1. Introduction
Bipolar disorder (BD) is a complex affective disorder integrating a group of clinical conditions characterized by biphasic episodes of mania or hypomania and depression expressed as recurrent episodic changes in mood, energy levels, thought, and behavior interpolated with periods of euthymia. BD includes bipolar I disorder, defined by the presence of a manic episode, and bipolar II disorder, defined by the presence of a hypomanic episode and a major depressive episode
[1][2][3].
The estimated lifetime prevalence is 0.6% for bipolar I disorder, 0.4% for bipolar II disorder, and 2.4% for the broader spectrum of bipolar disorders , explaining 0.4% of total disability-adjusted life years (DALYs) and 1.3% of total years lived with disability (YLDs) in 2013
[4]. BD usually begins in late adolescence or early adulthood, with around 75% of the cases having their first episode at that age period
[4]. This early-onset and the associated level of disability mean that BD is the 4th leading cause of global disease burden in adolescents and young adults
[4], disrupting the attainment of relationships and educational and occupational milestones. In addition, as bipolar disorders affect the economically active population, high costs to society are incurred in terms of direct healthcare costs and the costs of disability
[2].
A high prevalence of psychiatric and medical comorbidities is common in affected individuals. Bipolar disorders are comorbid with other psychiatric disorders, including anxiety disorders, substance use disorders, attention-deficit/hyperactivity disorders, and personality disorders. This comorbidity makes diagnosing and managing bipolar disorders more difficult and is associated with poorer outcomes
[1][2][3]. Cardiovascular disorders, diabetes, and obesity are highly comorbid with BD and arise earlier in the life course compared with the general population
[4][5]. The mortality gap between populations with BD and the general population is principally a result of excess deaths from cardiovascular disease and suicide, with a loss of approximately 10–20 potential years of life
[3].
Mental health services are increasingly adopting strategies to diagnose and treat BD as early as possible. The delay in diagnosis, as well as inappropriate treatment, can result in repeated mood episodes, persistent subthreshold symptoms, development of co-morbidities, and progression of the disease with cognitive impairment, functional decline
[6][7], and an excess of premature mortality
[8]. The high complexity, heterogeneity, and absence of biologically relevant diagnostic markers of BD are some of the factors that contribute to the misdiagnosis of the illness
[9].
Currently, the diagnosis of BD is made by a comprehensive clinical assessment with the administration of standardized diagnostic instruments, supplemented when possible with third-party information (e.g., family members, and clinical records). The identification of the various diagnostic categories that are part of the bipolar spectrum is made based on several operational criteria specified in international classification systems, such as the International Classification of Diseases, 11th revision (ICD-11)
[10], and the Diagnostic and Statistical Manual of Mental Disorders, 5th edition (DSM-5)
[11]. Additionally, clinical strategies applied to identify individuals at risk for BD remain mostly descriptive and have insufficient predictive validity, with only a minority of the individuals “at-risk” making the transition to BD
[12][13]. Given the progressive nature of BD, the delay in the correct diagnosis, and the risk of exposure to inappropriate pharmacological and psychosocial treatments, obtaining a timely and accurate diagnosis is extremely important. Thus, the identification of reliable biomarkers is becoming imperative.
Despite the extensive efforts to elucidate BD causes and mechanisms, these are still elusive
[14]. It is understood that the etiology and pathophysiology of BD involve several factors, including genetic, neurochemical, and environmental factors
[14][15]. Moreover, pathways underlying the neuroprogression in BD have been associated with immune dysfunction
[16][17], mitochondrial dysfunction
[18], oxidative stress
[19], neurotransmitter systems
[20][21], and impairment of cellular resilience and neural plasticity
[22]. In addition, emotional factors (e.g., maladaptive emotional processing and deficient emotion regulation) have been suggested as core factors in different psychopathologies, including bipolar disorders
[23].
To improve the knowledge about these complex disorders, omics approaches have arisen as relevant methodologies to achieve comprehensive information on disease pathogenesis and support the identification of reliable strategies for disease prediction and diagnoses
[24][25], towards the ultimate goal of improving patient care and outcome. However, the translation from research to successful clinical omics-based tests is still far from acceptable compared to the potential of these approaches.
