
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Filipa F. Vale | -- | 1829 | 2022-05-30 11:37:24 | | | |
| 2 | Dean Liu | -3 word(s) | 1826 | 2022-05-31 03:43:14 | | | | |
| 3 | Dean Liu | Meta information modification | 1826 | 2022-05-31 03:44:11 | | | | |
| 4 | Dean Liu | Meta information modification | 1826 | 2022-06-01 08:26:42 | | |
Video Upload Options
The Gram-negative bacterium Helicobacter pylori colonizes c.a. 50% of human stomachs worldwide and is the major risk factor for gastric adenocarcinoma. Its high genetic variability makes it difficult to identify biomarkers of early stages of infection that can reliably predict its outcome.
1. Introduction
2. Technologies Used for H. pylori Marker Detection
3. New Approaches to H. pylori Biomarker Detection
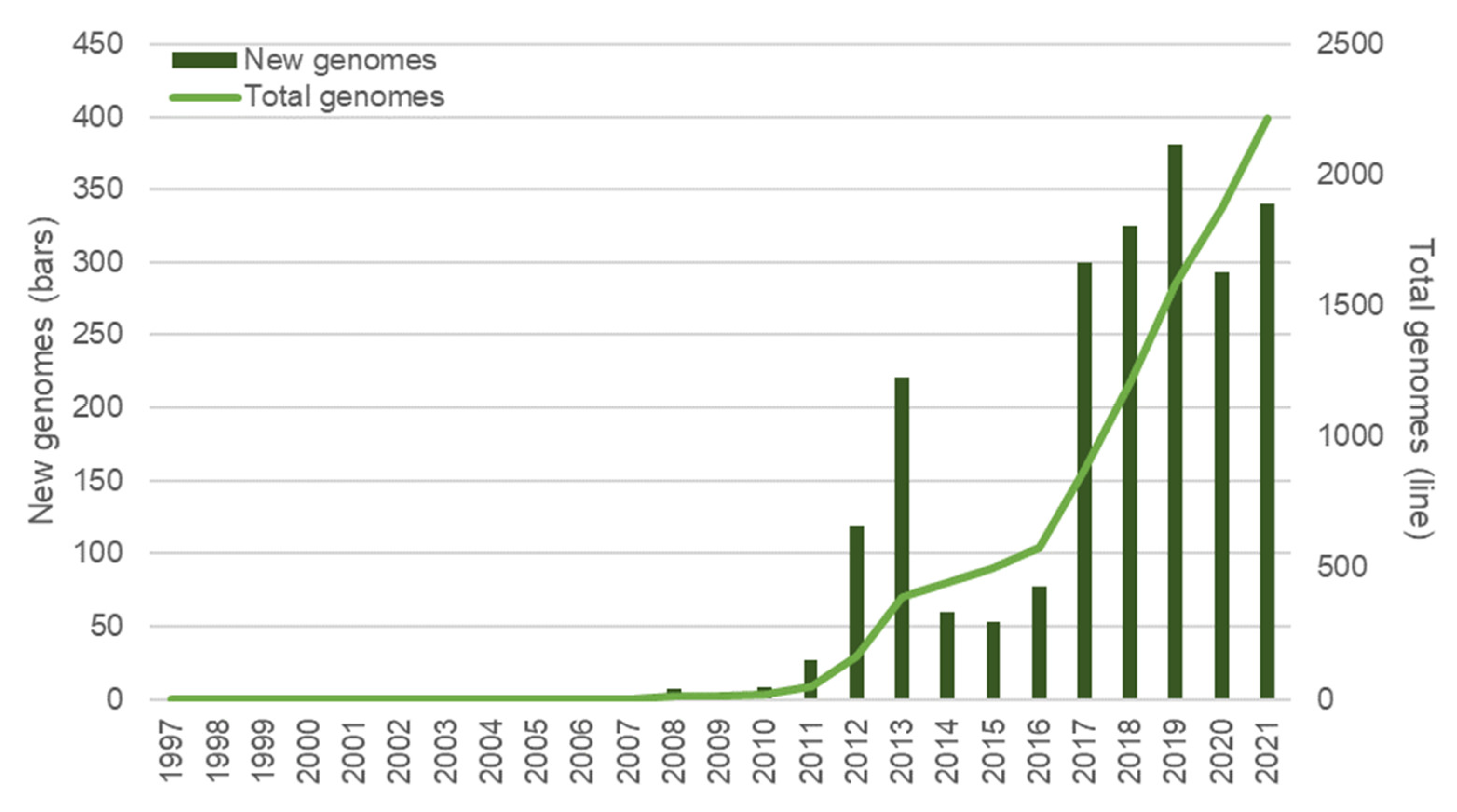
3.1. Sequencing Technologies
3.2. Virulence Potential Evaluation and Antibiotic-Resistance Profiling
Whole genome data provide the opportunity to evaluate the virulence potential of strains and to predict the resistance profile for several antibiotics from a single dataset, with no need for multiple cultures, PCR amplification or sequencing procedures. Regarding virulence factors, several studies have used whole genome data to establish an association between virulence factors and the pathogenic effect of strains, and to determine the pathogenic potential of H. pylori strains. The confirmation of virulence phenotype upon biomarker detection, however, is not simple due to the previously discussed complex relationship between virulence factors and phenotypic outcomes. Conversely, regarding antibiotic-resistance markers, several studies have addressed the coherence between genomic-based predictions of antibiotic resistance (with previously approached biomarkers) and the phenotypic evaluation of antibiotic resistance. Most studies found a very strong correlation between 23S rRNA markers for resistance and resistant phenotype [43][62][64], especially with the detection of the A2143G mutation [41], although not indisputably [65]. Regarding levofloxacin resistance, the literature reports a moderate to strong reliability for the detection of mutations on the gyrA gene [43][53][62]. Mutations on the pbp1 gene showed moderate to strong reliability for the prediction of amoxicillin resistance [62][65][66][67]. Less consistently, observations of metronidazole-resistance biomarkers