
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Gabriella Minchiotti | -- | 2397 | 2022-08-15 04:38:26 | | | |
| 2 | Vivi Li | Meta information modification | 2397 | 2022-08-15 05:36:28 | | |
Video Upload Options
An imbalance in proline metabolism shifts mouse ESCs toward a stable naïve-to-primed intermediate state of pluripotency. Proline-induced cells (PiCs), also named primitive ectoderm-like cells (EPLs), are phenotypically metastable, a trait linked to a rapid and reversible relocalization of E-cadherin from the plasma membrane to intracellular membrane compartments. The ESC-to-PiC transition relies on the activation of Erk and Tgfβ/Activin signaling pathways and is associated with extensive remodeling of the transcriptome, metabolome and epigenome. PiCs maintain several properties of naïve pluripotency (teratoma formation, blastocyst colonization and 3D gastruloid development) and acquire a few traits of primed cells (flat-shaped colony morphology, aerobic glycolysis metabolism and competence for primordial germ cell fate).
1. Introduction
2. Proline Levels Influence Pluripotency
2.1. Phenotypic Heterogeneity and Metastability of PiCs
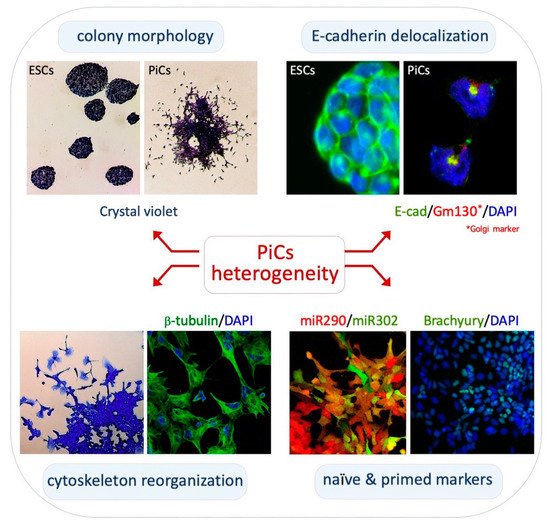
2.2. Transcriptomic and Epigenomic Landscapes of PiCs
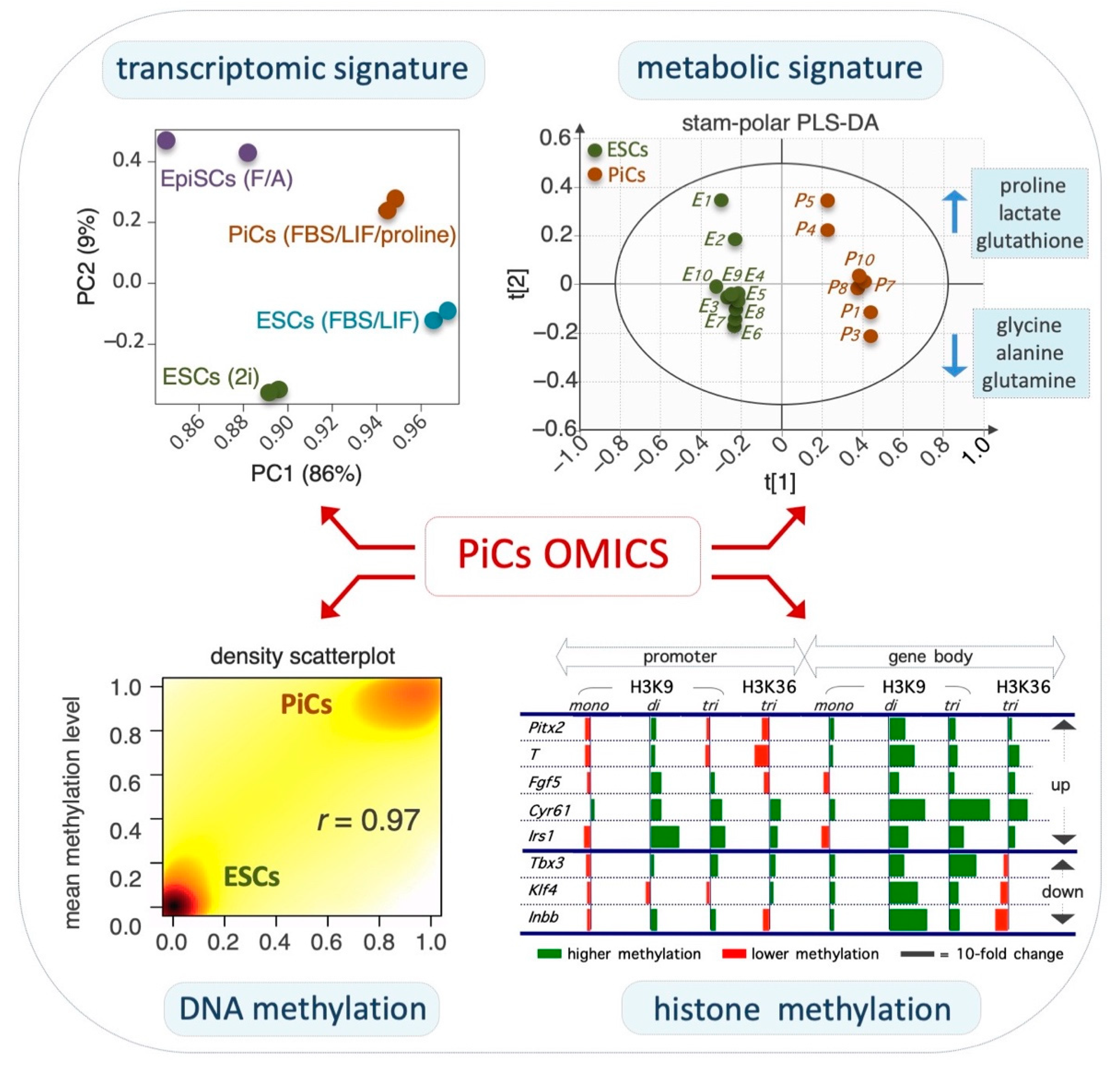
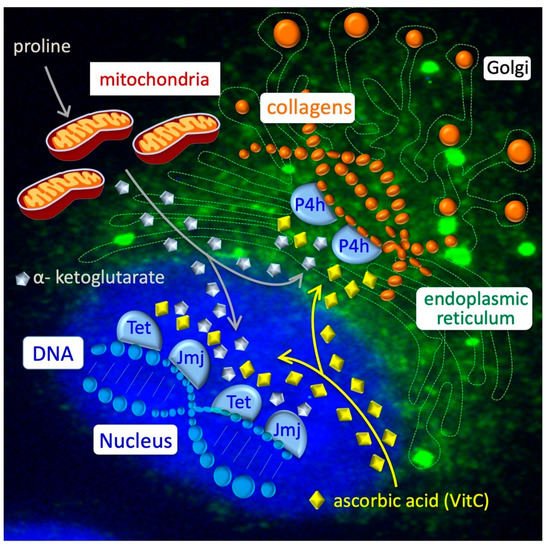
References
- Dundes, C.E.; Loh, K.M. Bridging naive and primed pluripotency. Nat. Cell Biol. 2020, 22, 513–515.
- Nichols, J.; Smith, A. Naive and primed pluripotent states. Cell Stem Cell 2009, 4, 487–492.
- Neagu, A.; van Genderen, E.; Escudero, I.; Verwegen, L.; Kurek, D.; Lehmann, J.; Stel, J.; Dirks, R.A.M.; van Mierlo, G.; Maas, A.; et al. In Vitro capture and characterization of embryonic rosette-stage pluripotency between naive and primed states. Nat. Cell Biol. 2020, 22, 534–545.
- Ying, Q.L.; Wray, J.; Nichols, J.; Batlle-Morera, L.; Doble, B.; Woodgett, J.; Cohen, P.; Smith, A. The ground state of embryonic stem cell self-renewal. Nature 2008, 453, 519–523.
