Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Pero Lučin | -- | 5519 | 2022-09-19 12:40:06 | | | |
| 2 | Conner Chen | -13 word(s) | 5506 | 2022-09-21 08:28:22 | | | | |
| 3 | Conner Chen | -6 word(s) | 5500 | 2022-09-21 08:31:45 | | | | |
| 4 | Conner Chen | + 20 word(s) | 5520 | 2022-09-22 10:28:07 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Lučin, H.M.; Zagorac, G.B.; Marcelić, M.; Lučin, P. Host Cell Signatures within Beta-Herpes Virions. Encyclopedia. Available online: https://encyclopedia.pub/entry/27302 (accessed on 24 May 2026).
Lučin HM, Zagorac GB, Marcelić M, Lučin P. Host Cell Signatures within Beta-Herpes Virions. Encyclopedia. Available at: https://encyclopedia.pub/entry/27302. Accessed May 24, 2026.
Lučin, Hana Mahmutefendić, Gordana Blagojević Zagorac, Marina Marcelić, Pero Lučin. "Host Cell Signatures within Beta-Herpes Virions" Encyclopedia, https://encyclopedia.pub/entry/27302 (accessed May 24, 2026).
Lučin, H.M., Zagorac, G.B., Marcelić, M., & Lučin, P. (2022, September 19). Host Cell Signatures within Beta-Herpes Virions. In Encyclopedia. https://encyclopedia.pub/entry/27302
Lučin, Hana Mahmutefendić, et al. "Host Cell Signatures within Beta-Herpes Virions." Encyclopedia. Web. 19 September, 2022.
Copy Citation
Beta-herpesviruses infect a large proportion of the human population and are associated with a variety of pathophysiological conditions. They are DNA viruses with a large genome that encodes a relatively large number of gene products for the construction of new viral progeny and the establishment of a complex series of interactions with infected cells.
beta-herpesviruses
beta-herpesvirus virions
proteome
1. Host Cell Signatures within Virions
CMV (cytomegalovirus) envelopment, either the membrane wrapping around the virions or the virions budding into membranous organelles, must involve the incorporation of the host cell proteins into the virions. Thus, by analyzing virion composition, people can learn about the origin of the membrane domain used for envelopment and the biochemical requirements for the envelopment process. However, because the virions are released from the cell together with extracellular vesicles (EVs), the analysis of the host cell signatures in the virions faces all the challenges that are strongly debated in the field of EVs. As discussed in a recent review [1], every cell generates various EVs using diverse mechanisms. Most EVs are developed directly at the PM or in the intracellular compartments, mainly multivesicular bodies (MVBs) and LEs, and are released from the cell through their fusion with the PM (Figure 1A). The heterogeneity of EVs is still not well understood and there is no consensus on their biogenesis and molecular composition [1]. EVs can also be released from the cell after autophagosomes have fused with MVB/LEs to form structures known as amphisomes, which deliver autophagocytosed cellular components to lysosomes for degradation but can also fuse with the PM and release the contents outside the cell (Figure 1A). This process, termed secretory autophagy, can contribute cellular contents to the secreted material in addition to increasing the heterogeneity of the extracellular material released. Thus, a major challenge in this field is the optimization of methods to isolate, separate, and characterize the different subpopulations of EVs.
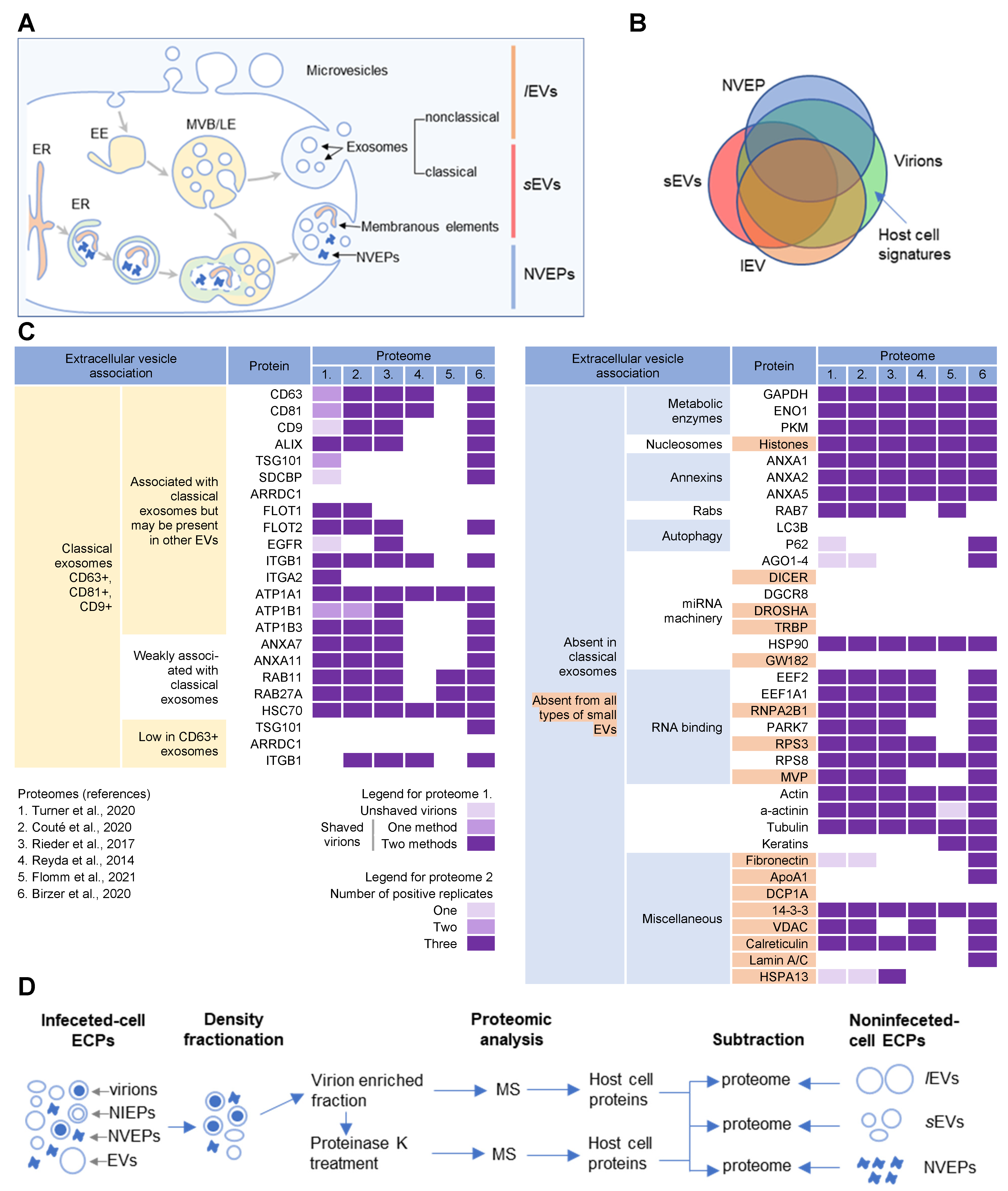
Figure 1. Extracellular nanoparticle content within preparations of beta-herpes virions. (A) Schematic representation of extracellular nanoparticle biogenesis via plasma membrane (PM), exosome, and amphisome-dependent pathways. Microvesicles and non-classical exosomes segregate as large (l) extracellular vesicles (lEVs), whereas classical exosomes segregate as small (s) extracellular vesicles (sEVs). Non-vesicular extracellular particles (NVEPs) and vesicular elements are released by amphisome-dependent and exosome-independent secretion [2][3][4]. (B) Schematic representation of the expected proteomic overlap between lEVs, sEVs, NVEPs, and extracellular virion preparations. (C) Reassessed nanoparticle content [2] identified in human cytomegalovirus (HCMV) and HSV-1 extracellular virion proteomes. Proteins were identified in five proteomes of HCMV virion preparations (proteome 1–5) and one proteome of HSV-1 heavy particles (proteome 6). Proteome 1 [5] presents data for untreated HCMV virion preparations (unshaved) and the same preparation treated with proteinase K (“shaved” virions). Identification of proteins in shaved virions is marked with different color codes depending on whether they were detected by one or two mass spectrometric methods. Proteome 2 [6] displays data in different color codes depending on whether a protein was identified in one, two, or three biological replicates. Proteomes 3–6 (proteomes 3 [7], 4 [8], 5 [9], and 6 [10]) indicate whether a protein is present or not. (D) Workflow for identification of host cell signatures from proteomes of virion preparations. All proteomes [5][6][7][8][9][10] generated by the mass spectrometry (MS) analysis of virion-enriched fractions after density-gradient separation contained many proteins that are present in EVs and NVEPs including one subjected to short proteinase K treatment [5]. Proteins identified in significant amounts in the proteomes of lEVs, sEVs, and NVEPs [2] are subtracted, which are defined as the lEV, sEV, and NVEP filters.
Several recent studies [2][3][4] have taken steps toward increasing the resolution of EVs’ purification and demonstrated that a large amount of secreted material is released from the cell by amphisomes as non-enveloped extracellular particles (NVEPs) that highly overlap in the content with large (l) EVs (lEVs) and small (s) EVs (sEVs) (Figure 1A,B). NVEPs [2] or exomeres [3][4] represent extracellular, non-membranous molecular assemblies released from cells after cytoplasmic components are engulfed in autophagosomes and fused with MVBs to form amphisomes. Most of the released proteins are the components of large cytoplasmic protein complexes, but components of the membranous organelles (ER, endosomes, and Golgi) can also be found.
2. Membrane-Trafficking Cargo Proteins
Cargo proteins are important markers that have been used over the years to learn about endosomal transport. They pass through different compartments within the membrane system and are usually retained in the intracellular localization with low exit rates. Cargo proteins are anchored in the membrane and therefore, when incorporated into the virions, represent the membrane fraction that becomes entrapped in the virions during envelopment, as well as indicate the transport pathway used by the virus for the envelopment process. The cargo proteins identified in the virion preparations were grouped according to the known transport pathways in the endosomal system (Figure 2).