have mostly been reported as fair predictors for metronidazole resistance [62][65][67], although some studies found only weak reliability for these observations [43][64][66]. Conversely, a good agreement between the truncated or mutated version of the rdxA gene and metronidazole-resistant phenotypes has been found [68]. A good correlation between the detection of 16S rRNA mutations and tetracycline resistance was also observed [65]. Furthermore, a close correlation between genotype data and phenotypic outcome has been observed [52]. Overall, existing studies suggest that genomic data can accurately predict antibiotic resistant phenotypes, especially for clarithromycin, levofloxacin and amoxicillin; however, it is necessary to highlight the importance of further research to uncover the complex phenotype-genotype correlation in H. pylori and to identify other antibiotic resistance mechanisms [52].
3.3. Current Tools as Possible Limiting Factors
3.4. New Applications: The Example of Methylome Analysis
References
- Marshall, B.J.; Warren, J.R. Unidentified curved bacilli in the stomach of patients with gastritis and peptic ulceration. Lancet 1984, 16, 1311–1315.
- Polk, D.B.; Peek, R.M. Helicobacter pylori: Gastric Cancer and Beyond. Nat. Rev. Cancer 2010, 10, 403–414.
- Oleastro, M.; Rocha, R.; Vale, F.F. Population Genetic Structure of Helicobacter pylori Strains from Portuguese-Speaking Countries. Helicobacter 2017, 22, e12382.
- Mégraud, F.; Lehours, P.; Vale, F.F. The History of Helicobacter pylori: From Phylogeography to Paleomicrobiology. Clin. Microbiol. Infect. 2016, 22, 922–927.
- Dastmalchi, N.; Safaralizadeh, R.; Banan Khojasteh, S.M. The Correlation between MicroRNAs and Helicobacter pylori in Gastric Cancer. Pathog. Dis. 2019, 77, ftz039.
- Ki, O.M.; Eun, J.S.; Hi, J.K.; Eui, J.L.; Won, I.K.; Chang, S.K.; Kim, K.M. Methylation of P16INK4A and P57KIP2 Are Involved in the Development and Progression of Gastric MALT Lymphomas. Mod. Pathol. 2006, 19, 141–148.
- International Agency for Research on Cancer; World Health Organization. Schistosomes, Liver Flukes and Helicobacter Pylori; International Agency for Research on Cancer (IARC): Lyon, France, 1994; ISBN 9283212614.
- Muhammad, J.; Zaidi, S.; Saeed, S.; Ishaq, M. Current Status of Helicobacter pylori Association with Haematological Andcardiovascular Diseases: A Mini Review. J. Pak. Med. Assoc. 2017, 67, 907–911.
