
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | ABHINAV JOSHI | -- | 1643 | 2022-06-29 19:49:09 | | | |
| 2 | Rita Xu | Meta information modification | 1643 | 2022-07-07 04:15:44 | | |
Video Upload Options
The HSP90 paralog TRAP1 was discovered more than 20 years ago, however, a detailed understanding of the function of this mitochondrial molecular chaperone remains elusive. The dispensable nature of TRAP1 in vitro and in vivo further complicates an understanding of its role in mitochondrial biology. TRAP1 is more homologous to the bacterial HSP90, HtpG, than to eukaryotic HSP90. Lacking co-chaperones, the unique structural features of TRAP1 likely regulate its temperature-sensitive ATPase activity and shed light on the alternative mechanisms driving the chaperone’s nucleotide-dependent cycle in a defined environment whose physiological temperature approaches 50 °C. TRAP1 has been shown to be an important bioregulator of mitochondrial respiration, mediating the balance between oxidative phosphorylation and glycolysis, while at the same time promoting mitochondrial homeostasis and displaying cytoprotective activity.
1. Molecular Chaperones, HSP90 and TRAP1
2. Is TRAP1 Cytoprotective or Pro-Neoplastic?
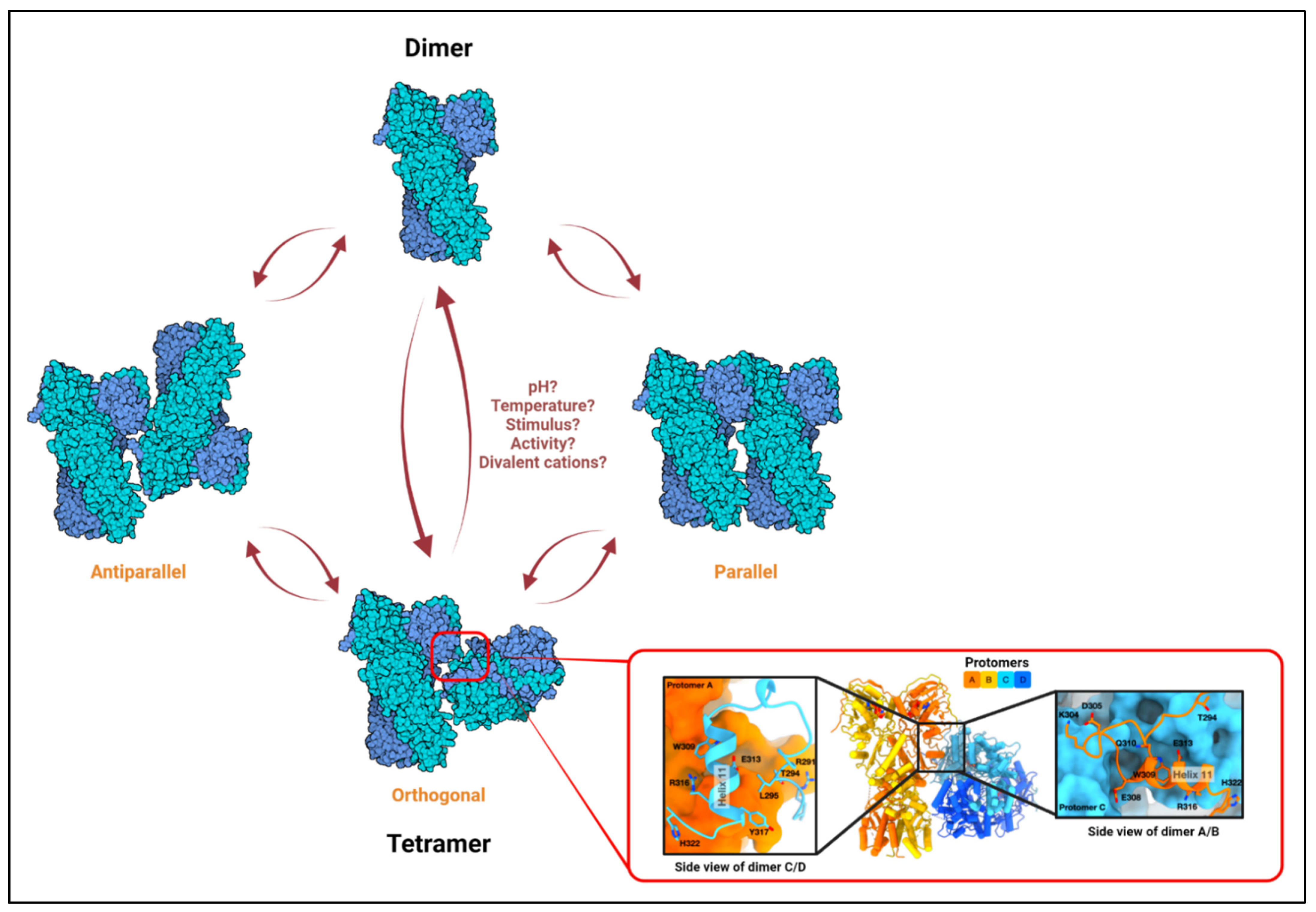
3. TRAP1 Structure, ATPase Cycle, Dimers, and Tetramers
References
- Hartl, F.U.; Bracher, A.; Hayer-Hartl, M. Molecular chaperones in protein folding and proteostasis. Nature 2011, 475, 324–332.