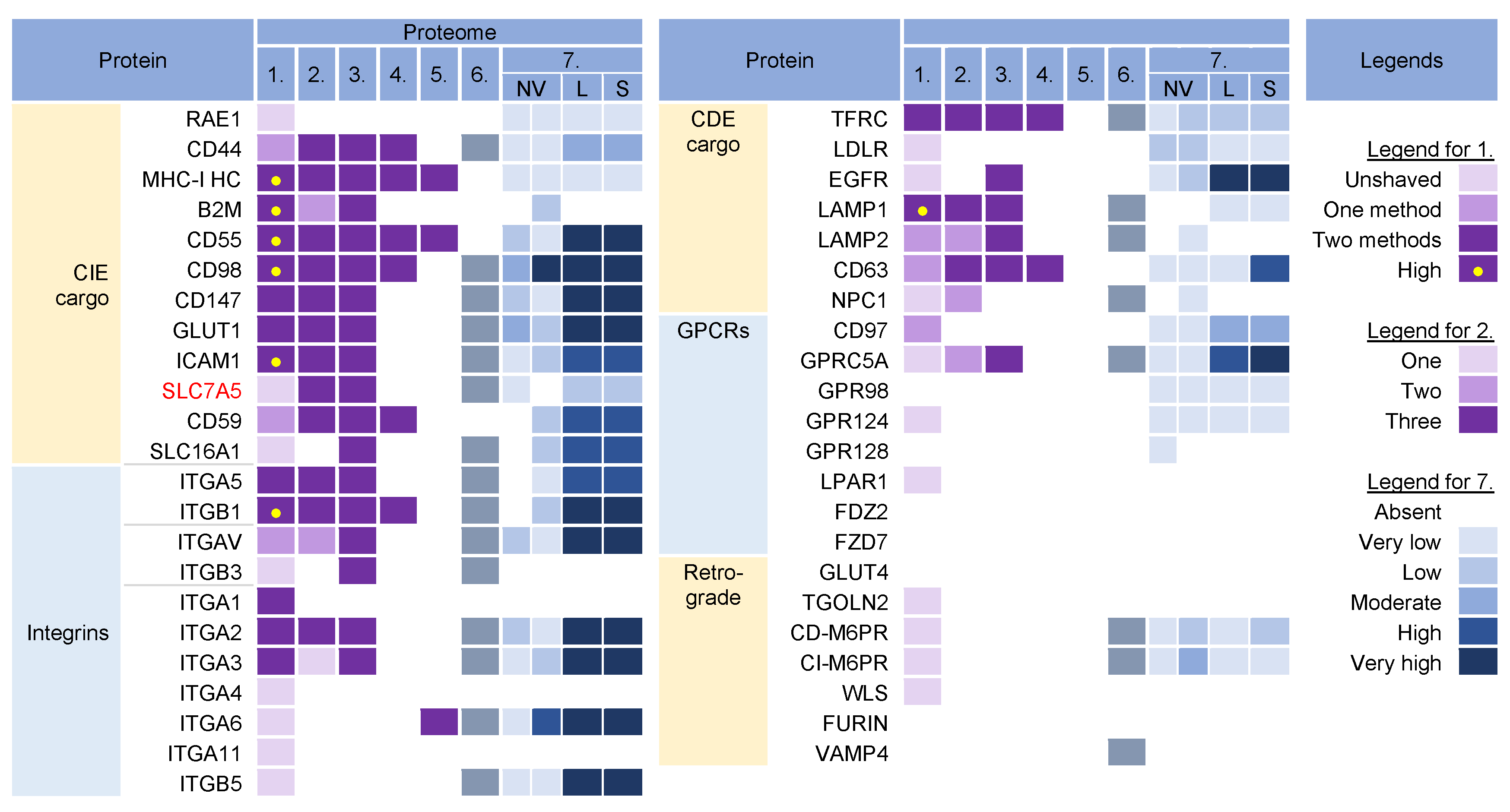
Figure 2. Identification of cargo proteins that travel the endosomal pathways in virion and extracellular nanoparticle preparations. Clathrin-independent endocytosis (CIE) [11][12][13][14] and clathrin-dependent endocytosis (CDE) [15][16][17][18][19][20] cargo proteins, integrins (both CIE and CDE proteins) [21], G protein-coupled receptors (GPCRs) [22], and cargo proteins of the retrograde EE-to-TGN (trans-Golgi network) pathway [16][23] were identified in five proteomes of HCMV virion preparations and one proteome of HSV-1 heavy particles as described in Figure 1. Proteome 7 [2] shows the abundance of a cargo protein in high-resolution density-gradient-purified non-vesicular (NV) samples of DKO-1 cells (left box) and Gli36 cells (right box), lEV (L), and sEV (S) samples of DKO-1 cells. A color code of very low (less than 2 × log2 abundance relative to average signal) was set up as a threshold [2]. Low means 2–3×, moderate 3–4×, high 4–5×, and very high ≥5×.
The virion preparations contained proteins that are internalized both clathrin-independently (CIE) and clathrin-dependently (CDE cargo) (Figure 2). Integrins and G protein-coupled receptors (CPCRs) were grouped separately because they can be internalized by both CIE and CDE mechanisms. All of these proteins have different itineraries for endosomal sorting after endocytic uptake into the early endosomes (EEs) and are returned to the PM by sorting into different recycling routes or diverted to the LEs and lysosomal degradation. A group of cargo proteins is sorted in the EEs and transported to the Golgi, which is called a retrograde route. The pathways of the known recycling and retrograde cargo proteins are shown in Figure 3. The recycling and retrograde pathways in the EEs are pleiotropic [12][24][25][26] and involve the development of tubular extensions at each stage of EE maturation unless the recycling and retrograde cargo is separated from the degrading cargo (Figure 3). Tubular extensions of the EEs generate transport intermediates that carry cargo directly to the PM (recycling) and Golgi (retrograde pathway) or travel to the cluster of pericentriolar tubular compartments known as the endosomal recycling compartment (ERC) for another round of sorting before returning to the PM or being diverted to the Golgi. This pleomorphic tubular traffic is apparently adapted to the type of cargo and final destination and requires the use of a complex array of sorting mechanisms (i.e., sorting nexins and retrieval complexes), a substantial portion of the Rab protein arsenal in cells, and the involvement of Arf GTPases. The major sites of recruitment of the Rab and Arf proteins, sorting nexins (SNXs), and retrieval complexes are shown in Figure 3.
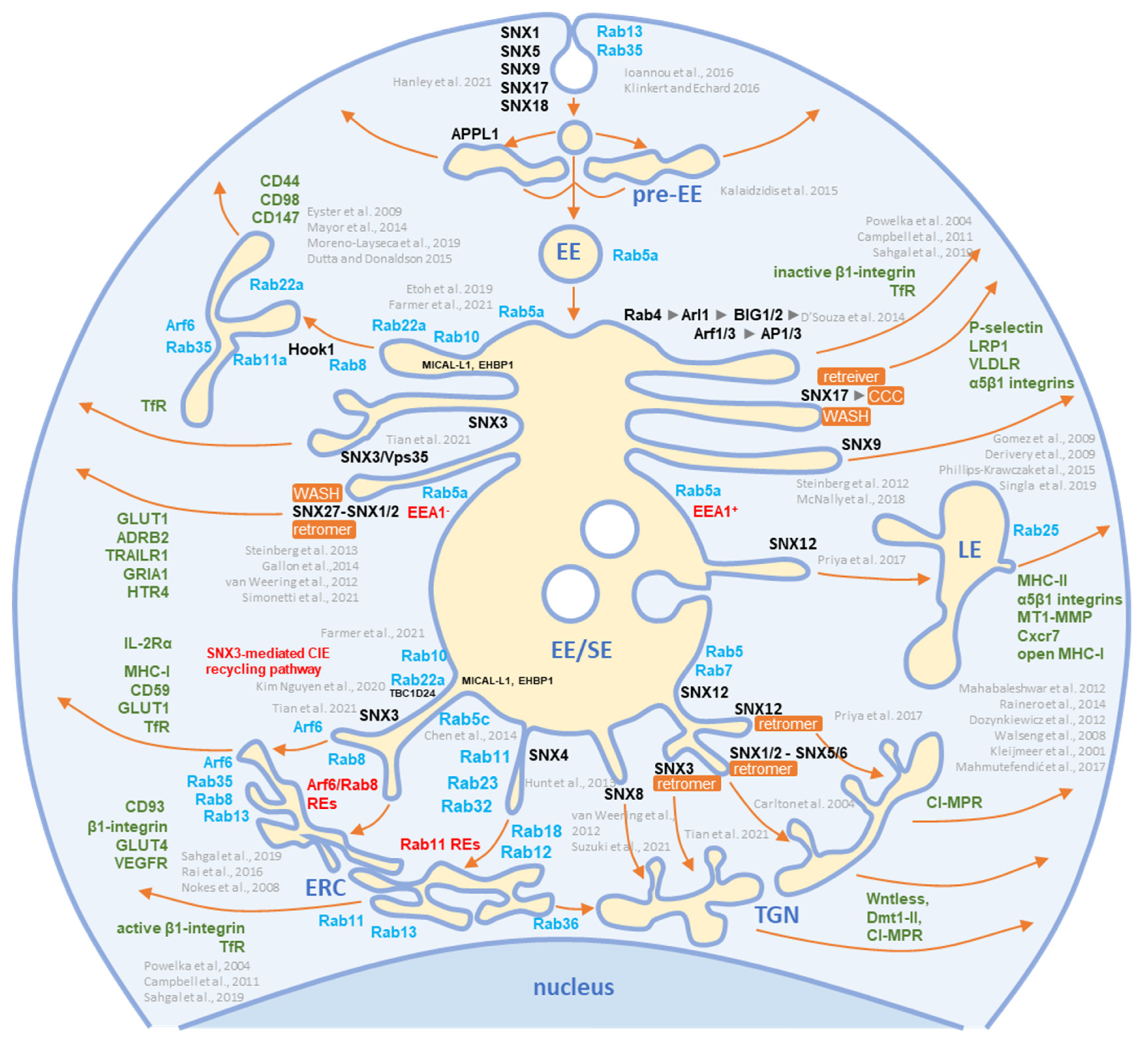
Figure 3. Schematic representation of known exit sites and recycling pathways from the early endosomal system. Cargo internalized by clathrin-dependent endocytosis (CDE) and clathrin-independent endocytosis (CIE) is collected in pre-early endosomes (pre-EEs) and either returned to the plasma membrane (PM) by rapid recycling or delivered to early endosomal (EEs) carriers. EEs undergo a series of fusion and maturation reactions that funnel cargo toward the cell center into enlarged early/sorting endosomes (EE/SE). The maturing EE/SEs sort transported cargo into tubular extensions forming a tubular endosomal network (TEN) that transports cargo directly to the plasma membrane (PM) for recycling, to pericentriolar clusters of tubular endosomes known as the endosomal recycling compartment (ERC) for indirect recycling to the PM, to the trans-Golgi network (TGN) for retrograde transport, or to late endosomes (LEs). EE/SEs also sort cargo by reverse budding into intraluminal vesicles (ILV). Upon completion of the sorting reactions, the remnants of EE /SEs enriched in ILVs become multivesicular endosomes (known as multivesicular bodies, MVBs), which undergo further LE maturation events and either fuse with lysosomes (degradation pathway) or release ILVs as exosomes (exosomal pathway). Cargo sorting into the tubular extension involves the complex and orchestrated recruitment of cellular proteins of the Rab and Arf families, their GEFs and GAPs, members of the sorting nexin (SNX) family, sorting complexes (i.e., retromer, retriever, CCC, and WASH), and other effector proteins required for endosomal tubulation and scission of transport carriers (e.g., dynamins, EHD proteins, AP complexes). The complex cascade of their recruitment is not yet fully elucidated, and the known locations in the EE/SE biogenesis are shown in the schematic. Rab and Arf family proteins are marked in blue, SNXs in black, and cargo molecules in green. Based on references [11][12][14][27][28][29][30][31][32][33][34][35][36][37][38][39][40][41][42][43][44][45][46][47][48][49][50][51][52][53][54][55][56][57][58][59][60][61][62].
The virion preparations were enriched in CIE cargo proteins (Figure 2, proteomes 1–6). Most of these proteins, which are enriched in PM fractions [63][64], were detected to varying degrees in NVEPs and were highly enriched in lEVs and sEVs (Figure 2, proteome 7). After internalization from the PM, the CIE proteins are collected together with the CDE proteins in the Rab5-positive EEs but at this stage, they are sorted into two main pathways [11][12][34]. One subset (CD44, CD98, and CD147) migrates directly from Rab5 endosomes into Rab22-dependent endosomal recycling tubules and does not reach the EEA1-positive compartments. The other subset (MHC-I, CD55, and CD59) returns to the PM indirectly via EEA1-positive endosomes or is sorted into lysosomes for degradation. These proteins are retained in the endosomes of HCMV- [65] and MCMV- [66][67] infected cells. Since MHC-I proteins were not enriched in NVEPs, lSVs, and sEVs (Figure 2, proteome 7), they can be considered as a cargo protein signature embedded in the virions. Their presence suggests that tubular membranes derived from EEs that recycle CIE cargo are used for envelopment.
The virion preparations were also rich in integrins (Figure 2, proteomes 1–6), which can use CIE and the CDE route for entry and may use multiple routes for recycling [22]. However, integrins were also highly enriched in lEVs and sEVs (Figure 2, proteome 7), and therefore cannot be considered a reliable cell signature within the virions.
The virion preparations were enriched in transferrin receptor (TFRC), a well-known CDE cargo; however, TFRC was also enriched in EVs and NVEPs (Figure 2, proteome 7) and cannot be considered a signature. The virion preparations, but not EVs and NVEPs, were also enriched in Lamp1 (Figure 2), a known LE protein that traverses the endosomal system via the CDE route and passes through the EEs/SEs [15]. The presence of Lamp1 suggests a detour of EE flux, as none of the other LE proteins could be detected in the virions. This is consistent with the observations that CMVs congregate in the endosomal compartments traversed by the CIE and CDE cargo, resulting in their retention in the assembly compartment (AC) [65].
