| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Haiyan Dai | + 914 word(s) | 914 | 2020-10-09 16:25:38 | | | |
| 2 | Conner Chen | Meta information modification | 914 | 2020-10-30 11:01:00 | | |
Video Upload Options
Myeloid-derived suppressor cells (MDSCs) are major immunosuppressive cells in the tumor microenvironment (TME). During the differentiation and development of MDSCs from myeloid progenitor cells, their functions are also affected by a series of regulatory factors in the TME, such as metabolic reprogramming, epigenetic modification, and cell signaling pathways. And there is a crosstalk between these regulatory factors. In cancer, there are important bidirectional regulatory mechanisms between metabolic remodeling and epigenome. Most chromatin modifying enzymes require cell metabolic intermediates as substrates or cofactors. In turn, epigenetic modifications regulate cell metabolism and function to some extent. The regulation between metabolism and epigenetic modification of MDSCs can be achieved through signaling pathways related to AMPK and HIF-1α,etc.
1. Connections between Metabolism and Epigenetic Modification in MDSCs
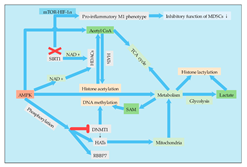
There are important bidirectional regulatory mechanisms between metabolic remodeling and the epigenome in cancer. Most chromatin modifying enzymes are required as substrates or cofactors for cell metabolic intermediates. Such metabolites, and the enzymes that normally produce them, can be transferred to the nucleus, directly linking metabolism to nuclear transcription [1]. Therefore, the regulation of MDSCs by epigenetic modification will not only change the differentiation and metabolism of MDSCs themselves but also affect the metabolism of other cells in the TME, enabling these cells to undergo metabolic reprogramming (Figure 1). The epigenome is also sensitive to metabolic states. Metabolites act as cofactors or substrates for several important enzyme reactions related to epigenetic modification and gene regulation [2]. For example, central metabolites are substrates that catalyze the deposition of covalently modified enzymes on histones, DNA, and RNA [3]. Lactate, for example, is an endogenous inhibitor of histone deacetylase, which regulates some genes at the transcriptional level [4]. Therefore, it is particularly important to study the relationship between metabolism and epigenetic modification of MDSCs. At present, epigenetics has been widely studied in the differentiation, proliferation, function, and other aspects of MDSCs, but there are few reports on its effect on metabolism. However, with the in-depth understanding of MDSC metabolism and the important role of cell metabolism on cell function, an increasing amount of studies have begun to focus on the regulation of MDSC metabolism by epigenetic modification.
Figure 1. Connections between epigenetic modification and metabolism in MDSCs. Acetyl-CoA, SAM, and lactate, as important substrates and group donors of epigenetic modification, are important pivot substances connecting metabolism and epigenetic modification and have important regulatory effects on epigenetic modification. AMPK, as an energy receptor, can not only regulate metabolism but also regulate some corresponding epigenetic modifications through the regulation of epigenetic modification enzymes such as DNMT, HATs, and HDACs. Moreover, AMPK regulates the metabolism of MDSCs by regulating histone acetylation and DNA methylation.
2. AMPK and HIF-1α Mediate the Association between Epigenetic Modification and Metabolism
2.1. AMPK
The tumorigenesis of MDSCs is mediated in part by the activation of AMPK [5], and this effect was achieved to some extent through the metabolic regulation of MDSCs by AMPK. One of the mechanisms by which AMPK regulates metabolism is histone acetylation. AMPK regulates histone acetylation through a variety of mechanisms. Activation of AMPK can increase the level of acetyl-CoA. Since acetyl-CoA is a substrate for lysine acetyltransferase (KATS), AMPK can affect the activity of KATS by regulating the cell level of acetyl-CoA. In addition, AMPK can activate HDACs by increasing the cell concentration of NAD+, a cofactor of HDACs [6]. AMPK also regulates DNA methylation and histone acetylation by phosphorylating epigenetic factors such as DNMT1, retinoblastoma binding protein 7 (RBBP7), and HAT1, thereby promoting mitochondrial biosynthesization and function. AMPK-mediated phosphorylation leads to the activation of HAT1 and inhibition of DNMT1 [7]. Therefore, the regulation of AMPK to regulate the metabolic pathways of MDSCs is achieved to a certain extent through epigenetic regulation, while epigenetic modification is also affected by the level of cell metabolism.
