
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Alyssa J. Matz | -- | 3331 | 2022-05-02 01:22:01 | | | |
| 2 | Amina Yu | + 51 word(s) | 3382 | 2022-05-05 04:26:53 | | |
Video Upload Options
Obesity-induced adipose tissue dysfunction is bolstered by chronic, low-grade inflammation and impairs systemic metabolic health. Adipose tissue macrophages (ATMs) perpetuate local inflammation but are crucial to adipose tissue homeostasis, exerting heterogeneous, niche-specific functions. Diversified macrophage actions are shaped through finely regulated factors, including microRNAs, which post-transcriptionally alter macrophage activation.
1. Introduction
1.1. Macrophages Are Crucial to Adipose Tissue Homeostasis and Perpetuate Inflammation
1.2. MicroRNA Biogenesis, Mechanism of Action, and Regulation of Macrophage Function
Heterogeneous macrophage functions are initiated by diverse, multilayered signals from their microenvironment and shaped by epigenetic regulatory elements such as microRNAs [11]. MicroRNAs are ~18–23 base pairs-long, non-coding RNAs that tune initiation, magnitude, and resolution of various cellular actions.
1.2.1. MicroRNA Mechanism of Action
In order for mature microRNAs to exert their function, they must be loaded onto an Argonaute (AGO) protein to form the RNA-induced silencing complex (RISC) [12]. The mature RISC is capable of binding target mRNAs, causing mRNA instability that can reduce both target gene mRNA and protein abundance. In humans, there are four distinct AGO proteins (AGO 1–4); all four can bind microRNAs to repress target mRNA translation. However, Ago2 can also slice target mRNAs to reduce expression further.
All these factors should be considered when assessing potential mRNA targets. Several algorithms have been developed to predict microRNA targets that incorporate these factors [14]. Further experimental validation is necessary to demonstrate RISC-mediated mRNA repression.
1.2.2. MicroRNA Regulation in Immune Cells
MicroRNAs provide an additional layer of post-transcriptional regulation of genes. More than 60% of human protein-coding genes contain at least one conserved microRNA-target site [15]. Multiple ones have demonstrated the importance of microRNA regulation in immune functions during homeostasis and under stress [11][16]. The concept that microRNA network motifs can regulate cellular processes through both positive and negative feedback loops provides further complexity to microRNA-regulated immune cell formation and function. For example, to ensure fine-tuned cellular actions, microRNAs can establish a threshold for master regulators to a narrow range, which is crucial for tightly controlled hematopoiesis or swift immune cell response [11].
Table 1. MicroRNAs and confirmed mRNA targets that regulate macrophage functions important to AT function.
| MicroRNA | Effects | Confirmed mRNA Targets | Source |
|---|---|---|---|
| miR-155 | Pro-inflammatory activation | Socs1, Ship1, IL7Rα1 | [17][18][19][20][21][22] |
| Pro-foam cell | Hbp1 | [23] | |
| miR-125b | Anti-inflammatory activation | TNF-α | [21][24] |
| miR-223 | Anti-inflammatory activation | Nfat5, Rasa1, PKNOX1 | [25][26][27][28] |
| miR-21 | Pro-fibroblast-activating interactions | [29][30] | |
| miR-142-5p | Pro-fibroblast-activating interactions | SOCS1 | [31] |
| miR-33 | Anti-cholesterol efflux/ Metabolic reprogramming |
ABCA1/Abcg1/PGC-1α/PDK4/SLC25A25 | [32][33][34][35][36][37] |
| miR-342-5p | Pro-foam cell | Akt1 | [38] |
| miR-146a | Anti-inflammatory activation | IRAK1, TRAF6 | [17][39][40] |
2. Adipose Tissue Changes in Obesity, Macrophage Responsibilities, and microRNA Regulation
2.1. Adipose Tissue Adipokines and Insulin Sensitivity
2.2. Adipogenesis and Angiogenesis
2.3. Extracellular Remodeling
2.4. Lipid Storage and Mobilization
2.5. Age-Related Changes
Obesity-induced inflammation in adolescence and middle age accelerates the onset of declining functionality characteristic of advanced age, increasing the risk for morbidity and mortality [103]. Reciprocally, age-related disorder in AT augments obesity-induced dysfunction. Both aging and obesity are marked by metabolic dysfunction linked to chronic, low-grade inflammation. Aging defines a gradual deterioration of functionality across tissues; likewise, age-related inflammation is juxtaposed to impaired immune function. One hallmark of aging is stem cell exhaustion, best exemplified in hematopoiesis. With each division, hematopoietic stem cells (HSCs) exhibit lower self-renewal capacity and myeloid-biased differentiation [104][105]. The increased proportion of macrophages exhibits dysfunction. In aged mice, overall macrophage capacity for autophagy and phagocytosis dissipates, and ATM populations produce greater levels of pro-inflammatory cytokines [39][106].
The level of miR-146a is much higher in aged compared to young peritoneal macrophages in mice [40]. MiR-146a imposes tolerance to pro-inflammatory cytokines and danger-associated molecular patterns (DAMPs) by targeting key cytokine-receptor and TLR adaptor molecules interleukin 1 receptor-associated kinase 1 (IRAK1) and TNF receptor-associated factor 6 (TRAF6) [17][37]. Chronic overexpression of miR-146a prevented M1 effector activation (IL-1β, IL-6, TNF-α production) in response to bacterial lipopolysaccharide (LPS) stimulation in vitro [40]. In addition, peritoneal macrophage miR-33 increases in aged mice, leading to reduced cholesterol efflux [107]. MicroRNA regulation in macrophage actions under chronic conditions like obesity and aging-related disorder has yielded valuable insight into epigenetic factors controlling disease risk.