The virion preparations were not enriched in the CDE cargo proteins that take the EE route to LEs/lysosomes (such as CD63 [19], LDLR, and EGFR [15]), traffic between the EE and TGN (such as CD63, [19]), and circulate between the EEs, LEs, and TGN via the retrograde recycling route (such as TGOLN2 [TGN38/42], GLUT4, M6PRs, WLS [Wntless], Furin, or Vamp4), or circulate at the interface of EE and LEs (such as NPC1) (Figure 2). The absence of these cargo proteins is consistent with the absence of retrieval machinery and suggests that the retrieval domains in EE are not used for envelopment. The virion preparations also lacked G protein-coupled receptors (GPCRs) (Figure 2), which are retrieved by SNX27 into tubular endosomes and returned to the PM in cooperation with the retromer and the WASH complex (Figure 3). This, together with the absence of SNX27 in the virions, suggests that the SNX27-dependent recycling domain does not contribute to envelopment. Thus, CIE and CDE cargo transport in the EEs of CMV-infected cells is arrested after the sorting of degradative and retrograde cargo, and envelopment likely occurs at the membranes traversed by the CIE cargo proteins.
3. Rab GTPases
Rab proteins are small GTPases that are master regulators of membrane flux in eukaryotic cells [68]. Similar to other small GTPases, they are activated by guanine nucleotide exchange factors (GEFs) to bind GTP (GTP-bound form) and inactivated by GTPase-activating proteins (GAPs) that facilitate the conversion of GTP to GDP (GDP-bound form). Activated Rab proteins are prenylated and inserted into the membrane, and inactivated Rab proteins are extracted from the membranes by GDP dissociation inhibitor proteins (GDI) and stored in the cytosol. The cycles of activation, followed by insertion into membranes, and inactivation, followed by extraction, determine membrane identity. More than 60 Rab proteins, together with seven phosphoinositides, create the navigation tags that control the recruitment of the effector proteins to the membranes, regulating biogenesis, transport, tethering, and the fusion of the membrane organelles. Membrane flux can be viewed as a sequential wave of Rab recruitment and de-recruitment that directs membrane flux in different directions within the cell. Therefore, Rab signatures in CMV virions should shed light on the history of membranes used for envelopment and their biochemical properties in the context of the known functions of Rab proteins in the membrane system.
Rab proteins are highly recruited to the AC area of CMV-infected cells [69], indicating a highly polymorphic membrane composition within the AC. This area is huge and may harbor thousands of membrane-like units in fibroblasts [69], as electron microscopy (EM) imaging showed that most membrane units are 50–200 nm in diameter [9][70][71][72][73][74]. Although several Rab proteins were expected in the virion preparations, the number of identified Rab proteins was surprisingly high, and at least 16 Rab proteins were detected in the virion preparations treated with proteinase K (Figure 4) Most of these Rab proteins were associated with the tubular domains of the REs, ERC, TGN, and ERGIC. Their presence in the virions would indicate that the virions are enveloped by a complex multidomain membrane structure. However, most of these Rab proteins were also enriched in late endosomes (LEs), sEVs, and even NVEPs (Figure 4, proteome 7). Although their enrichment in vesicular and non-vesicular extracellular components does not preclude their incorporation into the virions, most of these Rab proteins cannot be considered a reliable signature within the virions. Among the 16 Rabs detected in the proteinase K-treated virion preparations, Rabs 12, 18, 32, and 23 were either not detected or were present in low amounts in lEVs, sEVs, and NVEPs (Figure 4, proteome 7). Three of them (Rabs 12, 18, and 23) were consistently reported in other proteomic studies of HCMV virions (Figure 4, proteomes 2–5). Thus, these Rabs have arisen as the likely signature that may reveal the identity of the envelopment membrane.
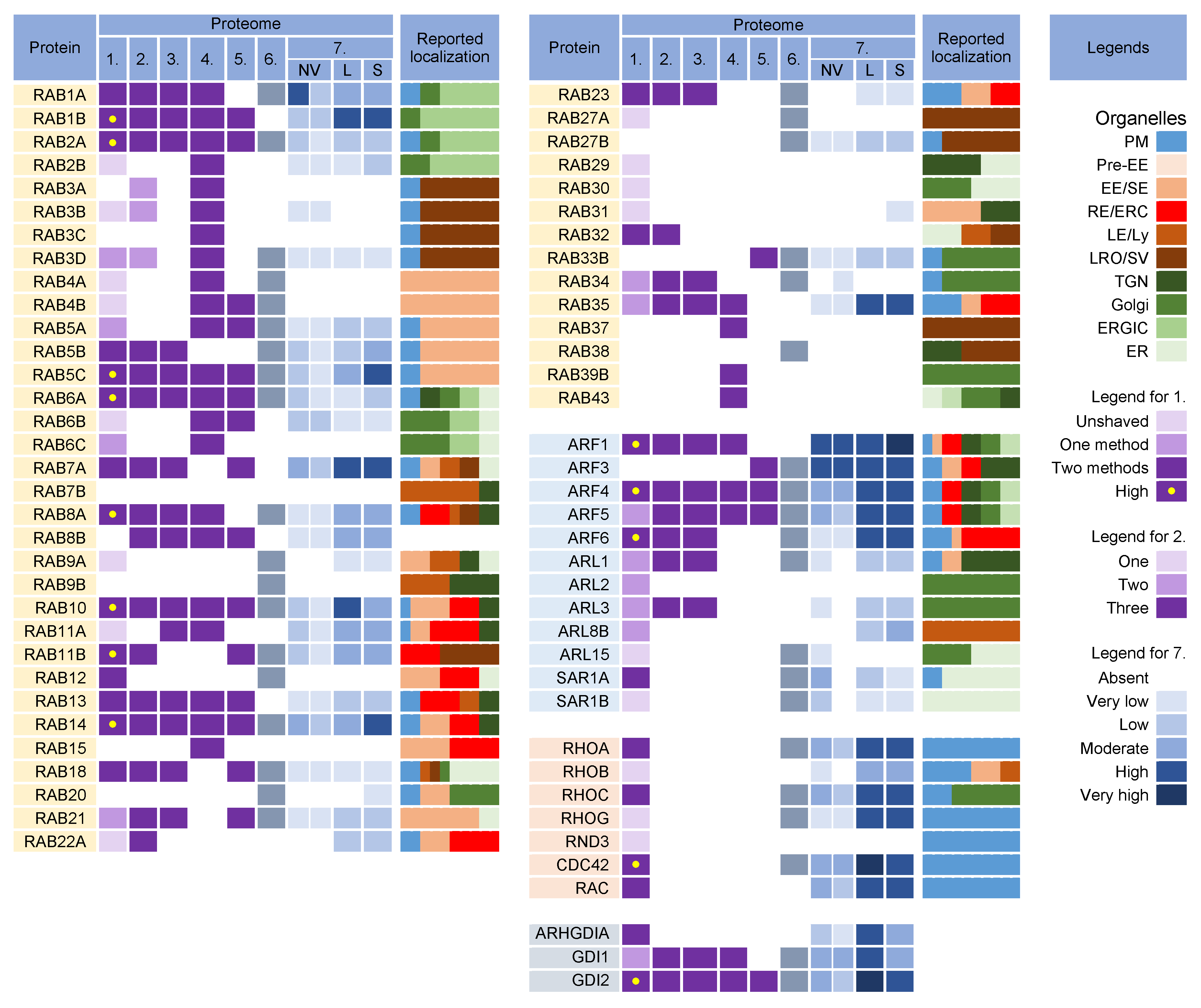
Figure 4. Membrane-trafficking-associated small GTPases in virion and extracellular nanoparticle preparations. Small GTPases were identified in proteomes of virion preparations (HCMV proteomes 1–5 and HSV-1 proteome 6), NVEPs (NV), lEVs (L), and sEVs (S) (proteome 7). The reported localization in membrane organelles of non-infected cells is shown by a color code to illustrate the preferred localization [17][27][28][37][63][64][67][68][74][75][76][77][78][79][80][81][82][83][84][85][86][87][88][89][90][91][92][93][94][95][96][97][98][99][100][101][102][103][104][105]. Proteomes used for analysis and legends are described in Figure 2.
3.1. Rab12, Rab18, Rab23, and Rab32
Rab12 localizes to the EEs and ERC and controls the transport of specific cargo, such as TfR, from the ERC to lysosomes [100] and stimulates autophagy [106], autophagosome trafficking [107], and retrograde transport from the ERC to TGN [108]. It has been suggested that Rab12 facilitates the direct or indirect fusion of RE/ERC-derived compartments with lysosomes [100] and links autophagosomes to motor proteins to facilitate autophagosome trafficking [107].
Rab18 has been localized in the ER and endosomes, and its activity is associated with several processes including autophagy, secretion, and lipid droplet biogenesis (reviewed in [102]). However, its contribution to these processes has not been well characterized, and existing evidence shows that it plays an important role in tethering to the ER and autophagy. Together with Rab10, it regulates ER tubulation, with Rab10 required for ER tubule expansion and Rab18 contributing to the tethering and fusion [109]. The best-established interaction partner of Rab18 is its GEF Rab3GAP [110], which also interacts with ER and the ERGIC protein ERGIC-53 [111], allowing its activation and binding to autophagosome membranes [112]. In addition to the ER, Rab18 is recruited to secretory granules [113], synaptic vesicles [114], and lysosomes [115] in neurons and neuroendocrine cells.
Rab23 has been found in tubular endosomes [35], colocalized with Rab5, and internalized Tf in later stages of TfR trafficking in the early recycling compartment, but not in early stages of the EE pathway, LEs, and Golgi [80]. It also contributes to autophagosome development [116].
Rab32 is localized in the EE /REs, mitochondria, ER, and lysosomes [83][84]. Its activity is associated with SNX6/retromer trafficking, trafficking from the EE/REs to melanosomes, the recycling of VAMP7 from melanosomes to early/recycling endosomes, and the regulation of lysosome-related organelles [84]. It is also localized to the ER and supports autophagic membrane formation and autophagy under basal, nutrient-rich conditions [117].
Overall, all four Rab proteins that pass through the NVEP, lEV, and sEV filters are associated with membrane system tubulation, ERC function, and autophagy. Their presence in the virions suggests that envelopment may occur on the tubular membranes derived from the CIE recycling pathway, consistent with the identification of the CIE cargo proteins in the virions (Figure 2). It is likely that these membranes belong to the downstream segments of the CIE recycling pathway, as the virions did not contain appreciable amounts of Rabs 22a, 35, and 36 (Figure 4), three GTPases reported to act at a proximal stage of maturation of the CIE recycling pathway (Figure 3) [118]. Rabs 12, 23, and 32 are involved in the exit pathways from the recycling endosomes distinct from the Rab8a/b pathway [35][80][84][100]. However, their role in the endosomal recycling circuit is insufficiently studied, and it is not possible to determine the functional sequence between them or to assign the effector functions associated with their recruitment. Interestingly, all Rab proteins that pass the NVEP filter are associated with autophagosomes, suggesting that the functions acquired by their recruitment may be important for the maturation of autophagosomes, including the development of phagophores. Thus, it is possible that these functions are redundantly activated by the dysregulated recruitment of a “package” of Rab proteins in the downstream tubular compartments of the modified (rearranged) tubular compartments of the inner AC. These membranes can be expanded and used for envelopment.
3.2. Rab Interactors and Effectors
Nine Rab proteins (Rabs 1a, 1b, 2a, 5c, 6a, 7a, 10, 11b, and 14) were consistently detected in all the CMV virion preparations (Figure 4, proteomes 1–5). However, these proteins did not pass the lEV, sEV, and NVEP filters (Figure 4, proteome 7), which does not allow conclusions about their incorporation into the virions. Given that these Rabs act as the master regulators of membrane flux and are highly recruited to the membranes of the AC [69], this group also analyzed the presence of their interactors and effectors in the virion preparations.
