Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Peiwen Zhang | -- | 1492 | 2022-07-25 09:07:00 | | | |
| 2 | Catherine Yang | + 6 word(s) | 1498 | 2022-07-27 05:13:23 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Zhang, P.; Wu, S.; He, Y.; Li, X.; Zhu, Y.; Lin, X.; Chen, L.; Zhao, Y.; Niu, L.; Zhang, S.; et al. LncRNA-Mediated Adipogenesis in Different Adipocytes. Encyclopedia. Available online: https://encyclopedia.pub/entry/25477 (accessed on 26 April 2026).
Zhang P, Wu S, He Y, Li X, Zhu Y, Lin X, et al. LncRNA-Mediated Adipogenesis in Different Adipocytes. Encyclopedia. Available at: https://encyclopedia.pub/entry/25477. Accessed April 26, 2026.
Zhang, Peiwen, Shuang Wu, Yuxu He, Xinrong Li, Yan Zhu, Xutao Lin, Lei Chen, Ye Zhao, Lili Niu, Shunhua Zhang, et al. "LncRNA-Mediated Adipogenesis in Different Adipocytes" Encyclopedia, https://encyclopedia.pub/entry/25477 (accessed April 26, 2026).
Zhang, P., Wu, S., He, Y., Li, X., Zhu, Y., Lin, X., Chen, L., Zhao, Y., Niu, L., Zhang, S., Li, X., Zhu, L., & Shen, L. (2022, July 25). LncRNA-Mediated Adipogenesis in Different Adipocytes. In Encyclopedia. https://encyclopedia.pub/entry/25477
Zhang, Peiwen, et al. "LncRNA-Mediated Adipogenesis in Different Adipocytes." Encyclopedia. Web. 25 July, 2022.
Copy Citation
Long-chain noncoding RNAs (lncRNAs) are RNAs that do not code for proteins, widely present in eukaryotes. They regulate gene expression at multiple levels through different mechanisms at epigenetic, transcription, translation, and the maturation of mRNA transcripts or regulation of the chromatin structure, and compete with microRNAs for binding to endogenous RNA.
long noncoding RNA
brown fat
ectopic fat
1. Introduction
About only 1–2% of human DNA is transcribed into mRNAs that code for proteins, whereas the rest is coded into noncoding RNAs (ncRNAs) [1]. The noncoding RNAs include snRNA (small nuclear RNA), siRNA (small interfering RNA), miRNA (microRNA), lncRNA (long noncoding RNAs), and circRNA (circular RNA), among others [2]. lncRNAs are noncoding RNAs present in numerous eukaryotes, first reported in 1990. Since a few lncRNAs can code for proteins, the naming and definition of long noncoding RNAs are still controversial [3]. For a long time, lncRNAs have been considered redundant transcripts, commonly called “noise sequences” [4]. However, emerging evidence shows that lncRNAs have strong temporal and spatial expression specificity and tissue specificity [5], are poorly conserved among species [6][7], and regulate numerous biological processes [8]. lncRNAs regulate gene expression at multiple levels [9]. Advances in high-throughput sequencing and other biotechnology have revealed the molecular mechanisms by which lncRNA regulates biological functions [10]. At present, the functions of lncRNAs are yet to be exhausted, and much remains to be discovered.
Obesity is currently a public health concern globally. According to the World Health Organization (WHO), 1.9 billion adults (39%) are overweight (BMI > 25) and 600 million people (13%) are obese (BMI > 30). Obesity metabolic syndrome is caused by excessive energy intake, in which the triglycerides that accumulate in the body are transformed into fat [11][12][13]. Obesity is a chronic recurrent disease characterized by excessive accumulation of fats in the body. The pathogenesis of obesity is quite complex, and many factors, including genetics, viral infections, insulin resistance, inflammation, gut microorganisms, and abnormal hormone secretion are implicated in obesity development. Chronic obesity causes systemic metabolic diseases, high blood pressure, hip cartilage, and other bone and joint diseases. Accumulation of large amounts of visceral fat can cause insulin resistance, fatty liver disease, type 2 diabetes, liver fibrosis, liver cancer, and other diseases. Similarly, high body fat can cause hypertriglyceridemia and irreversible chronic diseases such as atherosclerosis. At the same time, obesity predisposes childbearing age women to polycystic ovary syndrome or gestational diabetes mellitus, increases the maternal metabolic burden, and has irreversible effects on methylation of maternally imprinted genes in offspring. In addition, obesity appears to increase the severity of Corona Virus Disease 2019, (COVID-19) [14][15]. Obesity and vitamin deficiency are major risk factors for poor prognosis after COVID-19 infection. In addition, obesity, diabetes, and hypertension increase the risk of COVID-19 infection and severe COVID-19 [15].