Biomarker discovery, development, and application have been the center of extensive interest, especially with the recent emergence of new technologies such as proteomics-based approaches
[9]. Proteomics is considered a powerful tool in the omics field, examining the proteins expressed in a cell, tissue, or organism, enabling real-time evaluation of an individual state, health versus disease, and potentially predicting the susceptibility to develop a specific disorder
[26][27]. Extensive efforts have already been made toward the identification of biomarkers of several psychiatric disorders such as schizophrenia, depression, and BD
[28][29][30]. In mental health disorders, a biomarker, or a panel of biomarkers, may have several purposes: (i) to correctly diagnose and stratify a psychiatric patient in a field where several diseases may have overlapping clinical symptoms; (ii) to better classify at-risk individuals; (iii) to perform prognosis; (iv) to be used as therapeutic monitoring; and (v) to be used as predictive of therapy compliance
[13][31].
Proteomics strategies can generate several distinct levels of information, particularly (i) identification of proteins in a sample at a given moment; (ii) expression levels of proteins or quantitative proteomics; (iii) identification or quantification of post-translational modifications (such as phosphorylation, glycosylation, and acetylation, among others) of those proteins; (iv) determination of protein–protein interactions; and (v) proteomic functional studies
[9][32][33].
The increasing use of proteomics is closely related to the technological advances in mass spectrometry (MS), optimization in sample preparation, and computer data processing ability to deal with the large amount of information generated by the MS-based technologies
[33]. The success of MS in proteomics relies on its high specificity and sensitivity, mainly due to advances in liquid chromatography-tandem MS (LC-MS/MS), giving answers to different purposes and questions
[34].
Although in the beginning, MS-proteomics strategies were focused on screening approaches to qualitatively characterize proteins in complex matrices, over the last years, quantitative proteomics has become the analysis of choice when comparing proteomes, given that most of the biological changes of interest are slight differences in the amount of a protein present in a given situation and not an abrupt change testifying its presence or absence
[35]. Nowadays, proteomics tools usually combine qualitative and quantitative strategies, either using targeted or untargeted techniques to identify and quantify proteins in complex biological matrices
[36][37].
Numerous studies have employed MS-based techniques in BD, with most of the studies being initially performed in post-mortem brain tissue
[38][39][40]. Although very informative, brain tissue has some drawbacks, such as the susceptibility of confounding factors, namely age, chronicity of the disease and medication, and the fact that the tissue is static with no possibility of being manipulated, disturbed, or having longitudinal samplings
[41][42]. Although the search for biomarkers in psychiatric disorders started with brain tissue and cerebrospinal fluid samples, the whole body concept has emerged with great success based on the understanding that the brain and a variety of physiological conditions are reflected in the contents of body fluids
[43][44]. This link, created between the brain and the periphery, enhanced the search for biomarkers in body fluids that could be easily available, for example, in the blood (plasma, serum, and peripheral blood mononuclear cells—PBMCs)
[28][45][46], urine
[47], saliva
[48], and even tears
[49]. In this way, current proteomics studies have focused on peripheral fluids by integrating MS-based techniques towards a more comprehensive picture of BD concerning its onset, progression, and response to medication.
2. The Assessment of Protein Expression Differences Between BD Patients and Healthy Controls (CTR)
To evaluate the efficacy of MS proteomics applied to human peripheral fluids to assess BD biomarkers and identify relevant networks of biological pathways, a systematic review (following PRISMA Guidelines) and meta-analysis was performed. To do so, a search for studies using MS proteomics to identify proteomic differences between BD patients and healthy controls was performed. Overall, fourteen articles fulfilled the inclusion criteria, allowing the identification of 266 differentially expressed proteins in blood-related samples (plasma, serum, and PBMCs) and saliva.