- Franceschi, F.; Covino, M.; Roubaud Baudron, C. Review: Helicobacter pylori and Extragastric Diseases. Helicobacter 2019, 24, e12636.
- Tacconelli, E.; Carrara, E.; Savoldi, A.; Harbarth, S.; Mendelson, M.; Monnet, D.L.; Pulcini, C.; Kahlmeter, G.; Kluytmans, J.; Carmeli, Y.; et al. Discovery, Research, and Development of New Antibiotics: The WHO Priority List of Antibiotic-Resistant Bacteria and Tuberculosis. Lancet Infect. Dis. 2018, 18, 318–327.
- Achtman, M.; Azuma, T.; Berg, D.E.; Ito, Y.; Morelli, G.; Pan, Z.-J.; Suerbaum, S.; Thompson, S.A.; van der Ende, A.; van Doorn, L.-J. Recombination and Clonal Groupings within Helicobacter pylori from Different Geographical Regions. Mol. Microbiol. 1999, 32, 459–470.
- Björkholm, B.M.; Oh, J.D.; Falk, P.G.; Engstrand, L.G.; Gordon, J.I. Genomics and Proteomics Converge on Helicobacter pylori. Curr. Opin. Microbiol. 2001, 4, 237–245.
- Morelli, G.; Didelot, X.; Kusecek, B.; Schwarz, S.; Bahlawane, C.; Falush, D.; Suerbaum, S.; Achtman, M. Microevolution of Helicobacter pylori during Prolonged Infection of Single Hosts and within Families. PLoS Genet. 2010, 6, e1001036.
- Kojima, K.K.; Furuta, Y.; Yahara, K.; Fukuyo, M.; Shiwa, Y.; Nishiumi, S.; Yoshida, M.; Azuma, T.; Yoshikawa, H.; Kobayashi, I. Population Evolution of Helicobacter pylori through Diversification in DNA Methylation and Interstrain Sequence Homogenization. Mol. Biol. Evol. 2016, 33, 2848–2859.
- Yahara, K.; Lehours, P.; Vale, F.F. Analysis of Genetic Recombination and the Pan-Genome of a Highly Recombinogenic Bacteriophage Species. Microb. Genom. 2019, 5, e000282.
- Miftahussurur, M.; Fauzia, K.A.; Nusi, I.A.; Setiawan, P.B.; Syam, A.F.; Waskito, L.A.; Doohan, D.; Ratnasari, N.; Khomsan, A.; Adnyana, I.K.; et al. E-Test versus Agar Dilution for Antibiotic Susceptibility Testing of Helicobacter pylori: A Comparison Study. BMC Res. Notes 2020, 13, 22.
- Rhie, S.Y.; Park, J.Y.; Shin, T.S.; Kim, J.W.; Kim, B.J.; Kim, J.G. Discovery of a Novel Mutation in DNA Gyrase and Changes in the Fluoroquinolone Resistance of Helicobacter pylori over a 14-Year Period: A Single Center Study in Korea. Antibiotics 2020, 9, 287.
- Binh, T.T.; Shiota, S.; Suzuki, R.; Matsuda, M.; Trang, T.T.H.; Kwon, D.H.; Iwatani, S.; Yamaoka, Y. Discovery of Novel Mutations for Clarithromycin Resistance in Helicobacter pylori by Using Next-Generation Sequencing. J. Antimicrob. Chemother. 2014, 69, 1796–1803.
- Rimbara, E.; Noguchi, N.; Kawai, T.; Sasatsu, M. Correlation between Substitutions in Penicillin-Binding Protein 1 and Amoxicillin Resistance in Helicobacter pylori. Microbiol. Immunol. 2007, 51, 939–944.
- Mégraud, F.; Bénéjat, L.; Ontsira Ngoyi, E.N.; Lehours, P. Molecular Approaches to Identify Helicobacter pylori Antimicrobial Resistance. Gastroenterol. Clin. N. Am. 2015, 44, 577–596.
- Woo, H.-Y.; Park, D.I.; Park, H.; Kim, M.-K.; Kim, D.H.; Kim, I.-S.; Kim, Y.J. Dual-Priming Oligonucleotide-Based Multiplex PCR for the Detection of Helicobacter pylori and Determination of Clarithromycin Resistance with Gastric Biopsy Specimens. Helicobacter 2009, 14, 22–28.