- Kim, Y.E.; Hipp, M.S.; Bracher, A.; Hayer-Hartl, M.; Hartl, F.U. Molecular chaperone functions in protein folding and proteostasis. Annu. Rev. Biochem. 2013, 82, 323–355.
- Richter, K.; Haslbeck, M.; Buchner, J. The heat shock response: Life on the verge of death. Mol. Cell 2010, 40, 253–266.
- Gao, X.; Carroni, M.; Nussbaum-Krammer, C.; Mogk, A.; Nillegoda, N.B.; Szlachcic, A.; Guilbride, D.L.; Saibil, H.R.; Mayer, M.P.; Bukau, B. Human Hsp70 Disaggregase Reverses Parkinson’s-Linked α-Synuclein Amyloid Fibrils. Mol. Cell 2015, 59, 781–793.
- Nillegoda, N.B.; Bukau, B. Metazoan Hsp70-based protein disaggregases: Emergence and mechanisms. Front. Mol. Biosci. 2015, 2, 57.
- Fernandez-Funez, P.; Sanchez-Garcia, J.; de Mena, L.; Zhang, Y.; Levites, Y.; Khare, S.; Golde, T.E.; Rincon-Limas, D.E. Holdase activity of secreted Hsp70 masks amyloid-β42 neurotoxicity in Drosophila. Proc. Natl. Acad. Sci. USA 2016, 113, E5212–E5221.
- Thoma, J.; Burmann, B.M.; Hiller, S.; Müller, D.J. Impact of holdase chaperones Skp and SurA on the folding of β-barrel outer-membrane proteins. Nat. Struct. Mol. Biol. 2015, 22, 795–802.
- Demand, J.; Alberti, S.; Patterson, C.; Höhfeld, J. Cooperation of a ubiquitin domain protein and an E3 ubiquitin ligase during chaperone/proteasome coupling. Curr. Biol. 2001, 11, 1569–1577.
- Gamerdinger, M.; Hajieva, P.; Kaya, A.M.; Wolfrum, U.; Hartl, F.U.; Behl, C. Protein quality control during aging involves recruitment of the macroautophagy pathway by BAG3. EMBO J. 2009, 28, 889–901.
- Hartmann-Petersen, R.; Seeger, M.; Gordon, C. Transferring substrates to the 26S proteasome. Trends Biochem. Sci. 2003, 28, 26–31.
- Csermely, P.; Schnaider, T.; Soti, C.; Prohászka, Z.; Nardai, G. The 90-kDa molecular chaperone family: Structure, function, and clinical applications. A comprehensive review. Pharmacol. Ther. 1998, 79, 129–168.
- Schopf, F.H.; Biebl, M.M.; Buchner, J. The HSP90 chaperone machinery. Nat. Rev. Mol. Cell Biol. 2017, 18, 345–360.
- Radli, M.; Rüdiger, S.G.D. Dancing with the Diva: Hsp90–Client Interactions. J. Mol. Biol. 2018, 430, 3029–3040.
- Taipale, M.; Jarosz, D.F.; Lindquist, S. HSP90 at the hub of protein homeostasis: Emerging mechanistic insights. Nat. Rev. Mol. Cell Biol. 2010, 11, 515–528.