Apolipoproteins (APOs) were one of the groups of proteins most found in BD vs. control studies as differentially expressed. In fact, nine studies reported the dysregulation of apolipoproteins
[28][29][45][46][50][51][52][53][54]. Apolipoproteins are amphipathic molecules capable of interacting with both the lipids of the lipoprotein core and the aqueous environment of the plasma, having a well-established role in the transport and metabolism of lipids; and in inflammatory and immune response regulation
[55][56][57]. Several studies have indicated apolipoproteins as potential candidates for psychiatric biomarkers, with extensive evidence that the levels of cholesterol and APOs may be disturbed in psychiatric disorders
[44][56][58]. Accordingly, in the selected studies, APO alterations (in plasma, serum, peripheral blood mononuclear cells, and peripheral lymphocytes) were associated with inflammatory response
[28][46][54] and lipid metabolism [
63]. APO alterations were also related to cognitive decline and underlying morphological changes
[29] and fat and vitamin digestion and absorption.
Apolipoprotein A1, APOA1, a major protein associated with lipid metabolism, was the apolipoprotein found as significantly altered in BD patients in more studies (total of seven), three in plasma
[29][45][53], three in serum
[28][46][50] and one in PBMCs
[54]. APOA1 is a constituent of the HDL fraction, and regulates plasma levels of free fatty acids, and has an important role in HDL and triglyceride-rich lipoprotein metabolism and in the reverse cholesterol transport pathway
[50]. However, APOA1 showed a heterogeneous behavior in the selected studies, expressed in an increasing behavior in five studies
[28][29][50][53][54], and a decreasing one in two studies
[45][46]. APOE, a protein with a critical function in lipoprotein-mediated lipid transport, was identified as differentially expressed in four studies, three in serum and one in plasma, being increased in three studies
[32][53][59] and decreased in one study
[28]. APOC3, an apolipoprotein capable of inhibiting lipoprotein lipase and hepatic lipase with genes closely linked with APOA1 and APOA4 in the human genome, was also identified in three studies as significantly altered, being increased in two studies
[29][52] and decreased in one study
[46]. Four APOs were identified in two independent studies; APOA4 and APOC2