2. LncRNA
lncRNAs are small RNA transcripts greater than 200 nt. lncRNAs can be divided into seven types based on the transcription position from the genome (Figure 1). Intergenic lncRNAs (intergenic lncRNA), also called lincRNAs, are intergenic region lncRNAs, transcribed from the middle region of two coding genes and at least 1 kb away from the coding gene. They mainly regulate cellular activities. Intron lncRNAs (intronic lncRNAs), mainly produced in the intron region of the coding gene, have corresponding coding genes and the same expression pattern, and they primarily regulate the expression of genes. Antisense lncRNA (antisense lncRNA), mainly produced in the antisense strand of the coding strand, binds and regulates mRNAs’ expression. Sense lncRNAs (sense-overlapping lncRNA) are transcribed in a direction similar to that of adjacent mRNA and partial or complete overlap exons. They contain an open reading frame (ORF) for protein translation but do not code for protein, restricting the mRNA translation process by mediating the stop codon; promoter-associated lncRNA and untranslated-region overlapping lncRNA (untranslated-region overlapping lncRNA) bind to the promoter and untranslated region of the regulated mRNA. Enhancer sub-type lncRNA (enhancer lncRNA) mainly regulates the expression of neighboring genes through binding cis-regulatory sites or enhancers [16][17][18][19].
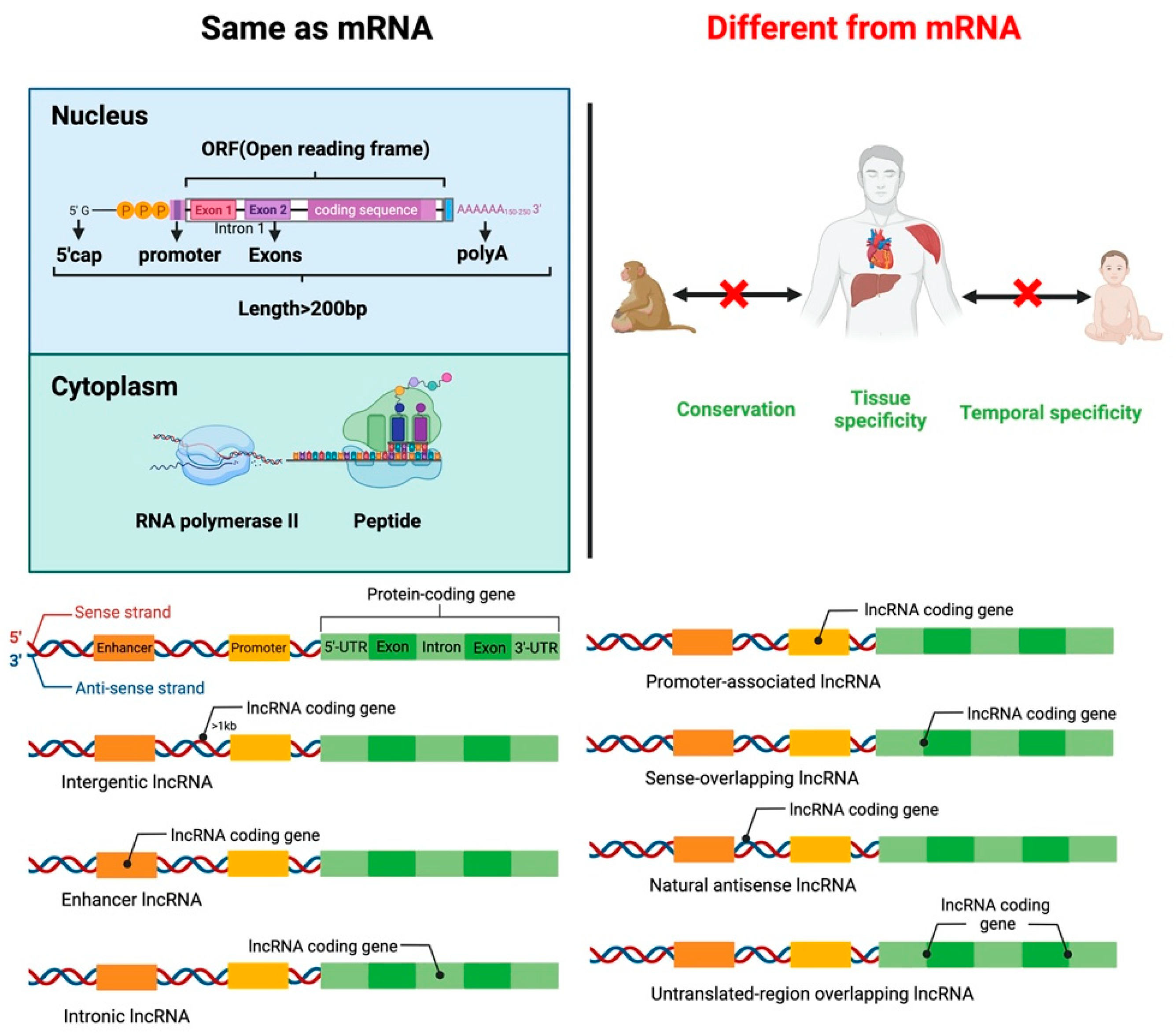
Figure 1. Similarities and differences between lncRNA and mRNA structure and function. Classification of lncRNAs according to the transcription position of the genome.
