uganda, reports of respiratory viruses of human origination infected two chimpanzee (Pan troglodytes schweinfurthii) communities in the same forest. It was later discovered to be caused by a human metapneumovirus (also known as MPV, Pneumoviridae, Metapneumovirus) and a human respirovirus 3 (also known as HRV3, Paramyxoviridae, Respirovirus, or formerly known as parainfluenza virus 3).[50]
Reverse zoonosis in gorillas
- Gorillas: Conservational areas subject to ecotourism in Uganda, Rwanda, and the Democratic Republic of the Congo, free-ranging gorillas have become increasingly accustomed to the presence of humans whether that be in the form of ranger guides, tourists, trackers, veterinarians, poachers, or researchers. Iodamoeba buetschlii, Giardia lamblia, Chilomastix sp., Endolimax nana, Entamoeba coli, and Entamoeba histolytica have been found in the feces of gorillas and promiscuous defecations left behind by humans encroaching on the habitat. Additionally, increased numbers of Cryptosporidium sp. and capillaria infections were found in gorillas that maintained more frequent contact with humans than those that did not. Together these findings suggest the occurrence of reverse zoonoses.[51]
References
- "Emerging human infectious diseases: anthroponoses, zoonoses, and sapronoses". Emerging Infectious Diseases 9 (3): 403–4. March 2003. doi:10.3201/eid0903.020208. PMID 12643844. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2958532
- Zoonoses, Joint FAO/WHO Expert Committee on; Organization, World Health; Nations, Food and Agriculture Organization of the United (1967) (in en). Joint FAO/WHO Expert Committee on Zoonoses [meeting held in Geneva from 6 to 12 December 1966] : third report. World Health Organization. ISBN 978-92-4-120378-4.
- Tuzio, H; Edwards, D; Elston, T; Jarboe, L; Kudrak, S; Richards, J; Rodan, I (August 2005). "Feline zoonoses guidelines from the American Association of Feline Practitioners" (in en). Journal of Feline Medicine & Surgery 7 (4): 243–274. doi:10.1016/j.jfms.2004.11.001. PMID 16130211. https://dx.doi.org/10.1016%2Fj.jfms.2004.11.001
- Prevention, CDC-Centers for Disease Control and (2019-01-28). "CDC - Malaria - About Malaria - Biology" (in en-us). https://www.cdc.gov/malaria/about/biology/index.html.
- Ramasamy, Ranjan (2014-08-18). "Zoonotic Malaria – Global Overview and Research and Policy Needs". Frontiers in Public Health 2: 123. doi:10.3389/fpubh.2014.00123. ISSN 2296-2565. PMID 25184118. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=4135302
- Lalremruata, Albert; Magris, Magda; Vivas-Martínez, Sarai; Koehler, Maike; Esen, Meral; Kempaiah, Prakasha; Jeyaraj, Sankarganesh; Perkins, Douglas Jay et al. (September 2015). "Natural infection of Plasmodium brasilianum in humans: Man and monkey share quartan malaria parasites in the Venezuelan Amazon". EBioMedicine 2 (9): 1186–1192. doi:10.1016/j.ebiom.2015.07.033. ISSN 2352-3964. PMID 26501116. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=4588399
- "CDC - African Trypanosomiasis - Biology" (in en-us). 2019-06-12. https://www.cdc.gov/parasites/sleepingsickness/biology.html.
- Funk, Sebastian; Nishiura, Hiroshi; Heesterbeek, Hans; Edmunds, W. John; Checchi, Francesco (2013-01-17). "Identifying Transmission Cycles at the Human-Animal Interface: The Role of Animal Reservoirs in Maintaining Gambiense Human African Trypanosomiasis". PLOS Computational Biology 9 (1): e1002855. doi:10.1371/journal.pcbi.1002855. ISSN 1553-734X. PMID 23341760. Bibcode: 2013PLSCB...9E2855F. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3547827
- Cordon-Obras, Carlos; Cano, Jorge; González-Pacanowska, Dolores; Benito, Agustin; Navarro, Miguel; Bart, Jean-Mathieu (2013-12-23). "Trypanosoma brucei gambiense Adaptation to Different Mammalian Sera Is Associated with VSG Expression Site Plasticity". PLOS ONE 8 (12): e85072. doi:10.1371/journal.pone.0085072. ISSN 1932-6203. PMID 24376866. Bibcode: 2013PLoSO...885072C. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3871602
- Kuno, G.; Chang, G. J.; Tsuchiya, K. R.; Karabatsos, N.; Cropp, C. B. (January 1998). "Phylogeny of the genus Flavivirus". Journal of Virology 72 (1): 73–83. doi:10.1128/JVI.72.1.73-83.1998. ISSN 0022-538X. PMID 9420202. PMC 109351. https://www.ncbi.nlm.nih.gov/pubmed?Db=pubmed&Cmd=ShowDetailView&TermToSearch=9420202.