Rabs 1a, 1b, and 2a are key organizers of the intermediate compartment (IC) [77], a vesiculo-tubular cluster in the secretory pathway between the ER and cis-Golgi that forms several pericentrosomal and peripheral subcompartments located in close proximity to the highly curved tubular-vesicular membrane domains of the ER, known as ER Exit Sites (ERES) [77]. The pericentrosomal IC is often positioned as a linker compartment of the Golgi ribbon with spatiotemporal dynamics that overlap with the ERC [119][120]. This topology and tubular phenotype make the IC a suitable structure for CMV envelopment. The virion preparations are also rich in several components of COPI complexes, which are recruited to the IC by both Rab1 and Rab2 [77] but are also abundant in lEVs and NVEPs (proteome 7). Since almost all tethering proteins of IC and other effector proteins of Rab1 and Rab2 [77] are absent in the virions, there is no clue to suggest that IC-derived membranes are used for envelopment.
Rab6a acts at the TGN, where it provides docking of endosome-derived vesicles [121]. It recruits the Golgin tethering factors and nucleates the tethering complexes required for the fusion of the arriving endosome-derived vesicles. Rab6 also regulates intra-Golgi trafficking and mediates retrograde transport from the cis-Golgi to ERGIC or ER via tubular carriers, known as the COPI-independent retrograde pathway [122]. The targeting and fusion of these retrograde carriers are mediated by active Rab18 at the ER membrane through interaction with ER-localized tethering factors [102][110][123]. Rab6a is also involved in the regulation of autophagy [124]. The trapping of Rab6 by enveloping the virions may suggest that envelopment also occurs at the Golgi-tract-derived membranes that form the inner (TGN) and outer (Golgi medial and cis- and trans-Golgi stacks) AC area of the CMV-infected cell [69][125]. However, none of the Golgin tethering factors and components of the COG and GARP tethering complexes and Rab6a interactors were found in the virions, suggesting that Rab6-positive TGN-derived membranes do not contribute to envelopment. The exception is the presence of BICD2, a known Rab6a effector at the Golgi–ERGIC–ER interface.
Rab7a, a key component of the LE pathway, controls transport to LEs and lysosomes and lysosomal biogenesis, positioning, and functions [126]. It is also localized to the ER and modulates ER morphology by controlling ER homeostasis and ER stress [127]. Rab7 is also found on autophagosomes [116]. On endosomes and lysosomes, Rab7a interacts with several effectors, but none of these proteins were found in the virions, suggesting that the envelopment does not occur at the membranes of endosomal or lysosomal origin.
Rab11b, a member of the Rab11 family, was highly enriched in the virion preparations. It recruits mainly to the membranes of the ERC and contributes to the control of endosomal recycling and the cell surface proteome [96]. It can also be localized in the EEs, TGN, and post-Golgi vesicles. Rab11b, like Rab11a, contributes to the control of the recycling of the CDE and CIE cargo proteins [11] but may act in different ways [128] and have different localization [129]. Rab11b can be recruited to peripheral lysosomes and regulate lysosome exocytosis [98]. However, none of the major regulatory components of lysosome exocytosis (Rab3a, Sec15, and GRAB) [98] were found in the virions.
Rab10 has a variety of functions and subcellular localizations and is involved in various activities in the ERC, Golgi/ TGN and endosomes [130], ER tubulation [131], autophagosome biogenesis, and autophagic flux [132]. Thus, the capture of Rab10 to the enveloping virions may be associated with any tubular compartment, as Rab10 contributes to the initiation of tubulation. The multiple functions of Rab10 have been linked to the recruitment of numerous effectors, but none of these proteins have been found in the virions.
Rab5c, similar to Rab5a and 5b isoforms, is an EE Rab essential for the cell survival and maintenance of EEs and LEs [133]. Rab5c can act semi-independently of the other Rab5 isoforms [134], can be recruited to the Arf6/integrin recycling pathway [135], and drives CD93 and active β1-integrin recycling [136]. Consistent with the capture of CIE cargo and several Rab proteins acting in the endosomal recycling pathway, Rab5c can also be captured by the enveloping virions in this pathway.
In addition to the known effectors and interactors, an important sign for Rab protein recruitment may be the identification of specific GEF and GAP proteins, which should remain as a signature. However, none of the GEF and GAP proteins reviewed by Müler and Goody [105] were found in the virions.
3.3. Why Were There So Many Rabs in the Virion Preparations?
The detection of nearly half of the Rab protein repertoire in the virion preparations may be related to their membrane-bound forms within heterogeneous EVs that copurify with the virions. Because of the physiology of the intracellular Rab cycling, Rab proteins could also be collected and detected in nonmembrane-bound forms either in EVs or NVEPs as adjacent cytosolic Rabs associated with chaperone proteins. The switch between the membrane and cytosol is an important mechanism for the regulation of Rabs [105][137]. After inactivation (conversion to a GDP-bound form), many Rabs are extracted from the membrane by GDI proteins and remain bound to the GDI near the membrane to await the next round of activation. Therefore, many Rab proteins detected in the virion preparations may be those associated with GDIs rather than membranes. Indeed, GDI2 was highly enriched in the virion preparations, as well as in lEVs, sEVs, and NVEPs (Figure 4).
3.4. What Did We Learn from Rab Analysis?
Analysis of the Rab proteins in the virions could not answer the question of the origin and identity of the envelopment organelle. Most of the identified Rab proteins passing the lEV, sEV, and NVEP filters (Rabs 12, 23, and 32) are associated with tubular REs harboring the CIE recycling cargo, consistent with the identification of CIE recycling cargo proteins in the virions. These Rabs suggest that the envelopment organelle develops from membranes derived from REs, consistent with the abundance of RE domains within the inner AC [69][138][139]. However, the identification of other Rab proteins that do not pass the lEV, sEV, and NVEP filters should not be neglected. Most of these Rabs are also associated with REs (Rabs 5c, 10, 11b, and 14) but some of them are also associated with tubulation at the ER and ERGIC (Rabs 6a, 7a, 10, and 18) and might suggest that these tubular domains are used for the establishment of the envelopment organelle. The bulk of the ER and ERGIC membranes are dislocated outside the inner AC [69][138][140][141], but there is insufficient information on the presence of ER- and ERGIC-derived membranes in the inner AC. The identified Rab proteins do not rule out a contribution from the TGN tubular domains and weakly support the possibility that the envelopment organelle could derive from the vacuolar EE domains abundant in the inner AC [69][139][140][142][143][144] or from LE domains, which are extruded from the inner AC [69][139][140][141].
All of the Rabs found in the virions are associated with autophagosome biogenesis and maturation, suggesting that autophagosomal membranes can be used for envelopment. However, as discussed later, none of the key autophagic factors were identified in the virions.
4. Arf GTPases
The Arf superfamily includes 6 Arfs, 22 Arls, and 2 Sars that control a wide range of cellular functions [89]. All members of the Arf subfamily were found in the virion preparations (Figure 4, proteomes 1–6). However, Arfs 1, 3, 4, and 5 were substantially present in NVEPs, lEVs, and sEVs, whereas Arf6 was highly enriched in lEVs and sEVs (Figure 4, proteome 7). Only five members of the Arl subfamily were detected in the viral proteomes (Figure 4, proteomes 1–6) but none of them were detected in convincing amounts after proteinase K treatment (Figure 4, proteome 1). Both members of the Sar subfamily were detected in the HCMV preparations and Sar1a was present in convincing amounts (Figure 4, proteome 1) but did not pass the NVEP and lEV filters (Figure 4, proteome 7).
Unfortunately, the incorporation of Arf proteins into the virions is not a suitable signature. All Arf proteins can be activated at different sites within the membrane system (Figure 4, [88][89][90][93]) and their recruitment into the virions can indicate any membrane structure within the system. In addition, Arf proteins are highly entrapped in EVs and NVEPs, which may complicate their detection in the virions. Nonetheless, Arf proteins appear to be of prominent importance in the biogenesis of the AC and possibly in the secondary envelopment of beta-herpesviruses and virion egress. All Arf proteins have been shown to be highly recruited to membranes within the AC of MCMV-infected cells: Arf3 at the outer and all Arfs at the inner AC membranes [69][145]. Class I (Arfs 1 and 3) and class II (Arfs 4 and 5) Arfs are also overexpressed in the early phase of MCMV infection and remain elevated in the later stages during virion assembly [145]. The suppression of Arf1, Arf3, Arf4, or Arf6 functions by siRNA prevented the establishment of the pre-AC in MCMV-infected cells [145], suggesting their significant contribution to membrane organelle remodeling. However, the suppression of Arf1 and Arf6 also inhibited the establishment of infection [145], suggesting that these GTPases are also involved in the earliest stages of infection.
Several studies on HCMV- [65][144][146] and MCMV- [66][67][69][145][147] infected cells have shown that the CIE and CDE cargo are retained inside the AC and their recycling is inhibited. These observations support the identification of the CDE and CIE cargo proteins within the virions Figure 2. The inhibition of the cargo exit from the endosomes of CMV-infected cells has been associated with the alteration of Arf6 function and may provide the basis for the envelopment organelle formation. Immunofluorescence studies of HCMV [65] and MCMV [66][69][145][147] showed excessive recruitment of Arf6 to the membrane units within the inner AC, beginning early in infection. This region of the AC is the site of the secondary envelopment of CMV and is composed mainly of EE -, ERC-, and TGN-derived elements. The HCMV study [65] demonstrated that the CIE cargo is stacked in enlarged Arf6 endosomes that retain Arf6 in the GDP-bound form.
Arf6 acts mainly on tubular endosomes that mediate endosomal recycling [148] and contributes to the regulation of autophagy [148][149][150]. These tubular endosomes, known as the tubular endosomal network (TEN) [35][36][151], recycle a subset of CIE cargo proteins to the PM (i.e., CD44, CD98, and CD147) [11][12]. The entire process requires the sequential and orchestrated activation of Rab5, Rab10, Rab22a, Rab35, and Rab11, culminating in the activation of Arf6 (ARF6:GTP) and its hydrolysis to Arf6:GDP before the recycling carriers are released toward the PM [11][12][34]. Another subset of cargo proteins (i.e., MHCI, CD55, and CD59) moves downstream in the EE tract, accumulates in EEA1-positive vacuolar endosomes, and is sorted into the tubular extension to be packaged into transport carriers [12][34][152]. Some of these carriers are returned to the PM via the TEN, whereas others are transported to the juxtanuclear cluster of tubular endosomes that form the ERC [12][14]. Transport to the ERC requires Rab11 activity and the CIE cargo proteins are sorted into Rab8/Arf6/Rab35 intermediates [153]. At the Rab8/Arf6/Rab35 endosomes, Rab35 can recruit multiple Rabs (i.e., Rabs 8, 13, 36) that move on in different directions [86]. Rab8 carriers with recruited Arf6 turn off Rab35 [86][154] and migrate to the PM to ensure the recycling of the CIE cargo from the ERC [86]. It is unclear to what extent the branching can occur at this stage. It is known that Rab36 is required for the delivery of carriers with cargo to the TGN [86], but also that Rab12 [100], Rab23 [35][80], and Rab32 [84] may be recruited and can contribute to different trafficking routes. Thus, multiple recycling paths may be associated with the Arf6-dependent recycling route, and there is still a long way to go to fully understand how the recycling cargo is sorted at this point [38]. In the end, Arf6:GTP hydrolysis is required for recycling and if Arf6 is not inactivated, the endosomal carriers do not reach the point of further progression, the cargo is not recycled, and the endosomal compartment enlarges and retains the endocytic cargo [11][12], as occurs in CMV-infected cells.
5. Rho Family GTPases
Some virion preparations were also enriched in RhoA, Rac1, and Cdc42 (Figure 4), the three canonical members of the Rho family GTPases, which consists of 22 genes encoding at least 25 proteins [101]. The Rho family is involved in all cellular processes that depend on the organization of the cytoskeleton and plays a role in the regulation of vesicle transport and endocytosis. However, they were also highly enriched in lEVs, sEVs, and NVEPs (Figure 4, proteome 7) and, therefore, cannot be identified as host cell signatures.