- Elviss, N.C.; Lawson, A.J.; Owen, R.J. Application of 3′-Mismatched Reverse Primer PCR Compared with Real-Time PCR and PCR-RFLP for the Rapid Detection of 23S rDNA Mutations Associated with Clarithromycin Resistance in Helicobacter pylori. Int. J. Antimicrob. Agents 2004, 23, 349–355.
- Ménard, A.; Santos, A.; Mégraud, F.; Oleastro, M. PCR-Restriction Fragment Length Polymorphism Can Also Detect Point Mutation A2142C in the 23S rRNA Gene, Associated with Helicobacter pylori Resistance to Clarithromycin. Antimicrob. Agents Chemother. 2002, 46, 1156–1157.
- Oleastro, M.; Ménard, A.; Santos, A.; Lamouliatte, H.; Monteiro, L.; Barthélémy, P.; Mégraud, F. Real-Time PCR Assay for Rapid and Accurate Detection of Point Mutations Conferring Resistance to Clarithromycin in Helicobacter pylori. J. Clin. Microbiol. 2003, 41, 397–402.
- Cambau, E.; Allerheiligen, V.; Coulon, C.; Corbel, C.; Lascols, C.; Deforges, L.; Soussy, C.J.; Delchier, J.C.; Megraud, F. Evaluation of a New Test, GenoType HelicoDR, for Molecular Detection of Antibiotic Resistance in Helicobacter pylori. J. Clin. Microbiol. 2009, 47, 3600–3607.
- Lee, J.W.; Kim, N.; Nam, R.H.; Park, J.H.; Choi, Y.J.; Kim, J.M.; Kim, J.S.; Jung, H.C. GenoType HelicoDR Test in the Determination of Antimicrobial Resistance of Helicobacter pylori in Korea. Scand. J. Gastroenterol. 2014, 49, 1058–1067.
- Pastukh, N.; Binyamin, D.; On, A.; Paritsky, M.; Peretz, A. GenoType® HelicoDR Test in Comparison with Histology and Culture for Helicobacter pylori Detection and Identification of Resistance Mutations to Clarithromycin and Fluoroquinolones. Helicobacter 2017, 22, e12447.
- Tomb, J.F.; White, O.; Kerlavage, A.R.; Clayton, R.A.; Sutton, G.G.; Fleischmann, R.D.; Ketchum, K.A.; Klenk, H.P.; Gill, S.; Dougherty, B.A.; et al. The Complete Genome Sequence of the Gastric Pathogen Helicobacter pylori. Nature 1997, 388, 539–547.
- Alm, R.A.; Ling, L.S.L.; Moir, D.T.; King, B.L.; Brown, E.D.; Doig, P.C.; Smith, D.R.; Noonan, B.; Guild, B.C.; DeJonge, B.L.; et al. Genomic-Sequence Comparison of Two Unrelated Isolates of the Human Gastric Pathogen Helicobacter pylori. Nature 1999, 397, 176–180.
- Whittam, T.S.; Bumbaugh, A.C. Inferences from Whole-Genome Sequences of Bacterial Pathogens. Curr. Opin. Genet. Dev. 2002, 12, 719–725.
- Berthenet, E.; Sheppard, S.; Vale, F.F. Recent “Omics” Advances in Helicobacter pylori. Helicobacter 2016, 21, 14–18.
- Davis, J.J.; Wattam, A.R.; Aziz, R.K.; Brettin, T.; Butler, R.; Butler, R.M.; Chlenski, P.; Conrad, N.; Dickerman, A.; Dietrich, E.M.; et al. The PATRIC Bioinformatics Resource Center: Expanding Data and Analysis Capabilities. Nucleic Acids Res. 2020, 48, D606–D612.
- Liu, L.; Li, Y.; Li, S.; Hu, N.; He, Y.; Pong, R.; Lin, D.; Lu, L.; Law, M. Comparison of Next-Generation Sequencing Systems. J. Biomed. Biotechnol. 2012, 2012, 251364.
- Pavlovic, S.; Klaassen, K.; Stankovic, B.; Stojiljkovic, M.; Zukic, B. Next-Generation Sequencing: The Enabler and the Way Ahead. In Microbiomics; Elsevier: Amsterdam, The Netherlands, 2020; pp. 175–200.