- Zuehlke, A.D.; Beebe, K.; Neckers, L.; Prince, T. Regulation and function of the human HSP90AA1 gene. Gene 2015, 570, 8–16.
- Marzec, M.; Eletto, D.; Argon, Y. GRP94: An HSP90-like protein specialized for protein folding and quality control in the endoplasmic reticulum. Biochim. Biophys. Acta 2012, 1823, 774–787.
- Felts, S.J.; Owen, B.A.; Nguyen, P.; Trepel, J.; Donner, D.B.; Toft, D.O. The hsp90-related protein TRAP1 is a mitochondrial protein with distinct functional properties. J. Biol. Chem. 2000, 275, 3305–3312.
- Cechetto, J.D.; Gupta, R.S. Immunoelectron microscopy provides evidence that tumor necrosis factor receptor-associated protein 1 (TRAP-1) is a mitochondrial protein which also localizes at specific extramitochondrial sites. Exp. Cell Res. 2000, 260, 30–39.
- Yoshida, S.; Tsutsumi, S.; Muhlebach, G.; Sourbier, C.; Lee, M.-J.; Lee, S.; Vartholomaiou, E.; Tatokoro, M.; Beebe, K.; Miyajima, N.; et al. Molecular chaperone TRAP1 regulates a metabolic switch between mitochondrial respiration and aerobic glycolysis. Proc. Natl. Acad. Sci. USA 2013, 110, E1604–E1612.
- Sciacovelli, M.; Guzzo, G.; Morello, V.; Frezza, C.; Zheng, L.; Nannini, N.; Calabrese, F.; Laudiero, G.; Esposito, F.; Landriscina, M.; et al. The Mitochondrial Chaperone TRAP1 Promotes Neoplastic Growth by Inhibiting Succinate Dehydrogenase. Cell Metab. 2013, 17, 988–999.
- Chae, Y.C.; Angelin, A.; Lisanti, S.; Kossenkov, A.V.; Speicher, K.D.; Wang, H.; Powers, J.F.; Tischler, A.S.; Pacak, K.; Fliedner, S.; et al. Landscape of the mitochondrial Hsp90 metabolome in tumours. Nat. Commun. 2013, 4, 2139.
- Joshi, A.; Dai, L.; Liu, Y.; Lee, J.; Ghahhari, N.M.; Segala, G.; Beebe, K.; Jenkins, L.M.; Lyons, G.C.; Bernasconi, L.; et al. The mitochondrial HSP90 paralog TRAP1 forms an OXPHOS-regulated tetramer and is involved in mitochondrial metabolic homeostasis. BMC Biol. 2020, 18, 10.
- Park, H.K.; Hong, J.H.; Oh, Y.T.; Kim, S.S.; Yin, J.; Lee, A.J.; Chae, Y.C.; Kim, J.H.; Park, S.H.; Park, C.K.; et al. Interplay between TRAP1 and Sirtuin-3 Modulates Mitochondrial Respiration and Oxidative Stress to Maintain Stemness of Glioma Stem Cells. Cancer Res. 2019, 79, 1369–1382.
- Rasola, A.; Neckers, L.; Picard, D. Mitochondrial oxidative phosphorylation TRAP(1)ped in tumor cells. Trends Cell Biol. 2014, 24, 455–463.
- Masgras, I.; Laquatra, C.; Cannino, G.; Serapian, S.A.; Colombo, G.; Rasola, A. The molecular chaperone TRAP1 in cancer: From the basics of biology to pharmacological targeting. Semin. Cancer Biol. 2021, 76, 45–53.
- Takamura, H.; Koyama, Y.; Matsuzaki, S.; Yamada, K.; Hattori, T.; Miyata, S.; Takemoto, K.; Tohyama, M.; Katayama, T. TRAP1 controls mitochondrial fusion/fission balance through Drp1 and Mff expression. PLoS ONE 2012, 7, e51912.
- Costa, A.C.; Loh, S.H.Y.; Martins, L.M. Drosophila Trap1 protects against mitochondrial dysfunction in a PINK1/parkin model of Parkinson’s disease. Cell Death Dis. 2013, 4, e467.