6. Adaptor Protein Complexes for Cargo Sorting
The formation of coated vesicles for transport between the membrane compartments is orchestrated by heterotetrameric adaptor protein complexes (AP) [155][156]. They recruit cargo proteins by binding to their cytoplasmic tails and facilitate the formation of clathrin-coated vesicles on membranes. They are activated by cargo and clathrin recruitment and by membrane-associated proteins such as Arf1 and FCH domain-only (FCHo) proteins [157]. Adaptor protein 1 (AP1) complexes can be activated at the TGN, EE, LE, and RE/ERC and can mediate the transport of cargo between the TGN and LE in both directions, from the TGN to the PM, and from the ERC/RE to the PM (recycling). AP2 complexes are associated with endocytic activities at the PM, AP3 likely mediates transport from the TGN to LEs and from LEs to lysosomes, AP4 is involved in transport from the TGN to EEs, and AP5 complexes regulate transport from LEs [155].
Almost all components of AP1 and AP2 complexes were found in the virion preparations (Figure 5, proteomes 1–6) and AP2 components were retained after the proteinase K treatment (Figure 5A, proteome 1). However, the components of AP1 and AP2 complexes were highly enriched in lEVs, sEVs, and NVEPs (Figure 5A, proteome 7). Similarly, clathrin heavy chains and dynamin 2 were highly enriched in most of the virion preparations [5][6][7] but also in lEVs, sEVs, and NVEPs [2] (data not shown). Nevertheless, the AP complexes, clathrin, and dynamin may play an important role in CMV assembly as they are enriched at the membranes of the inner AC of MCMV- [69][158] and HCMV- [70][139][159] infected cells.
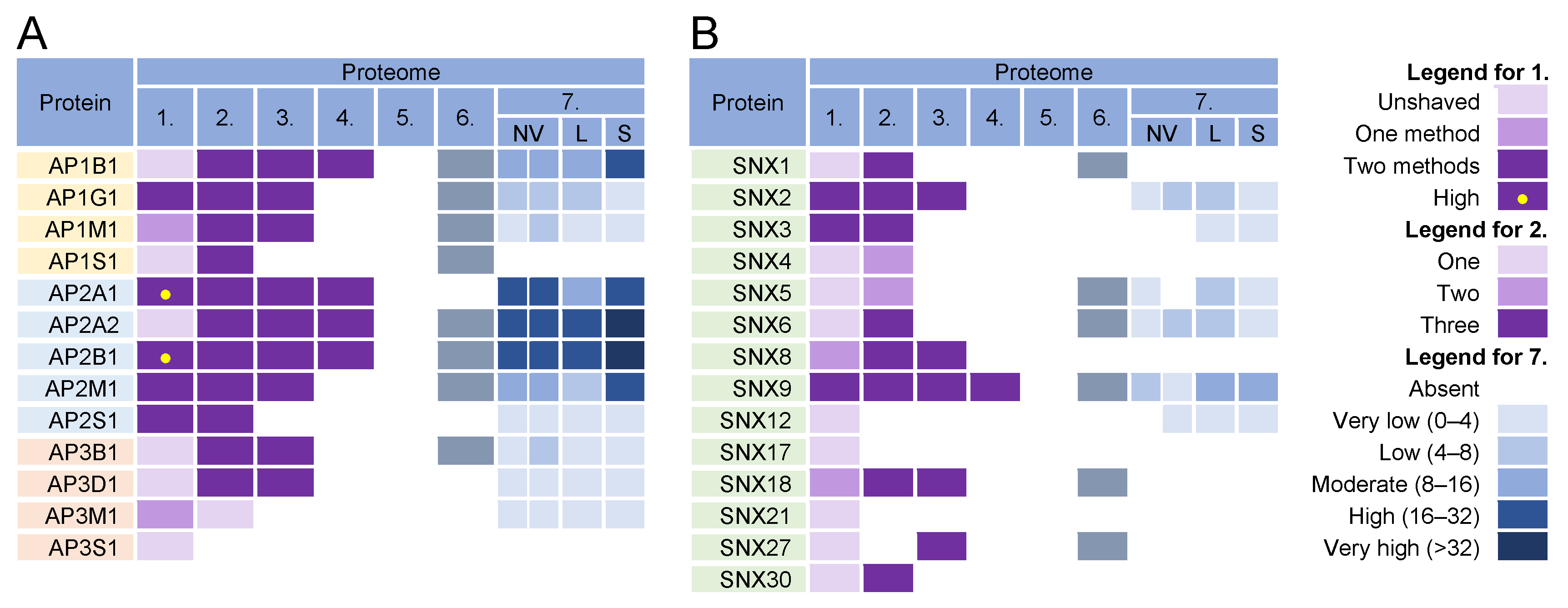
Figure 5. Identification of (A) adaptor protein (AP) complexes and (B) sorting nexins (SNX) in virions and NVEPs in extracellular vesicle preparations. The proteins are identified in five proteomes of HCMV virion preparations (proteome 1–5), proteomes of HSV-1 heavy particles (proteome 6), and NVEPs (proteome 7), as described in Figure 2.
In addition to classical adaptors, clathrin assembly can be facilitated by so-called monomeric adaptors; GGA complexes (Golgi-localized, γ-ear-containing, ADP-ribosylation factor-binding proteins) associate with the TGN and Hrs associate with endosomes [160]. Arrestins (ARRB1-2), epsins (EPN1-3), DAB2, ARH (autosomal recessive hypercholesterolemia protein), and several other proteins can also link cargo to clathrin and can therefore be classified as alternative adaptors [161]. None of these proteins were found in the proteinase K-treated virion preparations.
Overall, the analysis of the adaptor complexes provides no evidence that they can be used as a sorting mechanism for the concentration of viral glycoproteins at the envelopment membranes. Furthermore, this analysis suggests that the envelopment membranes are not active in cargo sorting based on clathrin recruitment.
References
- van Niel, G.; Carter, D.R.F.; Clayton, A.; Lambert, D.W.; Raposo, G.; Vader, P. Challenges and Directions in Studying Cell–Cell Communication by Extracellular Vesicles. Nat. Rev. Mol. Cell Biol. 2022, 23, 369–382.
- Jeppesen, D.K.; Fenix, A.M.; Franklin, J.L.; Higginbotham, J.N.; Zhang, Q.; Zimmerman, L.J.; Liebler, D.C.; Ping, J.; Liu, Q.; Evans, R.; et al. Reassessment of Exosome Composition. Cell 2019, 177, 428–445.e18.
- Zhang, Q.; Higginbotham, J.N.; Jeppesen, D.K.; Yang, Y.P.; Li, W.; McKinley, E.T.; Graves-Deal, R.; Ping, J.; Britain, C.M.; Dorsett, K.A.; et al. Transfer of Functional Cargo in Exomeres. Cell Rep. 2019, 27, 940–954.e6.
- Zhang, H.; Freitas, D.; Kim, H.S.; Fabijanic, K.; Li, Z.; Chen, H.; Mark, M.T.; Molina, H.; Martin, A.B.; Bojmar, L.; et al. Identification of Distinct Nanoparticles and Subsets of Extracellular Vesicles by Asymmetric Flow Field-Flow Fractionation. Nat. Cell Biol. 2018, 20, 332–343.
- Turner, D.L.; Korneev, D.V.; Purdy, J.G.; de Marco, A.; Mathias, R.A. The Host Exosome Pathway Underpins Biogenesis of the Human Cytomegalovirus Virion. eLife 2020, 9, e58288.
- Couté, Y.; Kraut, A.; Zimmermann, C.; Büscher, N.; Hesse, A.M.; Bruley, C.; de Andrea, M.; Wangen, C.; Hahn, F.; Marschall, M.; et al. Mass Spectrometry-Based Characterization of the Virion Proteome, Phosphoproteome, and Associated Kinase Activity of Human Cytomegalovirus. Microorganisms 2020, 8, 820.
- Rieder, F.J.J.; Kastner, M.T.; Hartl, M.; Puchinger, M.G.; Schneider, M.; Majdic, O.; Britt, W.J.; Djinović-Carugo, K.; Steininger, C. Human Cytomegalovirus Phosphoproteins Are Hypophosphorylated and Intrinsically Disordered. J. Gen. Virol. 2017, 98, 471–485.
- Reyda, S.; Büscher, N.; Tenzer, S.; Plachter, B. Proteomic Analyses of Human Cytomegalovirus Strain AD169 Derivatives Reveal Highly Conserved Patterns of Viral and Cellular Proteins in Infected Fibroblasts. Viruses 2014, 6, 172–188.
- Flomm, F.J.; Soh, T.K.; Schneider, C.; Britt, H.M.; Thalassinos, K.; Pfitzner, S.; Reimer, R.; Grünewald, K.; Bosse, J.B. Egress of Human Cytomegalovirus through Multivesicular Bodies. bioRxiv 2021.
- Birzer, A.; Kraner, M.E.; Heilingloh, C.S.; Mühl-Zürbes, P.; Hofmann, J.; Steinkasserer, A.; Popella, L. Mass Spectrometric Characterization of HSV-1 L-Particles From Human Dendritic Cells and BHK21 Cells and Analysis of Their Functional Role. Front. Microbiol. 2020, 11.
- Eyster, C.A.; Higginson, J.D.; Huebner, R.; Porat-Shliom, N.; Weigert, R.; Wu, W.W.; Shen, R.F.; Donaldson, J.G. Discovery of New Cargo Proteins That Enter Cells through Clathrin-Independent Endocytosis. Traffic 2009, 10, 590–599.
- Mayor, S.; Parton, R.G.; Donaldson, J.G. Clathrin-Independent Pathways of Endocytosis. Cold Spring Harb. Perspect. Biol. 2014, 6.
- Naslavsky, N.; Boehm, M.; Backlund, P.S.; Caplan, S. Rabenosyn-5 and EHD1 Interact and Sequentially Regulate Protein Recycling to the Plasma Membrane. Mol. Biol. Cell 2004, 15, 2410–2422.
- Maldonado-Báez, L.; Cole, N.B.; Krämer, H.; Donaldson, J.G. Microtubule-Dependent Endosomal Sorting of Clathrin-Independent Cargo by Hook1. J. Cell Biol. 2013, 201, 233–247.
- Lakadamyali, M.; Rust, M.J.; Zhuang, X. Ligands for Clathrin-Mediated Endocytosis Are Differentially Sorted into Distinct Populations of Early Endosomes. Cell 2006, 124, 997–1009.
- Maxfield, F.R.; McGraw, T.E. Endocytic Recycling. Nat. Rev. Mol. Cell Biol. 2004, 5, 121–132.
- Rink, J.; Ghigo, E.; Kalaidzidis, Y.; Zerial, M. Rab Conversion as a Mechanism of Progression from Early to Late Endosomes. Cell 2005, 122, 735–749.
- Staudt, C.; Puissant, E.; Boonen, M. Subcellular Trafficking of Mammalian Lysosomal Proteins: An Extended View. Int. J. Mol. Sci. 2017, 18, 47.
- Pols, M.S.; Klumperman, J. Trafficking and Function of the Tetraspanin CD63. Exp. Cell Research 2009, 315, 1584–1592.
- Poirier, S.; Mayer, G.; Murphy, S.R.; Garver, W.S.; Chang, T.Y.; Schu, P.; Seidah, N.G. The Cytosolic Adaptor AP-1A Is Essential for the Trafficking and Function of Niemann-Pick Type C Proteins. Traffic 2013, 14, 458–469.
- Moreno-Layseca, P.; Icha, J.; Hamidi, H.; Ivaska, J. Integrin Trafficking in Cells and Tissues. Nat. Cell Biol. 2019, 21, 122–132.
- Pavlos, N.J.; Friedman, P.A. GPCR Signaling and Trafficking: The Long and Short of It. Trends Endocrinol. Metab. 2017, 28, 213.
- Bonifacino, J.S.; Rojas, R. Retrograde Transport from Endosomes to the Trans-Golgi Network. Nat. Rev. Mol. Cell Biol. 2006, 7, 568–579.
- Cullen, P.J.; Steinberg, F. To Degrade or Not to Degrade: Mechanisms and Significance of Endocytic Recycling. Nat. Rev. Mol. Cell Biol. 2018, 19, 679–696.
- Naslavsky, N.; Caplan, S. The Enigmatic Endosome—Sorting the Ins and Outs of Endocytic Trafficking. J. Cell Sci. 2018, 131.