- Phuc, B.H.; Tuan, V.P.; Dung, H.D.Q.; Binh, T.T.; Tung, P.H.; Tri, T.D.; Thuan, N.P.M.; van Khien, V.; Trang, T.T.H.; Akada, J.; et al. Helicobacter pylori Type 4 Secretion Systems as Gastroduodenal Disease Markers. Sci. Rep. 2021, 11, 4584.
- Silva, B.; Nunes, A.; Vale, F.F.; Rocha, R.; Gomes, J.P.; Dias, R.; Oleastro, M. The Expression of Helicobacter pylori Tfs Plasticity Zone Cluster Is Regulated by pH and Adherence, and Its Composition Is Associated with Differential Gastric IL-8 Secretion. Helicobacter 2017, 22, e12390.
- Marques, A.T.; Vítor, J.M.B.; Santos, A.; Oleastro, M.; Vale, F.F. Trends in Helicobacter pylori Resistance to Clarithromycin: From Phenotypic to Genomic Approaches. Microb. Genom. 2020, 6, e000344.
- Palau, M.; Piqué, N.; José Ramírez-Lázaro, M.; Lario, S.; Calvet, X.; Miñana-Galbis, D. Whole-Genome Sequencing and Comparative Genomics of Three Helicobacter pylori Strains Isolated from the Stomach of a Patient with Adenocarcinoma. Pathogens 2021, 10, 331.
- Thorell, K.; Hosseini, S.; Palacios Gonzáles, R.V.P.; Chaotham, C.; Graham, D.Y.; Paszat, L.; Rabeneck, L.; Lundin, S.B.; Nookaew, I.; Sjöling, Å. Identification of a Latin American-Specific BabA Adhesin Variant through Whole Genome Sequencing of Helicobacter pylori Patient Isolates from Nicaragua. BMC Evol. Biol. 2016, 16, 53.
- Imkamp, F.; Lauener, F.N.; Pohl, D.; Lehours, P.; Vale, F.F.; Jehanne, Q.; Zbinden, R.; Keller, P.M.; Wagner, K. Rapid Characterization of Virulence Determinants in Helicobacter pylori Isolated from Non-Atrophic Gastritis Patients by Next-Generation Sequencing. J. Clin. Med. 2019, 8, 30.
- Gong, E.J.; Ahn, J.Y.; Kim, J.M.; Lee, S.M.; Na, H.K.; Lee, J.H.; Jung, K.W.; Choi, K.D.; Kim, D.H.; Song, H.J.; et al. Genotypic and Phenotypic Resistance to Clarithromycin in Helicobacter pylori Strains. J. Clin. Med. 2020, 9, 1930.
- Iwamoto, A.; Tanahashi, T.; Okada, R.; Yoshida, Y.; Kikuchi, K.; Keida, Y.; Murakami, Y.; Yang, L.; Yamamoto, K.; Nishiumi, S.; et al. Whole-Genome Sequencing of Clarithromycin Resistant Helicobacter pylori Characterizes Unidentified Variants of Multidrug Resistant Efflux Pump Genes. Gut Pathog. 2014, 6, 27.
- Camorlinga-Ponce, M.; Gómez-Delgado, A.; Aguilar-Zamora, E.; Torres, R.C.; Giono-Cerezo, S.; Escobar-Ogaz, A.; Torres, J. Phenotypic and Genotypic Antibiotic Resistance Patterns in Helicobacter pylori Strains from Ethnically Diverse Population in México. Front. Cell. Infect. Microbiol. 2021, 10, 539115.
- Watanabe, Y.; Oikawa, R.; Kodaka, Y.; Sato, Y.; Ono, S.; Kenmochi, T.; Suzuki, H.; Futagami, S.; Kato, M.; Yamamoto, H.; et al. Cancer-Related Genetic Variants of Helicobacter pylori Strains Determined Using Gastric Wash-Based Whole-Genome Analysis with Single-Molecule Real-Time Technology. Int. J. Cancer 2021, 148, 178–192.
- Suzuki, R.; Satou, K.; Shiroma, A.; Shimoji, M.; Teruya, K.; Matsumoto, T.; Akada, J.; Hirano, T.; Yamaoka, Y. Genome-Wide Mutation Analysis of Helicobacter pylori after Inoculation to Mongolian Gerbils. Gut Pathog. 2019, 11, 45.