3. How to Capture Dynamic Actions of Macrophages in Obese Adipose Tissue
3.1. Investigation Strategy
MicroRNAs’ regulatory power and potential as disease biomarkers have motivated rapid advancement in microRNA detection and experimentation. Strategies for microRNA detection can be found elsewhere [108]. The M1/M2 model of macrophage activation is an essential tool for in vitro validation of microRNA regulation of macrophage activation; however, more complex models are needed to understand ATMs and other in vivo macrophage functions. Investigations using the M1/M2 model have defined miR-155 and miR-223 as crucial regulators of macrophage activation towards pro-inflammatory or anti-inflammatory actions.
3.2. Incorporating Macrophages Diversity in Studies
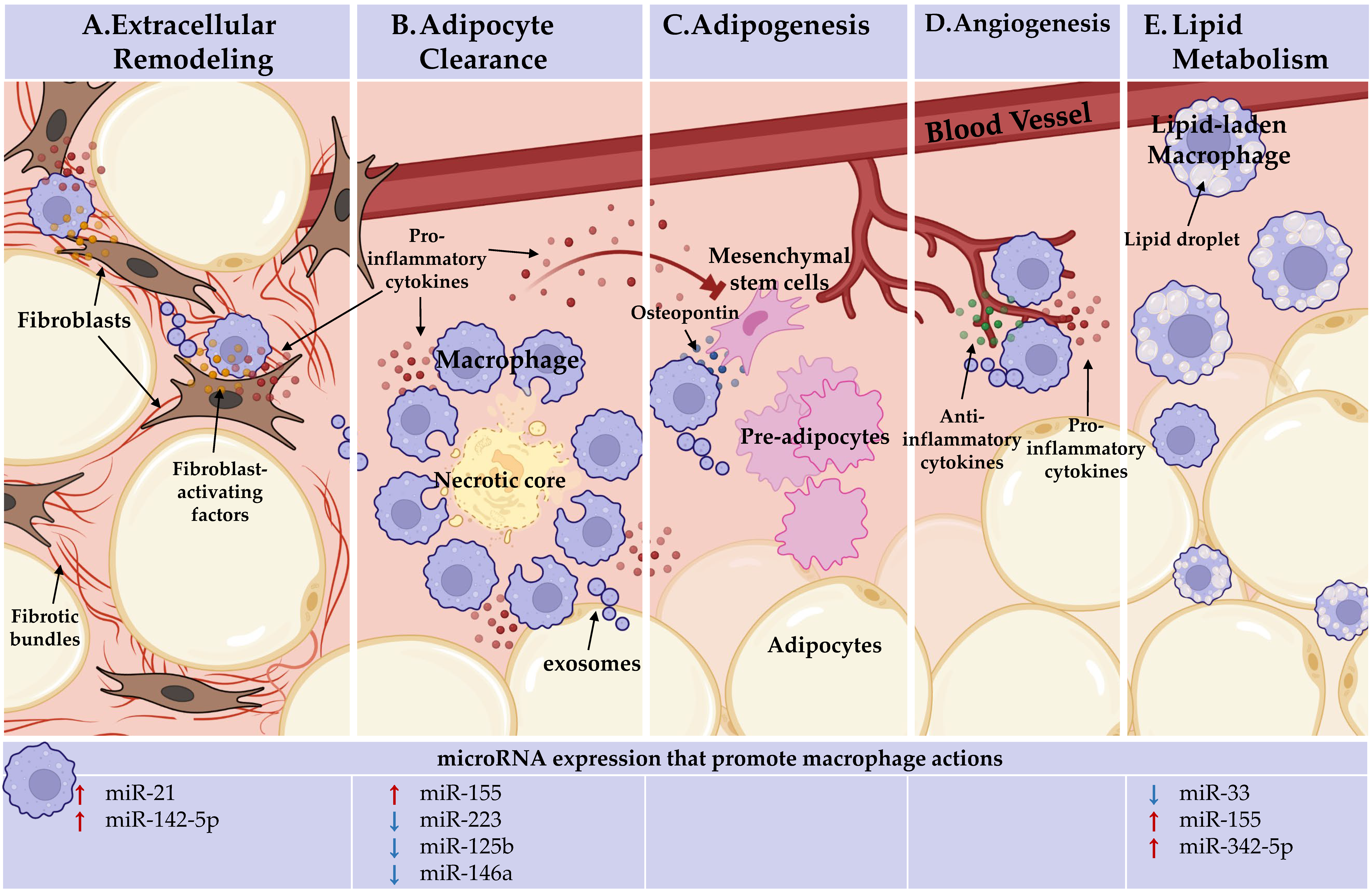
Figure 1. Adipose tissue macrophages (ATMs) perform heterogeneous, niche-specific functions and under obese stress perpetuate pathologic inflammation. In the top panel, ATM functions. In the bottom panel, the microRNAs known to endorse (red arrows) or inhibit (blue arrows) these macrophage actions are detailed. (A) Extracellular remodeling: ATMs support AT extracellular matrix remodeling. ATMs can activate fibroblasts through direct interactions and paracrine signaling, licensed by miR-21 and miR-142-5p, to increase tissue fibrosis under obesity. (B). Adipocyte clearance: pro-inflammatory ATMs clear dead and dying adipocytes in crown-like structures (CLS) and miR-155 supports, whereas miR-223, miR-125b, and miR-146a repress pro-inflammatory macrophage activation. (C) Adipogenesis: ATMs recruit progenitor cells through osteopontin secretion. Inflammatory ATM cytokine signaling within CLS, which miR-155, miR-223, miR-125b, and miR-146a regulate, impacts progenitors. ATMs may deliver microRNAs through exosomes to regulate adipogenesis. No endogenous role for microRNAs in ATM promotion of adipogenesis has been identified. (D) Angiogenesis: pro-inflammatory and anti-inflammatory-activated macrophages are required for de novo vessel outgrowth. Similar to adipogenesis, ATM may regulate angiogenesis through microRNA-containing exosomes. Beyond the importance of macrophage activation status, no endogenous role for microRNAs in ATM promotion of angiogenesis has been identified. (E) Lipid metabolism: ATMs uptake and metabolize lipids. Macrophage lipid efflux is repressed by miR-33. MiR-155 and miR-342-59 may play a role as they are upregulated during fatty plaque formation in vessels (atherosclerosis), but their networks towards lipid efflux has not been defined. Figure Created with BioRender.com (Accessed 13 February, 2022).