2.2. HIF-1α
Epigenetic reprogramming of myeloid cells, also known as training immunity. Induction of aerobic glycolysis by AKT-mTOR-HIF-1α is the metabolic basis of training immunity [8]. Therefore, HIF-1α is also a key regulatory point related to glucose metabolism and epigenetic modification, plays a very important role in metabolic reprogramming of MDSCs, and different cell signaling pathways play a certain role in this process. Since MDSCs exhibit an immature phenotype, the tumor may exhibit either a typically activated phenotype M1 or an alternatively activated phenotype M2. Studies have found that when MDSCs enter the periphery from the bone marrow, SIRT1 (an enzyme responsible for regulating the deacetylation of proteins) deficiency leads to a specific M1 lineage switch, which reduces the inhibitory function of MDSCs and is conducive to the pro-inflammatory M1 phenotype. Differentiation into M1 phenotype requires glycolysis activation of the mTOR-HIF-1 pathway to provide tumor protection. The study identified the nature of the SIRT1-mTOR/HIF-1 glycolysis pathway in determining the differentiation of MDSCs and suggested metabolic reprogramming as a cancer treatment. It is interesting that SIRT1 is a class of NAD+ dependent histone deacetylases that are widespread in life [9]. Therefore, phenotypic switching and metabolic reprogramming of MDSCs are likely to be strongly associated with histone deacetylation, and the association needs to be further studied.
3. Limitations
Although AMPK and HIF-1α can regulate epigenetic modifications such as DNA methylation and histone acetylation, the activation of AMPK and HIF-1α or related signaling pathways can regulate metabolism and function of MDSCs. However, few studies have pointed out that epigenetic modification directly regulates the metabolism of MDSCs. Perhaps in the future, with the in-depth study of MDSCs and the further development of epigenetics, epigenetic modification will make new breakthroughs in regulating the metabolism of MDSCs.
References
- Kinnaird, A.; Zhao, S.; Wellen, K.E.; Michelakis, E.D. Metabolic control of epigenetics in cancer. Nat. Rev. Cancer 2016, 16, 694–707, doi:10.1038/nrc.2016.82.
- Thakur, C.; Chen, F. Connections between metabolism and epigenetics in cancers. Semin. Cancer Biol. 2019, 57, 52–58, doi:10.1016/j.semcancer.2019.06.006.
- Sharma, U.; Rando, O.J. Metabolic Inputs into the Epigenome. Cell Metab. 2017, 25, 544–558, doi:10.1016/j.cmet.2017.02.003.
- Latham, T.; Mackay, L.; Sproul, D.; Karim, M.; Culley, J.; Harrison, D.J.; Hayward, L.; Langridge-Smith, P.; Gilbert, N.; Ramsahoye, B.H. Lactate, a product of glycolytic metabolism, inhibits histone deacetylase activity and promotes changes in gene expression. Nucleic Acids Res. 2012, 40, 4794–4803, doi:10.1093/nar/gks066.
- De Veirman, K.; Menu, E.; Maes, K.; De Beule, N.; De Smedt, E.; Maes, A.; Vlummens, P.; Fostier, K.; Kassambara, A.; Moreaux, J.; et al. Myeloid-derived suppressor cells induce multiple myeloma cell survival by activating the AMPK pathway. Cancer Lett. 2019, 442, 233–241, doi:10.1016/j.canlet.2018.11.002.
- Vancura, A.; Nagar, S.; Kaur, P.; Bu, P.; Bhagwat, M.; Vancurova, I. Reciprocal Regulation of AMPK/SNF1 and Protein Acetylation. Int. J. Mol. Sci. 2018, 19, 3314, doi:10.3390/ijms19113314.
- Marin, T.L.; Gongol, B.; Zhang, F.; Martin, M.; Johnson, D.A.; Xiao, H.; Wang, Y.; Subramaniam, S.; Chien, S.; Shyy, J.Y. AMPK promotes mitochondrial biogenesis and function by phosphorylating the epigenetic factors DNMT1, RBBP7, and HAT1. Sci. Signal. 2017, 10, aaf7478, doi:10.1126/scisignal.aaf7478.
- Cheng, S.C.; Quintin, J.; Cramer, R.A.; Shepardson, K.M.; Saeed, S.; Kumar, V.; Giamarellos-Bourboulis, E.J.; Martens, J.H.; Rao, N.A.; Aghajanirefah, A.; et al. mTOR- and HIF-1α-mediated aerobic glycolysis as metabolic basis for trained immunity. Science 2014, 345, 1250684, doi:10.1126/science.1250684.
- Liu, G.; Bi, Y.; Shen, B.; Yang, H.; Zhang, Y.; Wang, X.; Liu, H.; Lu, Y.; Liao, J.; Chen, X.; et al. SIRT1 limits the function and fate of myeloid-derived suppressor cells in tumors by orchestrating HIF-1α-dependent glycolysis. Cancer Res. 2014, 74, 727–737, doi:10.1158/0008-5472.Can-13-2584.