- Brons, I.G.; Smithers, L.E.; Trotter, M.W.; Rugg-Gunn, P.; Sun, B.; Chuva de Sousa Lopes, S.M.; Howlett, S.K.; Clarkson, A.; Ahrlund-Richter, L.; Pedersen, R.A.; et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature 2007, 448, 191–195.
- Tesar, P.J.; Chenoweth, J.G.; Brook, F.A.; Davies, T.J.; Evans, E.P.; Mack, D.L.; Gardner, R.L.; McKay, R.D. New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature 2007, 448, 196–199.
- Smith, A. Formative pluripotency: The executive phase in a developmental continuum. Development 2017, 144, 365–373.
- Morgani, S.; Nichols, J.; Hadjantonakis, A.K. The many faces of Pluripotency: In vitro adaptations of a continuum of in vivo states. BMC Dev. Biol. 2017, 17, 7.
- Pera, M.F.; Rossant, J. The exploration of pluripotency space: Charting cell state transitions in peri-implantation development. Cell Stem Cell 2021, 28, 1896–1906.
- Kinoshita, M.; Smith, A. Pluripotency Deconstructed. Dev. Growth Differ. 2018, 60, 44–52.
- Lackner, A.; Sehlke, R.; Garmhausen, M.; Giuseppe Stirparo, G.; Huth, M.; Titz-Teixeira, F.; van der Lelij, P.; Ramesmayer, J.; Thomas, H.F.; Ralser, M.; et al. Cooperative genetic networks drive embryonic stem cell transition from naive to formative pluripotency. EMBO J. 2021, 40, e105776.
- Kalkan, T.; Olova, N.; Roode, M.; Mulas, C.; Lee, H.J.; Nett, I.; Marks, H.; Walker, R.; Stunnenberg, H.G.; Lilley, K.S.; et al. Tracking the embryonic stem cell transition from ground state pluripotency. Development 2017, 144, 1221–1234.
- Ohinata, Y.; Ohta, H.; Shigeta, M.; Yamanaka, K.; Wakayama, T.; Saitou, M. A signaling principle for the specification of the germ cell lineage in mice. Cell 2009, 137, 571–584.
- Saitou, M. Mammalian Germ Cell Development: From Mechanism to In Vitro Reconstitution. Stem Cell Rep. 2021, 16, 669–680.
- Wang, X.; Xiang, Y.; Yu, Y.; Wang, R.; Zhang, Y.; Xu, Q.; Sun, H.; Zhao, Z.A.; Jiang, X.; Wang, X.; et al. Formative pluripotent stem cells show features of epiblast cells poised for gastrulation. Cell Res. 2021, 31, 526–541.
- Kinoshita, M.; Barber, M.; Mansfield, W.; Cui, Y.; Spindlow, D.; Stirparo, G.G.; Dietmann, S.; Nichols, J.; Smith, A. Capture of Mouse and Human Stem Cells with Features of Formative Pluripotency. Cell Stem Cell 2021, 28, 453–471.
- Yu, L.; Wei, Y.; Sun, H.X.; Mahdi, A.K.; Pinzon Arteaga, C.A.; Sakurai, M.; Schmitz, D.A.; Zheng, C.; Ballard, E.D.; Li, J.; et al. Derivation of Intermediate Pluripotent Stem Cells Amenable to Primordial Germ Cell Specification. Cell Stem Cell 2021, 28, 550–567.
- Han, D.W.; Tapia, N.; Joo, J.Y.; Greber, B.; Arauzo-Bravo, M.J.; Bernemann, C.; Ko, K.; Wu, G.; Stehling, M.; Do, J.T.; et al. Epiblast stem cell subpopulations represent mouse embryos of distinct pregastrulation stages. Cell 2010, 143, 617–627.
- Chang, K.H.; Li, M. Clonal isolation of an intermediate pluripotent stem cell state. Stem Cells 2013, 31, 918–927.
- Wei, M.; Chen, Y.; Zhao, C.; Zheng, L.; Wu, B.; Chen, C.; Li, X.; Bao, S. Establishment of Mouse Primed Stem Cells by Combination of Activin and LIF Signaling. Front. Cell Dev. Biol. 2021, 9, 713503.
