Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Hakim Manghwar | -- | 2853 | 2022-06-10 09:13:38 | | | |
| 2 | Amina Yu | -153 word(s) | 2700 | 2022-06-10 09:38:59 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Manghwar, H.; , .; Liu, F. Autophagy-Mediated Regulation of Different Meristems in Plants. Encyclopedia. Available online: https://encyclopedia.pub/entry/23908 (accessed on 01 June 2026).
Manghwar H, , Liu F. Autophagy-Mediated Regulation of Different Meristems in Plants. Encyclopedia. Available at: https://encyclopedia.pub/entry/23908. Accessed June 01, 2026.
Manghwar, Hakim, , Fen Liu. "Autophagy-Mediated Regulation of Different Meristems in Plants" Encyclopedia, https://encyclopedia.pub/entry/23908 (accessed June 01, 2026).
Manghwar, H., , ., & Liu, F. (2022, June 10). Autophagy-Mediated Regulation of Different Meristems in Plants. In Encyclopedia. https://encyclopedia.pub/entry/23908
Manghwar, Hakim, et al. "Autophagy-Mediated Regulation of Different Meristems in Plants." Encyclopedia. Web. 10 June, 2022.
Copy Citation
Autophagy is a highly conserved cell degradation process that widely exists in eukaryotic cells. In plants, autophagy helps maintain cellular homeostasis by degrading and recovering intracellular substances through strict regulatory pathways, thus helping plants respond to a variety of developmental and environmental signals. Autophagy is involved in plant growth and development, including leaf starch degradation, senescence, anthers development, regulation of lipid metabolism, and maintenance of peroxisome mass.
autophagy
stem meristem
plant stress
1. Meristem
Cell mass with the ability to divide continuously or periodically, located in the growing parts of plants, are called meristems. Root and shoot meristems are produced during embryogenesis [1]. Post-embryonically, branching is formed from AMs (axillary meristems) form in the leaf axil. During reproductive growth, IM (inflorescence meristem) are transformed from SAM for successful plant reproduction in Arabidopsis. FMs (floral meristems) are formed from the IM and generate floral organs, including carpels, sepals, stamens, and petals in Arabidopsis. Unlike IM, FMs only show transient stem cell activity. Secondary growth in gymnosperms and woody dicotyledons is dominated by lateral meristems, which is associated with rhizome thickening and re-formation of protective layers [2].
1.1. Root Apical Meristem (RAM)
The root apical meristem is adjacent to the root cap and consists of the root quiescent center (QC) and the root stem cell area, responsible for the formation of the main root. The QC is responsible for maintaining the surrounding stem cells. It also helps to maintain the properties of stem cells. Vascular stem cells are responsible for the formation of vascular tissue, root cap stem cells play a crucial role in forming root cap cells, and cortical/endodermis stem cells help to form cortex and endodermis [3].
TFs and hormones together play important regulatory functions in the root apical meristem. As a key regulator of the quiescent center of the root apical meristem, WOX5 (WUSCHEL RELATED HOMEOBOX 5) has been considered to be important for maintaining the balance of quiescent center cells and column cells [4][5]. The root cap stem cells of the wox5 mutant lose their stem cell activity and differentiate into root cap cells. Meanwhile, WOX5 can inhibit the cell division process of the quiescent center by affecting the activity of the cell cycle regulator (CYCD3) [6]. The TFs PLT1-3 (PLETHORA 1-3) [7][8], SCR (SCARECROW) [9], BBM/PLT4 (BABYBOOM) [10] and SHR (SHORT ROOT) [11][12] in the AIL (AINTEGUMENTA-Like) family are involved in the regulation of root apical meristem development. In Arabidopsis, plant hormones, such as cytokinin, auxin, ethylene, jasmonic acid, and gibberellin are also involved in regulating the development of root apical meristems. The information crossover between different hormone signaling pathways is important for plants to quickly respond to internal and external factors. It is also critical in performing real-time dynamic control. In addition, the expressions of some TFs that regulate the development of root apical meristems are also regulated by plant hormones [3][13][14].
1.2. Shoot Apical Meristem (SAM)
The plant shoot apical meristem is a hemispherical dome-like structure in which cell populations can undergo cell division to maintain self-renewal or develop into organs [15]. In the aerial parts of plants, the shoot apical meristem is responsible for producing and developing the leaves, floral meristems, and stems throughout the plant life cycle. At the histological level, the shoot apical meristem can be divided into the central, peripheral, and rib zone [16][17]. At the top of the stem-end meristem, the dome is the so-called central zone, where stem cells are located.