- Zhang, L.; Karsten, P.; Hamm, S.; Pogson, J.H.; Müller-Rischart, A.K.; Exner, N.; Haass, C.; Whitworth, A.J.; Winklhofer, K.F.; Schulz, J.B.; et al. TRAP1 rescues PINK1 loss-of-function phenotypes. Hum. Mol. Genet. 2013, 22, 2829–2841.
- Hua, G.; Zhang, Q.; Fan, Z. Heat Shock Protein 75 (TRAP1) Antagonizes Reactive Oxygen Species Generation and Protects Cells from Granzyme M-mediated Apoptosis. J. Biol. Chem. 2007, 282, 20553–20560.
- Montesano Gesualdi, N.; Chirico, G.; Pirozzi, G.; Costantino, E.; Landriscina, M.; Esposito, F. Tumor necrosis factor-associated protein 1 (TRAP-1) protects cells from oxidative stress and apoptosis. Stress 2007, 10, 342–350.
- Pridgeon, J.W.; Olzmann, J.A.; Chin, L.-S.; Li, L. PINK1 Protects against Oxidative Stress by Phosphorylating Mitochondrial Chaperone TRAP1. PLoS Biol. 2007, 5, e172.
- Ramos Rego, I.; Santos Cruz, B.; Ambrósio, A.F.; Alves, C.H. TRAP1 in Oxidative Stress and Neurodegeneration. Antioxidants 2021, 10, 1829.
- Kang, B.H.; Plescia, J.; Dohi, T.; Rosa, J.; Doxsey, S.J.; Altieri, D.C. Regulation of Tumor Cell Mitochondrial Homeostasis by an Organelle-Specific Hsp90 Chaperone Network. Cell 2007, 131, 257–270.
- Xu, L.; Voloboueva, L.A.; Ouyang, Y.; Emery, J.F.; Giffard, R.G. Overexpression of mitochondrial Hsp70/Hsp75 in rat brain protects mitochondria, reduces oxidative stress, and protects from focal ischemia. J. Cereb. Blood Flow Metab. 2009, 29, 365–374.
- Xiang, F.; Huang, Y.S.; Shi, X.H.; Zhang, Q. Mitochondrial chaperone tumour necrosis factor receptor-associated protein 1 protects cardiomyocytes from hypoxic injury by regulating mitochondrial permeability transition pore opening. FEBS J. 2010, 277, 1929–1938.
- Zhang, P.; Lu, Y.; Yu, D.; Zhang, D.; Hu, W. TRAP1 Provides Protection Against Myocardial Ischemia-Reperfusion Injury by Ameliorating Mitochondrial Dysfunction. Cell. Physiol. Biochem. Int. J. Exp. Cell. Physiol. Biochem. Pharmacol. 2015, 36, 2072–2082.
- Clarke, B.E.; Kalmar, B.; Greensmith, L. Enhanced Expression of TRAP1 Protects Mitochondrial Function in Motor Neurons under Conditions of Oxidative Stress. Int. J. Mol. Sci. 2022, 23, 1789.
- Zhang, L.; Su, N.; Luo, Y.; Chen, S.; Zhao, T. TRAP1 inhibits MIC60 ubiquitination to mitigate the injury of cardiomyocytes and protect mitochondria in extracellular acidosis. Cell Death Discov. 2021, 7, 389.
- Butler, E.K.; Voigt, A.; Lutz, A.K.; Toegel, J.P.; Gerhardt, E.; Karsten, P.; Falkenburger, B.; Reinartz, A.; Winklhofer, K.F.; Schulz, J.B. The mitochondrial chaperone protein TRAP1 mitigates α-Synuclein toxicity. PLoS Genet. 2012, 8, e1002488.
- Malpartida, A.B.; Williamson, M.; Narendra, D.P.; Wade-Martins, R.; Ryan, B.J. Mitochondrial Dysfunction and Mitophagy in Parkinson’s Disease: From Mechanism to Therapy. Trends Biochem. Sci. 2021, 46, 329–343.
- Chen, J.F.; Wu, Q.S.; Xie, Y.X.; Si, B.L.; Yang, P.P.; Wang, W.Y.; Hua, Q.; He, Q. TRAP1 ameliorates renal tubulointerstitial fibrosis in mice with unilateral ureteral obstruction by protecting renal tubular epithelial cell mitochondria. FASEB J. 2017, 31, 4503–4514.