[29][52] showed similar behavior in the two studies, expressing an increase in BD patients, whereas APOD and APOL1 showed a heterogeneous behavior. APOD was identified as up-regulated in the study of Smirnova et al.
[51], which was found in BD serum and absent in the controls and SCZ and down-regulated in the study of Knochel et al.
[29]. On the other hand, Knochel et al. found APOL1 up-regulated
[29], while Song et al. reported a down-regulation
[45].
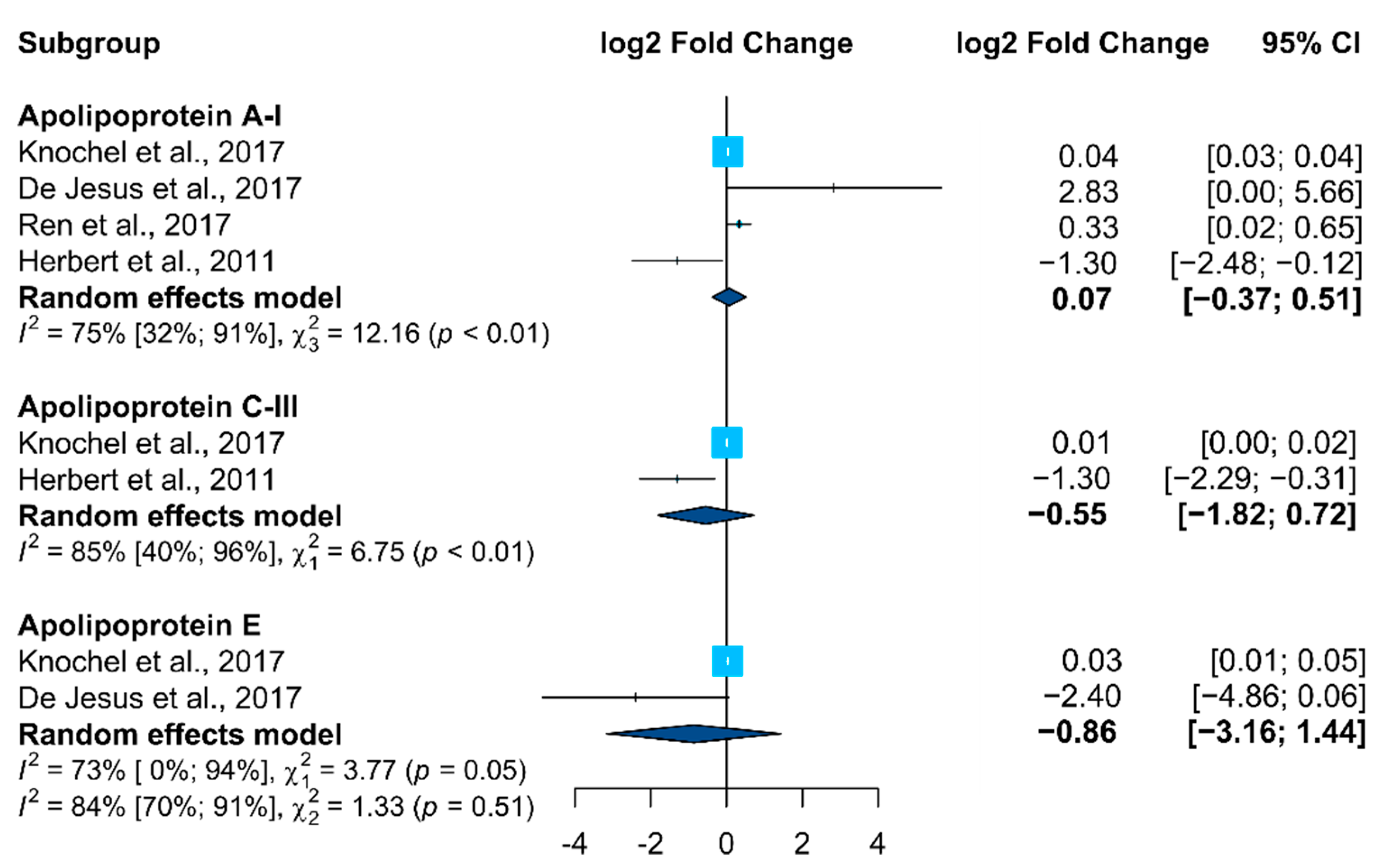
Only APOs (APOA1, APOC3, and APOE) fulfilled the required parameters (proteins in which it was possible to compute the effect size and corresponding significance in at least two research studies) to be included in the meta-analysis (Figure 1). In accordance, no clear alteration trend was observed for these proteins, with different behavior being identified in the selected studies.
Figure 1. Forest plot from the meta-analysis of proteins identified as altered in BD vs. control studies in at least two studies (95% CI, confidence intervals). Squares (whiskers represent 95% CI) indicate the effect sizes of the individual studies. The size of the squares reflects the sample size of each individual study. Diamonds represent summary statistics.
A set of four proteins was also identified in three selected studies as differentially expressed in BD patients, namely serotransferrin, TRFE
[46][60][61], complement C1q subcomponent subunit C, C1QC
[29][45][62], complement C4-A, CO4A
[28][45][63], and retinol-binding protein 4, RET4
[29][62][64]. However, these proteins did not fulfill the requirements to be added in the meta-analysis (lack of information related with their effect size and corresponding significance in the selected studies).
Transferrin (TRFE) is the main extracellular iron transport protein in blood that, when bound with iron, interacts with exofacial transferrin receptors allowing internalization of that complex of iron and proteins into cells
[65], being related to perturbations of iron uptake in subjects with a major depressive disorder associated with immune response and pro-inflammatory changes
[63][66]. TRFE was found in two studies to be significantly increased in BD patients
[45][63] and decreased in one study
[67], being associated with an inflammatory response in one of the studies
[63].
CO4A and C1Q are proteins involved in the complement system, which is a part of the innate immune system participating in the regulation of inflammation
[68][69].