- Goldenring, J.R. Recycling Endosomes. Curr. Opin. Cell Biol. 2015, 35, 117–122.
- Ioannou, M.S.; McPherson, P.S. Regulation of Cancer Cell Behavior by the Small GTPase Rab13. J. Biol. Chem. 2016, 291, 9929.
- Klinkert, K.; Echard, A. Rab35 GTPase: A Central Regulator of Phosphoinositides and F-Actin in Endocytic Recycling and Beyond. Traffic 2016, 17, 1063–1077.
- Hanley, S.E.; Cooper, K.F. Sorting Nexins in Protein Homeostasis. Cells 2021, 10, 17.
- Kalaidzidis, I.; Miaczynska, M.; Brewinska-Olchowik, M.; Hupalowska, A.; Ferguson, C.; Parton, R.G.; Kalaidzidis, Y.; Zerial, M. APPL Endosomes Are Not Obligatory Endocytic Intermediates but Act as Stable Cargo-Sorting Compartments. J. Cell Biol. 2015, 211, 123–144.
- Powelka, A.M.; Sun, J.; Li, J.; Gao, M.; Shaw, L.M.; Sonnenberg, A.; Hsu, V.W. Stimulation-Dependent Recycling of Integrin Β1 Regulated by ARF6 and Rab11. Traffic 2004, 5, 20–36.
- Campbell, I.D.; Humphries, M.J. Integrin Structure, Activation, and Interactions. Cold Spring Harb. Perspect. Biol. 2011, 3, a004994.
- Sahgal, P.; Alanko, J.; Icha, J.; Paatero, I.; Hamidi, H.; Arjonen, A.; Pietilä, M.; Rokka, A.; Ivaska, J. GGA2 and RAB13 Promote Activity-Dependent Β1-Integrin Recycling. J. Cell Sci. 2019, 132.
- Dutta, D.; Donaldson, J.G. Sorting of Clathrin-Independent Cargo Proteins Depends on Rab35 Delivered by Clathrin-Mediated Endocytosis. Traffic 2015, 16, 994–1009.
- Etoh, K.; Fukuda, M. Rab10 Regulates Tubular Endosome Formation through KIF13A and KIF13B Motors. J. Cell Sci. 2019, 132.
- Farmer, T.; Xie, S.; Naslavsky, N.; Stöckli, J.; James, D.E.; Caplan, S. Defining the Protein and Lipid Constituents of Tubular Recycling Endosomes. J. Biol. Chem. 2021, 296.
- D’Souza, R.S.; Semus, R.; Billings, E.A.; Meyer, C.B.; Conger, K.; Casanova, J.E. Rab4 Orchestrates a Small GTPase Cascade for Recruitment of Adaptor Proteins to Early Endosomes. Curr. Biol. 2014, 24, 1187–1198.
- Tian, Y.; Kang, Q.; Shi, X.; Wang, Y.; Zhang, N.; Ye, H.; Xu, Q.; Xu, T.; Zhang, R. SNX-3 Mediates Retromer-Independent Tubular Endosomal Recycling by Opposing EEA-1-Facilitated Trafficking. PLoS Genet. 2021, 17.
- Singla, A.; Fedoseienko, A.; Giridharan, S.S.P.; Overlee, B.L.; Lopez, A.; Jia, D.; Song, J.; Huff-Hardy, K.; Weisman, L.; Burstein, E.; et al. Endosomal PI(3)P Regulation by the COMMD/CCDC22/CCDC93 (CCC) Complex Controls Membrane Protein Recycling. Nat. Commun. 2019, 10, 4271.
- Phillips-Krawczak, C.A.; Singla, A.; Starokadomskyy, P.; Deng, Z.; Osborne, D.G.; Li, H.; Dick, C.J.; Gomez, T.S.; Koenecke, M.; Zhang, J.S.; et al. COMMD1 Is Linked to the WASH Complex and Regulates Endosomal Trafficking of the Copper Transporter ATP7A. Mol. Biol. Cell 2015, 26, 91–103.
- Gomez, T.S.; Billadeau, D.D. A FAM21-Containing WASH Complex Regulates Retromer-Dependent Sorting. Dev. Cell 2009, 17, 699–711.
- Derivery, E.; Sousa, C.; Gautier, J.J.; Lombard, B.; Loew, D.; Gautreau, A. The Arp2/3 Activator WASH Controls the Fission of Endosomes through a Large Multiprotein Complex. Dev. Cell 2009, 17, 712–723.
- Steinberg, F.; Heesom, K.J.; Bass, M.D.; Cullen, P.J. SNX17 Protects Integrins from Degradation by Sorting between Lysosomal and Recycling Pathways. J. Cell. Biol. 2012, 197, 219–230.
- McNally, K.E.; Cullen, P.J. Endosomal Retrieval of Cargo: Retromer Is Not Alone. Trends Cell Biol. 2018, 28, 807–822.
- Steinberg, F.; Gallon, M.; Winfield, M.; Thomas, E.C.; Bell, A.J.; Heesom, K.J.; Tavaré, J.M.; Cullen, P.J. A Global Analysis of SNX27-Retromer Assembly and Cargo Specificity Reveals a Function in Glucose and Metal Ion Transport. Nat. Cell Biol. 2013, 15, 461–471.
- Gallon, M.; Clairfeuille, T.; Steinberg, F.; Mas, C.; Ghai, R.; Sessions, R.B.; Teasdale, R.D.; Collins, B.M.; Cullen, P.J. A Unique PDZ Domain and Arrestin-like Fold Interaction Reveals Mechanistic Details of Endocytic Recycling by SNX27-Retromer. Proc. Natl. Acad. Sci. USA 2014, 111.
- van Weering, J.R.T.; Verkade, P.; Cullen, P.J. SNX-BAR-Mediated Endosome Tubulation Is Co-Ordinated with Endosome Maturation. Traffic 2012, 13, 94–107.
- Simonetti, B.; Guo, Q.; Gimenez-Andres, M.; Chen, K.-E.; Moody, E.R.; Evans, A.J.; Danson, C.M.; Williams, T.A.; Collins, B.M.; Cullen, P.J. Mechanistic Basis for SNX27-Retromer Coupling to ESCPE-1 in Promoting Endosomal Cargo Recycling. bioRxiv 2021, preprint.
- Priya, A.; Sugatha, J.; Parveen, S.; Lacas-Gervais, S.; Raj, P.; Gilleron, J.; Datta, S. Essential and Selective Role of SNX12 in Transport of Endocytic and Retrograde Cargo. J. Cell. Sci. 2017, 130, 2707–2721.
- Kim Nguyen, N.T.; Ohbayashi, N.; Kanaho, Y.; Funakoshi, Y. TBC1D24 Regulates Recycling of Clathrin-Independent Cargo Proteins Mediated by Tubular Recycling Endosomes. Biochem. Biophys. Res. Commun. 2020, 528, 220–226.
- Mahabaleshwar, H.; Tarbashevich, K.; Nowak, M.; Brand, M.; Raz, E. β-Arrestin Control of Late Endosomal Sorting Facilitates Decoy Receptor Function and Chemokine Gradient Formation. Development 2012, 139, 2897–2902.
- Rainero, E. CLIC3 Controls Recycling of Late Endosomal MT1-MMP and Dictates Invasion and Metastasis in Breast Cancer. J. Cell Sci. 2014.
- Dozynkiewicz, M.A.; Jamieson, N.B.; MacPherson, I.; Grindlay, J.; van den Berghe, P.V.E.; von Thun, A.; Morton, J.P.; Gourley, C.; Timpson, P.; Nixon, C.; et al. Rab25 and CLIC3 Collaborate to Promote Integrin Recycling from Late Endosomes/Lysosomes and Drive Cancer Progression. Dev. Cell 2012, 22, 131–145.
- Walseng, E.; Bakke, O.; Roche, P.A. Major Histocompatibility Complex Class II-Peptide Complexes Internalize Using a Clathrin- and Dynamin-Independent Endocytosis Pathway. J. Biol. Chem. 2008, 283, 14717–14727.
- Kleijmeer, M.; Ramm, G.; Schuurhuis, D.; Griffith, J.; Rescigno, M.; Ricciardi-Castagnoli, P.; Rudensky, A.Y.; Ossendorp, F.; Melief, C.J.M.; Stoorvogel, W.; et al. Reorganization of Multivesicular Bodies Regulates MHC Class II Antigen Presentation by Dendritic Cells. J. Cell Biol. 2001, 155, 53–63.
- Mahmutefendić, H.; Blagojević Zagorac, G.; Grabušić, K.; Karleuša, L.; Maćešić, S.; Momburg, F.; Lučin, P. Late Endosomal Recycling of Open MHC-I Conformers. J. Cell. Physiol. 2017, 232, 872–887.
- Chen, P.I.; Schauer, K.; Kong, C.; Harding, A.R.; Goud, B.; Stahl, P.D. Rab5 Isoforms Orchestrate a “Division of Labor” in the Endocytic Network; Rab5C Modulates Rac-Mediated Cell Motility. PLoS ONE 2014, 9, e90384.
- Hunt, S.D.; Townley, A.K.; Danson, C.M.; Cullen, P.J.; Stephens, D.J. Microtubule Motors Mediate Endosomal Sorting by Maintaining Functional Domain Organization. J. Cell Sci. 2013, 126, 2493–2501.
- Carlton, J.; Bujny, M.; Peter, B.J.; Oorschot, V.M.J.; Rutherford, A.; Mellor, H.; Klumperman, J.; McMahon, H.T.; Cullen, P.J. Sorting Nexin-1 Mediates Tubular Endosome-to-TGN Transport through Coincidence Sensing of High-Curvature Membranes and 3-Phosphoinositides. Curr. Biol. 2004, 14, 1791–1800.
- Rai, A.; Oprisko, A.; Campos, J.; Fu, Y.; Friese, T.; Itzen, A.; Goody, R.S.; Gazdag, E.M.; Müller, M.P. Bmerb Domains Are Bivalent Rab8 Family Effectors Evolved by Gene Duplication. eLife 2016, 5, e18675.
- Nokes, R.L.; Fields, I.C.; Collins, R.N.; Fölsch, H. Rab13 Regulates Membrane Trafficking between TGN and Recycling Endosomes in Polarized Epithelial Cells. J. Cell. Biol. 2008, 182, 845–853.
- Suzuki, S.W.; Oishi, A.; Nikulin, N.; Jorgensen, J.R.; Baile, M.G.; Emr, S.D. A Px-Bar Protein Mvp1/Snx8 and a Dynamin-like Gtpase Vps1 Drive Endosomal Recycling. eLife 2021, 10, e69883.
- Itzhak, D.N.; Tyanova, S.; Cox, J.; Borner, G.H.H. Global, Quantitative and Dynamic Mapping of Protein Subcellular Localization. eLife 2016, 5, 1–36.
- Weekes, M.P.; Tomasec, P.; Huttlin, E.L.; Fielding, C.A.; Nusinow, D.; Stanton, R.J.; Wang, E.C.Y.; Aicheler, R.; Murrell, I.; Wilkinson, G.W.G.; et al. Quantitative Temporal Viromics: An Approach to Investigate Host-Pathogen Interaction. Cell 2014, 157, 1460–1472.
- Zeltzer, S.; Zeltzer, C.A.; Igarashi, S.; Wilson, J.; Donaldson, J.G.; Goodrum, F. Virus Control of Trafficking from Sorting Endosomes. mBio 2018, 9, e00683-18.
- Karleuša, L.; Mahmutefendić, H.; Tomaš, M.I.; Zagorac, G.B.; Lučin, P. Landmarks of Endosomal Remodeling in the Early Phase of Cytomegalovirus Infection. Virology 2018, 515, 108–122.
- Tomaš, M.I.; Kučić, N.; Mahmutefendić, H.; Blagojević, G.; Lučin, P. Murine Cytomegalovirus Perturbs Endosomal Trafficking of Major Histocompatibility Complex Class I Molecules in the Early Phase of Infection. J. Virol. 2010, 84, 1101–1112.
- Homma, Y.; Hiragi, S.; Fukuda, M. Rab Family of Small GTPases: An Updated View on Their Regulation and Functions. FEBS J. 2021, 288, 36–55.