In the core of a complex regulatory network that determines the size and location of the central region, there is a signaling pathway involving the TF WUS (WUSCHEL), the receptor kinase CLV1 (CLAVATA 1), the receptor-like protein CLV2, and the ligand CLV3. The WUS controls the differentiation and maintenance of stem cells, which is expressed centrally in the tissue below the central zone. Ectopic expression of WUS can promote the transition of plants from vegetative to embryogenic growth [18]. Deletion of WUS will lead to the differentiation of stem cells and the loss of stem meristems [19]. In confirmation of previous conclusions regarding the importance of WUS in regulating the shoot meristem, recent reports further show that the miR156-SPL (squamosa promoter binding protein-like) pathway is directly or indirectly involved in the WUS mechanism regulating SAM size [20]. The small peptide of CLV3 expressed by stem cells negatively regulates WUS expression through the CLV1 receptor [21]. In the SAM, the CLV3-WUS feedback signaling affects stem cell proliferation and differentiation through an autoregulatory negative feedback loop comprising the stem cell-promoting TF WUS and the differentiation promoting peptide CLV3 [22][23]. Phytohormones are required for maintaining the homeostasis of shoot stem cells, and cross-talk exists between WUS function and cytokinin action in the SAM. Cytokinin signaling has been observed to play an important role in maintaining both shoot meristem activity and proliferation, by acting through AHK (Arabidopsis histidine kinase) receptors, which then pass on the signal to the two TF classes: ARRs (type-A Arabidopsis response regulators) and type-B ARRs [24][25][26].
1.3. Lateral Meristem
The meristem, which is distributed in a barrel shape on the near surface of some plant rhizome and other organs in the direction parallel to the long axis is called the lateral meristem. Lateral meristems include vascular cambium and cork cambium. Among them, plant vascular tissue is composed of xylem, phloem, and cambium/procambium, and exists radially in organs. The formation of vascular tissue patterns in Arabidopsis begins during embryonic development, along the root-hypocotyl axis, and differentiates from the precursor-procambium of vascular tissue.
The well-organized pattern of the plant vascular system is also regulated by different intercellular signals, and the regulatory mechanism of vascular system development is similar to that of shoot apical meristem and root apical meristem, both of which are controlled by plant hormones and genes. TDIF peptides (tracheary element differentiation inhibitory factor) were isolated from Zinnia elegans, and its C-terminal 12 amino acid motifs were identical to the conserved 12 motifs of CLE41/44 (CLV3/ESR-related 41/44) in Arabidopsis. The differentiation of stem cell–like procambial cells into xylem cells is inhibited by TDIF by participating in regulating the formation of vascular tissue and promoting the cell division [27]. The CLE41 and CLE44 genes of Arabidopsis encode TDIF, and according to previous studies on TDIF, the fate of vascular stem cells may also be controlled by the CLE receptor system [28]. TDIF/CLE41/CLE44 is expressed in the procambium/cambium and binds to the CLV1-related LRR-RLK (leucine-rich repeat receptor-like protein kinase) protein TDR/PXY (TDIF RECEPTOR/PHLOEM INTERCALATED WITH XYLEM). WOX4 and WOX14 are downstream factors of the TDIF/CLE41/CLE44-TDR/PXY signaling pathway and positively regulate the division of pro-cambial cells [29]. In addition, when TDR/PXY and WOX4 genes were deleted, the expression of some ERFs was induced, indicating that during the formation of vascular tissue, there is a link between the TDIF/CLE41/CLE44-TDR/PXY and the ethylene signaling pathway [30]. Studies have shown that the TDIF/CLE41/CLE44-TDR/PXY-WOX4 signaling pathway is a conserved mechanism in regulating plant vascular tissue development and can cooperate with the ER (endoplasmic reticulum) signaling pathway [31][32][33]. The xylogen polypeptide hormone is involved in the formation of plant vascular tissue. Arabidopsis atxyp1 atxyp2 mutants have defects in vascular tissue structure, which are manifested as discontinuous and thickened leaf veins, abnormal vascular connections, and simplified leaf vein structure [34], indicating that At XYP1 and At XYP2 proteins may play an important regulatory role in the development of vascular tissue. Rice (Oryza sativa L.) xylp7 mutants have significantly shortened internode spacing except for the lowest internode [35].
