| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Biao Jin | + 1778 word(s) | 1778 | 2020-11-11 07:27:22 | | | |
| 2 | Vicky Zhou | -63 word(s) | 1715 | 2020-11-24 05:17:17 | | |
Video Upload Options
The WRKY gene family is a plant-specific transcription factor (TF) group, playing important roles in many different response pathways of diverse abiotic stresses (drought, saline, alkali, temperature, and ultraviolet radiation, and so forth). In recent years, many studies have explored the role and mechanism of WRKY family members from model plants to agricultural crops and other species. Abiotic stress adversely affects the growth and development of plants.
1. Introduction
As a fixed-growth organism, plants are exposed to a variety of environmental conditions and may encounter many abiotic stresses, for example, drought, waterlogging, heat, cold, salinity, and Ultraviolet-B (UV-B) radiation. To adapt and counteract the effects of such abiotic stresses, plants have evolved several molecular mechanisms involving signal transduction and gene expression [1][2]. Transcription factors (TFs) are important regulators involved in the process of signal transduction and gene expression regulation under environmental stresses. TFs can be combined with cis-acting elements to regulate the transcriptional efficiency of target genes by inhibiting or enhancing their transcription [3][4]. Accordingly, plants may show corresponding responses to external stresses via TFs regulating target genes. Although some TF families (MYB, bZIP, AP2/EREBP, NAC) are associated with adversity [2][5], WRKY is the most extensively studied TF family in plant stress responses.
The WRKY family is a unique TF superfamily of higher plants and algae, which play important roles in many life processes, particularly in response against biotic and abiotic stress [6][7]. In 1994, the SWEET POTATO FACTOR1 (SPF1) gene of the WRKY family was first found in Impoea batatas [8]. Later, ABF1 and ABF2 were found in wild Avena sativa, and showed regulatory roles in seed germination [9]. A previous study successively cloned WRKY1, WRKY2, and WRKY3 from Petroselinum crispum, named the WRKY TF, and proved for the first time that WRKY protein can regulate plant responses to pathogens [10]. With an increase in available published genomes, many members of the WRKY TF family have been identified in various species, including 104 from Populus [11], 37 from Physcomitrella patens [12], 45 from Hordeum vulgare [13], 55 from Cucumis sativus [14], 74 from Arabidopsis thaliana [15], 83 from Pinus monticola [16], 81 