- Tsukiyama, T.; Ohinata, Y. A modified EpiSC culture condition containing a GSK3 inhibitor can support germline-competent pluripotency in mice. PLoS ONE 2014, 9, e95329.
- Bao, S.; Tang, W.W.; Wu, B.; Kim, S.; Li, J.; Li, L.; Kobayashi, T.; Lee, C.; Chen, Y.; Wei, M.; et al. Derivation of hypermethylated pluripotent embryonic stem cells with high potency. Cell Res. 2018, 28, 22–34.
- Du, P.; Pirouz, M.; Choi, J.; Huebner, A.J.; Clement, K.; Meissner, A.; Hochedlinger, K.; Gregory, R.I. An Intermediate Pluripotent State Controlled by MicroRNAs is Required for the Naive-to-Primed Stem Cell Transition. Cell Stem Cell 2018, 22, 851–864.
- Cornacchia, D.; Zhang, C.; Zimmer, B.; Chung, S.Y.; Fan, Y.; Soliman, M.A.; Tchieu, J.; Chambers, S.M.; Shah, H.; Paull, D.; et al. Lipid Deprivation Induces a Stable, Naive-to-Primed Intermediate State of Pluripotency in Human PSCs. Cell Stem Cell 2019, 25, 120–136.
- Zhou, W.; Choi, M.; Margineantu, D.; Margaretha, L.; Hesson, J.; Cavanaugh, C.; Blau, C.A.; Horwitz, M.S.; Hockenbery, D.; Ware, C.; et al. HIF1alpha induced switch from bivalent to exclusively glycolytic metabolism during ESC-to-EpiSC/hESC transition. EMBO J. 2012, 31, 2103–2116.
- Fiorenzano, A.; Pascale, E.; D’Aniello, C.; Acampora, D.; Bassalert, C.; Russo, F.; Andolfi, G.; Biffoni, M.; Francescangeli, F.; Zeuner, A.; et al. Cripto is essential to capture mouse epiblast stem cell and human embryonic stem cell pluripotency. Nat. Commun. 2016, 7, 12589.
- D’Aniello, C.; Cermola, F.; Patriarca, E.J.; Minchiotti, G. Metabolic-Epigenetic Axis in Pluripotent State Transitions. Epigenomes 2019, 3, 13.
- Reid, M.A.; Dai, Z.; Locasale, J.W. The impact of cellular metabolism on chromatin dynamics and epigenetics. Nat. Cell Biol. 2017, 19, 1298–1306.
- Ryall, J.G.; Cliff, T.; Dalton, S.; Sartorelli, V. Metabolic Reprogramming of Stem Cell Epigenetics. Cell Stem Cell 2015, 17, 651–662.
- Wang, J.; Alexander, P.; Wu, L.; Hammer, R.; Cleaver, O.; McKnight, S.L. Dependence of mouse embryonic stem cells on threonine catabolism. Science 2009, 325, 435–439.
- Shiraki, N.; Shiraki, Y.; Tsuyama, T.; Obata, F.; Miura, M.; Nagae, G.; Aburatani, H.; Kume, K.; Endo, F.; Kume, S. Methionine metabolism regulates maintenance and differentiation of human pluripotent stem cells. Cell Metab. 2014, 19, 780–794.
- Casalino, L.; Comes, S.; Lambazzi, G.; De Stefano, B.; Filosa, S.; De Falco, S.; De Cesare, D.; Minchiotti, G.; Patriarca, E.J. Control of embryonic stem cell metastability by L-proline catabolism. J. Mol. Cell Biol. 2011, 3, 108–122.
- Glover, H.J.; Shparberg, R.A.; Morris, M.B. L-Proline Supplementation Drives Self-Renewing Mouse Embryonic Stem Cells to a Partially Primed Pluripotent State: The Early Primitive Ectoderm-Like Cell. Methods Mol. Biol. 2022, 2490, 11–24.