CO4A is an essential component of the effector arm of the humoral immune response, playing a central role in activating the classical pathway of the complement system and mediating inflammatory processes. CO4A was found as increased in two studies
[45][63]. In Song et al., higher levels of complement factors, such as plasma C3 or C4 concentrations, as well as acute-phase proteins and a moderate increase of pro-inflammatory cytokines, were identified in BD patients suggesting that chronic, mild inflammatory processes in the peripheral and central nervous system (neuroinflammation) are involved in BD pathophysiology.
C1Q is a complex glycoprotein that mediates a variety of immunoregulatory functions considered important in preventing autoimmunities, such as enhancing phagocytosis, regulating cytokine production by antigen-presenting cells and subsequent alteration in T-lymphocyte maturation
[70]. C1Q was found to increase in two studies
[45][62], whereas in another study, it was found to decrease, although slightly (fold change = 0.99) and not statistically significant
[29]. In Song et al.
[45], a comparative proteomic study was performed to identify differentially expressed proteins in various BD mood states (depressed BD, manic BD, and euthymic BD) relative to healthy controls, an increased behavior of C1Q was only found in the maniac state, suggesting that C1Q may be involved in the pathophysiology of maniac episodes.
RET4 is mainly expressed in the liver with a primary function of transporting retinol (vitamin A) from the liver to peripheral tissues, with retinol being essential for the brain to facilitate learning, memory, and cognition
[71]. The RET4-retinol complex interacts with transthyretin and prealbumin, increasing the serum half-life of RET4
[72]. It is known that retinoid signaling plays a significant role in immune cell function; thus, it is suggested that factors affecting this system could have important implications for schizophrenia and other psychiatric disorders-associated inflammatory stress
[72]. RET4 was found as increased in three studies
[29][62][64]. The information was consistent; however, only one of those studies could compute the effect size
[29], highlighting the lack of complete statistical information in some selected studies (six out of 14) in a format amenable to a meta-analysis.
Following the same trend observed in APOs results, the trends for these proteins were also heterogeneous, with the different studies reporting a distinct protein behavior. This is an important observation, highlighting the urgent need to standardize proteomics strategies to obtain meaningful and consistent information about BD.