- Satou, K.; Shiroma, A.; Teruya, K.; Shimoji, M.; Nakano, K.; Juan, A.; Tamotsu, H.; Terabayashi, Y.; Aoyama, M.; Teruya, M.; et al. Complete Genome Sequences of Eight Helicobacter pylori Strains with Different Virulence Factor Genotypes and Methylation Profiles, Isolated from Patients with Diverse Gastrointestinal Diseases on Okinawa Island, Japan, Determined Using PacBio Single-Molecule Real-Time Technology. Genome Announc. 2014, 2, e00286-14.
- Mannion, A.; Dzink-Fox, J.; Shen, Z.; Piazuelo, M.B.; Wilson, K.T.; Correa, P.; Peek, R.M.; Constanza Camargo, M.; Fox, J.G. Helicobacter pylori Antimicrobial Resistance and Gene Variants in High-and Low-Gastric-Cancer-Risk Populations. J. Clin. Microbiol. 2021, 59, e03203-20.
- Mwangi, C.; Njoroge, S.; Tshibangu-Kabamba, E.; Moloo, Z.; Rajula, A.; Devani, S.; Matsumoto, T.; Nyerere, K.; Kariuki, S.; Revathi, G.; et al. Whole Genome Sequencing Reveals Virulence Potentials of Helicobacter pylori Strain KE21 Isolated from a Kenyan Patient with Gastric Signet Ring Cell Carcinoma. Toxins 2020, 12, 556.
- Duncan, S.S.; Valk, P.L.; McClain, M.S.; Shaffer, C.L.; Metcalf, J.A.; Bordenstein, S.R.; Cover, T.L. Comparative Genomic Analysis of East Asian and Non-Asian Helicobacter pylori Strains Identifies Rapidly Evolving Genes. PLoS ONE 2013, 8, e0055120.
- Mehrotra, T.; Devi, T.B.; Kumar, S.; Talukdar, D.; Karmakar, S.P.; Kothidar, A.; Verma, J.; Kumari, S.; Alexander, S.M.; Retnakumar, R.J.; et al. Antimicrobial Resistance and Virulence in Helicobacter pylori: Genomic Insights. Genomics 2021, 113, 3951–3966.
- Hayashi, H.; Inoue, J.; Oyama, K.; Matsuoka, K.; Nishiumi, S.; Yoshida, M.; Yano, Y.; Kodama, Y. Detection of Novel Amino Acid Polymorphisms in the East Asian CagA of Helicobacter pylori with Full Sequencing Data. Kobe J. Med. Sci. 2020, 66, E22–E31.
- Domanovich-Asor, T.; Craddock, H.A.; Motro, Y.; Khalfin, B.; Peretz, A.; Moran-Gilad, J. Unraveling Antimicrobial Resistance in Helicobacter pylori: Global Resistome Meets Global Phylogeny. Helicobacter 2021, 26, e12782.
- Cui, R.; Song, Z.; Suo, B.; Tian, X.; Xue, Y.; Meng, L.; Niu, Z.; Jin, Z.; Zhang, H.; Zhou, L. Correlation Analysis among Genotype Resistance, Phenotype Resistance and Eradication Effect of Helicobacter pylori. Infect. Drug Resist. 2021, 14, 1747–1756.
- Lang, J.; Zhu, R.; Sun, X.; Zhu, S.; Li, T.; Shi, X.; Sun, Y.; Yang, Z.; Wang, W.; Bing, P.; et al. Evaluation of the MGISEQ-2000 Sequencing Platform for Illumina Target Capture Sequencing Libraries. Front. Genet. 2021, 12, 730519.
- Rizzato, C.; Torres, J.; Plummer, M.; Muñoz, N.; Franceschi, S.; Camorlinga-Ponce, M.; Fuentes-Pananá, E.M.; Canzian, F.; Kato, I. Variations in Helicobacter pylori Cytotoxin-Associated Genes and Their Influence in Progression to Gastric Cancer: Implications for Prevention. PLoS ONE 2012, 7, e0029605.
- Nezami, B.G.; Jani, M.; Alouani, D. Helicobacter pylori Mutations Detected by Next-Generation-Sequencing in Formalin-Fixed, Paraffin-Embedded Gastric Biopsy Specimens Are Associated with Treatment Failure. J. Clin. Microbiol. 2019, 57, e01834-18.