- Forrester, N. L.; Palacios, G.; Tesh, R. B.; Savji, N.; Guzman, H.; Sherman, M.; Weaver, S. C.; Lipkin, W. I. (March 2012). "Genome-scale phylogeny of the alphavirus genus suggests a marine origin". Journal of Virology 86 (5): 2729–2738. doi:10.1128/JVI.05591-11. ISSN 1098-5514. PMID 22190718. PMC 3302268. https://www.ncbi.nlm.nih.gov/pubmed?Db=pubmed&Cmd=ShowDetailView&TermToSearch=22190718.
- Figueiredo, Luiz Tadeu Moraes (2019). "Human Urban Arboviruses Can Infect Wild Animals and Jump to Sylvatic Maintenance Cycles in South America" (in English). Frontiers in Cellular and Infection Microbiology 9: 259. doi:10.3389/fcimb.2019.00259. ISSN 2235-2988. PMID 31380302. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=6653809
- Ndenga, Bryson Alberto; Mutuku, Francis Maluki; Ngugi, Harun Njenga; Mbakaya, Joel Omari; Aswani, Peter; Musunzaji, Peter Siema; Vulule, John; Mukoko, Dunstan et al. (2017-12-19). "Characteristics of Aedes aegypti adult mosquitoes in rural and urban areas of western and coastal Kenya" (in en). PLOS ONE 12 (12): e0189971. doi:10.1371/journal.pone.0189971. ISSN 1932-6203. PMID 29261766. Bibcode: 2017PLoSO..1289971N. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=5736227
- Hanley, Kathryn A.; Monath, Thomas P.; Weaver, Scott C.; Rossi, Shannan L.; Richman, Rebecca L.; Vasilakis, Nikos (October 2013). "Fever versus fever: the role of host and vector susceptibility and interspecific competition in shaping the current and future distributions of the sylvatic cycles of dengue virus and yellow fever virus". Infection, Genetics and Evolution: Journal of Molecular Epidemiology and Evolutionary Genetics in Infectious Diseases 19: 292–311. doi:10.1016/j.meegid.2013.03.008. ISSN 1567-7257. PMID 23523817. PMC 3749261. https://www.ncbi.nlm.nih.gov/pubmed?Db=pubmed&Cmd=ShowDetailView&TermToSearch=23523817.
- "Zika Virus Epidemiology and Countries at Risk of Zika Transmisison" (in en-gb). http://www.zikavirusnet.com/epidemiology.html.
- "HHSF223201610542P Final Report". May 22, 2017. http://wayback.archive-it.org/7993/20180126133803/https://www.fda.gov/downloads/EmergencyPreparedness/Counterterrorism/MedicalCountermeasures/MCMLegalRegulatoryandPolicyFramework/UCM565405.pdf.
- Bryant, Juliet E; Holmes, Edward C; Barrett, Alan D. T (May 2007). "Out of Africa: A Molecular Perspective on the Introduction of Yellow Fever Virus into the Americas". PLOS Pathogens 3 (5): e75. doi:10.1371/journal.ppat.0030075. ISSN 1553-7366. PMID 17511518. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1868956
- Couto-Lima, Dinair; Madec, Yoann; Bersot, Maria Ignez; Campos, Stephanie Silva; Motta, Monique de Albuquerque; Santos, Flávia Barreto Dos; Vazeille, Marie; Vasconcelos, Pedro Fernando da Costa et al. (July 7, 2017). "Potential risk of re-emergence of urban transmission of Yellow Fever virus in Brazil facilitated by competent Aedes populations". Scientific Reports 7 (1): 4848. doi:10.1038/s41598-017-05186-3. ISSN 2045-2322. PMID 28687779. PMC 5501812. Bibcode: 2017NatSR...7.4848C. https://www.ncbi.nlm.nih.gov/pubmed?Db=pubmed&Cmd=ShowDetailView&TermToSearch=28687779.