from Solanum lycopersicum [17], and 102 from Oryza sativa [18]. WRKY TFs exist as gene families in plants, and the number of WRKY TFs varies among species. In plants exposed to abiotic stresses (salt, drought, temperature, and so forth), WRKY family members play important roles in diverse stress responses. In addition, these TFs affect the growth and development of plants [19][20]. Therefore, WRKY TFs have attracted broad attention. The Structural Characteristics of WRKY TFs can be seen in Figure 1.

2. WRKY TF Involved in Abiotic Stress Responses
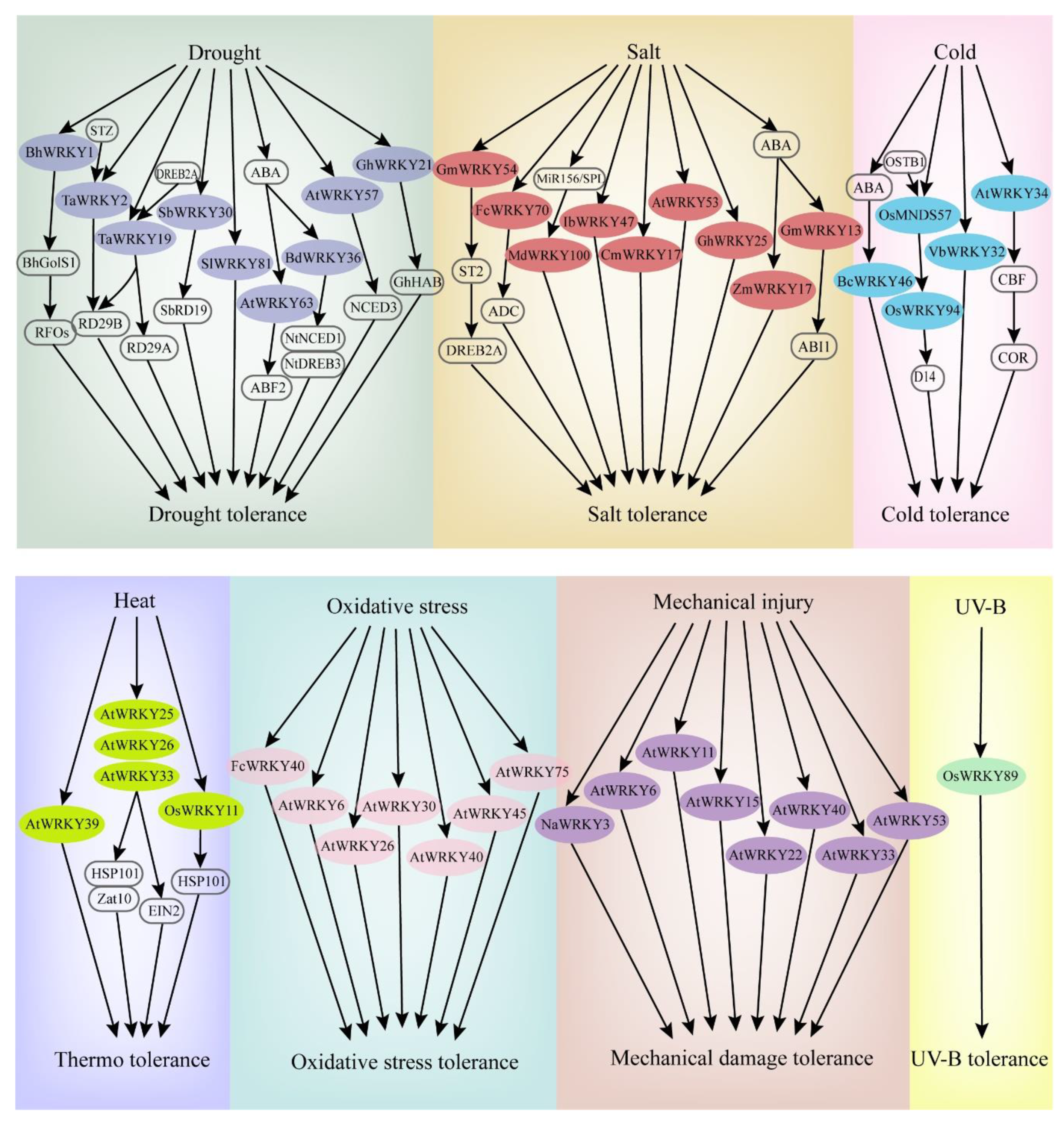
When plants sense stress, the corresponding signaling is activated and transferred to the cell interior. Reactive oxygen species (ROS) and Ca2+ ions are usually exchanged as the signal transduction in the cell. Protein kinases such as MPKs are subsequently activated to regulate the activities of related TFs. Consequently, the plant presents a stress response [21][22]. In response to abiotic stresses, some WRKY TFs can be rapidly differentially expressed, promoting signal transduction and regulating the expression of related genes [23]. The expression patterns and functional identifications of WRKYs in most studies are generally based on transcriptome analyses, real-time fluorescence quantitative PCR, gene chip analyses, and genetic transformation. Hence, WRKY genes can function effectively in most abiotic stress responses or tolerances in various plants (Figure 2).
Figure 2. Some WRKY genes involved in the response pathways of major abiotic stresses (drought, salt, cold, heat, oxidative stress, mechanical injury, UV-B).
2.1. WRKY TFs and Drought Stress
Drought has a major impact on plant growth and development, resulting in a significant decrease in grain and other types of crop yield [24]. Under drought stress, drought-tolerant plants can accumulate oligosaccharides through sucrose metabolism to improve drought resistance. For example, when Arabidopsis is subjected to drought stress, the expression of AtWRKY53 combined with the Qua-Quine Starch (QQS) promoter sequence is rapidly induced, hydrogen peroxide content is reduced, and the glucose metabolism pathway is significantly enhanced, thereby regulating stomatal opening and ultimately affecting drought tolerance [25]. In Boea hygrometrica, BhWRKY1 effectively regulates the expression of the BhGolS1 gene, and the overexpression of BhGolS1 and BhWRKY1 induces the accumulation of raffinose family oligosaccharides (RFOs) in transgenic Nicotiana tabacum, thus improving the ability of seedlings to resist drought [26].