Like mRNA-encoding protein, lncRNA has a 5’cap and a 3’poly A tail. lncRNAs use the same gene as a transcription template to form different lncRNA transcripts by variable shearing. Unlike mRNA, lncRNA has strong tissue specificity, and its abundance is lower than that of mRNA. lncRNA exists in the nucleus, cytoplasm, and organelles, but they are more abundant in the nucleus than in the cytoplasm and organelles [20][21][22]. lncRNA regulates proliferation, differentiation, apoptosis of cells, and the development of tissues and organs. For example, lncLSTR (liver-specific triglyceride regulator RNA) is an lncRNA specifically expressed in mouse liver and regulates energy metabolism and lipid metabolism in the liver by directly regulating one of the rate-limiting enzymes responsible for bile acid synthesis, Cyp8b1 [23].
3. LncRNA and Adipose Tissue
High-throughput sequencing technology has revealed many adipogenesis and development-related lncRNAs. However, many regulatory pathways related to obesity and lipid deposition have not been fully uncovered. Studies have shown that excessive lipid accumulation in adipose tissue is associated with dysregulated lncRNA expression. The amount of fats also affects the quality of livestock and poultry meat products [24]. Therefore, clarifying how lncRNA participates in fat development can further research in fat development-related diseases and the production of quality meat products. The biological functions of white, brown, and beige fats and lncRNAs related to this process are discussed (Figure 2, Figure 3 and Figure 4). As shown in Figure 2, Figure 3 and Figure 4, lncRNAs are regulated in a variety of ways in adipocytes to regulate lipid deposition or to regulate white fat browning. The most commonly reported mode of regulation is that of ceRNA regulation, where lncRNAs regulate the transcription or translation of target genes by competitively binding miRNAs or proteins, such as lncGAS5 [25]. lncRNAs also regulate lipogenic differentiation by acting as decoy molecules, RNA-RNA dimers, histone modifications, regulation of target-gene-promoter transcriptional activity and signaling pathway activity, and white fat browning, among other processes. In addition, it is worth noting that adipose-derived MSCs, as a class of pluripotent stem cells, are an excellent cell model for studying the biological processes of lipogenic differentiation, but most studies have been conducted on mature adipocytes. However, most of the studies are based on mature adipocytes. There are very few studies on adipose-derived MSCs and adipose-progenitor-cell-related lncRNAs and the regulatory mechanisms are not well studied, such as how lncRNAs are involved in the transduction of signaling pathways and the regulatory relationship with the host gene.