2. Autophagy in Plant Meristems
2.1. Autophagy in Root Apical Meristem (RAM) under Stress Conditions
Root growth depends on apical meristem cell division, followed by cell elongation and differentiation. When plants suffer from nutrient starvation, the number of autophagosomes increases in the elongation and differentiation regions, and numerous autophagosomes appear in plant cells near the apical meristem [36]. In addition, the canonical traits of the plant-type autophagosomes-autolysosomes pathway was observed in the root tip cells that received systemic PCD induced by TMV (tobacco mosaic virus) local infection for cell contents removal and recovery [37]. ATG genes are required for autophagy in root tip cells. In Arabidopsis root tip cells, both ATG2 and ATG5 proteins are necessary for autophagy, while the ATG9 protein contributes to but is not required for autophagy [38]. Autophagy is involved not only in nutrient recycling under nutrient-limited conditions, but also in root hair formation and cell growth in plant root cells. Plants respond to nutrient deficiency in the soil by increasing the number of lateral roots (LR) to increase the surface area of the root. Consistent with this, phosphate availability was found to have a significant effect on the root system architecture [39].
The root meristem is responsible for optimal root growth and architecture, and its stem cell activity is sustained by a variety of factors, such as nutrient availability (e.g., glucose), hormonal levels (e.g., auxin) and ROS species homeostasis (e.g., superoxide and H2O2) [40][41][42]. Glucose produced by photosynthesis is a hormone-like signaling molecule that regulates plant development and physiological activities. Through genetic, genomic, and systematic analyses, photosynthesis-driven glucose-TOR (target of rapamycin) signaling was found to stimulate and maintain meristem activity for unlimited root growth [43]. In plants, TOR can integrate multiple exogenous and endogenous signals to coordinate several downstream processes, such as cell division and elongation, nutrient transport and metabolism, biological rhythm, and stress response [44]. In addition to regulating root development, glucose-TOR also promotes hypocotyl elongation. Studies have shown that under dark conditions, glucose-TOR signaling stabilizes the key TF BZR1 in the BRs signaling pathway by inhibiting the autophagy process, thereby promoting hypocotyl elongation [45]. After further studies found that autophagy-deficient mutants exhibit greater tolerance to glucose, and accumulate ROS, induced by less glucose at the root tips, Arabidopsis sensors may signal the constitutive autophagy system through the direct or indirect action of TOR after sensing high glucose stress in roots; these results indicate that autophagy regulates the activity of glucose-mediated root meristems in Arabidopsis by modulating ROS production [46] (Figure 1). TOR signaling can modulate glucose-suppressed root meristem activity and negatively regulates autophagy in plants [40][47]. Low levels of ROS, which are produced as byproducts of cellular metabolism, act as critical secondary messengers, controlling a number of vital developmental processes, such as root meristem maintenance [43][48]. However, excessive ROS may cause severe oxidative damage to cells. ROS are actively produced in the root tips, and their concentrations control the extent and direction of root growth [49].
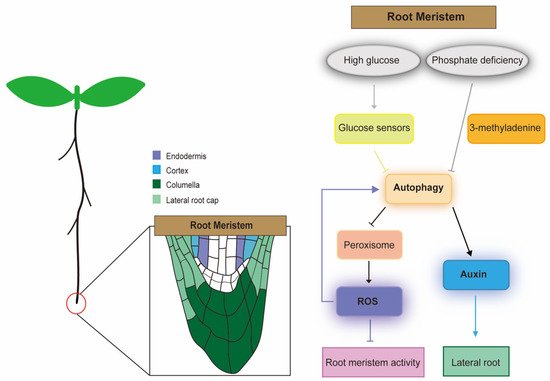
Figure 1. Schematic diagram of the regulatory network of autophagy in the Arabidopsis root meristem. Under high glucose stress, the glucose sensor signals to the autophagy system. Autophagy further regulates peroxisomes and contributes to the production of ROS. ROS accumulation under the induction of high levels of glucose impairs root meristem activity, and high levels of ROS also enhance autophagy mechanisms to maintain root meristem function under stress conditions. Under the phosphate-deficient condition, LR development and auxin accumulation in root meristems are inhibited when autophagy is inhibited by treatment with the autophagy inhibitor 3-methyladenine. Arrow indicates positive interactions; barred arrows indicate repressive interactions.