- Moreira-Soto, A.; Torres, M. C.; Lima de Mendonça, M. C.; Mares-Guia, M. A.; Dos Santos Rodrigues, C. D.; Fabri, A. A.; Dos Santos, C. C.; Machado Araújo, E. S. et al. (September 2018). "Evidence for multiple sylvatic transmission cycles during the 2016-2017 yellow fever virus outbreak, Brazil". Clinical Microbiology and Infection 24 (9): 1019.e1–1019.e4. doi:10.1016/j.cmi.2018.01.026. ISSN 1469-0691. PMID 29427798. https://dx.doi.org/10.1016%2Fj.cmi.2018.01.026
- Tsetsarkin, Konstantin A; Chen, Rubing; Weaver, Scott C (February 2016). "Interspecies transmission and chikungunya virus emergence". Current Opinion in Virology 16: 143–150. doi:10.1016/j.coviro.2016.02.007. ISSN 1879-6257. PMID 26986235. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=4824623
- Hanley, Kathryn A.; Monath, Thomas P.; Weaver, Scott C.; Rossi, Shannan L.; Richman, Rebecca L.; Vasilakis, Nikos (October 2013). "Fever versus Fever: the role of host and vector susceptibility and interspecific competition in shaping the current and future distributions of the sylvatic cycles of dengue virus and yellow fever virus". Infection, Genetics and Evolution : Journal of Molecular Epidemiology and Evolutionary Genetics in Infectious Diseases 0: 292–311. doi:10.1016/j.meegid.2013.03.008. ISSN 1567-1348. PMID 23523817. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3749261
- de Thoisy, Benoît; Lacoste, Vincent; Germain, Adeline; Muñoz-Jordán, Jorge; Colón, Candimar; Mauffrey, Jean-François; Delaval, Marguerite; Catzeflis, François et al. (April 2009). "Dengue infection in neotropical forest mammals". Vector Borne and Zoonotic Diseases (Larchmont, N.Y.) 9 (2): 157–170. doi:10.1089/vbz.2007.0280. ISSN 1557-7759. PMID 18945183. https://www.ncbi.nlm.nih.gov/pubmed?Db=pubmed&Cmd=ShowDetailView&TermToSearch=18945183.
- Calderón, Alfonso; Guzmán, Camilo; Mattar, Salim; Rodriguez, Virginia; Martínez, Caty; Violet, Lina; Martínez, Jairo; Figueiredo, Luiz Tadeu Moraes (October 2019). "Dengue Virus in Bats from Córdoba and Sucre, Colombia". Vector Borne and Zoonotic Diseases (Larchmont, N.Y.) 19 (10): 747–751. doi:10.1089/vbz.2018.2324. ISSN 1557-7759. PMID 31211661. PMC 6765209. https://www.ncbi.nlm.nih.gov/pubmed?Db=pubmed&Cmd=ShowDetailView&TermToSearch=31211661.
- de Figueiredo, Mario LG; de C Gomes, Almério; Amarilla, Alberto A.; de S Leandro, André; de S Orrico, Agnaldo; de Araujo, Renato F.; do SM Castro, Jesuína; Durigon, Edison L. et al. (2010-07-12). "Mosquitoes infected with dengue viruses in Brazil". Virology Journal 7 (1): 152. doi:10.1186/1743-422X-7-152. ISSN 1743-422X. PMID 20624314. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2913956
- Ash, A.; Lymbery, A.; Lemon, J.; Vitali, S.; Thompson, R. C. A. (2010-12-15). "Molecular epidemiology of Giardia duodenalis in an endangered carnivore – The African painted dog" (in en). Veterinary Parasitology 174 (3): 206–212. doi:10.1016/j.vetpar.2010.08.034. ISSN 0304-4017. PMID 20851525. http://www.sciencedirect.com/science/article/pii/S0304401710004899.