WRKY protein can directly regulate the expression of drought-resistant genes. For example, in sorghum, SbWRKY30 regulates the drought stress response gene SbRD19 by binding with W-box elements of the SbRD19 promoter, and protects plant cells from the damage of reactive oxygen species by improving ROS scavenging capability, enhancing drought tolerance [27]. TaWRKY2 of wheat can bind to STZ and downstream drought-resistant gene RD29B promoter, with a positive regulatory effect on the expression of RD29B [28]. DREB2A regulates the expression of dehydration stress-related genes [29], while TaWRKY19 can bind to DREB2A promoter, ultimately activating the expression of RD29A, RD29B, and Cor6.6 in transgenic Arabidopsis plants [28]. Similarly, Arabidopsis AtWRKY57 positively regulates the expression of RD29A and NCED3 genes by binding their W-box elements in the promoter regions [30]. In addition, WRKY protein can act on other TFs to play regulatory roles in drought tolerance. For example, TcWRKY53 of Thlaspi arvense significantly inhibits the expression of NtERF5 and NterEBp-1 of the AP2/ERF TF family, thus improving plant resistance to drought stress [31].
WRKY TFs also regulate plant tolerance through abscisic acid (ABA) and ROS-related signaling pathways. During drought stress, higher ABA levels were accumulated in plants, and leaf stomata were closed to reduce transpiration rate, thus regulating water balance in plants. ABA accumulation in cells, integrated with a variety of stress signals, regulates the expression of downstream genes, consequently sensing and responding to the adverse environment [40]. Arabidopsis AtWRKY63 has a specific effect on ABA-mediated stomatal closure and other signal transduction pathways, thus affecting the drought response [32]. GhWRKY21 regulates ABA-mediated cotton drought tolerance by promoting the expression of GhHAB [33]. Overexpression of BdWRKY36 in tobacco reduces the accumulation of ROS, activated NtLEA5, NtNCED1, and NtDREB3 in the ABA biosynthetic pathway, and significantly enhances the drought resistance of plants [34]. In Solanum lycopersicum, SlWRKY81 increases the drought tolerance of plants by inhibiting the accumulation of H2O2, playing a negative regulation role of stomatal closure [35].
2.2. WRKY TFs and Salt Stress
Salt stress is an important abiotic stress affecting crop productivity, particularly in arid and semiarid regions. WRKY TFs play essential roles in regulating the response to salt stress. To date, a total of 47 WRKY genes have been found to be expressed under salt stress in the wheat genome [36]. STZ is a protein related to ZPT2, which acts as a transcriptional inhibitor to downregulate the deactivation of other transcription factors. GmWRKY54 of Glycine max inhibits STZ expression and responds to salt stress by positively regulating the DREB2A-mediated pathway [37]. FcWRKY70 promotes the upregulation of arginine decarboxylase (ADC) expression, which is heterologously expressed in tobacco, and the content of lemon putrescine is significantly increased, thus enhancing the salt tolerance of plants [38]. The IbWRKY47 gene positively regulates stress resistance-related genes and significantly improves the salt tolerance of Ipomoea batatas [39]. MiR156/SPL modulates salt tolerance by upregulation of Malus domestica salt tolerance gene MdWRKY100 [40]. In Sorghum bicolor, SbWRKY50 could directly bind to the upstream promoter of SOS1 and HKT1 and participate in plant salt response by controlling ion homeostasis [41].
In addition, some WRKY genes function as negative regulation factors involved in salt stress resistance. Arabidopsis RPD3-like histone deacetylase HDA9 inhibits salt stress tolerance by regulating the DNA binding and transcriptional activity of WRKY53 [42]. Chrysanthemum CmWRKY17 overexpressed in Arabidopsis allows the plants to be more sensitive to salt stress. The expression level of stress resistance-related genes in transgenic Arabidopsis is lower than that in wild-type plants, indicating that CmWRKY17 may be involved in negatively regulating the salt stress response in Chrysanthemum [43]. The expression of GhWRKY68 is strongly induced in upland cotton and decreases salt tolerance [44]. In contrast, a high expression level of GhWRKY25 enhances the salt tolerance of upland cotton, while transgenic tobacco shows a relatively weaker tolerance to drought stress [45], indicating that the regulatory effects of different WRKY TFs involved in drought response are different.
