
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Craig J Marshall | + 1195 word(s) | 1195 | 2020-11-04 10:02:12 | | | |
| 2 | Vicky Zhou | Meta information modification | 1195 | 2020-11-16 10:15:18 | | |
Video Upload Options
DNA repair is essential across all organisms to maintain levels genomic stability suitable for life. Archaea often thrive in environmental extremes, enduring levels of heat, pressure, salinity, pH, and radiation that prove intolerable to most life. Many environmental extremes raise the propensity for DNA damaging events and thus, impact DNA stability, placing greater reliance on molecular mechanisms that recognize DNA damage and initiate accurate repair. Archaea can presumably prosper in harsh and DNA-damaging environments in part due to robust DNA repair pathways but surprisingly, no DNA repair pathways unique to Archaea have been described.
1. DNA Repair in Archaeal Species
Aside from the intrinsic instability of DNA, genomes are threatened by a plethora of endogenous and exogenous insults. Left unrepaired, DNA damage increases mutation rates, causing adverse effects on cellular health, with often drastic consequences to cellular and organismal fitness. Endogenous damage has many sources: genomic material can spontaneously undergo base hydrolysis or deaminate, and torsional stresses brought about by information processing systems can bring about genomic instability. Cellular machineries will occasionally incorporate mismatch errors or ribonucleotide monophosphates (rNMPs) into newly synthesized DNA, and many metabolic enzymes produce reactive oxygen species (ROS) which may oxidize DNA bases. Cells must also tolerate exogenous sources of DNA damage which vary depending on the external environment. Chemical crosslinkers, environmentally generated ROS, ultraviolet light, and ionizing radiations, from within or which penetrate the atmosphere, all have mutagenic effects on DNA.
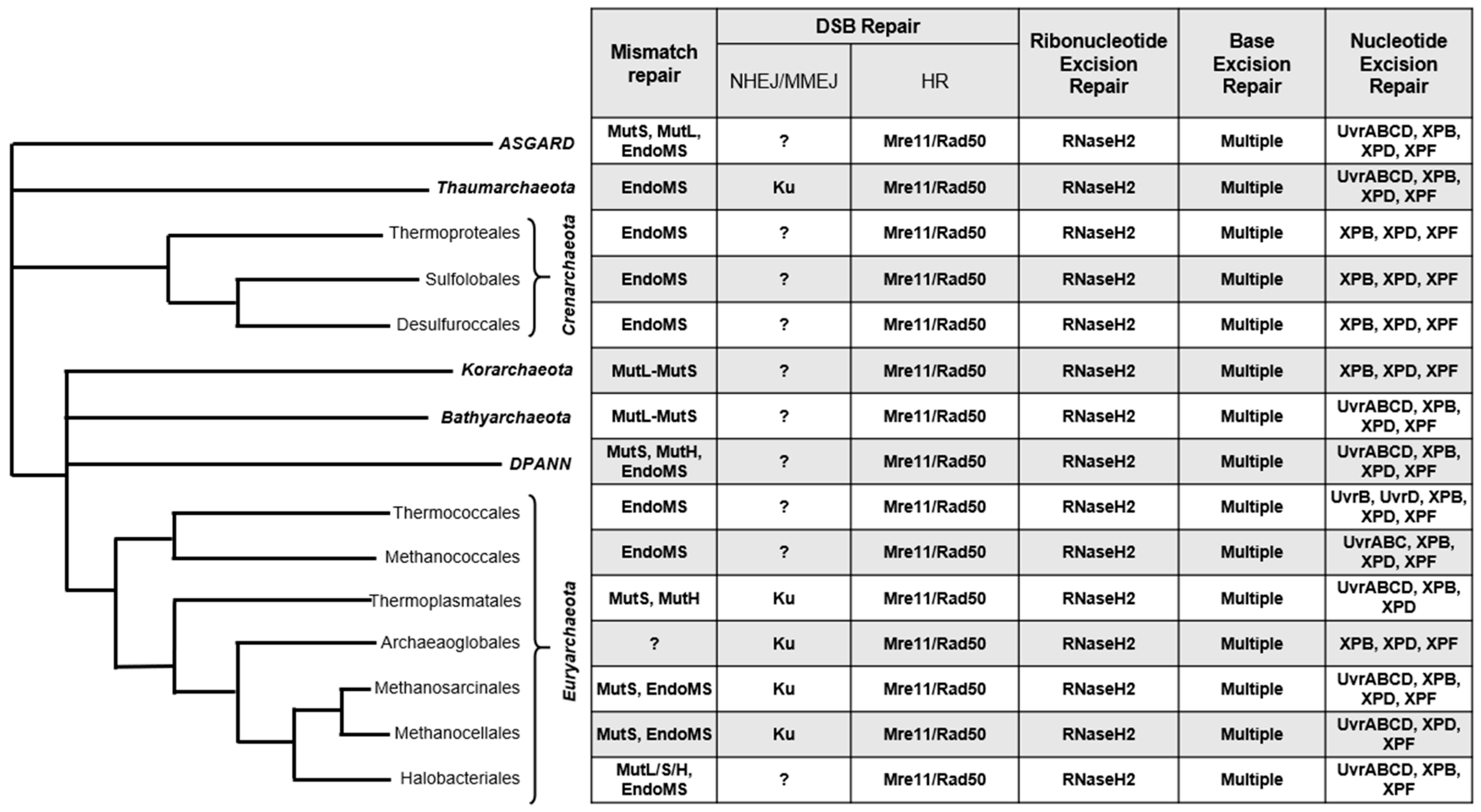