References
- Weisberg, S.P.; McCann, D.; Desai, M.; Rosenbaum, M.; Leibel, R.L.; Ferrante, A.W. Obesity is associated with macrophage accumulation in adipose tissue. J. Clin. Invest. 2003, 112, 1796–1808.
- Lumeng, C.N.; Bodzin, J.L.; Saltiel, A.R. Obesity induces a phenotypic switch in adipose tissue macrophage polarization. J. Clin. Investig. 2007, 117, 175–184.
- Coats, B.R.; Schoenfelt, K.Q.; Barbosa-Lorenzi, V.C.; Peris, E.; Cui, C.; Hoffman, A.; Zhou, G.; Fernandez, S.; Zhai, L.; Hall, B.A.; et al. Metabolically Activated Adipose Tissue Macrophages Perform Detrimental and Beneficial Functions during Diet-Induced Obesity. Cell Rep. 2017, 20, 3149–3161.
- Thomas, D.; Apovian, C. Macrophage functions in lean and obese adipose tissue. Metabolism 2017, 72, 120–143.
- Kanda, H.; Tateya, S.; Tamori, Y.; Kotani, K.; Hiasa, K.-I.; Kitazawa, R.; Kitazawa, S.; Miyachi, H.; Maeda, S.; Egashira, K.; et al. MCP-1 contributes to macrophage infiltration into adipose tissue, insulin resistance, and hepatic steatosis in obesity. J. Clin. Investig. 2006, 116, 1494–1505.
- Amano, S.U.; Cohen, J.L.; Vangala, P.; Tencerova, M.; Nicoloro, S.M.; Yawe, J.C.; Shen, Y.; Czech, M.P.; Aouadi, M. Local Proliferation of Macrophages Contributes to Obesity-Associated Adipose Tissue Inflammation. Cell Metab. 2014, 19, 162–171.
- Lavin, Y.; Mortha, A.; Rahman, A.; Merad, M. Regulation of macrophage development and function in peripheral tissues. Nat. Rev. Immunol. 2015, 15, 731–744.
- Scott, C.; Zheng, F.; De Baetselier, P.; Martens, L.; Saeys, Y.; De Prijck, S.; Lippens, S.; Abels, C.; Schoonooghe, S.; Raes, G.; et al. Bone marrow-derived monocytes give rise to self-renewing and fully differentiated Kupffer cells. Nat. Commun. 2016, 7, 10321.
- Odegaard, J.I.; Ricardo-Gonzalez, R.R.; Goforth, M.H.; Morel, C.R.; Subramanian, V.; Mukundan, L.; Eagle, A.R.; Vats, D.; Brombacher, F.; Ferrante, A.W.; et al. Macrophage-specific PPARgamma controls alternative activation and improves insulin resistance. Nature 2007, 447, 1116–1120.
- Patsouris, D.; Li, P.-P.; Thapar, D.; Chapman, J.; Olefsky, J.M.; Neels, J.G. Ablation of CD11c-Positive Cells Normalizes Insulin Sensitivity in Obese Insulin Resistant Animals. Cell Metab. 2008, 8, 301–309.
- Mehta, A.; Baltimore, D. MicroRNAs as regulatory elements in immune system logic. Nat. Rev. Immunol. 2016, 16, 279–294.
- Hammond, S.M.; Boettcher, S.; Caudy, A.A.; Kobayashi, R.; Hannon, G.J. Argonaute2, a Link Between Genetic and Biochemical Analyses of RNAi. Science 2001, 293, 1146–1150.
- Lewis, B.P.; Shih, I.-H.; Jones-Rhoades, M.W.; Bartel, D.P.; Burge, C.B. Prediction of Mammalian MicroRNA Targets. Cell 2003, 115, 787–798.
- Bartel, D.P. MicroRNAs: Target Recognition and Regulatory Functions. Cell 2009, 136, 215–233.
- Friedman, R.C.; Farh, K.K.-H.; Burge, C.B.; Bartel, D.P. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 2009, 19, 92–105.
- Lodish, H.F.; Zhou, B.; Liu, G.; Chen, C.-Z. Micromanagement of the immune system by microRNAs. Nat. Rev. Immunol. 2008, 8, 120–130.
- Taganov, K.D.; Boldin, M.P.; Chang, K.-J.; Baltimore, D. NF-κB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc. Natl. Acad. Sci. USA 2006, 103, 12481–12486.