2.3. WRKY TFs and Temperature Stress
Both low- and high-temperature stress can reduce crop yield and quality in plants. WRKY TFs play a role in the stress response through different signal transduction pathways. For example, in Verbena bonariensis, VbWRKY32 as a positive regulator, upregulates the transcriptional level of cold response genes, which increases the antioxidant activity, maintains membrane stability, and enhances osmotic regulation ability, thereby improving the survival ability under cold stress [46]. The BcWRKY46 gene of Brassica campestris is strongly induced by low temperature and ABA, activating related genes in the ABA signaling pathway to improve the low-temperature tolerance of plants [47]. CBF TFs regulate the expression of COR, and the overexpressed transgenic lines of CBF1, CBF2, and CBF3 show stronger cold resistance [48]. AtWRKY34 has a negative regulatory effect on the CBF-mediated cold response pathway; it is specifically expressed in mature pollen grains after exposure to low temperatures, resulting in resistance to low temperatures [49]. In addition, plants respond to temperature changes by coordinating organ development in an adverse environment. At low temperatures, rice MADS-Box TF OsMADS57 and its interacting protein OsTB1 synergistically activate the transcriptional regulation of OsWRKY94, preventing tillering by inhibiting transcription of the organ development gene D14 [50].
2.4. WRKY TFs and Other Abiotic Stresses
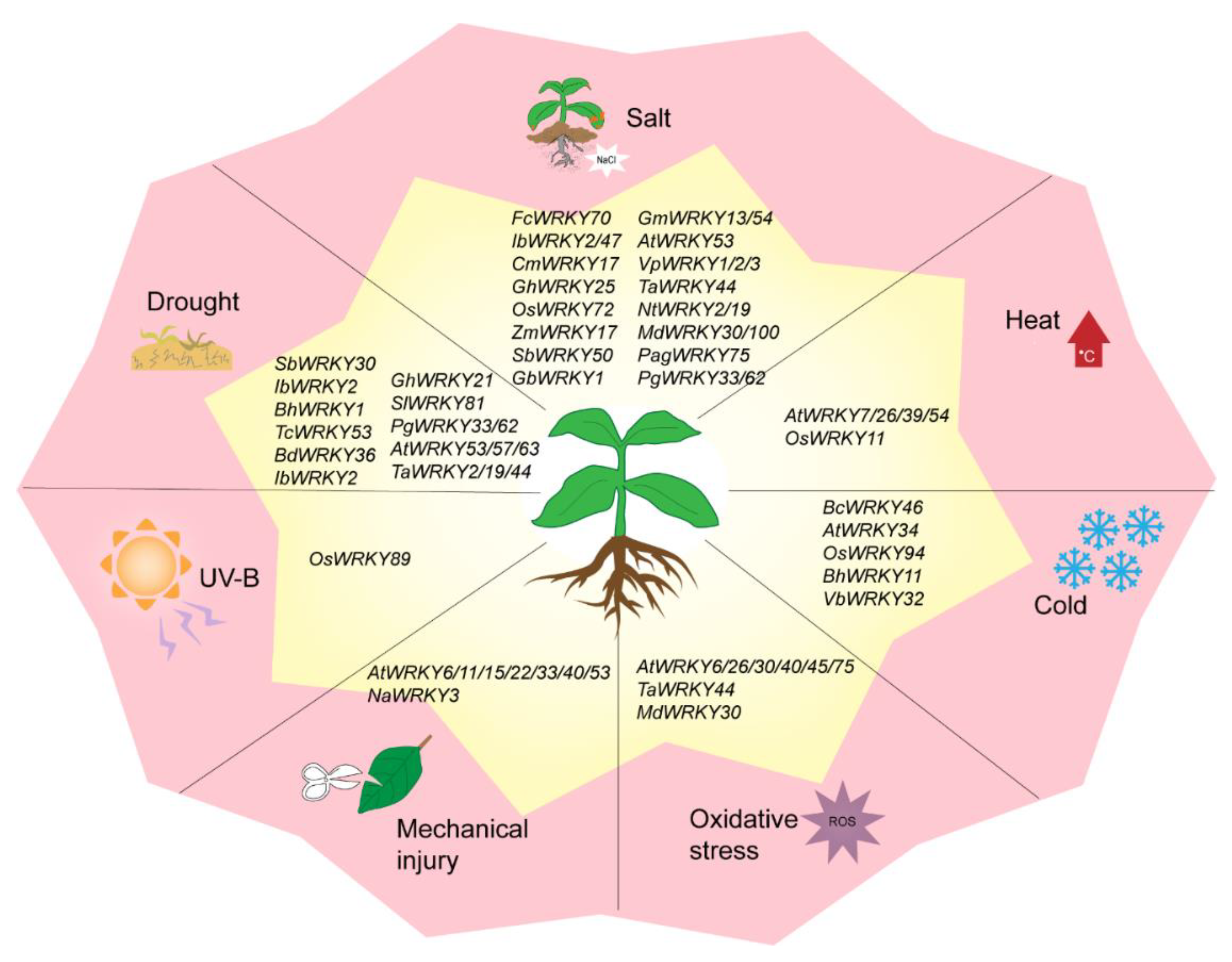
WRKY TFs are also involved in oxidative stress, mechanical damage, UV radiation, and other abiotic stresses (Figure 3). FcWRKY40 overexpression can significantly enhance the resistance of transgenic tobacco to oxidative stress [51]. When Arabidopsis is treated with ROS, the expressions of AtWRKY30, AtWRKY40, AtWRKY75, AtWRKY6, AtWRKY26, and AtWRKY45 are significantly upregulated [52]. After mechanical injury, the expression levels of AtWRKY11, AtWRKY15, AtWRKY22, AtWRKY33, AtWRKY40, AtWRKY53 [53] and AtWRKY6 [54] are upregulated. Similarly, NaWRKY3 is strongly expressed in tobacco. By contrast, the sensitivity of transgenic plants is increased when NaWRKY3 is knocked out [54]. In two previous studies, UV-B radiation treatment induced three WRKY genes in Arabidopsis and the OsWRKY89 gene in rice, resulting in a thick waxy substance on the leaf surface and improved tolerance to heat [55][56].