- Canzian, F.; Rizzato, C.; Obazee, O.; Stein, A.; Flores-Luna, L.; Camorlinga-Ponce, M.; Mendez-Tenorio, A.; Vivas, J.; Trujillo, E.; Jang, H.; et al. Genetic Polymorphisms in the Cag Pathogenicity Island of Helicobacter pylori and Risk of Stomach Cancer and High-Grade Premalignant Gastric Lesions. Int. J. Cancer 2020, 147, 2437–2445.
- Aftab, H.; Miftahussurur, M.; Subsomwong, P.; Ahmed, F.; Khan, A.K.A.; Matsumoto, T.; Suzuki, R.; Yamaoka, Y. Two Populations of Less-Virulent Helicobacter pylori Genotypes in Bangladesh. PLoS ONE 2017, 12, e0182947.
- Miftahussurur, M.; Tuda, J.; Suzuki, R.; Kido, Y.; Kawamoto, F.; Matsuda, M.; Tantular, I.S.; Pusarawati, S.; Nasronudin; Harijanto, P.N.; et al. Extremely Low Helicobacter pylori Prevalence in North Sulawesi, Indonesia and Identification of a Maori-Tribe Type Strain: A Cross Sectional Study. Gut Pathog. 2014, 6, 42.
- Binh, T.T.; Suzuki, R.; Trang, T.T.H.; Kwon, D.H.; Yamaoka, Y. Search for Novel Candidate Mutations for Metronidazole Resistance in Helicobacter pylori Using Next-Generation Sequencing. Antimicrob. Agents Chemother. 2015, 59, 2343–2348.
- Hashinaga, M.; Suzuki, R.; Akada, J.; Matsumoto, T.; Kido, Y.; Okimoto, T.; Kodama, M.; Murakami, K.; Yamaoka, Y. Differences in Amino Acid Frequency in CagA and VacA Sequences of Helicobacter pylori Distinguish Gastric Cancer from Gastric MALT Lymphoma. Gut Pathog. 2016, 8, 54.
- Hulten, K.G.; Genta, R.M.; Kalfus, I.N.; Zhou, Y.; Zhang, H.; Graham, D.Y. Comparison of Culture With Antibiogram to Next-Generation Sequencing Using Bacterial Isolates and Formalin-Fixed, Paraffin-Embedded Gastric Biopsies. Gastroenterology 2021, 161, 1433–1442.e2.
- American Molecular Laboratories Inc. PyloriARTM/AmHPR® H. pylori Antibiotic Resistance Next Generation Sequencing Panel. Available online: http://amlaboratories.com/testing-services/helicobacter-pylori-detection-antibiotic-resistant-analysis/pyloriar-amhpr-h-pylori-antibiotic-resistance-next-generation-sequencing-panel/ (accessed on 28 March 2022).
- Lauener, F.N.; Imkamp, F.; Lehours, P.; Buissonnière, A.; Benejat, L.; Zbinden, R.; Keller, P.M.; Wagner, K. Genetic Determinants and Prediction of Antibiotic Resistance Phenotypes in Helicobacter pylori. J. Clin. Med. 2019, 8, 53.
- Azzaya, D.; Gantuya, B.; Oyuntsetseg, K.; Davaadorj, D.; Matsumoto, T.; Akada, J.; Yamaoka, Y. High Antibiotic Resistance of Helicobacter pylori and Its Associated Novel Gene Mutations among the Mongolian Population. Microorganisms 2020, 8, 1062.
- Tuan, V.P.; Narith, D.; Tshibangu-Kabamba, E.; Dung, H.D.Q.; Viet, P.T.; Sokomoth, S.; Binh, T.T.; Sokhem, S.; Tri, T.D.; Ngov, S.; et al. A Next-Generation Sequencing-Based Approach to Identify Genetic Determinants of Antibiotic Resistance in Cambodian Helicobacter pylori Clinical Isolates. J. Clin. Med. 2019, 8, 858.
- Tshibangu-Kabamba, E.; de Jesus Ngoma-Kisoko, P.; Tuan, V.P.; Matsumoto, T.; Akada, J.; Kido, Y.; Tshimpi-Wola, A.; Tshiamala-Kashala, P.; Ahuka-Mundeke, S.; Ngoy, D.M.; et al. Next-Generation Sequencing of the Whole Bacterial Genome for Tracking Molecular Insight into the Broad-Spectrum Antimicrobial Resistance of Helicobacter pylori Clinical Isolates from the Democratic Republic of Congo. Microorganisms 2020, 8, 887.