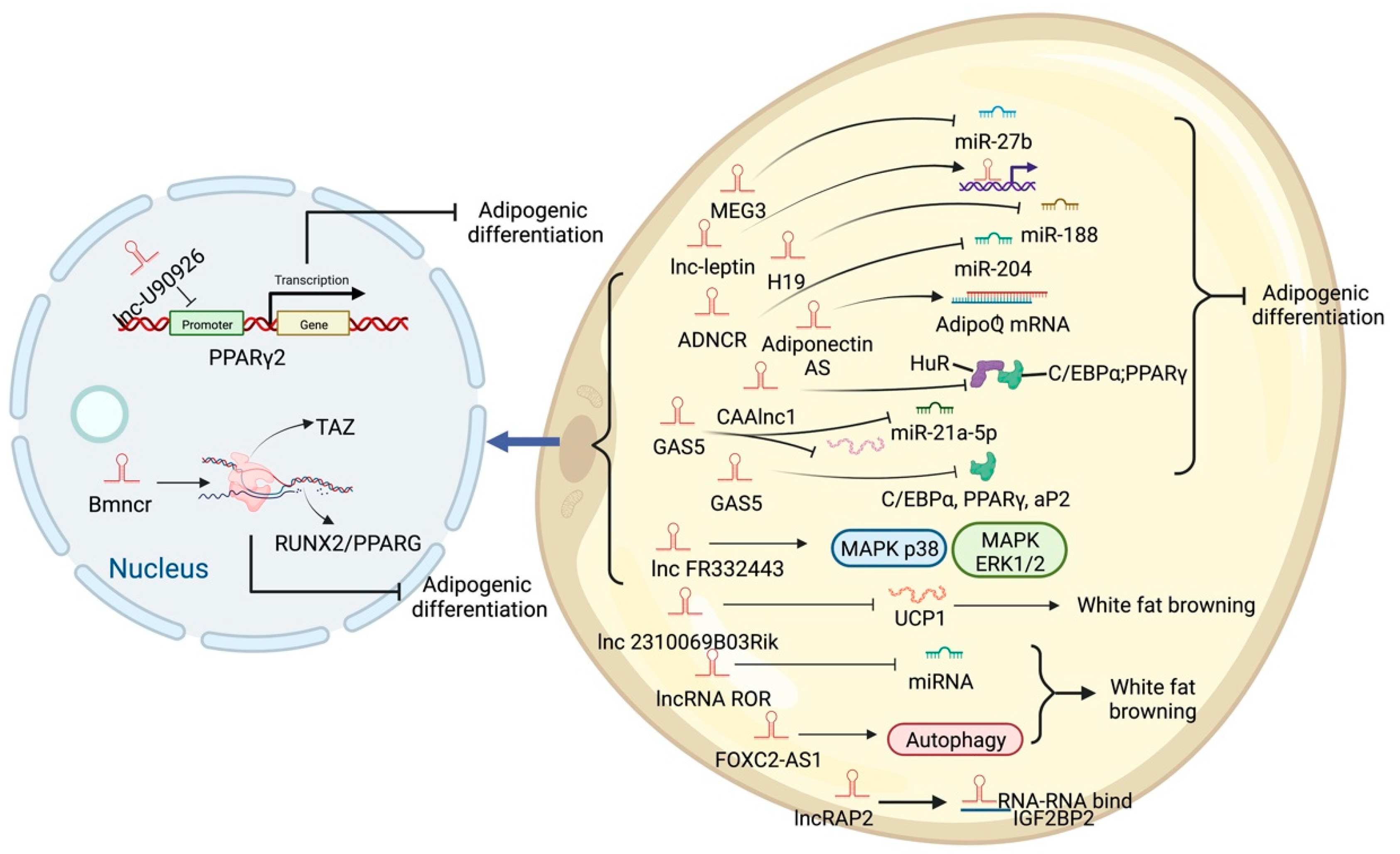
Figure 2. Schematic diagram of the biological function of lncRNA that promotes lipid deposition of white fat.
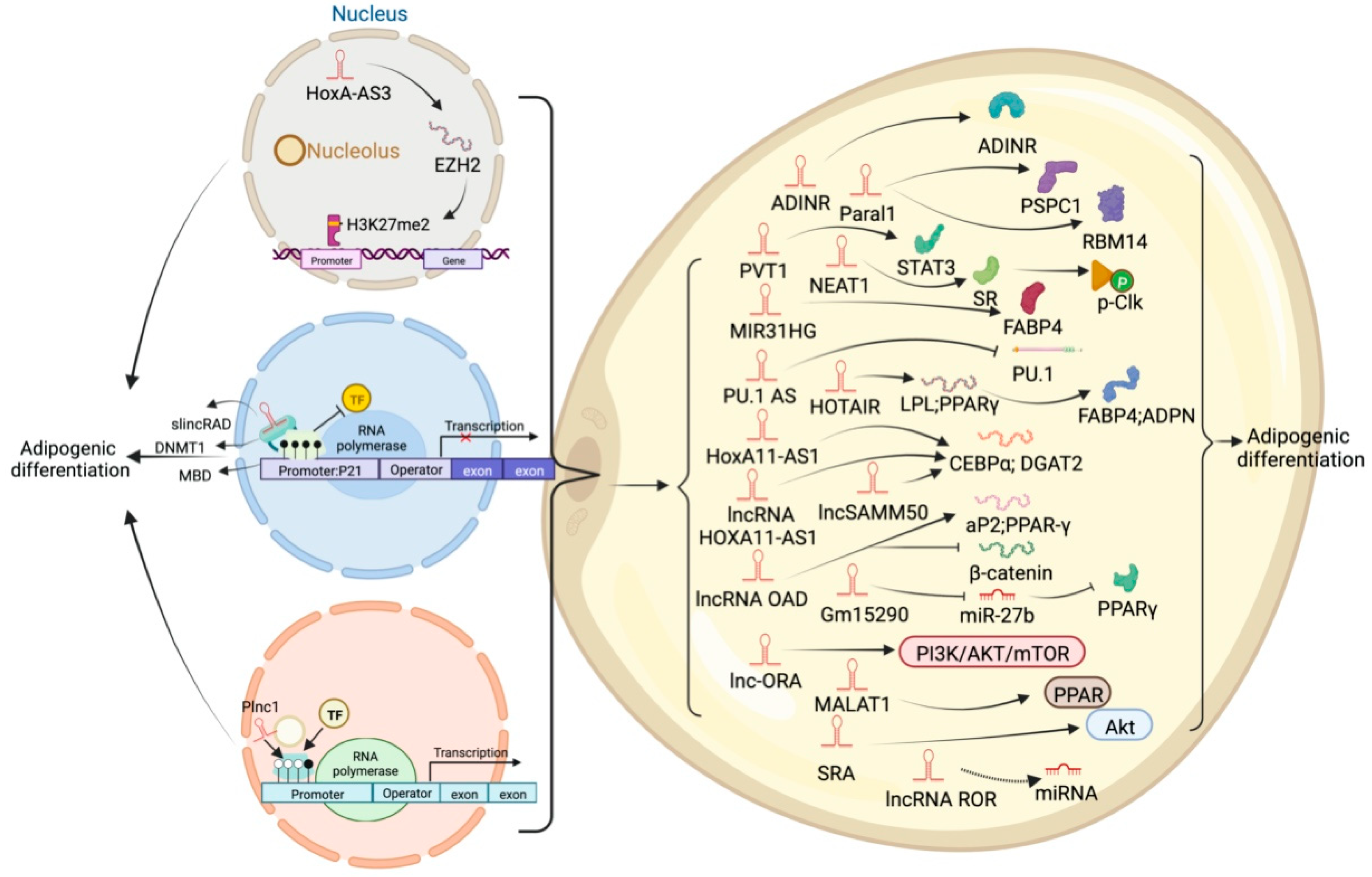
Figure 3. Schematic diagram of the biological function of lncRNA that inhibits or participates in the deposition of white fat lipids.
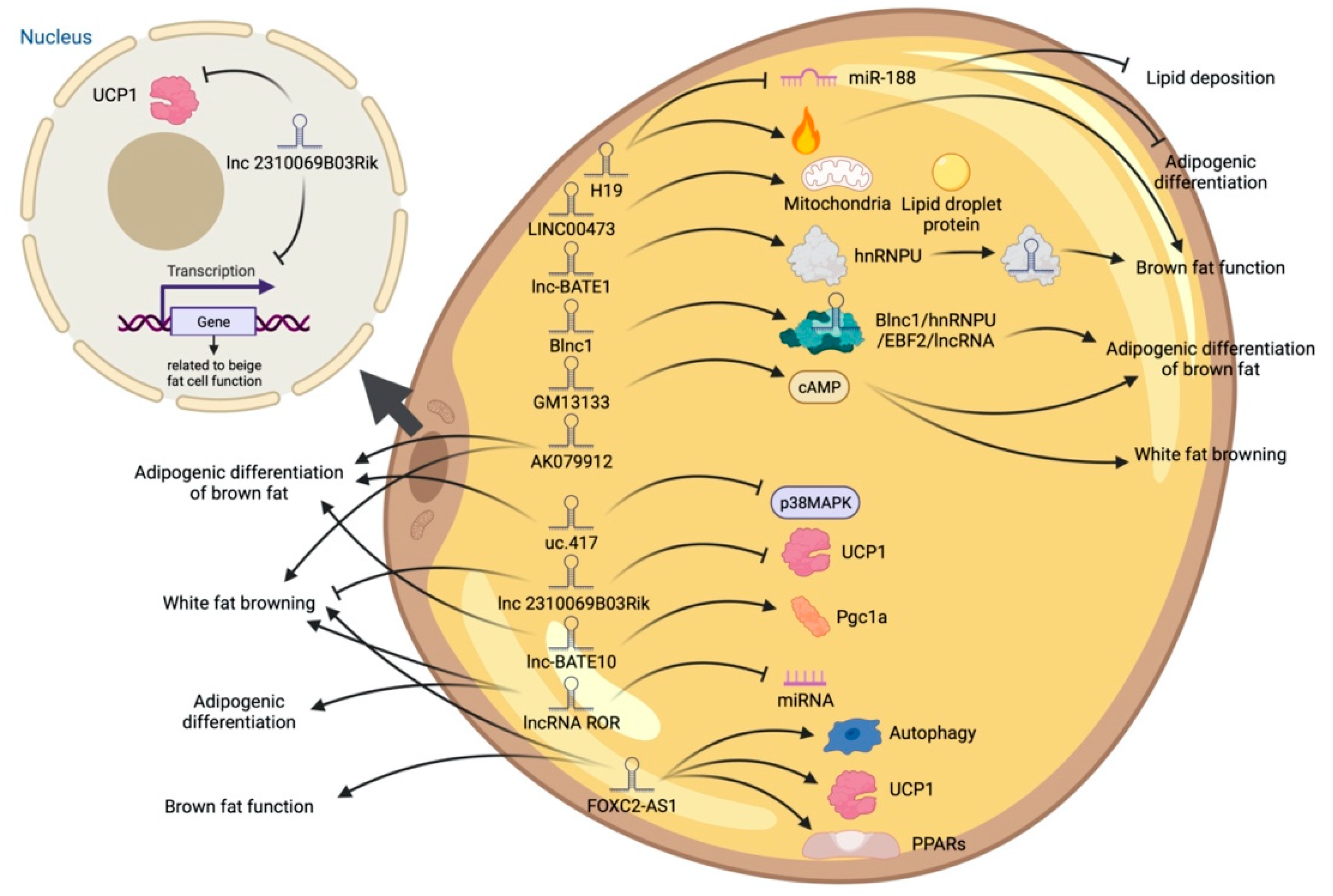
Figure 4. Schematic diagram of the biological function of lncRNA involved in the differentiation of brown adipocytes.
4. LncRNA-Mediated Adipogenesis Dysregulation and Disease
There is no doubt that lncRNAs are an important source of potential targets against obesity and other related metabolic diseases. However, the molecular mechanisms underlying the cooperative regulation of lncRNAs with some nutrients and with classical signaling pathways are still unclear. In addition, most studies have used mouse 3T3-L1 preadipocytes or human-derived, adipose-derived mesenchymal stem cells as models for in vitro experiments, and many of the experimental effects of lncRNAs in vivo have not been validated. Therefore, it is important to investigate