Many archaea thrive within niche and extreme environments which can increase rates of DNA damage. Many halophilic archaea, for example, thrive in shallow salt plains and endure extreme levels of UV radiation [1], while some hyperthermophilic species persist at temperatures that would easily denature purified DNA [2][3], and yet, the presumed increased rates of deamination, depurination, and oxidation are somehow tolerated [4][5][6]. In addition to growth in the extremes, many archaeal species maintain genomic stability levels to display similar rates of spontaneous mutation to mesophilic prokaryotes such as Escherichia coli [7][8][9]. Perhaps surprisingly, no unique DNA repair pathways have been described in Archaea (Figure 1), nor extremophilic Bacteria, i.e., Deinococcus radiodurans. Insight into how Archaea detect and convert damaged DNA bases into repairable substrates has begun to reveal how genomic integrity is preserved in extremis.
A multitude of strategies to identify modified nucleotides or damaged DNA structures (here, collectively termed recognition) and initiate repair is encoded in most genomes, with processes for recognition and repair perhaps best studied within mesophilic bacteria and eukarya. Some DNA damage repair can be directly reversed, i.e., photoreactivation of thymine–thymine dimers by photolyases and repair of methylation adducts by alkyltransferases such as AlkB [11][12]. However, DNA repair more commonly involves pathways which require several specialized enzymes through steps of damage recognition, initiation of repair, and final polymerization/ligation of resynthesized DNA. Collectively, the cycle of recognition-, initiation-, and ligation-based DNA repair (Figure 2) dominates the conserved DNA repair pathways that account for the majority of DNA repair, be it double-strand break (DSB) repair, mismatch repair (MMR), ribonucleotide excision repair (RER), base excision repair (BER), or both global genomic and transcription coupled nucleotide excision repair (GG-NER, TC-NER). The core DNA repair pathways generally consist of recognition factors that more often than not cleave the DNA backbone and or glycosidic linkage to the nucleotide base, a repair DNA polymerase (DNAP) for strand resynthesis, a nuclease (or the exonuclease activity of DNAP) for removal of damaged bases/strands displaced during resynthesis, and DNA ligase to seal nicks generated during repair.