- Lučin, P.; Jug Vučko, N.; Karleuša, L.; Lučin, M.H.; Zagorac, B.G.; Lisnić, B.; Pavišić, V.; Marcelić, M.; Grabušić, K.; Brizić, I.; et al. Cytomegalovirus Generates Assembly Compartment in the Early Phase of Infection by Perturbation of Host-Cell Factors Recruitment at the Early Endosome/Endosomal Recycling Compartment/Trans-Golgi Interface. Front. Cell Dev. Biol. 2020, 8, 914.
- Archer, M.A.; Brechtel, T.M.; Davis, L.E.; Parmar, R.C.; Hasan, M.H.; Tandon, R. Inhibition of Endocytic Pathways Impacts Cytomegalovirus Maturation. Sci. Rep. 2017, 7, 46069.
- Schauflinger, M.; Villinger, C.; Mertens, T.; Walther, P.; von Einem, J. Analysis of Human Cytomegalovirus Secondary Envelopment by Advanced Electron Microscopy. Cell. Microbiol. 2013, 15, 305–314.
- Buser, C.; Walther, P.; Mertens, T.; Michel, D. Cytomegalovirus Primary Envelopment Occurs at Large Infoldings of the Inner Nuclear Membrane. J. Virol. 2007, 81, 3042–3048.
- Maninger, S.; Bosse, J.B.; Lemnitzer, F.; Pogoda, M.; Mohr, C.A.; von Einem, J.; Walther, P.; Koszinowski, U.H.; Ruzsics, Z. M94 Is Essential for the Secondary Envelopment of Murine Cytomegalovirus. J. Virol. 2011, 85, 9254–9267.
- Homman-Loudiyi, M.; Hultenby, K.; Britt, W.; Söderberg-Nauclér, C. Envelopment of Human Cytomegalovirus Occurs by Budding into Golgi-Derived Vacuole Compartments Positive for GB, Rab 3, Trans-Golgi Network 46, and Mannosidase II. J. Virol. 2003, 77, 3191–3203.
- Goud, B.; Liu, S.; Storrie, B. Rab Proteins as Major Determinants of the Golgi Complex Structure. Small GTPases 2018, 9, 66–75.
- Martínez-Menárguez, J.Á.; Martínez-Alonso, E.; Cara-Esteban, M.; Tomás, M. Focus on the Small GTPase Rab1: A Key Player in the Pathogenesis of Parkinson’s Disease. Int. J. Mol. Sci. 2021, 22, 12087.
- Saraste, J. Spatial and Functional Aspects of ER-Golgi Rabs and Tethers. Front. Cell Dev. Biol. 2016, 4.
- Hutagalung, A.H.; Novick, P.J. Role of Rab GTPases in Membrane Traffic and Cell Physiology. Physiol. Rev. 2011, 91, 119–149.
- Shakya, S.; Sharma, P.; Bhatt, A.M.; Jani, R.A.; Delevoye, C.; Gangi Setty, S.R. Rab22A Recruits BLOC -1 and BLOC -2 to Promote the Biogenesis of Recycling Endosomes. EMBO Rep. 2018, 19, e45918.
- Evans, T.M.; Ferguson, C.; Wainwright, B.J.; Parton, R.G.; Wicking, C. Rab23, a Negative Regulator of Hedgehog Signaling, Localizes to the Plasma Membrane and the Endocytic Pathway. Traffic 2003, 4, 869–884.
- Fukuda, M. Rab27 Effectors, Pleiotropic Regulators in Secretory Pathways. Traffic 2013, 14, 949–963.
- Wang, S.; Ma, Z.; Xu, X.; Wang, Z.; Sun, L.; Zhou, Y.; Lin, X.; Hong, W.; Wang, T. A Role of Rab29 in the Integrity of the Trans-Golgi Network and Retrograde Trafficking of Mannose-6-Phosphate Receptor. PLoS ONE 2014, 9, e96242.
- Drizyte-Miller, K.; Chen, J.; Cao, H.; Schott, M.B.; McNiven, M.A. The Small GTPase Rab32 Resides on Lysosomes to Regulate MTORC1 Signaling. J. Cell Sci. 2020, 133.
- Ohbayashi, N.; Fukuda, M.; Kanaho, Y. Rab32 Subfamily Small GTPases: Pleiotropic Rabs in Endosomal Trafficking. J. Biochem. 2017, 162, 65–71.
- Sönnichsen, B.; de Renzis, S.; Nielsen, E.; Rietdorf, J.; Zerial, M. Distinct Membrane Domains on Endosomes in the Recycling Pathway Visualized by Multicolor Imaging of Rab4, Rab5, and Rab11. J. Cell. Biol. 2000, 149, 901–913.
- Kobayashi, H.; Etoh, K.; Ohbayashi, N.; Fukuda, M. Rab35 Promotes the Recruitment of Rab8, Rab13 and Rab36 to Recycling Endosomes through MICAL-L1 during Neurite Outgrowth. Biol. Open 2014, 3, 803–814.
- Liu, S.; Storrie, B. How Rab Proteins Determine Golgi Structure; Elsevier: Amsterdam, The Netherlands, 2015; Volume 315.
- Adarska, P.; Wong-Dilworth, L.; Bottanelli, F. ARF GTPases and Their Ubiquitous Role in Intracellular Trafficking Beyond the Golgi. Front. Cell Dev. Biol. 2021, 9, 1977.
- Sztul, E.; Chen, P.W.; Casanova, J.E.; Cherfils, J.; Dacks, J.B.; Lambright, D.G.; Lee, F.J.S.; Randazzo, P.A.; Santy, L.C.; Schürmann, A.; et al. Arf GTPases and Their GEFs and GAPS: Concepts and Challenges. Mol. Biol. Cell 2019, 30, 1249–1271.
- Donaldson, J.G.; Jackson, C.L. ARF Family G Proteins and Their Regulators: Roles in Membrane Transport, Development and Disease. Nat. Rev. Mol. Cell Biol. 2011, 12, 362–375.
- Vetter, M.; Wang, J.; Lorentzen, E.; Deretic, D. Novel Topography of the Rab 11-Effector Interaction Network within a Ciliary Membrane Targeting Complex. Small GTPases 2015, 6, 165–173.
- Barbero, P.; Bittova, L.; Pfeffer, S.R. Visualization of Rab9-Mediated Vesicle Transport from Endosomes to the Trans-Golgi in Living Cells. J. Cell Biol. 2002, 156, 511–518.
- D’Souza-Schorey, C.; Chavrier, P. ARF Proteins: Roles in Membrane Traffic and Beyond. Nat. Rev. Mol. Cell Biol. 2006, 7, 347–358.
- Zhou, C.; Cunningham, L.; Marcus, A.I.; Li, Y.; Kahn, R.A. Arl2 and Arl3 Regulate Different Microtubule-Dependent Processes. Mol. Biol. Cell 2006, 17, 2476–2487.
- Takahashi, S.; Kubo, K.; Waguri, S.; Yabashi, A.; Shin, H.W.; Katoh, Y.; Nakayama, K. Rab11 Regulates Exocytosis of Recycling Vesicles at the Plasma Membrane. J. Cell Sci. 2012, 125, 4049–4057.
- Welz, T.; Wellbourne-Wood, J.; Kerkhoff, E. Orchestration of Cell Surface Proteins by Rab11. Trends Cell Biol. 2014, 24, 407–415.
- Hofmann, I.; Munro, S. An N-Terminally Acetylated Arf-like GTPase Is Localised to Lysosomes and Affects Their Motility. J. Cell Sci. 2006, 119, 1494–1503.
- Escrevente, C.; Bento-Lopes, L.; Ramalho, J.S.; Barral, D.C. Rab11 Is Required for Lysosome Exocytosis through the Interaction with Rab3a, Sec15 and GRAB. J. Cell Sci. 2021, 134.
- Zulkefli, K.L.; Houghton, F.J.; Gosavi, P.; Gleeson, P.A. A Role for Rab11 in the Homeostasis of the Endosome-Lysosomal Pathway. Exp. Cell Res. 2019, 380, 55–68.
- Matsui, T.; Itoh, T.; Fukuda, M. Small GTPase Rab12 Regulates Constitutive Degradation of Transferrin Receptor. Traffic 2011, 12, 1432–1443.
- Mosaddeghzadeh, N.; Ahmadian, M.R. The RHO Family GTPases: Mechanisms of Regulation and Signaling. Cells 2021, 10, 1831.
- Dejgaard, S.Y.; Presley, J.F. Rab18: New Insights into the Function of an Essential Protein. Cell. Mol. Life Sci. 2019, 76, 1935–1945.
- Wheeler, A.P.; Ridley, A.J. Why Three Rho Proteins? RhoA, RhoB, RhoC, and Cell Motility. Exp. Cell Res. 2004, 301, 43–49.
- Simpson, J.C.; Griffiths, G.; Wessling-Resnick, M.; Fransen, J.A.M.; Bennett, H.; Jones, A.T. A Role for the Small GTPase Rab21 in the Early Endocytic Pathway. J. Cell Sci. 2004, 117, 6297–6311.
- Müller, M.P.; Goody, R.S. Molecular Control of Rab Activity by GEFs, GAPs and GDI. Small GTPases 2018, 9, 5–21.
- Matsui, T.; Fukuda, M. Rab12 Regulates MTORC1 Activity and Autophagy through Controlling the Degradation of Amino-Acid Transporter PAT4. EMBO Rep. 2013, 14, 450–457.
- Xu, J.; McPherson, P.S. DENND3: A Signaling/Trafficking Interface in Autophagy. Cell Cycle 2015, 14, 2717.
- Omar, J.; Rosenbaum, E.; Efergan, A.; Sneineh, B.A.; Yeheskel, A.; Maruta, Y.; Fukuda, M.; Sagi-Eisenberg, R. Biochemical and Structural Insights into Rab12 Interactions with RILP and Its Family Members. Sci. Rep. 2021, 11, 10317.
- Feldmann, A.; Bekbulat, F.; Huesmann, H.; Ulbrich, S.; Tatzelt, J.; Behl, C.; Kern, A. The RAB GTPase RAB18 Modulates Macroautophagy and Proteostasis. Biochem. Biophys. Res. Commun. 2017, 486, 738–743.
- Gerondopoulos, A.; Bastos, R.N.; Yoshimura, S.I.; Anderson, R.; Carpanini, S.; Aligianis, I.; Handley, M.T.; Barr, F.A. Rab18 and a Rab18 GEF Complex Are Required for Normal ER Structure. J. Cell Biol. 2014, 205, 707–720.
- Haines, D.S.; Lee, J.E.; Beauparlant, S.L.; Kyle, D.B.; den Besten, W.; Sweredoski, M.J.; Graham, R.L.J.; Hess, S.; Deshaies, R.J. Protein Interaction Profiling of the P97 Adaptor UBXD1 Points to a Role for the Complex in Modulating ERGIC-53 Trafficking. Mol. Cell. Proteom. 2012, 11.
- Takáts, S.; Lévay, L.; Boda, A.; Tóth, S.; Simon-Vecsei, Z.; Rubics, A.; Varga, Á.; Lippai, M.; Lőrincz, P.; Glatz, G.; et al. The Warburg Micro Syndrome-Associated Rab3GAP-Rab18 Module Promotes Autolysosome Maturation through the Vps34 Complex I. FEBS J. 2021, 288, 190–211.
- Vazquez-Martinez, R.; Cruz-Garcia, D.; Duran-Prado, M.; Peinado, J.R.; Castaño, J.P.; Malagon, M.M. Rab18 Inhibits Secretory Activity in Neuroendocrine Cells by Interacting with Secretory Granules. Traffic 2007, 8, 867–882.
- Takamori, S.; Holt, M.; Stenius, K.; Lemke, E.A.; Grønborg, M.; Riedel, D.; Urlaub, H.; Schenck, S.; Brügger, B.; Ringler, P.; et al. Molecular Anatomy of a Trafficking Organelle. Cell 2006, 127, 831–846.