- O’Connell, R.M.; Taganov, K.D.; Boldin, M.P.; Cheng, G.; Baltimore, D. MicroRNA-155 is induced during the macrophage inflammatory response. Proc. Natl. Acad. Sci. USA 2007, 104, 1604–1609.
- Androulidaki, A.; Iliopoulos, D.; Arranz, A.; Doxaki, C.; Schworer, S.; Zacharioudaki, V.; Margioris, A.N.; Tsichlis, P.N.; Tsatsanis, C. The Kinase Akt1 Controls Macrophage Response to Lipopolysaccharide by Regulating MicroRNAs. Immunity 2009, 31, 220–231.
- O’Connell, R.M.; Chaudhuri, A.A.; Rao, D.S.; Baltimore, D. Inositol phosphatase SHIP1 is a primary target of miR-155. Proc. Natl. Acad. Sci. USA 2009, 106, 7113–7118.
- Rajaram, M.V.S.; Ni, B.; Morris, J.D.; Brooks, M.N.; Carlson, T.K.; Bakthavachalu, B.; Schoenberg, D.R.; Torrelles, J.B.; Schlesinger, L.S. Mycobacterium tuberculosis lipomannan blocks TNF biosynthesis by regulating macrophage MAPK-activated protein kinase 2 (MK2) and microRNA miR-125b. Proc. Natl. Acad. Sci. USA 2011, 108, 17408–17413.
- Martinez-Nunez, R.T.; Louafi, F.; Sanchez-Elsner, T. The Interleukin 13 (IL-13) Pathway in Human Macrophages Is Modulated by MicroRNA-155 via Direct Targeting of Interleukin 13 Receptor α1 (IL13Rα1). J. Biol. Chem. 2011, 286, 1786–1794.
- Tian, F.; An, L.-N.; Wang, G.-K.; Zhu, J.-Q.; Li, Q.; Zhang, Y.-Y.; Zeng, A.; Zou, J.; Zhu, R.-F.; Han, X.-S.; et al. Elevated microRNA-155 promotes foam cell formation by targeting HBP1 in atherogenesis. Cardiovasc. Res. 2014, 103, 100–110.
- Rückerl, D.; Jenkins, S.; Laqtom, N.N.; Gallagher, I.; Sutherland, T.; Duncan, S.; Buck, A.; Allen, J. Induction of IL-4Rα–dependent microRNAs identifies PI3K/Akt signaling as essential for IL-4–driven murine macrophage proliferation in vivo. Blood 2012, 120, 2307–2316.
- Li, T.; Morgan, M.J.; Choksi, S.; Zhang, Y.; Kim, Y.-S.; Liu, Z.-G. MicroRNAs modulate the noncanonical transcription factor NF-κB pathway by regulating expression of the kinase IKKα during macrophage differentiation. Nat. Immunol. 2010, 11, 799–805.
- Zhuang, G.; Meng, C.; Guo, X.; Cheruku, P.S.; Shi, L.; Xu, H.; Li, H.; Wang, G.; Evans, A.R.; Safe, S.; et al. A novel regulator of macrophage activation: miR-223 in obesity-associated adipose tissue inflammation. Circulation 2012, 125, 2892–2903.
- Ying, W.; Tseng, A.; Chang, R.C.-A.; Morin, A.; Brehm, T.; Triff, K.; Nair, V.; Zhuang, G.; Song, H.; Kanameni, S.; et al. MicroRNA-223 is a crucial mediator of PPARγ-regulated alternative macrophage activation. J. Clin. Investig. 2015, 125, 4149–4159.
- Zhou, H.; Xiao, J.; Wu, N.; Liu, C.; Xu, J.; Liu, F.; Wu, L. MicroRNA-223 Regulates the Differentiation and Function of Intestinal Dendritic Cells and Macrophages by Targeting C/EBPβ. Cell Rep. 2015, 13, 1149–1160.
- Pan, T.; Jia, P.; Chen, N.; Fang, Y.; Liang, Y.; Guo, M.; Ding, X. Delayed Remote Ischemic Preconditioning ConfersRenoprotection against Septic Acute Kidney Injury via Exosomal miR-21. Theranostics 2019, 9, 405–423.
- Ramanujam, D.; Schön, A.P.; Beck, C.; Vaccarello, P.; Felician, G.; Dueck, A.; Esfandyari, D.; Meister, G.; Meitinger, T.; Schulz, C.; et al. MicroRNA-21–Dependent Macrophage-to-Fibroblast Signaling Determines the Cardiac Response to Pressure Overload. Circulation 2021, 143, 1513–1525.
- Su, S.; Zhao, Q.; He, C.; Huang, D.; Liu, J.; Chen, F.; Chen, J.; Liao, J.-Y.; Cui, X.; Zeng, Y.; et al. miR-142-5p and miR-130a-3p are regulated by IL-4 and IL-13 and control profibrogenic macrophage program. Nat. Commun. 2015, 6, 8523.
- Najafi-Shoushtari, S.H.; Kristo, F.; Li, Y.; Shioda, T.; Cohen, D.E.; Gerszten, R.E.; Näär, A.M. MicroRNA-33 and the SREBP Host Genes Cooperate to Control Cholesterol Homeostasis. Science 2010, 328, 1566–1569.
- Rayner, K.; Sheedy, F.; Esau, C.C.; Hussain, F.N.; Temel, R.E.; Parathath, S.; van Gils, J.; Rayner, A.J.; Chang, A.N.; Suarez, Y.; et al. Antagonism of miR-33 in mice promotes reverse cholesterol transport and regression of atherosclerosis. J. Clin. Investig. 2011, 121, 2921–2931.