The immune system and inflammatory response were the most identified biological processes being altered in BD patients
[28][45][46][48][60][62][63][64][73][74]. These findings are consistent with current knowledge about BD, with the immune system and inflammatory response being strongly associated with the BD pathophysiology
[17][75][76]. Several mechanisms have been identified to explain the relationship between BD and immune dysfunction, including cytokine-induced monoamine changes, increased oxidative stress, pathological microglial over-activation, hypothalamic–pituitary–adrenal (HPA) axis over-activation, alterations of the microbiome–gut–brain axis and sleep-related immune change
[17]. In accordance, the regulation of the biological processes known as transport, morphological changes, cognitive impairment, oxidative stress lipid metabolism and activation of HPG (hypothalamic–pituitary–gonadal) axis hormones have been identified as altered in BD patients
[77][78][79].
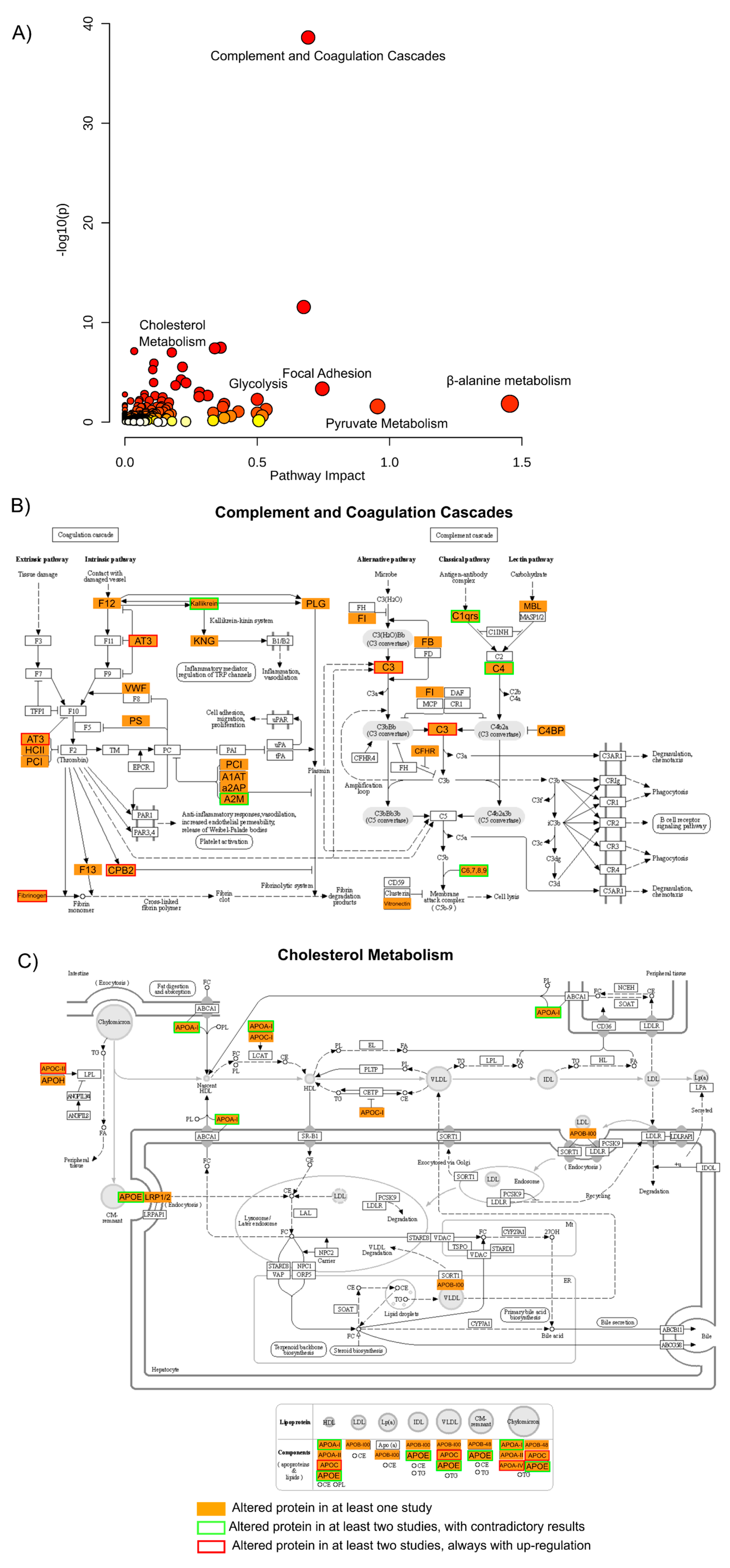
In order to validate these findings and integrate the biological meaning of the results from all the studies presented, a gene ontology (GO) analysis was performed where all proteins that were found to be altered in any of these studies were used (Figure 2).
Figure 2. Gene ontology analysis of all altered proteins. (
A) A gene ontology approach was used to assess pathway impact and enrichment (here presented by the
p-value and color scheme) of the 264 proteins described as altered in BD vs. CTR in at least one study, represented here as a scatter plot
[55]. From the pathways shown as enriched by this list of proteins, two were selected and their visual representation obtained through KEGG Mapper Color tool
[56]: (
B) complement and coagulation cascades, and (
C) cholesterol metabolism. In these last two panels, the proteins found in any of the studies are shown in orange, and proteins found to be altered in at least two studies are highlighted in red when the protein is always found to be up-regulated in BD cases or highlighted in green when the results from the two or more studies are contradictory.
From this ontological analysis, it is possible to observe that many proteins were found to be altered throughout the studies (264 proteins, two of the proteins identified as altered in the saliva study
[48] were not included in the gene ontology analysis as no accession number was available), but only 29 proteins (roughly 14%) were found to be altered in more than one study. From these 29 proteins, only 12 proteins are reported as having similar regulation trends in BD, in this case, up-regulation for all 12. The other 17 proteins are reported with contradictory regulations between the studies.