In Arabidopsis mutants abo6 (ABA overly sensitive 6) and abo8 (ABA overly sensitive 8), AT5G04895/ABO6 and AT4G11690/ABO8 lack a pentatricopeptide repeat protein and DExH box RNA helicase, respectively, resulting in increased ABA sensitivity and decreased meristem activity in roots [50][51]. In addition, the root meristem phenotype in these mutants could be partially rescued by the addition of the reducing agent glutathione (GSH), suggesting that mitochondrial ROS in the root tip is a crucial retrograde signal to maintain its meristem activity [50][51]. A large number of studies have shown that ROS from different sources can regulate autophagy in plant cells. Exogenous H2O2 treatment causes severe oxidative stress in Arabidopsis, which induces the process of autophagy [52]. In Arabidopsis atg2 and atg5 mutants, massive accumulation of H2O2 was observed [53], similar to the results of H2O2 treatment. ROS plays a key role in maintaining root meristem activity, and deletion mutations in ATGs lead to ROS accumulation. Thus, it can be seen that autophagy and ROS may regulate root meristem activity, but how the two co-regulate this process is still unknown. Besides glucose and ROS, auxin is an important hormone that regulates root growth and plays a crucial role in maintaining the root meristem [54][55]. Auxin accumulates in primary root tips via the action of polar transporters, such as the pin-formed proteins (PINs), and it contributes to the root patterning and helps in regulating the root cell division [56]. Recent evidence indicates a link between regulating auxin-dependent lateral root development and autophagy under the conditions of phosphate starvation in Arabidopsis. Under phosphate-deficient conditions, LR development and auxin accumulation in root meristems are inhibited when autophagy is inhibited by treatment with the autophagy inhibitor 3-methyladenine, indicating that autophagy plays an important role in regulating LR development [57][58] (Figure 1).
2.2. Autophagy in Shoot Apical Meristem (SAM) under Stress Conditions
The cells at the tip of the stem, in the SAM, are responsible for deriving the aerial structure of higher plants. The SAM produces lateral organs and stem tissues and also regenerates itself throughout the plant life. Therefore, development of plant under stress is dependent on the meristem, and the SAM of seedlings plays a crucial role in plant defense signaling. Ultrastructural changes in the shoot apical meristem and formation of autophagic vacuoles in canola (Brassica napus cv. Symbol) have been observed under salinity conditions [59]. Recently, it was revealed that autophagy was involved in thermopriming-induced defense mechanism in SAM of young Arabidopsis. Most of the ATG expressions were increased, and autophagy was induced after the thermopriming treatment which caused autophagosome formation in the shoot apices [60]. This means autophagy is recommended as a potential protection system for the homeostasis of SAM.
The relationship between plant autophagy and deficiency of micronutrients in SAM has recently been discovered in Arabidopsis. Although nutrient deficiency interferes with plant growth, an excess of certain elements can also be problematic. The concentration of some important nutrients in the soil environment must remain within a certain range for healthy growth, and the optimal range for micronutrients is narrower than for macronutrients. Plant growth can be inhibited by excess Zn in the rhizosphere. Plants display growth defects in both roots and shoots, and chlorosis in shoots occurs when Zn is in excess. Excessive Zn competes with other divalent metal elements, which is one of its toxic effects. Due to the fact that divalent metal ions are transported into plant cells via the same transporter, excess Zn can result in a deficiency of other divalent metal ions [61]. In the presence of Zn excess, autophagy is activated, and autophagy-deficient Arabidopsis plants exhibit obvious growth defects and chlorosis. However, the excess supply of Fe markedly alleviates growth defects and chlorosis due to an excess of Zn, implying that symptoms caused by excess Zn in atg mutants are due to Fe starvation. In consequence, autophagy has been discussed in the distribution of Fe3+ to juvenile leaves to promote healthy plant growth in the presence of excess Zn [62].