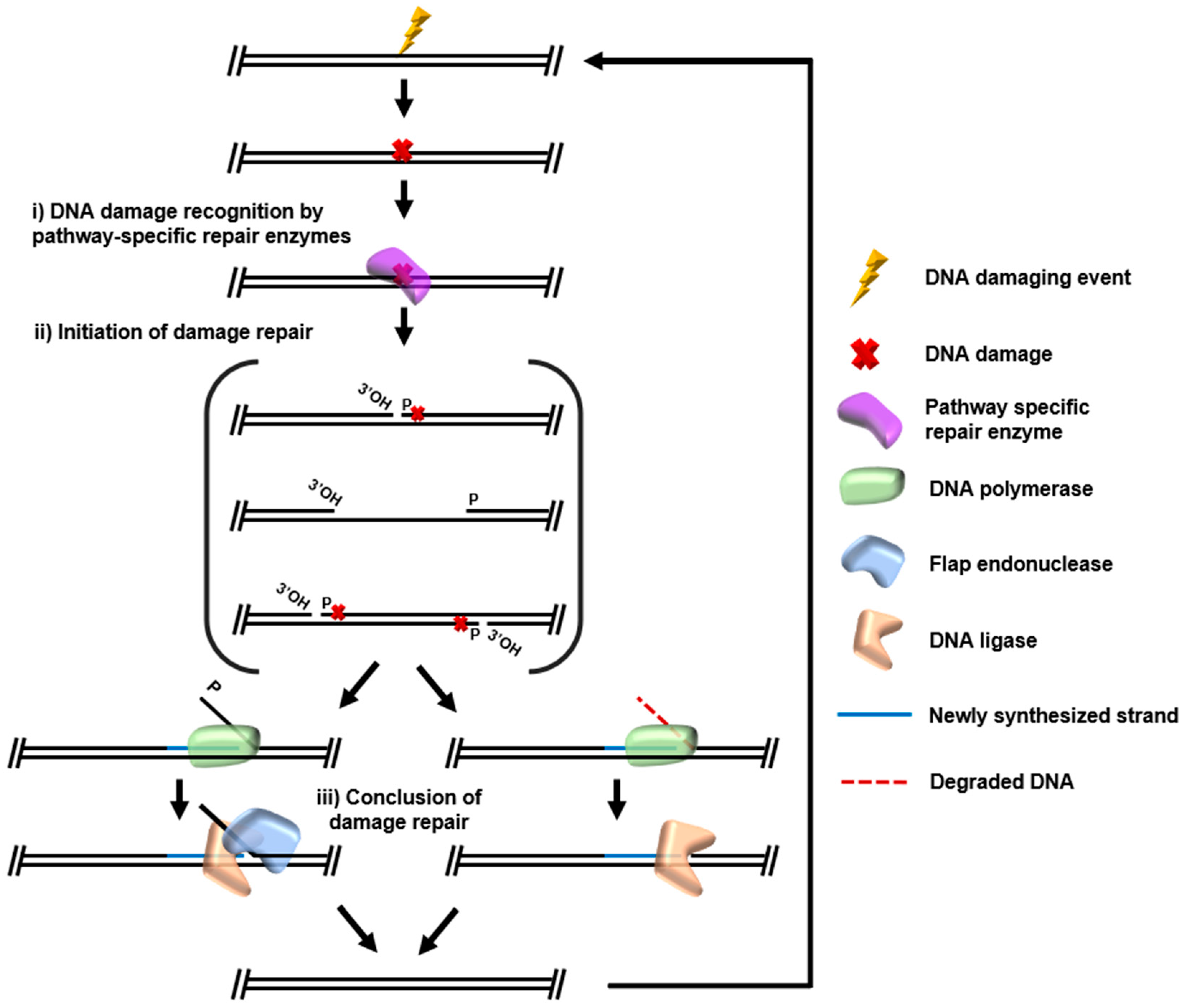
2. Future perspectives on archaeal DNA Repair
Archaeal DNA repair-based research has offered inspiring mechanistic insight into the strategies of preserving DNA stability in extremes once thought inhospitable. Surprisingly, such strategies are not unique and resemble those found in mesophilic Eukarya and Bacteria. How then, do extremophilic species maintain low mutation rates in extreme conditions using the “same tools”? Clues may lie in the apparent crosstalk of archaeal DNA repair pathways or their intimate links with replisome components, or perhaps extremophilic species successfully protect their genomes, avoiding DNA damages to begin with. At face value, strategies of recognition of DNA damages and their preparation for the core resynthesis machinery (i.e., DNAP, Fen1, DNA Ligase) are intrinsically fascinating, but perhaps the most alluring facet of archaeal DNA repair has recently been the development of new techniques at the protein and whole-genome level as archaeal species have become more genetically accessible. Novel archaeal DNA repair enzymes will likely continue to be characterized and find new roles in the exponentially growing biotechnology world. Bioinformatic approaches, such as RADAR-seq, will continue to provide population-/genomic-level DNA repair activities, and the continuously developing knowledge of DSBR in relation to CRISPR systems will surely yield more tools for geneticists. Super-resolution microscopy, once thought over encumbered by the small size of archaeal cells, has recently become optimized and used to image foci of DSB sites in H. volcanii [13], offering the DNA repair research as a platform for development of more broadly applicable procedures. The continued development of these (and new) technologies, however, will only be progressed alongside our understanding of archaeal DNA repair as a whole—and thus, identifying and answering the most pressing questions in the field must remain a priority.