References
- Yoon, Y.; Seo, D.H.; Shin, H.; Kim, H.J.; Kim, C.M.; Jang, G. The role of stress-responsive transcription factors in modulating abiotic stress tolerance in plants. Agronomy 2020, 10, 788.
- Ma, Q.; Xia, Z.; Cai, Z.; Li, L.; Cheng, Y.; Liu, J.; Nian, H. GmWRKY16 enhances drought and salt tolerance through an ABA-mediated pathway in Arabidopsis thaliana. Front. Plant Sci. 2019, 9, 9.
- Liu, Y.; Yang, T.; Lin, Z.; Guo, B.; Xing, C.; Zhao, L.; Dong, H.; Gao, J.; Xie, Z.; Zhang, S.-L.; et al. A WRKY transcription factor PbrWRKY53 from Pyrus betulaefolia is involved in drought tolerance and AsA accumulation. Plant Biotechnol. J. 2019, 17, 1770–1787.
- Shrestha, A.; Khan, A.; Dey, N. cis-trans Engineering: Advances and perspectives on customized transcriptional regulation in plants. Mol. Plant 2018, 11, 886–898.
- Yamasaki, K.; Kigawa, T.; Inoue, M.; Watanabe, S.; Tateno, M.; Seki, M.; Shinozaki, K.; Yokoyama, S. Structures and evolutionary origins of plant-specific transcription factor DNA-binding domains. Plant Physiol. Biochem. 2008, 46, 394–401.
- Zhang, Y.; Wang, L. The WRKY transcription factor superfamily: Its origin in eukaryotes and expansion in plants. BMC Evol. Biol. 2005, 5, 1.
- Pandey, S.P.; Somssich, I.E. The role of WRKY transcription factors in plant immunity. Plant Physiol. 2009, 150, 1648–1655.
- Ishiguro, S.; Nakamura, K. Characterization of a cDNA encoding a novel DNA-binding protein, SPF1, that recognizes SP8 sequences in the 5′ upstream regions of genes coding for sporamin and beta-amylase from sweet potato. Mol Gen Genet 1994, 244, 563–571.
- Rushton, P.J.; Macdonald, H.; Huttly, A.K.; Lazarus, C.M.; Hooley, R. Members of a new family of DNA-binding proteins bind to a conserved cis-element in the promoters of Amy2 genes. Plant Mol. Biol. 1995, 29, 691–702.
- Rushton, P.J.; Torres, J.T.; Parniske, M.; Wernert, P.; Hahlbrock, K.; Somssich, I.E. Interaction of elicitor-induced DNA-binding proteins with elicitor response elements in the promoters of parsley PR1 genes. EMBO J. 1996, 15, 5690–5700.
- He, H.; Dong, Q.; Shao, Y.; Jiang, H.; Zhu, S.; Cheng, B.; Xiang, Y. Genome-wide survey and characterization of the WRKY gene family in Populus trichocarpa. Plant Cell Rep. 2012, 31, 1199–1217.
- Rensing, S.A.; Lang, D.; Zimmer, A.D.; Terry, A.; Salamov, A.A.; Shapiro, H.; Nishiyama, T.; Perroud, P.-F.; Lindquist, E.; Kamisugi, Y.; et al. The Physcomitrella genome reveals evolutionary insights into the conquest of land by plants. Science 2007, 319, 64–69.