- Osterhaus, A. D.; Rimmelzwaan, G. F.; Martina, B. E.; Bestebroer, T. M.; Fouchier, R. A. (2000-05-12). "Influenza B virus in seals". Science 288 (5468): 1051–1053. doi:10.1126/science.288.5468.1051. ISSN 0036-8075. PMID 10807575. Bibcode: 2000Sci...288.1051O. https://dx.doi.org/10.1126%2Fscience.288.5468.1051
- Barasona, J. A.; Vicente, J.; Díez-Delgado, I.; Aznar, J.; Gortázar, C.; Torres, M. J. (August 2017). "Environmental Presence of Mycobacterium tuberculosis Complex in Aggregation Points at the Wildlife/Livestock Interface". Transboundary and Emerging Diseases 64 (4): 1148–1158. doi:10.1111/tbed.12480. ISSN 1865-1682. PMID 26865411. https://dx.doi.org/10.1111%2Ftbed.12480
- Ewers, Christa; Grobbel, Mirjam; Stamm, Ivonne; Kopp, Peter A.; Diehl, Ines; Semmler, Torsten; Fruth, Angelika; Beutlich, Janine et al. (April 2010). "Emergence of human pandemic O25:H4-ST131 CTX-M-15 extended-spectrum-beta-lactamase-producing Escherichia coli among companion animals". The Journal of Antimicrobial Chemotherapy 65 (4): 651–660. doi:10.1093/jac/dkq004. ISSN 1460-2091. PMID 20118165. https://dx.doi.org/10.1093%2Fjac%2Fdkq004
- Erwin, Paul C.; Bemis, David A.; McCombs, Scott B.; Sheeler, Lorinda L.; Himelright, Inga M.; Halford, Sandy K.; Diem, Lois; Metchock, Beverly et al. (December 2004). "Mycobacterium tuberculosis transmission from human to canine". Emerging Infectious Diseases 10 (12): 2258–2210. doi:10.3201/eid1012.040094. ISSN 1080-6040. PMID 15672533. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3323378
- Swenson, Sabrina L.; Koster, Leo G.; Jenkins-Moore, Melinda; Killian, Mary L.; DeBess, Emilio E.; Baker, Rocky J.; Mulrooney, Donna; Weiss, Robin et al. (September 2010). "Natural cases of 2009 pandemic H1N1 Influenza A virus in pet ferrets". Journal of Veterinary Diagnostic Investigation 22 (5): 784–788. doi:10.1177/104063871002200525. ISSN 1040-6387. PMID 20807944. https://dx.doi.org/10.1177%2F104063871002200525
- Shi, Jianzhong; Wen, Zhiyuan; Zhong, Gongxun; Yang, Huanliang; Wang, Chong; Huang, Baoying; Liu, Renqiang; He, Xijun et al. (2020-04-08). "Susceptibility of ferrets, cats, dogs, and other domesticated animals to SARS–coronavirus 2" (in en). Science: eabb7015. doi:10.1126/science.abb7015. ISSN 0036-8075. PMID 32269068. PMC 7164390. https://science.sciencemag.org/content/early/2020/04/07/science.abb7015.