- Nian, F.S.; Li, L.L.; Cheng, C.Y.; Wu, P.C.; Lin, Y.T.; Tang, C.Y.; Ren, B.S.; Tai, C.Y.; Fann, M.J.; Kao, L.; et al. Rab18 Collaborates with Rab7 to Modulate Lysosomal and Autophagy Activities in the Nervous System: An Overlapping Mechanism for Warburg Micro Syndrome and Charcot-Marie-Tooth Neuropathy Type 2B. Mol. Neurobiol. 2019, 56, 6095–6105.
- Ao, X.; Zou, L.; Wu, Y. Regulation of Autophagy by the Rab GTPase Network. Cell Death Differ. 2014, 21, 348.
- Hirota, Y.; Tanaka, Y. A Small GTPase, Human Rab32, Is Required for the Formation of Autophagic Vacuoles under Basal Conditions. Cell. Mol. Life Sci. 2009, 66, 2913–2932.
- Hokanson, D.E.; Bretscher, A.P. EPI64 Interacts with Slp1/JFC1 to Coordinate Rab8a and Arf6 Membrane Trafficking. Mol. Biol. Cell 2012, 23, 701–715.
- Saraste, J.; Prydz, K. Assembly and Cellular Exit of Coronaviruses: Hijacking an Unconventional Secretory Pathway from the Pre-Golgi Intermediate Compartment via the Golgi Ribbon to the Extracellular Space. Cells 2021, 10, 503.
- Saraste, J.; Prydz, K. A New Look at the Functional Organization of the Golgi Ribbon. Front. Cell Dev. Biol. 2019, 7.
- Pfeffer, S.R. Entry at the Trans-Face of the Golgi. Cold Spring Harb. Perspect. Biol. 2017, 3, a005272.
- Heffernan, L.F.; Simpson, J.C. The Trials and Tubule-Ations of Rab6 Involvement in Golgi-to-ER Retrograde Transport. Biochem. Soc. Trans. 2014, 42, 1453–1459.
- Gillingham, A.K.; Munro, S. Transport Carrier Tethering—How Vesicles Are Captured by Organelles. Curr. Opin. Cell Biol. 2019, 59, 140–146.
- Lu, Q.; Wang, P.S.; Yang, L. Golgi-Associated Rab GTPases Implicated in Autophagy. Cell Biosci. 2021, 11, 35.
- Indran, S.v.; Britt, W.J. A Role for the Small GTPase Rab6 in Assembly of Human Cytomegalovirus. J. Virol. 2011, 85, 5213–5219.
- Langemeyer, L.; Fröhlich, F.; Ungermann, C. Rab GTPase Function in Endosome and Lysosome Biogenesis. Trends Cell Biol. 2018, 28, 957–970.
- Mateus, D.; Marini, E.S.; Progida, C.; Bakke, O. Rab7a Modulates ER Stress and ER Morphology. Biochim. Biophys. Acta Mol. Cell Res. 2018, 1865, 781–793.
- Grimsey, N.J.; Coronel, L.J.; Cordova, I.C.; Trejo, J. Recycling and Endosomal Sorting of Protease-Activated Receptor-1 Is Distinctly Regulated by Rab11A and Rab11B Proteins. J. Biol. Chem. 2016, 291, 2223–2236.
- Lapierre, L.A.; Dorn, M.C.; Zimmerman, C.F.; Navarre, J.; Burnette, J.O.; Goldenring, J.R. Rab11b Resides in a Vesicular Compartment Distinct from Rab11a in Parietal Cells and Other Epithelial Cells. Exp. Cell Res. 2003, 290, 322–331.
- Chua, C.E.L.; Tang, B.L. Rab 10—A Traffic Controller in Multiple Cellular Pathways and Locations. J. Cell. Physiol. 2018, 233, 6483–6494.
- English, A.R.; Voeltz, G.K. Endoplasmic Reticulum Structure and Interconnections with Other Organelles. Cold Spring Harb. Perspect. Biol. 2013, 5, a013227.
- Palmisano, N.J.; Rosario, N.; Wysocki, M.; Hong, M.; Grant, B.; Meléndez, A. The Recycling Endosome Protein RAB-10 Promotes Autophagic Flux and Localization of the Transmembrane Protein ATG-9. Autophagy 2017, 13, 1742–1753.
- Hatoyama, Y.; Homma, Y.; Hiragi, S.; Fukuda, M. Establishment and Analysis of Conditional Rab1- and Rab5-Knockout Cells Using the Auxin-Inducible Degron System. J. Cell Sci. 2021, 134, jcs259184.
- Mendoza, P.; Ortiz, R.; Dýáz, J.; Quest, A.F.G.; Leyton, L.; Stupack, W.; Torres, V.A. Rab5 Activation Promotes Focal Adhesion Disassembly, Migration and Invasiveness in Tumor Cells. J. Cell Sci. 2013, 126, 3835–3847.
- Chen, P.W.; Luo, R.; Jian, X.; Randazzo, P.A. The Arf6 GTPase-Activating Proteins ARAP2 and ACAP1 Define Distinct Endosomal Compartments That Regulate Integrin A5βS1 Traffic. J. Biol. Chem. 2014, 289, 30237–30248.
- Barbera, S.; Nardi, F.; Elia, I.; Realini, G.; Lugano, R.; Santucci, A.; Tosi, G.M.; Dimberg, A.; Galvagni, F.; Orlandini, M. The Small GTPase Rab5c Is a Key Regulator of Trafficking of the CD93/Multimerin-2/Β1 Integrin Complex in Endothelial Cell Adhesion and Migration. Cell Commun. Signal. 2019, 17, 1–15.
- Cherfils, J.; Zeghouf, M. Regulation of Small GTPases by GEFs, GAPs, and GDIs. Physiol. Rev. 2013, 93, 269–309.
- Das, S.; Vasanji, A.; Pellett, P.E. Three-Dimensional Structure of the Human Cytomegalovirus Cytoplasmic Virion Assembly Complex Includes a Reoriented Secretory Apparatus. J. Virol. 2007, 81, 11861–11869.
- Momtaz, S.; Molina, B.; Mlera, L.; Goodrum, F.; Wilson, J.M. Cell Type-Specific Biogenesis of Novel Vesicles Containing Viral Products in Human Cytomegalovirus Infection. J. Virol. 2021, 95, e02358-20.
- Das, S.; Pellett, P.E. Spatial Relationships between Markers for Secretory and Endosomal Machinery in Human Cytomegalovirus-Infected Cells versus Those in Uninfected Cells. J. Virol. 2011, 85, 5864–5879.
- Sanchez, V.; Greis, K.D.; Sztul, E.; Britt, W.J. Accumulation of Virion Tegument and Envelope Proteins in a Stable Cytoplasmic Compartment during Human Cytomegalovirus Replication: Characterization of a Potential Site of Virus Assembly. J. Virol. 2000, 74, 975–986.
- Cepeda, V.; Esteban, M.; Fraile-Ramos, A. Human Cytomegalovirus Final Envelopment on Membranes Containing Both Trans-Golgi Network and Endosomal Markers. Cell. Microbiol. 2010, 12, 386–404.
- Taisne, C.; Lussignol, M.; Hernandez, E.; Moris, A.; Mouna, L.; Esclatine, A. Human Cytomegalovirus Hijacks the Autophagic Machinery and LC3 Homologs in Order to Optimize Cytoplasmic Envelopment of Mature Infectious Particles. Sci. Rep. 2019, 9, 4560.
- Hook, L.M.; Grey, F.; Grabski, R.; Tirabassi, R.; Doyle, T.; Hancock, M.; Landais, I.; Jeng, S.; McWeeney, S.; Britt, W.; et al. Cytomegalovirus MiRNAs Target Secretory Pathway Genes to Facilitate Formation of the Virion Assembly Compartment and Reduce Cytokine Secretion. Cell Host Microbe 2014, 15, 363–373.
- Pavišić, V.; Lučin, H.M.; Zagorac, G.B.; Lučin, P. Arf Gtpases Are Required for the Establishment of the Pre-Assembly Compartment in the Early Phase of Cytomegalovirus Infection. Life 2021, 11, 867.
- Krzyzaniak, M.A.; Mach, M.; Britt, W.J. HCMV-Encoded Glycoprotein M (UL100) Interacts with Rab11 Effector Protein FIP4. Traffic 2009, 10, 1439–1457.
- Lučin, P.; Kareluša, L.; Zagorac, B.G.; Lučin, M.H.; Pavišić, V.; Vučko, J.N.; Jurić, L.S.; Marcelić, M.; Lisnić, B.; Jonjić, S. Cytomegaloviruses Exploit Recycling Rab Proteins in the Sequential Establishment of the Assembly Compartment. Front. Cell Dev. Biol. 2018, 6, 165.
- Van Acker, T.; Tavernier, J.; Peelman, F. The Small GTPase Arf6: An Overview of Its Mechanisms of Action and of Its Role in Host- Pathogen Interactions and Innate Immunity. Int. J. Mol. Sci. 2019, 20, 2209.
- Moreau, K.; Ravikumar, B.; Puri, C.; Rubinsztein, D.C. Arf6 Promotes Autophagosome Formation via Effects on Phosphatidylinositol 4,5-Bisphosphate and Phospholipase D. J. Cell Biol. 2012, 196, 483–496.
- George, A.A.; Hayden, S.; Stanton, G.R.; Brockerhoff, S.E. Arf6 and the 5’phosphatase of Synaptojanin 1 Regulate Autophagy in Cone Photoreceptors. Bioessays 2016, 38 (Suppl. S1), S119–S135.
- Tooze, J.; Hollinshead, M. Tubular Early Endosomal Networks in AtT20 and Other Cells. J. Cell Biol. 1991, 115, 635–653.
- Eyster, C.A.; Cole, N.B.; Petersen, S.; Viswanathan, K.; Fruḧ, K.; Donaldson, J.G. MARCH Ubiquitin Ligases Alter the Itinerary of Clathrin-Independent Cargo from Recycling to Degradation. Mol. Biol. Cell 2011, 22, 3218–3230.
- Homma, Y.; Fukuda, M. Rabin8 Regulates Neurite Outgrowth in Both GEF Activity-Dependent and -Independent Manners. Mol. Biol. Cell 2016, 27, 2107–2118.
- Chesneau, L.; Dambournet, D.; MacHicoane, M.; Kouranti, I.; Fukuda, M.; Goud, B.; Echard, A. An ARF6/Rab35 GTPase Cascade for Endocytic Recycling and Successful Cytokinesis. Curr. Biol. 2012, 22, 147–153.
- Park, S.Y.; Guo, X. Adaptor Protein Complexes and Intracellular Transport. Biosci. Rep. 2014, 34, 381–390.
- Sanger, A.; Hirst, J.; Davies, A.K.; Robinson, M.S. Adaptor Protein Complexes and Disease at a Glance. J. Cell Sci. 2019, 132.
- Mettlen, M.; Chen, P.; Srinivasan, S.; Danuser, G.; Schmid, S.L. Regulation of Clathrin-Mediated Endocytosis. Annu. Rev. Biochem. 2018, 87, 871–896.
- Štimac, I.; Vučko, N.J.; Zagorac, G.B.; Marcelić, M.; Lučin, H.M.; Lučin, P. Dynamin Inhibitors Prevent the Establishment of the Cytomegalovirus Assembly Compartment in the Early Phase of Infection. Life 2021, 11, 876.
- Hasan, M.H.; Davis, L.E.; Bollavarapu, R.K.; Mitra, D.; Parmar, R.; Tandon, R. Dynamin Is Required for Efficient Cytomegalovirus Maturation and Envelopment. J. Virol. 2018, 92, e01418-18.
- McMahon, H.T.; Mills, I.G. COP and Clathrin-Coated Vesicle Budding: Different Pathways, Common Approaches. Curr. Opin. Cell Biol. 2004, 16, 379–391.
- Tehran, D.A.; López-Hernández, T.; Maritzen, T. Endocytic Adaptor Proteins in Health and Disease: Lessons from Model Organisms and Human Mutations. Cells 2019, 8, 1345.
More
Information
Subjects:
Virology; Cell Biology
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
914
Revisions:
4 times
(View History)
Update Date:
22 Sep 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