the regulatory mechanisms of lncRNAs and the regulation of lipogenic differentiation under pathophysiological conditions, or to explore the association between lncRNAs and certain disease models of lipid metabolism imbalance, which could better facilitate the development of lncRNA-targeted drugs and other related research.
Metabolism-related lipid disorders include atherosclerosis, obesity, Alzheimer’s disease, type II diabetes, tumors, and other diseases [26]. Numerous studies have shown that lncRNAs are involved in biological processes such as metabolic regulation, inflammation, immunity, and vascular function [27][28].
Lipid metabolism in cancer stem cells is important for drug sensitivity and maintenance of stemness in cancer cells. Liu [29] et al. showed that lncROPM enhances the stability of PLA2G16 mRNA by directly binding to the 3’-UTR of PLA2G16, thereby activating active PI3K/AKT, Wnt/β-catenin, and Hippo/YAP signaling, ultimately involved in the maintenance of stemness and the enhancement in chemoresistance in BCSCs. In a mechanistically similar fashion, Hilnc promotes PPARγ expression through the formation of an RNA-RNA stable dimer with IGFBP2 and promotes lipid deposition in the liver [30]. In atherosclerosis, kcnq1ot1 enhances HDAC3 expression by competitively binding to miR-452-3p, thereby inhibiting ABCA1 expression as well as cholesterol efflux. kcnq1ot1 promotes macrophage lipid accumulation and accelerates the development of atherosclerosis via the miR-452-3p/HDAC3/ABCA1 pathway [31]. In addition, as mentioned above, lncRNA is also a good potential biomarker. lnc-P3134 was significantly upregulated in the blood exosomes of patients with type 2 diabetes [32] and, in addition, lncH19 was upregulated in the serum of patients [33]. lncRNAs have a very important impact on lipid metabolism homeostasis, but the specific regulatory mechanisms and biological processes such as lncRNA regulation of lipid production, β-oxidation, and lipid catabolism need to be further investigated. In-depth investigation of the regulatory functions of lncRNAs in lipid metabolism homeostasis has important biological implications for lncRNAs to become biomarkers for certain diseases.