- Sjurseth, Siri Kulberg; Gjerset, Britt; Bragstad, Karoline; Hungnes, Olav; Wisløff, Helene; Er, Chiek; Valheim, Mette; Løtvedt, Siri M. et al. (2017). "Human to animal transmission of influenza A(H1N1)pdm09 in a turkey breeder flock in Norway". Infection Ecology & Epidemiology 7 (1): 1416249. doi:10.1080/20008686.2017.1416249. ISSN 2000-8686. PMID 29296243. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=5738641
- Howden, Krista J.; Brockhoff, Egan J.; Caya, Francois D.; McLeod, Laura J.; Lavoie, Martin; Ing, Joan D.; Bystrom, Janet M.; Alexandersen, Soren et al. (November 2009). "An investigation into human pandemic influenza virus (H1N1) 2009 on an Alberta swine farm". The Canadian Veterinary Journal 50 (11): 1153–1161. ISSN 0008-5286. PMID 20119537. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2764467
- Song, Min-Suk; Lee, Jun Han; Pascua, Philippe Noriel Q.; Baek, Yun Hee; Kwon, Hyeok-il; Park, Kuk Jin; Choi, Hwan-Woon; Shin, Yeun-Kyung et al. (September 2010). "Evidence of Human-to-Swine Transmission of the Pandemic (H1N1) 2009 Influenza Virus in South Korea". Journal of Clinical Microbiology 48 (9): 3204–3211. doi:10.1128/JCM.00053-10. ISSN 0095-1137. PMID 20610681. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2937722
- Nelson, Martha I.; Gramer, Marie R.; Vincent, Amy L.; Holmes, Edward C. (October 2012). "Global transmission of influenza viruses from humans to swine". The Journal of General Virology 93 (Pt 10): 2195–2203. doi:10.1099/vir.0.044974-0. ISSN 0022-1317. PMID 22791604. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3541789
- Chastagner, Amélie; Enouf, Vincent; Peroz, David; Hervé, Séverine; Lucas, Pierrick; Quéguiner, Stéphane; Gorin, Stéphane; Beven, Véronique et al. (October 2019). "Bidirectional Human–Swine Transmission of Seasonal Influenza A(H1N1)pdm09 Virus in Pig Herd, France, 2018". Emerging Infectious Diseases 25 (10): 1940–1943. doi:10.3201/eid2510.190068. ISSN 1080-6040. PMID 31538914. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=6759248
- Seguin, Jennifer C.; Walker, Robert D.; Caron, John P.; Kloos, Wesley E.; George, Carol G.; Hollis, Richard J.; Jones, Ronald N.; Pfaller, Michael A. (May 1999). "Methicillin-Resistant Staphylococcus aureus Outbreak in a Veterinary Teaching Hospital: Potential Human-to-Animal Transmission". Journal of Clinical Microbiology 37 (5): 1459–1463. doi:10.1128/JCM.37.5.1459-1463.1999. ISSN 0095-1137. PMID 10203505. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=84801
- Price, Lance B.; Stegger, Marc; Hasman, Henrik; Aziz, Maliha; Larsen, Jesper; Andersen, Paal Skytt; Pearson, Talima; Waters, Andrew E. et al. (2012-02-21). "Staphylococcus aureus CC398: Host Adaptation and Emergence of Methicillin Resistance in Livestock". mBio 3 (1). doi:10.1128/mBio.00305-11. ISSN 2150-7511. PMID 22354957. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3280451
- Holt, Nathalia (2015-03-24). "We Are in the Midst of an Elephant Tuberculosis Epidemic" (in en). https://slate.com/technology/2015/03/elephant-tuberculosis-epidemic-zoo-and-circus-animals-passing-tb-to-humans-and-one-another.html.
- Crossley, Beate M.; Mock, Richard E.; Callison, Scott A.; Hietala, Sharon K. (2012-12-12). "Identification and Characterization of a Novel Alpaca Respiratory Coronavirus Most Closely Related to the Human Coronavirus 229E". Viruses 4 (12): 3689–3700. doi:10.3390/v4123689. ISSN 1999-4915. PMID 23235471. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3528286
- "Preliminary Investigation of Zooanthroponosis in a Nigerian Zoological Garden". https://medwelljournals.com/abstract/?doi=vr.2008.38.41.
- Willy, M. E.; Woodward, R. A.; Thornton, V. B.; Wolff, A. V.; Flynn, B. M.; Heath, J. L.; Villamarzo, Y. S.; Smith, S. et al. (February 1999). "Management of a measles outbreak among Old World nonhuman primates". Laboratory Animal Science 49 (1): 42–48. ISSN 0023-6764. PMID 10090093. http://www.worldcat.org/issn/0023-6764
- Every, Alison L.; Selwood, Lynne; Castano-Rodriguez, Natalia; Lu, Wei; Windsor, Helen M.; Wee, Janet LK; Swierczak, Agnieszka; Marshall, Barry J. et al. (2011-02-07). "Did transmission of Helicobacter pylori from humans cause a disease outbreak in a colony of Stripe-faced Dunnarts (Sminthopsis macroura)?". Veterinary Research 42 (1): 26. doi:10.1186/1297-9716-42-26. ISSN 1297-9716. PMID 21314909. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3042409
- The Chimpanzees of the Taï Forest: Behavioural Ecology and Evolution. Oxford, New York: Oxford University Press. 2000-06-15. ISBN 978-0-19-850507-5. https://global.oup.com/academic/product/the-chimpanzees-of-the-ta-forest-9780198505075?cc=us&lang=en&.