- Saranathan, R.; Levi, M.H.; Wattam, A.R.; Malek, A.; Asare, E.; Behin, D.S.; Pan, D.H.; Jacobs, W.R.; Szymczak, W.A. Helicobacter pylori Infections in the Bronx, New York: Surveying Antibiotic Susceptibility and Strain Lineage by Whole-Genome Sequencing. J. Clin. Microbiol. 2020, 58, e01591-19.
- Boolchandani, M.; D’Souza, A.W.; Dantas, G. Sequencing-Based Methods and Resources to Study Antimicrobial Resistance. Nat. Rev. Genet. 2019, 20, 356–370.
- Mégraud, F.H. pylori Antibiotic Resistance: Prevalence, Importance, and Advances in Testing. Gut 2004, 53, 1374–1384.
- Yusibova, M.; Hasman, H.; Clausen, P.T.L.C.; Imkamp, F.; Wagner, K.; Andersen, L.P. CRHP Finder, a Webtool for the Detection of Clarithromycin Resistance in Helicobacter pylori from Whole-Genome Sequencing Data. Helicobacter 2020, 25, e12752.
- Zankari, E.; Allesøe, R.; Joensen, K.G.; Cavaco, L.M.; Lund, O.; Aarestrup, F.M. PointFinder: A Novel Web Tool for WGS-Based Detection of Antimicrobial Resistance Associated with Chromosomal Point Mutations in Bacterial Pathogens. J. Antimicrob. Chemother. 2017, 72, 2764–2768.
- Ye, L.; Meng, F.; Mao, X.; Zhang, Y.; Wang, J.; Liu, Y.; Zhu, W.; Gu, B.; Huang, Q. Using Next-Generation Sequencing to Analyze Helicobacter pylori Clones with Different Levofloxacin Resistances from a Patient with Eradication Failure. Medicine 2020, 99, e20761.
- Alcock, B.P.; Raphenya, A.R.; Lau, T.T.Y.; Tsang, K.K.; Bouchard, M.; Edalatmand, A.; Huynh, W.; Nguyen, A.L.V.; Cheng, A.A.; Liu, S.; et al. CARD 2020: Antibiotic Resistome Surveillance with the Comprehensive Antibiotic Resistance Database. Nucleic Acids Res. 2020, 48, D517–D525.
- Gorrell, R.; Kwok, T. The Helicobacter pylori Methylome: Roles in Gene Regulation and Virulence. In Current Topics in Microbiology and Immunology; Springer: Berlin/Heidelberg, Germany, 2017; Volume 400, pp. 105–127. ISBN 9783319505206.
- Roberts, R.J.; Vincze, T.; Posfai, J.; Macelis, D. REBASE—Enzymes and Genes for DNA Restriction and Modification. Nucleic Acids Res. 2007, 35, D269–D270.
- Krebes, J.; Morgan, R.D.; Bunk, B.; Spröer, C.; Luong, K.; Parusel, R.; Anton, B.P.; König, C.; Josenhans, C.; Overmann, J.; et al. The Complex Methylome of the Human Gastric Pathogen Helicobacter pylori. Nucleic Acids Res. 2014, 42, 2415–2432.
- Gauntlett, J.C.; Nilsson, H.O.; Fulurija, A.; Marshall, B.J.; Benghezal, M. Phase-Variable Restriction/Modification Systems Are Required for Helicobacter pylori Colonization. Gut Pathog. 2014, 6, 35–40.
- Lee, W.C.; Anton, B.P.; Wang, S.; Baybayan, P.; Singh, S.; Ashby, M.; Chua, E.G.; Tay, C.Y.; Thirriot, F.; Loke, M.F.; et al. The Complete Methylome of Helicobacter pylori UM032. BMC Genom. 2015, 16, 424.
- Lamichhane, B.; Chua, E.G.; Wise, M.J.; Laming, C.; Marshall, B.J.; Tay, C.Y. The Complete Genome and Methylome of Helicobacter pylori HpNEAfrica Strain HP14039. Gut Pathog. 2019, 11, 7.