Several pathways were highlighted by performing this functional analysis, as shown in
Figure 2. The result of this analysis shows that metabolic pathways such as β-alanine or pyruvate metabolism (along with glycolysis or gluconeogenesis and the TCA cycle) are among the pathways with higher enrichment
p-value and pathway impact. This metabolic dysregulation of BD patients has been consistently reported, although with sometimes conflicting results, along with oxidative stress and the dysregulation of antioxidant systems
[80][81] also highlighted in this list with terms as glutathione or riboflavin metabolism.
Another pathway that is enriched by the proteins considered altered within all the studies is the complement and coagulation cascades, which present the lowest
p-value (and FDR corrected
p-value) for the enrichment of the pathway. Some of the proteins considered as altered in this pathway, with special emphasis on the complement cascade, have already been discussed above and are among the proteins that are consistently found to be altered in several studies. Many of these proteins are also related to immune response, and these pathways are also highlighted by this gene ontology analysis with terms related to bacterial infection or auto-immune disorders. Interestingly, proteins implicated in the complement and coagulation cascade have been highlighted in recently proposed proteomic prediction models of transition to clinical psychosis
[82].
As it has been discussed above, among the proteins most widely reported as altered in BD are apolipoproteins being the cholesterol metabolism one of the pathways presented with a lower enrichment
p-value. Several apolipoproteins are reported altered in more than one study, and although some are reported with inconsistent results (see meta-analysis above), others are reported as up-regulated in all studies. In particular, lipid metabolism and cholesterol metabolism have long since been associated with brain disorders, including BD, and some supplementation to the medication has already been reported as beneficial for the treatment of the disorder
[83].
Another pathway highlighted by this analysis, with high impact and low enrichment
p-value, is focal adhesion, where several key proteins were found to be altered throughout the studies, but only actin was reported in common in more than one study (with inconsistent results). This pathway has already been related to BD in other studies
[84], and here in this analysis, it has a high impact. Thus, it may be interesting to investigate and validate the potential of these proteins in BD diagnosis or physiology.
This ontological analysis focused primarily on proteins that were found to be altered between BD and control individuals, as this is the most widely reported comparison. Nonetheless, other comparisons are performed throughout the studies with as much or even more interest in investigating BD biomarkers and pathophysiology.
The recent interest in the comparisons of BD vs. SCZ and BD vs. other disorders studies reflects the increasing concern on the definition of disorder-specific biomarkers and the understanding of the associated altered biological pathways to improve diagnostic specificity.
Five studies had a comparison between BD and SCZ
[28][29][48][64][67], showing a strong heterogeneity of the type of biological sample analyzed, serum
[28][64][67], plasma
[29], and saliva
[48]. Following the same trend observed in BD vs. CTR studies, the immune system and inflammatory response were the most highlighted biological pathways
[28][48][64].
In the study performed by Smirnova et al.
[64], the definition of the proteome profiles of different groups revealed 27 proteins specific for schizophrenia (not present in BD) and 18 for BD. The specific proteins of schizophrenia were mostly associated with immune response, cell communication, cell growth and maintenance, protein metabolism and regulation of nucleic acid metabolism, whereas the protein set in BD was mostly associated with immune response, regulating transport processes across the cell membrane and cell communication, development of neurons and oligodendrocytes and cell growth.
In de Jesus et al.
[28], three unique proteins (CO4A, CO4-B, and SAMP) were identified as differentially abundant between BD vs. SCZ, being associated with the inflammatory response. All these proteins were found with lower abundance in the serum of BD patients when compared with serum from SCZ patients.
In the study performed by Knochel et al.
[29], the plasma protein expression in BD and SCZ patients was associated with cognitive deficits and their underlying brain structures. BD patients were found to have higher concentrations of the proteins A2AP, ANT3, ApoB, ApoD, and ApoF (all with
p < 0.05). The results suggested that detecting molecular patterns in association with cognitive performance and its underlying brain morphology was important to better understand the pathological mechanisms of BD and SCZ and, consequently, to support the diagnosis and treatment of both disorders.