- Nicolas, E.; Demidova, E.V.; Iqbal, W.; Serebriiskii, I.G.; Vlasenkova, R.; Ghatalia, P.; Zhou, Y.; Rainey, K.; Forman, A.F.; Dunbrack, R.L., Jr.; et al. Interaction of germline variants in a family with a history of early-onset clear cell renal cell carcinoma. Mol. Genet. Genom. Med. 2019, 7, e556.
- Fitzgerald, J.C.; Zimprich, A.; Carvajal Berrio, D.A.; Schindler, K.M.; Maurer, B.; Schulte, C.; Bus, C.; Hauser, A.K.; Kübler, M.; Lewin, R.; et al. Metformin reverses TRAP1 mutation-associated alterations in mitochondrial function in Parkinson’s disease. Brain J. Neurol. 2017, 140, 2444–2459.
- Skinner, S.J.; Doonanco, K.R.; Boles, R.G.; Chan, A.K.J. Homozygous TRAP1 sequence variant in a child with Leigh syndrome and normal kidneys. Kidney Int. 2014, 86, 860.
- Boles, R.G.; Hornung, H.A.; Moody, A.E.; Ortiz, T.B.; Wong, S.A.; Eggington, J.M.; Stanley, C.M.; Gao, M.; Zhou, H.; McLaughlin, S.; et al. Hurt, tired and queasy: Specific variants in the ATPase domain of the TRAP1 mitochondrial chaperone are associated with common, chronic “functional” symptomatology including pain, fatigue and gastrointestinal dysmotility. Mitochondrion 2015, 23, 64–70.
- Saisawat, P.; Kohl, S.; Hilger, A.C.; Hwang, D.Y.; Yung Gee, H.; Dworschak, G.C.; Tasic, V.; Pennimpede, T.; Natarajan, S.; Sperry, E.; et al. Whole-exome resequencing reveals recessive mutations in TRAP1 in individuals with CAKUT and VACTERL association. Kidney Int. 2014, 85, 1310–1317.
- Megger, D.A.; Bracht, T.; Kohl, M.; Ahrens, M.; Naboulsi, W.; Weber, F.; Hoffmann, A.-C.; Stephan, C.; Kuhlmann, K.; Eisenacher, M.; et al. Proteomic differences between hepatocellular carcinoma and nontumorous liver tissue investigated by a combined gel-based and label-free quantitative proteomics study. Mol. Cell. Proteom. 2013, 12, 2006–2020.
- Zhang, B.; Wang, J.; Huang, Z.; Wei, P.; Liu, Y.; Hao, J.; Zhao, L.; Zhang, F.; Tu, Y.; Wei, T. Aberrantly upregulated TRAP1 is required for tumorigenesis of breast cancer. Oncotarget 2015, 6, 44495.
- Li, S.; Lv, Q.; Sun, H.; Xue, Y.; Wang, P.; Liu, L.; Li, Z.; Li, Z.; Tian, X.; Liu, Y.H. Expression of TRAP1 predicts poor survival of malignant glioma patients. J. Mol. Neurosci. 2015, 55, 62–68.
- Lee, J.H.; Kang, K.W.; Kim, J.-E.; Hwang, S.W.; Park, J.H.; Kim, S.-H.; Ji, J.H.; Kim, T.G.; Nam, H.-Y.; Roh, M.S.; et al. Differential expression of heat shock protein 90 isoforms in small cell lung cancer. Int. J. Clin. Exp. Pathol. 2015, 8, 9487–9493.
- Gao, C.; Li, M.; Jiang, A.-L.; Sun, R.; Jin, H.-L.; Gui, H.-W.; Xiao, F.; Ding, X.-W.; Fu, Z.-M.; Feng, J.-P. Overexpression of the mitochondrial chaperone tumor necrosis factor receptor-associated protein 1 is associated with the poor prognosis of patients with colorectal cancer. Oncol. Lett. 2018, 15, 5451–5458.
- Gao, J.-Y.; Song, B.-R.; Peng, J.-J.; Lu, Y.-M. Correlation between mitochondrial TRAP-1 expression and lymph node metastasis in colorectal cancer. World J. Gastroenterol. 2012, 18, 5965–5971.