- Comes, S.; Gagliardi, M.; Laprano, N.; Fico, A.; Cimmino, A.; Palamidessi, A.; De Cesare, D.; De Falco, S.; Angelini, C.; Scita, G.; et al. L-Proline induces a mesenchymal-like invasive program in embryonic stem cells by remodeling H3K9 and H3K36 methylation. Stem Cell Rep. 2013, 1, 307–321.
- Cermola, F.; D’Aniello, C.; Tate, R.; De Cesare, D.; Martinez-Arias, A.; Minchiotti, G.; Patriarca, E.J. Gastruloid Development Competence Discriminates Different States of Pluripotency. Stem Cell Rep. 2021, 16, 354–369.
- D’Aniello, C.; Habibi, E.; Cermola, F.; Paris, D.; Russo, F.; Fiorenzano, A.; Di Napoli, G.; Melck, D.J.; Cobellis, G.; Angelini, C.; et al. Vitamin C and l-Proline Antagonistic Effects Capture Alternative States in the Pluripotency Continuum. Stem Cell Rep. 2017, 8, 1–10.
- D’Aniello, C.; Fico, A.; Casalino, L.; Guardiola, O.; Di Napoli, G.; Cermola, F.; De Cesare, D.; Tate, R.; Cobellis, G.; Patriarca, E.J.; et al. A novel autoregulatory loop between the Gcn2-Atf4 pathway and (L)-Proline metabolism controls stem cell identity. Cell Death Differ. 2015, 22, 1094–1105.
- Fico, A.; Fiorenzano, A.; Pascale, E.; Patriarca, E.J.; Minchiotti, G. Long non-coding RNA in stem cell pluripotency and lineage commitment: Functions and evolutionary conservation. Cell Mol. Life Sci. 2019, 76, 1459–1471.
- Fiorenzano, A.; Pascale, E.; Patriarca, E.J.; Minchiotti, G.; Fico, A. LncRNAs and PRC2: Coupled Partners in Embryonic Stem Cells. Epigenomes 2019, 3, 14.
- Fiorenzano, A.; Pascale, E.; Gagliardi, M.; Terreri, S.; Papa, M.; Andolfi, G.; Galasso, M.; Tagliazucchi, G.M.; Taccioli, C.; Patriarca, E.J.; et al. An Ultraconserved Element Containing lncRNA Preserves Transcriptional Dynamics and Maintains ESC Self-Renewal. Stem Cell Rep. 2018, 10, 1102–1114.
- Pascale, E.; Beclin, C.; Fiorenzano, A.; Andolfi, G.; Erni, A.; De Falco, S.; Minchiotti, G.; Cremer, H.; Fico, A. Long Non-coding RNA T-UCstem1 Controls Progenitor Proliferation and Neurogenesis in the Postnatal Mouse Olfactory Bulb through Interaction with miR-9. Stem Cell Rep. 2020, 15, 836–844.
- Parchem, R.J.; Ye, J.; Judson, R.L.; LaRussa, M.F.; Krishnakumar, R.; Blelloch, A.; Oldham, M.C.; Blelloch, R. Two miRNA clusters reveal alternative paths in late-stage reprogramming. Cell Stem Cell 2014, 14, 617–631.
- Habibi, E.; Stunnenberg, H.G. Transcriptional and epigenetic control in mouse pluripotency: Lessons from in vivo and in vitro studies. Curr. Opin. Genet. Dev. 2017, 46, 114–122.
- Lu, F.; Liu, Y.; Jiang, L.; Yamaguchi, S.; Zhang, Y. Role of Tet proteins in enhancer activity and telomere elongation. Genes Dev. 2014, 28, 2103–2119.
- Pandolfini, L.; Luzi, E.; Bressan, D.; Ucciferri, N.; Bertacchi, M.; Brandi, R.; Rocchiccioli, S.; D’Onofrio, M.; Cremisi, F. RISC-mediated control of selected chromatin regulators stabilizes ground state pluripotency of mouse embryonic stem cells. Genome Biol. 2016, 17, 94.