The study using whole saliva samples
[48] confirmed a schizophrenia-associated dysregulation of the immune pathway of peripheral white blood cells and suggested that the dysregulation in the BD group could involve the activation of more specific cell types than that of the SCZ group. The significant increase in salivary concentrations of the eight proteins reported in SCZ and BD subjects confirms that these patients display strong innate immune system activation. Moreover, the very significant and higher correlations found between the levels of these proteins in BD subjects with respect to the SCZ group strongly suggest that the immune system activation in BD subjects could be linked to more specific cell types than in SCZ subjects.
In Pessoa et al.
[67], a metalloproteomics study was performed, allowing the identification of the proteins IGHG1 (both BD and SCZ), IGKV2D-28 (only in BD), and Ig lambda chain V-IV region Hil and ApoH (only in SCZ) as altered in BD and SCZ comparing to a healthy group and the identification of different concentrations of Li, Mg, Mn, and Zn in BD patients and high levels of Cu for SCZ patients, indicating an imbalance in the homeostasis of important micronutrients.
The comparison of BD vs. other disorders (OD) was assessed in five studies, reflecting comparisons with major depressive disorder (MDD)
[62][73][74], major depressive episode (MDE)
[63], and one study described as other mental disorders
[28]. Consistently with the previously mentioned comparisons, a strong heterogeneity of biological samples was also used, with the analysis of serum
[28][62], plasma
[73][74], and PBMCs
[63].
In Ren et al. studies
[73], the proteomics profiles of plasma samples were compared to differentiate between BD and MDD in the first depressive episode because of the potential treatment implications. Nine proteins were identified as significantly altered between MDD and BP, most of them related to the immune system. Particularly relevant, given the important role of B2RAN2 (highly similar to vanin-1) and endoglin in oxidative stress and the immune system, they may play roles in depression, being suggested as potential candidate biomarkers to distinguish between MDD and BP.
The association between vitamin D and inflammatory markers in adolescents with BD and MDD was studied by Petrov et al.
[62], allowing the identification of higher levels of D-binding protein (DBP) in BD patients and its suggestion as a marker candidate for BD.
BD is closely related to inflammatory processes, which were thoroughly investigated for their role in BD and MDD in the context of major inflammatory markers such as IL-6, MCP1, and TNFα. Accordingly, with the relevance of inflammatory response identified by de Jesus et al.
[28], the distinction between BD and OD was assessed, allowing to identify four proteins (VTN, ALB, CO4A, and APOC3) significantly altered and also associated with the inflammatory response.
3. Directions For Future Research
The recent advances in mass spectrometry proteomics approaches applied to human peripheral fluids allow the establishment of a robust platform for proteome profiling of clinical samples with an unprecedented depth. In fact, the MS ability to generate different levels of information about the individual proteome may lead to the comprehensive characterization of the biological network of pathways involved in BD, seeking the identification of reliable biomarkers of the disorder to improve prediction and diagnosis towards the ultimate goal of improving patient care and outcome.
However, for more specific clinical proteomics studies, a standardization of the studies’ characteristics is required, seeking to minimize the confounding factors. In fact, the clear definition of the study’s objectives and standardization of sociodemographic, clinical, and cognitive variables across the studied groups would make them more objective and specific, allowing a more comprehensive understanding of BD pathophysiology and increasing the possibility of identifying specific biomarkers of BD. This will minimize the confounding factors, leading to improvements in the statistical power and, consequently, to the efficiency of translating biomarker candidates and drug targets to the clinical application associated with the disorder.
The use of proteomics pipelines combining (i) standardized study conditions; (ii) high-throughput sample preparation techniques; (iii) high computational power for data processing and analysis will lead to a rapid expansion of clinical cohort sizes and consequently to more robust studies. An extra effort should be made to provide data in an open format so the community can re-analyze and perform larger studies based on data analysis from multiple centers. After full implementation of those proteomics pipelines, their application in extended clinical cohorts will allow taking into account the different variables (such as gender, comorbidities, illness duration, and treatment), leading to a more comprehensive understanding of BD pathophysiology and, consequently, increasing the possibility of identifying specific biomarkers of BD, seeking to improve prediction and diagnosis towards the ultimate goal of improving patient care and outcome.