2.3. Autophagy in Other Types of Meristems under Stress Conditions
The function of autophagy in other plant meristems has also been revealed. It was performed in potato (Solanum tuberosum) tuber apical bud meristem (TAB-meristem) revealed the role of autophagy in controlling cold-induced apical dominance [63]. GAPC1, GAPC2, and GAPC3 have the ability to interact with ATG3 both in vitro and in vivo in potatoes. In plants, the interaction between GAPC and ATG3 negatively regulates autophagy [64]. Therefore, cell death in TAB-meristem could be regulated by the ATG3-dependent process. The interactions between ATG3 and GAPCs could be an important factor for the apical dominance of potato tubers [63]. By combining the data from RNA-seq analysis and ERF-VII targeted analysis, it was found that bud populations on leafless vines are gradually starved during natural dormancy cycles, and that starvation-induced ABA acts to temporarily inhibit meristem growth [61]. While starvation-triggered autophagy may be the primary catabolic resource, renewal of energy availability induces degradation of ABA and restoration of meristem activity [65]. In wheat, basal florets are gradually approaching the fertile stage while distal floret rot is observed. For the mechanism of fertile flower number in wheat, autophagy was found to play a role in whole flower decay as a mechanism for regulating the number of fertile florets [66]. However, there is no direct evidence that this phenomenon is caused by the effect of autophagy on the floral meristem.
References
- Zhang, Y.; Jiao, Y.; Jiao, H.; Zhao, H.; Zhu, Y.X. Two-Step Functional Innovation of the Stem-Cell Factors WUS/WOX5 during Plant Evolution. Mol. Biol. Evol. 2017, 34, 640–653.
- Doerner, P. Plant meristems: A merry-go-round of signals review. Curr. Biol. 2003, 13, R368–R374.
- Drisch, R.C.; Stahl, Y. Function and regulation of transcription factors involved in root apical meristem and stem cell maintenance. Front. Plant Sci. 2015, 6, 505.
- Pi, L.; Aichinger, E.; van der Graaff, E.; Llavata-Peris, C.I.; Weijers, D.; Hennig, L.; Groot, E.; Laux, T. Organizer-Derived WOX5 Signal Maintains Root Columella Stem Cells through Chromatin-Mediated Repression of CDF4 Expression. Dev. Cell 2015, 33, 576–588.
- Berckmans, B.; Kirschner, G.; Gerlitz, N.; Stadler, R.; Simon, R. CLE40 Signaling Regulates Root Stem Cell Fate. Plant Physiol. 2020, 182, 1776–1792.
- Forzani, C.; Aichinger, E.; Sornay, E.; Willemsen, V.; Laux, T.; Dewitte, W.; Murray, J.A. WOX5 suppresses CYCLIN D activity to establish quiescence at the center of the root stem cell niche. Curr. Biol. 2014, 24, 1939–1944.
- Aida, M.; Beis, D.; Heidstra, R.; Willemsen, V.; Blilou, I.; Galinha, C.; Nussaume, L.; Noh, Y.S.; Amasino, R.; Scheres, B. The PLETHORA genes mediate patterning of the Arabidopsis root stem cell niche. Cell 2004, 119, 109–120.
- Xiong, F.; Zhang, B.K.; Liu, H.H.; Wei, G.; Wu, J.H.; Wu, Y.N.; Zhang, Y.; Li, S. Transcriptional Regulation of PLETHORA1 in the Root Meristem Through an Importin and Its Two Antagonistic Cargos. Plant Cell 2020, 32, 3812–3824.
- Heidstra, R.; Sabatini, S. Plant and animal stem cells: Similar yet different. Nat. Rev. Mol. Cell Biol. 2014, 15, 301–312.
- Jha, P.; Kumar, V. BABY BOOM (BBM): A candidate transcription factor gene in plant biotechnology. Biotechnol. Lett. 2018, 40, 1467–1475.
- Sozzani, R.; Cui, H.; Moreno-Risueno, M.A.; Busch, W.; Van Norman, J.M.; Vernoux, T.; Brady, S.M.; Dewitte, W.; Murray, J.A.; Benfey, P.N. Spatiotemporal regulation of cell-cycle genes by SHORTROOT links patterning and growth. Nature 2010, 466, 128–132.
- Hakoshima, T. Structural basis of the specific interactions of GRAS family proteins. FEBS Lett. 2018, 592, 489–501.
- Ding, Z.; Friml, J. Auxin regulates distal stem cell differentiation in Arabidopsis roots. Proc. Natl. Acad. Sci. USA 2010, 107, 12046–12051.
- Vilarrasa-Blasi, J.; González-García, M.-P.; Frigola, D.; Fàbregas, N.; Alexiou, K.G.; López-Bigas, N.; Rivas, S.; Jauneau, A.; Lohmann, J.U.; Benfey, P.N. Regulation of plant stem cell quiescence by a brassinosteroid signaling module. Dev. Cell 2014, 30, 36–47.
- Bowman, J.L.; Eshed, Y. Formation and maintenance of the shoot apical meristem. Trends Plant Sci. 2000, 5, 110–115.