- Mangelsen, E.; Kilian, J.; Berendzen, K.W.; Kolukisaoglu, Ü.; Harter, K.; Jansson, C.; Wanke, D. Phylogenetic and comparative gene expression analysis of barley (Hordeum vulgare) WRKY transcription factor family reveals putatively retained functions between monocots and dicots. BMC Genom. 2008, 9, 194.
- Ling, J.; Jiang, W.; Zhang, Y.; Yu, H.; Mao, Z.; Gu, X.; Huang, S.; Xie, B. Genome-wide analysis of WRKY gene family in Cucumis sativus. BMC Genom. 2011, 12, 1–20.
- Berri, S.; Abbruscato, P.; Faivre-Rampant, O.; Brasileiro, A.C.M.; Fumasoni, I.; Satoh, K.; Kikuchi, S.; Mizzi, L.; Morandini, P.; Pè, M.E.; et al. Characterization of WRKY co-regulatory networks in rice and Arabidopsis. BMC Plant Biol. 2009, 9, 1–22.
- Liu, J.-J.; Ekramoddoullah, A.K. Identification and characterization of the WRKY transcription factor family in Pinus monticola. Genome 2009, 52, 77–88.
- Huang, S.; Gao, Y.; Liu, J.; Peng, X.; Niu, X.; Fei, Z.; Cao, S.; Liu, Y. Genome-wide analysis of WRKY transcription factors in Solanum lycopersicum. Mol. Genet. Genom. 2012, 287, 495–513.
- Ross, C.A.; Liu, Y.; Shen, Q.J. The WRKY gene family in rice (Oryza sativa). J. Integr. Plant Biol. 2007, 49, 827–842.
- Rushton, P.J.; Somssich, I.E.; Ringler, P.; Shen, Q.J. WRKY transcription factors. Trends Plant Sci. 2010, 15, 247–258.
- Ülker, B.; Somssich, E.I. WRKY transcription factors: From DNA binding towards biological function. Curr. Opin. Plant Biol. 2004, 7, 491–498.
- Grierson, C.; Du, J.-S.; Zabala, M.D.T.; Beggs, K.; Smith, C.; Holdsworth, M.; Bevan, M.W. Separate cis sequences and trans factors direct metabolic and developmental regulation of a potato tuber storage protein gene. Plant J. 1994, 5, 815–826.
- Sun, C.; Palmqvist, S.; Olsson, H.; Borén, M.; Ahlandsberg, S.; Jansson, C. A novel WRKY transcription factor, SUSIBA2, participates in sugar signaling in barley by binding to the sugar-responsive elements of the iso1 promoter. Plant Cell 2003, 15, 2076–2092.
- Jiang, Y.; Deyholos, M.K. Functional characterization of Arabidopsis NaCl-inducible WRKY25 and WRKY33 transcription factors in abiotic stresses. Plant Mol. Biol. 2008, 69, 91–105.
- Anjum, S.A.; Xie, X.-Y.; Wang, L.-C.; Saleem, M.F.; Man, C.; Lei, W. Morphological, physiological and biochemical responses of plants to drought stress. Afr. J. Agr. Res. 2011, 6, 2026–2032.
- Sun, Y.; Yu, D. Activated expression of AtWRKY53 negatively regulates drought tolerance by mediating stomatal movement. Plant Cell Rep. 2015, 34, 1295–1306.
- Wang, Z.; Zhu, Y.; Wang, L.; Liu, X.; Liu, Y.; Phillips, J.; Deng, X. A WRKY transcription factor participates in dehydration tolerance in Boea hygrometrica by binding to the W-box elements of the galactinol synthase (BhGolS1) promoter. Planta 2009, 230, 1155–1166.
- Yang, Z.; Chi, X.; Guo, F.; Jin, X.; Luo, H.; Hawar, A.; Chen, Y.; Feng, K.; Wang, B.; Qi, J.; et al. SbWRKY30 enhances the drought tolerance of plants and regulates a drought stress-responsive gene, SbRD19, in sorghum. J. Plant Physiol. 2020, 246, 153142.
- Niu, C.-F.; Wei, W.; Zhou, Q.-Y.; Tian, A.-G.; Hao, Y.-J.; Zhang, W.; Ma, B.; Lin, Q.; Zhang, Z.-B.; Zhang, J.-S.; et al. Wheat WRKY genes TaWRKY2 and TaWRKY19 regulate abiotic stress tolerance in transgenic Arabidopsis plants. Plant Cell Environ. 2012, 35, 1156–1170.