- Rayner, K.J.; Suárez, Y.; Dávalos, A.; Parathath, S.; Fitzgerald, M.L.; Tamehiro, N.; Fisher, E.A.; Moore, K.J.; Fernández-Hernando, C. MiR-33 Contributes to the Regulation of Cholesterol Homeostasis. Science 2010, 328, 1570–1573.
- Horie, T.; Ono, K.; Horiguchi, M.; Nishi, H.; Nakamura, T.; Nagao, K.; Kinoshita, M.; Kuwabara, Y.; Marusawa, H.; Iwanaga, Y.; et al. MicroRNA-33 encoded by an intron of sterol regulatory element-binding protein 2 ( Srebp2 ) regulates HDL in vivo. Proc. Natl. Acad. Sci. USA 2010, 107, 17321–17326.
- Karunakaran, D.; Thrush, A.B.; Nguygen, M.-A.; Richards, L.; Geoffrion, M.; Singaravelu, R.; Ramphos, E.; Shangari, P.; Ouimet, M.; Pezacki, J.P.; et al. Macrophage Mitochondrial Energy Status Regulates Cholesterol Efflux and Is Enhanced by Anti-miR33 in Atherosclerosis. Circ. Res. 2015, 117, 266–278.
- Dai, R.; Phillips, R.A.; Zhang, Y.; Khan, D.; Crasta, O.; Ahmed, S.A. Suppression of LPS-induced Interferon-γ and nitric oxide in splenic lymphocytes by select estrogen-regulated microRNAs: A novel mechanism of immune modulation. Blood 2008, 112, 4591–4597.
- Wei, Y.; Nazari-Jahantigh, M.; Chan, L.; Zhu, M.; Heyll, K.; Corbalán-Campos, J.; Hartmann, P.; Thiemann, A.; Weber, C.; Schober, A. The microRNA-342-5p Fosters Inflammatory Macrophage Activation Through an Akt1- and microRNA-155 –Dependent Pathway During Atherosclerosis. Circulation 2013, 127, 1609–1619.
- Inomata, M.; Xu, S.; Chandra, P.; Meydani, S.N.; Takemura, G.; Philips, J.A.; Leong, J.M. Macrophage LC3-associated phagocytosis is an immune defense against Streptococcus pneumoniae that diminishes with host aging. Proc. Natl. Acad. Sci. USA 2020, 117, 33561–33569.
- Jiang, M.; Xiang, Y.; Wang, D.; Gao, J.; Liu, D.; Liu, Y.; Liu, S.; Zheng, D. Dysregulated expression of miR-146a contributes to age-related dysfunction of macrophages. Aging Cell 2011, 11, 29–40.
- Hotamisligil, G.S.; Murray, D.L.; Choy, L.N.; Spiegelman, B.M. Tumor necrosis factor alpha inhibits signaling from the insulin receptor. Proc. Natl. Acad. Sci. USA 1994, 91, 4854–4858.
- Hotamisligil, G.S.; Peraldi, P.; Budavari, A.; Ellis, R.; White, M.F.; Spiegelman, B.M. IRS-1-Mediated Inhibition of Insulin Receptor Tyrosine Kinase Activity in TNF-α- and Obesity-Induced Insulin Resistance. Science 1996, 271, 665–670.
- Laurencikiene, J.; van Harmelen, V.; Arvidsson Nordström, E.; Lennart, B.; Erik, N.; Dominique, L.; Peter, A.; Mikael, R. NF-kappaB is important for TNF-alpha-induced lipolysis in human adipocytes. J. Lipid Res. 2007, 48, 1069–1077.
- Hotamisligil, G.S. Endoplasmic Reticulum Stress and the Inflammatory Basis of Metabolic Disease. Cell 2010, 140, 900–917.
- Ranjit, S.; Boutet, E.; Gandhi, P.; Prot, M.; Tamori, Y.; Chawla, A.; Greenberg, A.S.; Puri, V.; Czech, M.P. Regulation of fat specific protein 27 by isoproterenol and TNF-α to control lipolysis in murine adipocytes. J. Lipid Res. 2011, 52, 221–236.
- Gastaldelli, A.; Gaggini, M.; DeFronzo, R.A. Role of Adipose Tissue Insulin Resistance in the Natural History of Type 2 Diabetes: Results From the San Antonio Metabolism Study. Diabetes 2017, 66, 815–822.
- Zhu, L.; Yang, T.; Li, L.; Sun, L.; Hou, Y.; Hu, X.; Zhang, L.; Tian, H.; Zhao, Q.; Peng, J.; et al. TSC1 controls macrophage polarization to prevent inflammatory disease. Nat. Commun. 2014, 5, 4696.
- Tili, E.; Michaille, J.-J.; Cimino, A.; Costinean, S.; Dumitru, C.D.; Adair, B.; Fabbri, M.; Alder, H.; Liu, C.G.; Calin, G.; et al. Modulation of miR-155 and miR-125b Levels following Lipopolysaccharide/TNF-α Stimulation and Their Possible Roles in Regulating the Response to Endotoxin Shock. J. Immunol. 2007, 179, 5082–5089.
- Koch, M.; Mollenkopf, H.-J.; Klemm, U.; Meyer, T.F. Induction of microRNA-155 is TLR- and type IV secretion system-dependent in macrophages and inhibits DNA-damage induced apoptosis. Proc. Natl. Acad. Sci. USA 2012, 109, E1153–E1162.
- Valadi, H.; Ekström, K.; Bossios, A.; Sjöstrand, M.; Lee, J.J.; Lötvall, J.O. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654–659.