- Perales, M.; Reddy, G.V. Stem cell maintenance in shoot apical meristems. Curr. Opin. Plant Biol. 2012, 15, 10–16.
- Xie, M.; Tataw, M.; Venugopala Reddy, G. Towards a functional understanding of cell growth dynamics in shoot meristem stem-cell niche. Semin. Cell Dev. Biol. 2009, 20, 1126–1133.
- Zuo, J.; Niu, Q.W.; Frugis, G.; Chua, N.H. The WUSCHEL gene promotes vegetative-to-embryonic transition in Arabidopsis. Plant J. 2002, 30, 349–359.
- van den Berg, C.; Willemsen, V.; Hage, W.; Weisbeek, P.; Scheres, B. Cell fate in the Arabidopsis root meristem determined by directional signalling. Nature 1995, 378, 62–65.
- Fouracre, J.P.; Poethig, R.S. Role for the shoot apical meristem in the specification of juvenile leaf identity in Arabidopsis. Proc. Natl. Acad. Sci. USA 2019, 116, 10168–10177.
- Yadav, R.K.; Perales, M.; Gruel, J.; Girke, T.; Jonsson, H.; Reddy, G.V. WUSCHEL protein movement mediates stem cell homeostasis in the Arabidopsis shoot apex. Genes Dev. 2011, 25, 2025–2030.
- Schoof, H.; Lenhard, M.; Haecker, A.; Mayer, K.F.; Jurgens, G.; Laux, T. The stem cell population of Arabidopsis shoot meristems in maintained by a regulatory loop between the CLAVATA and WUSCHEL genes. Cell 2000, 100, 635–644.
- Somssich, M.; Je, B.I.; Simon, R.; Jackson, D. CLAVATA-WUSCHEL signaling in the shoot meristem. Development 2016, 143, 3238–3248.
- Kieber, J.J.; Schaller, G.E. Cytokinin signaling in plant development. Development 2018, 145, dev149344.
- Muller, B.; Sheen, J. Advances in cytokinin signaling. Science 2007, 318, 68–69.
- Gordon, S.P.; Chickarmane, V.S.; Ohno, C.; Meyerowitz, E.M. Multiple feedback loops through cytokinin signaling control stem cell number within the Arabidopsis shoot meristem. Proc. Natl. Acad. Sci. USA 2009, 106, 16529–16534.
- Ito, Y.; Nakanomyo, I.; Motose, H.; Iwamoto, K.; Sawa, S.; Dohmae, N.; Fukuda, H. Dodeca-CLE peptides as suppressors of plant stem cell differentiation. Science 2006, 313, 842–845.
- Hirakawa, Y.; Shinohara, H.; Kondo, Y.; Inoue, A.; Nakanomyo, I.; Ogawa, M.; Sawa, S.; Ohashi-Ito, K.; Matsubayashi, Y.; Fukuda, H. Non-cell-autonomous control of vascular stem cell fate by a CLE peptide/receptor system. Proc. Natl. Acad. Sci. USA 2008, 105, 15208–15213.
- Hirakawa, Y.; Kondo, Y.; Fukuda, H. TDIF peptide signaling regulates vascular stem cell proliferation via the WOX4 homeobox gene in Arabidopsis. Plant Cell 2010, 22, 2618–2629.
- Etchells, J.P.; Turner, S.R. The PXY-CLE41 receptor ligand pair defines a multifunctional pathway that controls the rate and orientation of vascular cell division. Development 2010, 137, 767–774.
- Sieburth, L.E.; Deyholos, M.K. Vascular development: The long and winding road. Curr. Opin. Plant Biol. 2006, 9, 48–54.
- Strabala, T.J.; Phillips, L.; West, M.; Stanbra, L. Bioinformatic and phylogenetic analysis of the CLAVATA3/EMBRYO-SURROUNDING REGION (CLE) and the CLE-LIKE signal peptide genes in the Pinophyta. BMC Plant Biol. 2014, 14, 47.
- Etchells, J.P.; Provost, C.M.; Mishra, L.; Turner, S.R. WOX4 and WOX14 act downstream of the PXY receptor kinase to regulate plant vascular proliferation independently of any role in vascular organisation. Development 2013, 140, 2224–2234.
- Motose, H.; Sugiyama, M.; Fukuda, H. A proteoglycan mediates inductive interaction during plant vascular development. Nature 2004, 429, 873–878.