References
- Stefani, G.; Slack, F.J. Small non-coding RNAs in animal development. Nat. Rev. Mol. Cell. Biol. 2008, 9, 219–230.
- Ravasi, T. Experimental validation of the regulated expression of large numbers of non-coding RNAs from the mouse genome. Genome Res. 2006, 16, 11–19.
- Liu, Z.J.; Cao, Y.; Wureli, H.; Wei, N.I.; Wang, D.W.; Sheng-Wei, H.U. Cloning and Protein Encoding Ability Analysis of Sheep lncRNA-MRLN Gene. Prog. Vet. Med. 2018.
- Necsulea, A.; Soumillon, M.; Warnefors, M.; Liechti, A.; Daish, T.; Zeller, U.; Baker, J.C.; Grützner, F.; Kaessmann, H. The evolution of lncRNA repertoires and expression patterns in tetrapods. Nature 2014, 505, 635–640.
- Iwakiri, J.; Terai, G.; Hamada, M. Computational prediction of lncRNA-mRNA interactionsby integrating tissue specificity in human transcriptome. Biol. Direct 2017, 12, 1–8.
- Kosinska-Selbi, B.; Mielczarek, M.; Szyda, J. Review: Long non-coding RNA in livestock. Animal 2020, 14, 2003–2013.
- Bridges, M.C.; Daulagala, A.C.; Kourtidis, A. LNCcation: lncRNA localization and function. J. Cell Biol. 2021, 220.
- Rey, F.; Urrata, V.; Gilardini, L.; Bertoli, S.; Calcaterra, V.; Zuccotti, G.V.; Cancello, R.; Carelli, S. Role of long non-coding RNAs in adipogenesis: State of the art and implications in obesity and obesity-associated diseases. Obes. Rev. 2021, 22, e13203.
- Zhu, J.J.; Fu, H.J.; Wu, Y.G.; Zheng, X.F. Function of lncRNAs and approaches to lncRNA-protein interactions. Sci. China Life Sci. 2013, 56, 876–885.
- Chen, X.; Yan, G.-Y. Novel human lncRNA–disease association inference based on lncRNA expression profiles. Bioinformatics 2013, 29, 2617–2624.
- Verma, M.; Rajput, M.; Kishore, K.; Kathirvel, S. Asian BMI criteria are better than WHO criteria in predicting Hypertension: A cross-sectional study from rural India. J. Fam. Med. Prim. Care 2019, 8, 2095–2100.
- Després, J.; Lemieux, I.; Alméras, N. Abdominal Obesity and the Metabolic Syndrome; Springer: New York, NY, USA, 2006.
- Fernandez, X.; Monin, G.; Talmant, A.; Mourot, J.; Lebret, B. Influence of intramuscular fat content on the quality of pig meat—1. Composition of the lipid fraction and sensory characteristics of m. longissimus lumborum. Meat Sci. 1999, 53, 59–65.
- Karampela, Ι.; Vallianou, N.; Magkos, F.; Apovian, C.M.; Dalamaga, Μ. Obesity, Hypovitaminosis D, and COVID-19: The Bermuda Triangle in Public Health. Curr. Obes. Rep. 2022, 1–10.
- Dalamaga, M.; Christodoulatos, G.S.; Karampela, I.; Vallianou, N.; Apovian, C.M. Understanding the Co-Epidemic of Obesity and COVID-19: Current Evidence, Comparison with Previous Epidemics, Mechanisms, and Preventive and Therapeutic Perspectives. Curr. Obes. Rep. 2021, 10, 214–243.
- Yoon, J.H.; Abdelmohsen, K.; Srikantan, S.; Yang, X.; Martindale, J.L.; De, S.; Huarte, M.; Zhan, M.; Becker, K.G.; Gorospe, M. LincRNA-p21 Suppresses Target mRNA Translation. Mol. Cell 2012, 47, 648–655.
- Zhang, C.L.; Zhu, K.P.; Ma, X.L. Antisense lncRNA FOXC2-AS1 promotes doxorubicin resistance in osteosarcoma by increasing the expression of FOXC2. Cancer Lett. 2017, 396, 66–75.
- Hamazaki, N.; Nakashima, K.; Hayashi, K.; Imamura, T. Detection of Bidirectional Promoter-Derived lncRNAs from Small-Scale Samples Using Pre-Amplification-Free Directional RNA-seq Method. Methods Mol. Biol. 2017, 1605, 83.
- Alvarez-Dominguez, J.R.; Knoll, M.; Gromatzky, A.A.; Lodish, H.F. The Super-Enhancer-Derived alncRNA-EC7/Bloodlinc Potentiates Red Blood Cell Development in trans. Cell Rep. 2017, 19, 2503.