- Leav, I.; Plescia, J.; Goel, H.L.; Li, J.; Jiang, Z.; Cohen, R.J.; Languino, L.R.; Altieri, D.C. Cytoprotective mitochondrial chaperone TRAP-1 as a novel molecular target in localized and metastatic prostate cancer. Am. J. Pathol. 2010, 176, 393–401.
- Lv, Q.; Sun, H.; Cao, C.; Gao, B.; Qi, Y. Overexpression of tumor necrosis factor receptor-associated protein 1 (TRAP1) are associated with poor prognosis of epithelial ovarian cancer. Tumor Biol. J. Int. Soc. Oncodev. Biol. Med. 2016, 37, 2721–2727.
- Ou, Y.; Liu, L.; Xue, L.; Zhou, W.; Zhao, Z.; Xu, B.; Song, Y.; Zhan, Q. TRAP1 shows clinical significance and promotes cellular migration and invasion through STAT3/MMP2 pathway in human esophageal squamous cell cancer. J. Genet. Genom. Yi Chuan Xue Bao 2014, 41, 529–537.
- Pak, M.G.; Koh, H.J.; Roh, M.S. Clinicopathologic significance of TRAP1 expression in colorectal cancer: A large scale study of human colorectal adenocarcinoma tissues. Diagn. Pathol. 2017, 12, 6.
- Si, T.; Yang, G.; Qiu, X.; Luo, Y.; Liu, B.; Wang, B. Expression of tumor necrosis factor receptor-associated protein 1 and its clinical significance in kidney cancer. Int. J. Clin. Exp. Pathol. 2015, 8, 13090–13095.
- Chen, R.; Pan, S.; Lai, K.; Lai, L.A.; Crispin, D.A.; Bronner, M.P.; Brentnall, T.A. Up-regulation of mitochondrial chaperone TRAP1 in ulcerative colitis associated colorectal cancer. World J. Gastroenterol. 2014, 20, 17037–17048.
- Kowalik, M.A.; Guzzo, G.; Morandi, A.; Perra, A.; Menegon, S.; Masgras, I.; Trevisan, E.; Angioni, M.M.; Fornari, F.; Quagliata, L.; et al. Metabolic reprogramming identifies the most aggressive lesions at early phases of hepatic carcinogenesis. Oncotarget 2016, 7, 32375–32393.
- Wallace, D.C. Mitochondria and cancer. Nat. Rev. Cancer 2012, 12, 685–698.
- Vartholomaiou, E.; Madon-Simon, M.; Hagmann, S.; Mühlebach, G.; Wurst, W.; Floss, T.; Picard, D. Cytosolic Hsp90α and its mitochondrial isoform Trap1 are differentially required in a breast cancer model. Oncotarget 2017, 8, 17428.
- Chen, B.; Piel, W.H.; Gui, L.; Bruford, E.; Monteiro, A. The HSP90 family of genes in the human genome: Insights into their divergence and evolution. Genomics 2005, 86, 627–637.
- Lisanti, S.; Tavecchio, M.; Chae, Y.C.; Liu, Q.; Brice, A.K.; Thakur, M.L.; Languino, L.R.; Altieri, D.C. Deletion of the Mitochondrial Chaperone TRAP-1 Uncovers Global Reprogramming of Metabolic Networks. Cell Rep. 2014, 8, 671–677.
- Jackson, S.E. Hsp90: Structure and function. Top. Curr. Chem. 2013, 328, 155–240.
- Lavery, L.A.; Partridge, J.R.; Ramelot, T.A.; Elnatan, D.; Kennedy, M.A.; Agard, D.A. Structural asymmetry in the closed state of mitochondrial Hsp90 (TRAP1) supports a two-step ATP hydrolysis mechanism. Mol. Cell 2014, 53, 330–343.
- Kang, B.H. TRAP1 regulation of mitochondrial life or death decision in cancer cells and mitochondria-targeted TRAP1 inhibitors. BMB Rep. 2012, 45, 1–6.
- Tsutsumi, S.; Mollapour, M.; Prodromou, C.; Lee, C.T.; Panaretou, B.; Yoshida, S.; Mayer, M.P.; Neckers, L.M. Charged linker sequence modulates eukaryotic heat shock protein 90 (Hsp90) chaperone activity. Proc. Natl. Acad. Sci. USA 2012, 109, 2937–2942.