- Ying, W.; Riopel, M.; Bandyopadhyay, G.; Dong, Y.; Birmingham, A.; Seo, J.B.; Ofrecio, J.M.; Wollam, J.; Hernandez-Carretero, A.; Fu, W.; et al. Adipose Tissue Macrophage-Derived Exosomal miRNAs Can Modulate In Vivo and In Vitro Insulin Sensitivity. Cell 2017, 171, 372–384.e12.
- Fazi, F.; Racanicchi, S.; Zardo, G.; Starnes, L.M.; Mancini, M.; Travaglini, L.; Diverio, D.; Ammatuna, E.; Cimino, G.; Lo-Coco, F.; et al. Epigenetic silencing of the myelopoiesis regulator microRNA-223 by the AML1/ETO oncoprotein. Cancer Cell 2007, 12, 457–466.
- Johnnidis, J.B.; Harris, M.H.; Wheeler, R.T.; Stehling-Sun, S.; Lam, M.H.; Kirak, O.; Brummelkamp, T.R.; Fleming, M.; Camargo, F.D. Regulation of progenitor cell proliferation and granulocyte function by microRNA-223. Nature 2008, 451, 1125–1129.
- Liu, G.; Abraham, E. MicroRNAs in Immune Response and Macrophage Polarization. Arter. Thromb. Vasc. Biol. 2013, 33, 170–177.
- Morris, D.L.; Cho, K.W.; DelProposto, J.L.; Oatmen, K.E.; Geletka, L.M.; Martinez-Santibanez, G.; Singer, K.; Lumeng, C.N. Adipose Tissue Macrophages Function As Antigen-Presenting Cells and Regulate Adipose Tissue CD4+ T Cells in Mice. Diabetes 2013, 62, 2762–2772.
- Zeyda, M.; Farmer, D.; Todoric, J.; Aszmann, O.; Speiser, M.; Györi, G.; Zlabinger, G.; Stulnig, T. Human adipose tissue macrophages are of an anti-inflammatory phenotype but capable of excessive pro-inflammatory mediator production. Int. J. Obes. 2007, 31, 1420–1428.
- Dunand-Sauthier, I.; Santiago-Raber, M.-L.; Capponi, L.P.; Vejnar, C.; Schaad, O.; Irla, M.; Estévez, M.Q.S.; Descombes, P.; Zdobnov, E.M.; Acha-Orbea, H.; et al. Silencing of c-Fos expression by microRNA-155 is critical for dendritic cell maturation and function. Blood 2011, 117, 4490–4500.
- Mari, L.; Hoefnagel, S.J.; Zito, D.; van de Meent, M.; van Endert, P.; Calpe, S.; Serra, M.D.C.S.; Heemskerk, M.H.; van Laarhoven, H.W.; Hulshof, M.C.; et al. microRNA 125a Regulates MHC-I Expression on Esophageal Adenocarcinoma Cells, Associated With Suppression of Antitumor Immune Response and Poor Outcomes of Patients. Gastroenterology 2018, 155, 784–798.
- Pischon, T.; Boeing, H.; Hoffmann, K.; Bergmann, M.; Schulze, M.B.; Overvad, K.; Van Der Schouw, Y.T.; Spencer, E.; Moons, K.G.M.; Tjønneland, A.; et al. General and Abdominal Adiposity and Risk of Death in Europe. N. Engl. J. Med. 2008, 359, 2105–2120.
- Shungin, D.; Winkler, T.W.; Croteau-Chonka, D.C.; Ferreira, T.; Locke, A.E.; Mägi, R.; Strawbridge, R.J.; Pers, T.H.; Fischer, K.; Justice, A.E.; et al. New genetic loci link adipose and insulin biology to body fat distribution. Nature 2015, 518, 187–196.
- Wang, Q.A.; Tao, C.; Gupta, R.K.; Scherer, P.E. Tracking adipogenesis during white adipose tissue development, expansion and regeneration. Nat. Med. 2013, 19, 1338–1344.
- Jeffery, E.; Church, C.D.; Holtrup, B.; Colman, L.; Rodeheffer, M.S. Rapid depot-specific activation of adipocyte precursor cells at the onset of obesity. Nat. Cell Biol. 2015, 17, 376–385.
- Wueest, S.; Rapold, R.A.; Rytka, J.M.; Schoenle, E.J.; Konrad, D. Basal lipolysis, not the degree of insulin resistance, differentiates large from small isolated adipocytes in high-fat fed mice. Diabetologia 2009, 52, 541–546.
- Gao, H.; Mejhert, N.; Fretz, J.; Arner, E.; Lorente-Cebrián, S.; Ehrlund, A.; Dahlman-Wright, K.; Gong, X.; Strömblad, S.; Douagi, I.; et al. Early B Cell Factor 1 Regulates Adipocyte Morphology and Lipolysis in White Adipose Tissue. Cell Metab. 2014, 19, 981–992.
- Lee, Y.-H.; Petkova, A.P.; Granneman, J.G. Identification of an Adipogenic Niche for Adipose Tissue Remodeling and Restoration. Cell Metab. 2013, 18, 355–367.
- Hill, D.A.; Lim, H.-W.; Kim, Y.H.; Ho, W.Y.; Foong, Y.H.; Nelson, V.L.; Nguyen, H.C.B.; Chegireddy, K.; Kim, J.; Habertheuer, A.; et al. Distinct macrophage populations direct inflammatory versus physiological changes in adipose tissue. Proc. Natl. Acad. Sci. USA 2018, 115, E5096–E5105.