- Ma, T.; Ma, H.; Zhao, H.; Qi, H.; Zhao, J. Identification, characterization, and transcription analysis of xylogen-like arabinogalactan proteins in rice (Oryza sativa L.). BMC Plant Biol. 2014, 14, 299.
- Yano, K.; Suzuki, T.; Moriyasu, Y. Constitutive autophagy in plant root cells. Autophagy 2007, 3, 360–362.
- Zhou, S.; Hong, Q.; Li, Y.; Li, Q.; Li, R.; Zhang, H.; Wang, M.; Yuan, X. Macroautophagy occurs in distal TMV-uninfected root tip tissue of tomato taking place systemic PCD. Protoplasma 2018, 255, 3–9.
- Inoue, Y.; Suzuki, T.; Hattori, M.; Yoshimoto, K.; Ohsumi, Y.; Moriyasu, Y. AtATG genes, homologs of yeast autophagy genes, are involved in constitutive autophagy in Arabidopsis root tip cells. Plant Cell Physiol. 2006, 47, 1641–1652.
- Williamson, L.C.; Ribrioux, S.P.; Fitter, A.H.; Leyser, H.M. Phosphate availability regulates root system architecture in Arabidopsis. Plant Physiol. 2001, 126, 875–882.
- Xiong, Y.; McCormack, M.; Li, L.; Hall, Q.; Xiang, C.; Sheen, J. Glucose-TOR signalling reprograms the transcriptome and activates meristems. Nature 2013, 496, 181–186.
- Sanchez-Fernandez, R.; Fricker, M.; Corben, L.B.; White, N.S.; Sheard, N.; Leaver, C.J.; Van Montagu, M.; Inze, D.; May, M.J. Cell proliferation and hair tip growth in the Arabidopsis root are under mechanistically different forms of redox control. Proc. Natl. Acad. Sci. USA 1997, 94, 2745–2750.
- Pacifici, E.; Polverari, L.; Sabatini, S. Plant hormone cross-talk: The pivot of root growth. J. Exp. Bot. 2015, 66, 1113–1121.
- Dunand, C.; Crevecoeur, M.; Penel, C. Distribution of superoxide and hydrogen peroxide in Arabidopsis root and their influence on root development: Possible interaction with peroxidases. New Phytol. 2007, 174, 332–341.
- Burkart, G.M.; Brandizzi, F. A tour of TOR complex signaling in plants. Trends Biochem. Sci. 2021, 46, 417–428.
- Zhang, Z.; Zhu, J.-Y.; Roh, J.; Marchive, C.; Kim, S.-K.; Meyer, C.; Sun, Y.; Wang, W.; Wang, Z.-Y. TOR signaling promotes accumulation of BZR1 to balance growth with carbon availability in Arabidopsis. Curr. Biol. 2016, 26, 1854–1860.
- Huang, L.; Yu, L.J.; Zhang, X.; Fan, B.; Wang, F.Z.; Dai, Y.S.; Qi, H.; Zhou, Y.; Xie, L.J.; Xiao, S. Autophagy regulates glucose-mediated root meristem activity by modulating ROS production in Arabidopsis. Autophagy 2019, 15, 407–422.
- Liu, Y.; Bassham, D.C. TOR is a negative regulator of autophagy in Arabidopsis thaliana. PLoS ONE 2010, 5, e11883.
- Del Pozo, J.C. Reactive Oxygen Species: From Harmful Molecules to Fine-Tuning Regulators of Stem Cell Niche Maintenance. PLoS Genet. 2016, 12, e1006251.
- Liszkay, A.; van der Zalm, E.; Schopfer, P. Production of reactive oxygen intermediates (O(2)(.−), H(2)O(2), and (.)OH) by maize roots and their role in wall loosening and elongation growth. Plant Physiol. 2004, 136, 3114–3123, discussion 3001.
- He, J.; Duan, Y.; Hua, D.; Fan, G.; Wang, L.; Liu, Y.; Chen, Z.; Han, L.; Qu, L.-J.; Gong, Z. DEXH box RNA helicase–mediated mitochondrial reactive oxygen species production in Arabidopsis mediates crosstalk between abscisic acid and auxin signaling. Plant Cell 2012, 24, 1815–1833.
- Yang, L.; Zhang, J.; He, J.; Qin, Y.; Hua, D.; Duan, Y.; Chen, Z.; Gong, Z. ABA-mediated ROS in mitochondria regulate root meristem activity by controlling PLETHORA expression in Arabidopsis. PLoS Genet. 2014, 10, e1004791.