- Sakuma, Y.; Maruyama, K.; Osakabe, Y.; Qin, F.; Seki, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Functional analysis of an Arabidopsis transcription factor, DREB2A, involved in drought-responsive gene expression. Plant Cell 2006, 18, 1292–1309.
- Jiang, Y.; Liang, G.; Yu, D. Activated expression of WRKY57 confers drought tolerance in Arabidopsis. Mol. Plant 2012, 5, 1375–1388.
- Wei, W.; Zhang, Y.; Han, L.; Guan, Z.; Chai, T. A novel WRKY transcriptional factor from Thlaspi caerulescens negatively regulates the osmotic stress tolerance of transgenic tobacco. Plant Cell Rep. 2008, 27, 795–803.
- Ren, X.; Chen, Z.; Liu, Y.; Zhang, H.; Zhang, M.; Liu, Q.; Hong, X.; Zhu, J.-K.; Gong, Z. ABO3, a WRKY transcription factor, mediates plant responses to abscisic acid and drought tolerance in Arabidopsis. Plant J. 2010, 63, 417–429.
- Wang, J.; Wang, L.; Yan, Y.; Zhang, S.; Li, H.; Gao, Z.; Wang, C.; Guo, X. GhWRKY21 regulates ABA-mediated drought tolerance by fine-tuning the expression of GhHAB in cotton. Plant Cell Rep. 2020, 39.
- Sun, J.; Hu, W.; Zhou, R.; Wang, L.; Wang, X.; Wang, Q.; Feng, Z.-J.; Yu, H.; Qiu, D.; He, G.; et al. The Brachypodium distachyon BdWRKY36 gene confers tolerance to drought stress in transgenic tobacco plants. Plant Cell Rep. 2014, 34, 23–35.
- Ahammed, G.J.; Li, X.; Yang, Y.; Liu, C.; Zhou, G.; Wan, H.; Cheng, Y. Tomato WRKY81 acts as a negative regulator for drought tolerance by modulating guard cell H2O2–mediated stomatal closure. Environ. Exp. Bot. 2020, 171, 103960.
- Hassan, S.; Lethin, J.; Blomberg, R.; Mousavi, H.; Aronsson, H. In silico based screening of WRKY genes for identifying functional genes regulated by WRKY under salt stress. Comput. Biol. Chem. 2019, 83, 107131.
- Zhou, Q.-Y.; Tian, A.-G.; Zou, H.-F.; Xie, Z.-M.; Lei, G.; Huang, J.; Wang, C.-M.; Wang, H.-W.; Zhang, J.-S.; Chen, S.-Y. Soybean WRKY-type transcription factor genes, GmWRKY13, GmWRKY21, and GmWRKY54, confer differential tolerance to abiotic stresses in transgenic Arabidopsis plants. Plant Biotechnol. J. 2008, 6, 486–503.
- Gong, X.; Zhang, J.; Hu, J.; Wang, W.; Wu, H.; Zhang, Q.; Liu, J.-H. FcWRKY70, a WRKY protein of Fortunella crassifolia, functions in drought tolerance and modulates putrescine synthesis by regulating arginine decarboxylase gene. Plant Cell Environ. 2015, 38, 2248–2262.
- Qin, Z.; Hou, F.; Li, A.; Dong, S.; Wang, Q.; Zhang, L. Transcriptome-wide identification of WRKY transcription factor and their expression profiles under salt stress in sweetpotato (Ipomoea batatas L.). Plant Biotechnol. Rep. 2020, 14, 599–611.
- Ma, Y.; Xue, H.; Zhang, F.; Jiang, Q.; Yang, S.; Yue, P.; Wang, F.; Zhang, Y.; Li, L.; He, P.; et al. The miR156/SPL module regulates apple salt stress tolerance by activating MdWRKY100 expression. Plant Biotechnol. J. 2020, 18.
- Song, Y.; Li, J.; Sui, Y.; Han, G.; Zhang, Y.; Guo, S.; Sui, N. The sweet sorghum SbWRKY50 is negatively involved in salt response by regulating ion homeostasis. Plant Mol. Biol. 2020, 102, 603–614.