- Cinti, S.; Mitchell, G.; Barbatelli, G.; Murano, I.; Ceresi, E.; Faloia, E.; Wang, S.; Fortier, M.; Greenberg, A.S.; Obin, M.S. Adipocyte death defines macrophage localization and function in adipose tissue of obese mice and humans. J. Lipid Res. 2005, 46, 2347–2355.
- Meng, L.; Zhou, J.; Sasano, H.; Suzuki, T.; Zeitoun, K.M.; Bulun, S.E. Tumor necrosis factor alpha and interleukin 11 secreted by malignant breast epithelial cells inhibit adipocyte differentiation by selectively down-regulating CCAAT/enhancer binding protein alpha and peroxisome proliferator-activated receptor gamma: Mechanism of desmoplastic reaction. Cancer Res. 2001, 61, 2250–2255.
- Cawthorn, W.P.; Heyd, F.; Hegyi, K.; Sethi, J.K. Tumour necrosis factor-alpha inhibits adipogenesis via a beta-catenin/TCF4(TCF7L2)-dependent pathway. Cell Death Differ. 2007, 14, 1361–1373.
- Orr, J.S.; Kennedy, A.; Anderson-Baucum, E.K.; Webb, C.D.; Fordahl, S.C.; Erikson, K.M.; Zhang, Y.; Etzerodt, A.; Moestrup, S.K.; Hasty, A.H. Obesity Alters Adipose Tissue Macrophage Iron Content and Tissue Iron Distribution. Diabetes 2014, 63, 421–432.
- Hubler, M.J.; Peterson, K.R.; Hasty, A.H. Iron homeostasis: A new job for macrophages in adipose tissue? Trends Endocrinol. Metab. 2015, 26, 101–109.
- Brandão, B.B.; Guerra, B.A.; Mori, M.A. Shortcuts to a functional adipose tissue: The role of small non-coding RNAs. Redox Biol. 2017, 12, 82–102.
- Zhai, K.; Duan, H.; Wang, W.; Zhao, S.; Khan, G.J.; Wang, M.; Zhang, Y.; Thakur, K.; Fang, X.; Wu, C.; et al. Ginsenoside Rg1 ameliorates blood–brain barrier disruption and traumatic brain injury via attenuating macrophages derived exosomes miR-21 release. Acta Pharm. Sin. B 2021, 11, 3493–3507.
- Liu, S.; Chen, J.; Shi, J.; Zhou, W.; Wang, L.; Fang, W.; Zhong, Y.; Chen, X.; Chen, Y.; Sabri, A.; et al. M1-like macrophage-derived exosomes suppress angiogenesis and exacerbate cardiac dysfunction in a myocardial infarction microenvironment. Basic Res. Cardiol. 2020, 115, 1–17.
- Rupnick, M.A.; Panigrahy, D.; Zhang, C.-Y.; Dallabrida, S.M.; Lowell, B.B.; Langer, R.; Folkman, M.J. Adipose tissue mass can be regulated through the vasculature. Proc. Natl. Acad. Sci. USA 2002, 99, 10730–10735.
- Cao, Y. Angiogenesis and Vascular Functions in Modulation of Obesity, Adipose Metabolism, and Insulin Sensitivity. Cell Metab. 2013, 18, 478–489.
- Cho, C.-H.; Koh, Y.J.; Han, J.; Sung, H.-K.; Lee, H.J.; Morisada, T.; Schwendener, R.A.; Brekken, R.A.; Kang, G.; Oike, Y.; et al. Angiogenic Role of LYVE-1–Positive Macrophages in Adipose Tissue. Circ. Res. 2007, 100, e47–e57.
- Spiller, K.L.; Anfang, R.R.; Spiller, K.J.; Ng, J.; Nakazawa, K.R.; Daulton, J.W.; Vunjak-Novakovic, G. The role of macrophage phenotype in vascularization of tissue engineering scaffolds. Biomaterials 2014, 35, 4477–4488.
- Landskroner-Eiger, S.; Moneke, I.; Sessa, W.C. miRNAs as Modulators of Angiogenesis. Cold Spring Harb. Perspect. Med. 2013, 3, a006643.
- Yang, Y.; Guo, Z.; Chen, W.; Wang, X.; Cao, M.; Han, X.; Zhang, K.; Teng, B.; Cao, J.; Wu, W.; et al. M2 Macrophage-Derived Exosomes Promote Angiogenesis and Growth of Pancreatic Ductal Adenocarcinoma by Targeting E2F2. Mol. Ther. 2021, 29, 1226–1238.
- Nicoli, S.; Knyphausen, C.-P.; Zhu, L.; Lakshmanan, A.; Lawson, N.D. miR-221 Is Required for Endothelial Tip Cell Behaviors during Vascular Development. Dev. Cell 2012, 22, 418–429.
- Pankratz, F.; Bemtgen, X.; Zeiser, R.; Leonhardt, F.; Kreuzaler, S.; Hilgendorf, I.; Smolka, C.; Helbing, T.; Hoefer, I.; Esser, J.S.; et al. MicroRNA-155 Exerts Cell-Specific Antiangiogenic but Proarteriogenic Effects During Adaptive Neovascularization. Circulation 2015, 131, 1575–1589.
- Divoux, A.; Tordjman, J.; Lacasa, D.; Veyrie, N.; Hugol, D.; Aissat, A.; Basdevant, A.; Guerre-Millo, M.; Poitou, C.; Zucker, J.-D.; et al. Fibrosis in Human Adipose Tissue: Composition, Distribution, and Link With Lipid Metabolism and Fat Mass Loss. Diabetes 2010, 59, 2817–2825.