Once thought a detriment to cellular health, the double-strand break is appearing more of an essential intermediate to many metabolic processes outside of replication, potentially altering our view of archaeal metabolic biology. How Archaea deal with such an intermediate has been resolved through multiple pathways (NHEJ, MMEJ, HR), but the next challenge is understanding how cells “decide” which of these pathways is most appropriate in a given context, and if the ploidy state influences rate of HR. One such context may be resultant DSBs from EndoMS activity during archaeal MMR which, if verified, will allow us to probe how cells use DSB substrates purposefully outside of the replisome. Generating depth to our current understanding is of great importance—but there still remain significant “unknowns” in the field which have yet to be resolved. Are there alternative pathways for MMR or BER yet undiscovered—and can recognition enzymes be repurposed? Does NER or the transcription coupled sub-pathway (TCR) exist in Archaea, and do they more closely mirror a prokaryotic or eukaryotic system? Finally, as interconnectedness and crosstalk between repair and replication systems becomes more apparent, how are repair pathways regulated, segregated, or organized in the context of the prokaryotic cell? The answers to these questions will not only provide a clearer picture of DNA maintenance in extremis, but likely hold intriguing insights into our own ancestral metabolic history.
References
- Jones, D.L.; Baxter, B.K. DNA Repair and Photoprotection: Mechanisms of Overcoming Environmental Ultraviolet Radiation Exposure in Halophilic Archaea. Front. Microbiol. 2017, 8, 1882.
- Atomi, H.; Reeve, J.N. Microbe Profile: Thermococcus kodakarensis: The model hyperthermophilic archaeon. Microbiology 2019, 165, 1166–1168.
- Vieille, C.; Zeikus, J.G. Hyperthermophilic Enzymes: Sources, Uses, and Molecular Mechanisms for Thermostability. Microbiol. Mol. Biol. Rev. 2001, 65, 1–43.
- Lindahl, T. Instability and decay of the primary structure of DNA. Nat. Cell Biol. 1993, 362, 709–715.
- Lewis, J.C.A.; Crayle, J.; Zhou, S.; Swanstrom, R.; Wolfenden, R. Cytosine deamination and the precipitous decline of spontaneous mutation during Earth’s history. Proc. Natl. Acad. Sci. USA 2016, 113, 8194–8199.
- Ehrlich, M.; Norris, K.F.; Wang, R.Y.; Kuo, K.C.; Gehrke, C.W. DNA cytosine methylation and heat-induced deamination. Biosci. Rep. 1986, 6, 387–393.
- Ishino, Y.; Narumi, I. DNA repair in hyperthermophilic and hyperradioresistant microorganisms. Curr. Opin. Microbiol. 2015, 25, 103–112.
- Grogan, D.W.; Carver, G.T.; Drake, J.W. Genetic fidelity under harsh conditions: Analysis of spontaneous mutation in the thermoacidophilic archaeon Sulfolobus acidocaldarius. Proc. Natl. Acad. Sci. USA 2001, 98, 7928–7933.
- Palud, A.; Villani, G.; L’Haridon, S.; Querellou, J.; Henneke, G.; Raffin, J.-P. Intrinsic properties of the two replicative DNA polymerases of Pyrococcus abyssi in replicating abasic sites: Possible role in DNA damage tolerance? Mol. Microbiol. 2008, 70, 746–761.
- Baker, B.J.; De Anda, V.; Seitz, K.W.; Dombrowski, N.; Santoro, A.E.; Lloyd, K.G. Diversity, ecology and evolution of Archaea. Nat. Microbiol. 2020, 5, 887–900.
- Yi, C.; He, C. DNA Repair by Reversal of DNA Damage. Cold Spring Harb. Perspect. Biol. 2013, 5, a012575.
- Sancar, A. Mechanisms of DNA Repair by Photolyase and Excision Nuclease (Nobel Lecture). Angew. Chem. Int. Ed. 2016, 55, 8502–8527.
- Delpech, F.; Collien, Y.; Mahou, P.; Beaurepaire, E.; Myllykallio, H.; Lestini, R. Snapshots of archaeal DNA replication and repair in living cells using super-resolution imaging. Nucleic Acids Res. 2018, 46, 10757–10770.