- Partridge, J.R.; Lavery, L.A.; Elnatan, D.; Naber, N.; Cooke, R.; Agard, D.A. A novel N-terminal extension in mitochondrial TRAP1 serves as a thermal regulator of chaperone activity. eLife 2014, 3, e03487.
- Sung, N.; Lee, J.; Kim, J.-H.; Chang, C.; Joachimiak, A.; Lee, S.; Tsai, F.T.F. Mitochondrial Hsp90 is a ligand-activated molecular chaperone coupling ATP binding to dimer closure through a coiled-coil intermediate. Proc. Natl. Acad. Sci. USA 2016, 113, 2952–2957.
- Leskovar, A.; Wegele, H.; Werbeck, N.D.; Buchner, J.; Reinstein, J. The ATPase cycle of the mitochondrial Hsp90 analog Trap1. J. Biol. Chem. 2008, 283, 11677–11688.
- Elnatan, D.; Betegon, M.; Liu, Y.; Ramelot, T.; Kennedy, M.A.; Agard, D.A. Symmetry broken and rebroken during the ATP hydrolysis cycle of the mitochondrial Hsp90 TRAP1. eLife 2017, 6, e25235.
- Elnatan, D.; Agard, D.A. Calcium binding to a remote site can replace magnesium as cofactor for mitochondrial Hsp90 (TRAP1) ATPase activity. J. Biol. Chem. 2018, 293, 13717–13724.
- Liu, Y.; Sun, M.; Elnatan, D.; Larson, A.G.; Agard, D.A. Cryo-EM analysis of human mitochondrial Hsp90 in multiple tetrameric states. bioRxiv 2020.
- Shiau, A.K.; Harris, S.F.; Southworth, D.R.; Agard, D.A. Structural Analysis of E. coli hsp90 reveals dramatic nucleotide-dependent conformational rearrangements. Cell 2006, 127, 329–340.
- Nemoto, T.; Sato, N. Oligomeric forms of the 90-kDa heat shock protein. Biochem. J. 1998, 330, 989–995.
- Lee, C.-C.; Lin, T.-W.; Ko, T.-P.; Wang, A.H.J. The Hexameric Structures of Human Heat Shock Protein 90. PLoS ONE 2011, 6, e19961.
- Chadli, A.; Ladjimi, M.M.; Baulieu, E.-E.; Catelli, M.G. Heat-induced Oligomerization of the Molecular Chaperone Hsp90: Inhibition by atp and geldanamycin and activation by transition metal oxyanions. J. Biol. Chem. 1999, 274, 4133–4139.
- Moullintraffort, L.; Bruneaux, M.; Nazabal, A.; Allegro, D.; Giudice, E.; Zal, F.; Peyrot, V.; Barbier, P.; Thomas, D.; Garnier, C. Biochemical and Biophysical Characterization of the Mg2+-induced 90-kDa Heat Shock Protein Oligomers. J. Biol. Chem. 2010, 285, 15100–15110.
- Yonehara, M.; Minami, Y.; Kawata, Y.; Nagai, J.; Yahara, I. Heat-induced Chaperone Activity of HSP90. J. Biol. Chem. 1996, 271, 2641–2645.
- Jakob, U.; Meyer, I.; Bügl, H.; André, S.; Bardwell, J.C.A.; Buchner, J. Structural Organization of Procaryotic and Eucaryotic Hsp90. Influence of divalent cations on structure and function. J. Biol. Chem. 1995, 270, 14412–14419.
- Chrétien, D.; Bénit, P.; Ha, H.-H.; Keipert, S.; El-Khoury, R.; Chang, Y.-T.; Jastroch, M.; Jacobs, H.T.; Rustin, P.; Rak, M. Mitochondria are physiologically maintained at close to 50 °C. PLoS Biol. 2018, 16, e2003992.
- Lepvrier, E.; Thomas, D.; Garnier, C. Hsp90 Quaternary Structures and the Chaperone Cycle: Highly Flexible Dimeric and Oligomeric Structures and Their Regulation by Co-Chaperones. Curr. Proteom. 2019, 16, 5–11.