- Mackay, Ian M.; Arden, Katherine E.; Speicher, David J.; O’Neil, Nicholas T.; McErlean, Peter K.; Greer, Ristan M.; Nissen, Michael D.; Sloots, Theo P. (2012-04-23). "Co-circulation of Four Human Coronaviruses (HCoVs) in Queensland Children with Acute Respiratory Tract Illnesses in 2004". Viruses 4 (4): 637–653. doi:10.3390/v4040637. ISSN 1999-4915. PMID 22590689. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3347326
- Corman, Victor M.; Muth, Doreen; Niemeyer, Daniela; Drosten, Christian (2018). "Hosts and Sources of Endemic Human Coronaviruses". Advances in Virus Research 100: 163–188. doi:10.1016/bs.aivir.2018.01.001. ISBN 9780128152010. ISSN 0065-3527. PMID 29551135. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=7112090
- Patrono, Livia V.; Samuni, Liran; Corman, Victor M.; Nourifar, Leila; Röthemeier, Caroline; Wittig, Roman M.; Drosten, Christian; Calvignac-Spencer, Sébastien et al. (2018-06-27). "Human coronavirus OC43 outbreak in wild chimpanzees, Côte d´Ivoire, 2016". Emerging Microbes & Infections 7 (1): 118. doi:10.1038/s41426-018-0121-2. ISSN 2222-1751. PMID 29950583. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=6021434
- Scully, Erik J.; Basnet, Sarmi; Wrangham, Richard W.; Muller, Martin N.; Otali, Emily; Hyeroba, David; Grindle, Kristine A.; Pappas, Tressa E. et al. (February 2018). "Lethal Respiratory Disease Associated with Human Rhinovirus C in Wild Chimpanzees, Uganda, 2013". Emerging Infectious Diseases 24 (2): 267–274. doi:10.3201/eid2402.170778. ISSN 1080-6040. PMID 29350142. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=5782908
- Miller, Michele A.; Buss, Peter; Roos, Eduard O.; Hausler, Guy; Dippenaar, Anzaan; Mitchell, Emily; van Schalkwyk, Louis; Robbe-Austerman, Suelee et al. (2019-02-06). "Fatal Tuberculosis in a Free-Ranging African Elephant and One Health Implications of Human Pathogens in Wildlife". Frontiers in Veterinary Science 6: 18. doi:10.3389/fvets.2019.00018. ISSN 2297-1769. PMID 30788347. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=6373532
- Negrey, Jacob D.; Reddy, Rachna B.; Scully, Erik J.; Phillips-Garcia, Sarah; Owens, Leah A.; Langergraber, Kevin E.; Mitani, John C.; Emery Thompson, Melissa et al. (2019-01-21). "Simultaneous outbreaks of respiratory disease in wild chimpanzees caused by distinct viruses of human origin". Emerging Microbes & Infections 8 (1): 139–149. doi:10.1080/22221751.2018.1563456. ISSN 2222-1751. PMID 30866768. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=6455141
- Nizeyi, John Bosco; Innocent, Rwego B.; Erume, Joseph; Kalema, Gladys R. N. N.; Cranfield, Michael R.; Graczyk, Thaddeus K. (April 2001). "Campylobacteriosis, Salmonellosis, and Shigellosis in Free-Ranging Human-Habituated Mountain Gorillas of Uganda" (in en). Journal of Wildlife Diseases 37 (2): 239–244. doi:10.7589/0090-3558-37.2.239. ISSN 0090-3558. PMID 11310873. https://dx.doi.org/10.7589%2F0090-3558-37.2.239