- Xiong, Y.; Contento, A.L.; Bassham, D.C. Disruption of autophagy results in constitutive oxidative stress in Arabidopsis. Autophagy 2007, 3, 257–258.
- Yoshimoto, K.; Jikumaru, Y.; Kamiya, Y.; Kusano, M.; Consonni, C.; Panstruga, R.; Ohsumi, Y.; Shirasu, K. Autophagy negatively regulates cell death by controlling NPR1-dependent salicylic acid signaling during senescence and the innate immune response in Arabidopsis. Plant Cell 2009, 21, 2914–2927.
- Overvoorde, P.; Fukaki, H.; Beeckman, T. Auxin control of root development. Cold Spring Harb. Perspect. Biol. 2010, 2, a001537.
- Lee, Y.; Lee, W.S.; Kim, S.H. Hormonal regulation of stem cell maintenance in roots. J. Exp. Bot. 2013, 64, 1153–1165.
- Petricka, J.J.; Winter, C.M.; Benfey, P.N. Control of Arabidopsis root development. Annu. Rev. Plant Biol. 2012, 63, 563–590.
- Deb, S.; Sankaranarayanan, S.; Wewala, G.; Widdup, E.; Samuel, M.A. The S-Domain Receptor Kinase Arabidopsis Receptor Kinase2 and the U Box/Armadillo Repeat-Containing E3 Ubiquitin Ligase9 Module Mediates Lateral Root Development under Phosphate Starvation in Arabidopsis. Plant Physiol. 2014, 165, 1647–1656.
- Sankaranarayanan, S.; Samuel, M.A. A proposed role for selective autophagy in regulating auxin-dependent lateral root development under phosphate starvation in Arabidopsis. Plant Signal Behav. 2015, 10, e989749.
- Mahmoodzadeh, H. Ultrastructural changes in shoot apical meristem of canola (Brassica napus cv. Symbol) treated with sodium chloride. Pak. J. Biol. Sci. PJBS 2008, 11, 1161–1164.
- Taheri Sedeh, H.; Bazgir, E. Thermopriming-Induced Autophagy in Shoot Apical Meristem of Arabidopsis. Iran J. Biotechnol. 2021, 19, e2901.
- Fukao, Y.; Ferjani, A.; Tomioka, R.; Nagasaki, N.; Kurata, R.; Nishimori, Y.; Fujiwara, M.; Maeshima, M. iTRAQ analysis reveals mechanisms of growth defects due to excess zinc in Arabidopsis. Plant Physiol. 2011, 155, 1893–1907.
- Shinozaki, D.; Tanoi, K.; Yoshimoto, K. Optimal Distribution of Iron to Sink Organs via Autophagy Is Important for Tolerance to Excess Zinc in Arabidopsis. Plant Cell Physiol. 2021, 62, 515–527.
- Liu, T.; Fang, H.; Liu, J.; Reid, S.; Hou, J.; Zhou, T.; Tian, Z.; Song, B.; Xie, C. Cytosolic glyceraldehyde-3-phosphate dehydrogenases play crucial roles in controlling cold-induced sweetening and apical dominance of potato (Solanum tuberosum L.) tubers. Plant Cell Environ. 2017, 40, 3043–3054.
- Han, S.; Wang, Y.; Zheng, X.; Jia, Q.; Zhao, J.; Bai, F.; Hong, Y.; Liu, Y. Cytoplastic Glyceraldehyde-3-Phosphate Dehydrogenases Interact with ATG3 to Negatively Regulate Autophagy and Immunity in Nicotiana benthamiana. Plant Cell 2015, 27, 1316–1331.
- Sienko, K.; Poormassalehgoo, A.; Yamada, K.; Goto-Yamada, S. Microautophagy in Plants: Consideration of Its Molecular Mechanism. Cells 2020, 9, 887.
- Ghiglione, H.O.; Gonzalez, F.G.; Serrago, R.; Maldonado, S.B.; Chilcott, C.; Cura, J.A.; Miralles, D.J.; Zhu, T.; Casal, J.J. Autophagy regulated by day length determines the number of fertile florets in wheat. Plant J. 2008, 55, 1010–1024.
More
Information
Subjects:
Plant Sciences
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
912
Revisions:
2 times
(View History)
Update Date:
10 Jun 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