- Blythe, A.J.; Fox, A.; Bond, C.S. The ins and outs of lncRNA structure: How, why and what comes next? Biochim. Biophys. Acta 2016, 1859, 46–58.
- Chengtao, D.U.; Wang, H.; Yang, S.; Xiangchen, L.I.; Zhou, X.; Zhao, A. Study on the Role of lncRNA in the Process of Japanese Encephalitis Virus Infecting PK15 Cells. China Anim. Husb. Vet. Med. 2019, 46, 2045–2052.
- Zhang, K.; Shi, Z.M.; Chang, Y.N.; Hu, Z.M.; Qi, H.X.; Hong, W. The ways of action of long non-coding RNAs in cytoplasm and nucleus. Gene 2014, 547, 1–9.
- Li, P.; Ruan, X.; Yang, L.; Kiesewetter, K.; Zhao, Y.; Luo, H.; Chen, Y.; Gucek, M.; Zhu, J.; Cao, H. A liver-enriched long non-coding RNA, lncLSTR, regulates systemic lipid metabolism in mice. Cell Metab. 2015, 21, 455–467.
- Renaville, B.; Bacciu, N.; Lanzoni, M.; Corazzin, M.; Piasentier, E. Polymorphism of fat metabolism genes as candidate markers for meat quality and production traits in heavy pigs. Meat Sci. 2015, 110, 220–223.
- Hu, G.; Lou, Z.; Mamta, G.; Ki, K.Y. The Long Non-Coding RNA GAS5 Cooperates with the Eukaryotic Translation Initiation Factor 4E to Regulate c-Myc Translation. PLoS ONE 2014, 9, e107016.
- Lawrence, V.J.; Kopelman, P.G. Medical consequences of obesity. Clin. Dermatol. 2004, 22, 296–302.
- Sun, L.; Goff, L.A.; Trapnell, C.; Alexander, R.; Lo, K.A.; Hacisuleyman, E.; Sauvageau, M.; Tazon-Vega, B.; Kelley, D.R.; Hendrickson, D.G.; et al. Long noncoding RNAs regulate adipogenesis. Proc. Natl. Acad. Sci. USA 2013, 110, 3387–3392.
- Liu, S.; Sheng, L.; Miao, H.; Saunders, T.L.; MacDougald, O.A.; Koenig, R.J.; Xu, B. SRA gene knockout protects against diet-induced obesity and improves glucose tolerance. J. Biol. Chem. 2014, 289, 13000–13009.
- Liu, S.; Sun, Y.; Hou, Y.; Yang, L.; Wan, X.; Qin, Y.; Liu, Y.; Wang, R.; Zhu, P.; Teng, Y.; et al. A novel lncRNA ROPM-mediated lipid metabolism governs breast cancer stem cell properties. J. Hematol. Oncol. 2021, 14, 178.
- Jiang, Y.; Peng, J.; Song, J.; He, J.; Jiang, M.; Wang, J.; Ma, L.; Wang, Y.; Lin, M.; Wu, H.; et al. Loss of Hilnc prevents diet-induced hepatic steatosis through binding of IGF2BP2. Nat. Metab. 2021, 3, 1569–1584.
- Yu, X.H.; Deng, W.Y.; Chen, J.J.; Xu, X.D.; Liu, X.X.; Chen, L.; Shi, M.W.; Liu, Q.X.; Tao, M.; Ren, K. LncRNA kcnq1ot1 promotes lipid accumulation and accelerates atherosclerosis via functioning as a ceRNA through the miR-452-3p/HDAC3/ABCA1 axis. Cell Death Dis. 2020, 11, 1043.
- Ruan, Y.; Lin, N.; Ma, Q.; Chen, R.; Zhang, Z.; Wen, W.; Chen, H.; Sun, J. Circulating LncRNAs Analysis in Patients with Type 2 Diabetes Reveals Novel Genes Influencing Glucose Metabolism and Islet β-Cell Function. Cell Physiol. Biochem. 2018, 46, 335–350.
- Fawzy, M.S.; Abdelghany, A.A.; Toraih, E.A.; Mohamed, A.M. Circulating long noncoding RNAs H19 and GAS5 are associated with type 2 diabetes but not with diabetic retinopathy: A preliminary study. Bosn. J. Basic Med. Sci. 2020, 20, 365–371.
More
Information
Subjects:
Endocrinology & Metabolism
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.3K
Revisions:
2 times
(View History)
Update Date:
01 Aug 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