- Zheng, Y.; Ge, J.; Bao, C.; Chang, W.; Liu, J.; Shao, J.; Liu, X.; Su, L.; Pan, L.; Zhou, D.-X. Histone deacetylase HDA9 and WRKY53 transcription factor are mutual antagonists in regulation of plant stress response. Mol. Plant 2020, 13, 598–611.
- Linxiao, W.; Song, A.; Gao, C.; Wang, L.; Wang, Y.; Sun, J.; Jiang, J.; Chen, F.; Chen, S. Chrysanthemum WRKY gene CmWRKY17 negatively regulates salt stress tolerance in transgenic chrysanthemum and Arabidopsis plants. Plant Cell Rep. 2015, 34, 1365–1378.
- Jia, H.; Wang, C.; Wang, F.; Liu, S.; Li, G.; Guo, X. GhWRKY68 reduces resistance to salt and drought in transgenic Nicotiana benthamiana. PLoS ONE 2015, 10, e0120646.
- Liu, X.; Song, Y.; Xing, F.; Wang, N.; Wen, F.; Zhu, C. GhWRKY25, a group I WRKY gene from cotton, confers differential tolerance to abiotic and biotic stresses in transgenic Nicotiana benthamiana. Protoplasma 2015, 253, 1265–1281.
- Wang, M.-Q.; Huang, Q.-X.; Lin, P.; Zeng, Q.-H.; Li, Y.; Liu, Q.-L.; Zhang, L.; Pan, Y.-Z.; Jiang, B.-B.; Zhang, F. The overexpression of a transcription factor gene VbWRKY32 enhances the cold tolerance in Verbena bonariensis. Front. Plant Sci. 2020, 10, 1746.
- Wang, F.; Hou, X.; Tang, J.; Wang, Z.; Wang, S.; Jiang, F.; Li, Y. A novel cold-inducible gene from Pak-choi (Brassica campestris ssp. chinensis), BcWRKY46, enhances the cold, salt and dehydration stress tolerance in transgenic tobacco. Mol. Biol. Rep. 2011, 39, 4553–4564.
- Jaglo-Ottosen, K.R. Arabidopsis CBF1 overexpression induces COR genes and enhances freezing tolerance. Science 1998, 280, 104–106.
- Zou, C.; Jiang, W.; Yu, D. Male gametophyte-specific WRKY34 transcription factor mediates cold sensitivity of mature pollen in Arabidopsis. J. Exp. Bot. 2010, 61, 3901–3914.
- Chen, L.; Zhao, Y.; Xu, S.; Zhang, Z.; Xu, Y.; Zhang, J.; Chong, K. OsMADS57 together with OsTB1 coordinates transcription of its target OsWRKY94 and D14 to switch its organogenesis to defense for cold adaptation in rice. New Phytol. 2018, 218, 219–231.
- Gong, X.-Q.; Hu, J.-B.; Liu, J.-H. Cloning and characterization of FcWRKY40, A WRKY transcription factor from Fortunella crassifolia linked to oxidative stress tolerance. Plant Cell Tissue Organ Cult. PCTOC 2014, 119, 197–210.
- Cheong, Y.H.; Chang, H.-S.; Gupta, R.; Wang, X.; Zhu, T.; Luan, S. Transcriptional profiling reveals novel interactions between wounding, pathogen, abiotic stress, and hormonal responses in Arabidopsis. Plant Physiol. 2002, 129, 661–677.
- Robatzek, S.; Somssich, I.E. A new member of the Arabidopsis WRKY transcription factor family, AtWRKY6, is associated with both senescence- and defence-related processes. Plant J. 2001, 28, 123–133.
- Skibbe, M.; Qu, N.; Galis, I.; Baldwin, I.T. Induced plant defenses in the natural environment: Nicotiana attenuata WRKY3 and WRKY6 coordinate responses to herbivory. Plant Cell 2008, 20, 1984–2000.
- Wang, H.; Hao, J.; Chen, X.; Hao, Z.; Wang, X.; Lou, Y.; Peng, Y.; Guo, Z. Overexpression of rice WRKY89 enhances ultraviolet B tolerance and disease resistance in rice plants. Plant Mol. Biol. 2007, 65, 799–815.
- Kilian, J.; Whitehead, D.; Horak, J.; Wanke, D.; Weinl, S.; Batistic, O.; D’Angelo, C.; Bornberg-Bauer, E.; Kudla, J.; Harter, K. The AtGenExpress global stress expression data set: Protocols, evaluation and model data analysis of UV-B light, drought and cold stress responses. Plant J. 2007, 50, 347–363.