- Lackey, D.E.; Burk, D.H.; Ali, M.R.; Mostaedi, R.; Smith, W.H.; Park, J.; Scherer, P.E.; Seay, S.A.; McCoin, C.; Bonaldo, P.; et al. Contributions of adipose tissue architectural and tensile properties toward defining healthy and unhealthy obesity. Am. J. Physiol. Metab. 2014, 306, E233–E246.
- Hasegawa, Y.; Ikeda, K.; Chen, Y.; Alba, D.L.; Stifler, D.; Shinoda, K.; Hosono, T.; Maretich, P.; Yang, Y.; Ishigaki, Y.; et al. Repression of Adipose Tissue Fibrosis through a PRDM16-GTF2IRD1 Complex Improves Systemic Glucose Homeostasis. Cell Metab. 2018, 27, 180–194.e6.
- Tanaka, M.; Ikeda, K.; Suganami, T.; Komiya, C.; Ochi, K.; Shirakawa, I.; Hamaguchi, M.; Nishimura, S.; Manabe, I.; Matsuda, T.; et al. Macrophage-inducible C-type lectin underlies obesity-induced adipose tissue fibrosis. Nat. Commun. 2014, 5, 4982.
- Dewald, O.; Zymek, P.; Winkelmann, K.; Koerting, A.; Ren, G.; Abou-Khamis, T.; Michael, L.H.; Rollins, B.J.; Entman, M.L.; Frangogiannis, N.G. CCL2/monocyte chemoattractant protein-1 regulates inflammatory responses critical to healing myocardial infarcts. Circ. Res. 2005, 96, 881–889.
- Frangogiannis, N.G.; Dewald, O.; Xia, Y.; Ren, G.; Haudek, S.; Leucker, T.; Kraemer, D.; Taffet, G.; Rollins, B.J.; Entman, M.L. Critical role of monocyte chemoattractant protein-1/CC chemokine ligand 2 in the pathogenesis of ischemic cardiomyopathy. Circulation 2007, 115, 584–592.
- Juban, G.; Saclier, M.; Yacoub-Youssef, H.; Kernou, A.; Arnold, L.; Boisson, C.; Ben Larbi, S.; Magnan, M.; Cuvellier, S.; Théret, M.; et al. AMPK Activation Regulates LTBP4-Dependent TGF-β1 Secretion by Pro-inflammatory Macrophages and Controls Fibrosis in Duchenne Muscular Dystrophy. Cell Rep. 2018, 25, 2163–2176.e6.
- Thum, T.; Gross, C.; Fiedler, J.; Fischer, T.; Kissler, S.; Bussen, M.; Galuppo, P.; Just, S.; Rottbauer, W.; Frantz, S.; et al. MicroRNA-21 contributes to myocardial disease by stimulating MAP kinase signalling in fibroblasts. Nature 2008, 456, 980–984.
- Liu, G.; Friggeri, A.; Yang, Y.; Milosevic, J.; Ding, Q.; Thannickal, V.J.; Kaminski, N.; Abraham, E. miR-21 mediates fibrogenic activation of pulmonary fibroblasts and lung fibrosis. J. Exp. Med. 2010, 207, 1589–1597.
- Chau, B.N.; Xin, C.; Hartner, J.; Ren, S.; Castano, A.P.; Linn, G.; Li, J.; Tran, P.T.; Kaimal, V.; Huang, X.; et al. MicroRNA-21 Promotes Fibrosis of the Kidney by Silencing Metabolic Pathways. Sci. Transl. Med. 2012, 4, 121ra18.
- Ardite, E.; Perdiguero, E.; Vidal, B.; Gutarra, S.; Serrano, A.L.; Muñoz-Cánoves, P. PAI-1–regulated miR-21 defines a novel age-associated fibrogenic pathway in muscular dystrophy. J. Cell Biol. 2012, 196, 163–175.
- Hinkel, R.; Ramanujam, D.P.; Kaczmarek, V.; Howe, A.; Klett, K.; Beck, C.; Dueck, A.; Thum, T.; Laugwitz, K.-L.; Maegdefessel, L.; et al. AntimiR-21 Prevents Myocardial Dysfunction in a Pig Model of Ischemia/Reperfusion Injury. J. Am. Coll. Cardiol. 2020, 75, 1788–1800.
- Cui, H.; He, Y.; Chen, S.; Zhang, D.; Yu, Y.; Fan, C. Macrophage-derived miRNA-containing exosomes induce peritendinous fibrosis after tendon injury through the miR-21-5p/Smad7 pathway. Mol. Ther. Nucleic Acids 2019, 14, 114–130.
- Mildner, A.; Chapnik, E.; Manor, O.; Yona, S.; Kim, K.-W.; Aychek, T.; Varol, D.; Beck, G.; Itzhaki, Z.B.; Feldmesser, E.; et al. Mononuclear phagocyte miRNome analysis identifies miR-142 as critical regulator of murine dendritic cell homeostasis. Blood 2013, 121, 1016–1027.
- Xu, X.; Grijalva, A.; Skowronski, A.; van Eijk, M.; Serlie, M.J.; Ferrante, A.W. Obesity Activates a Program of Lysosomal-Dependent Lipid Metabolism in Adipose Tissue Macrophages Independently of Classic Activation. Cell Metab. 2013, 18, 816–830.
- Jaitin, D.A.; Adlung, L.; Thaiss, C.A.; Weiner, A.; Li, B.; Descamps, H.; Lundgren, P.; Bleriot, C.; Liu, Z.; Deczkowska, A.; et al. Lipid-Associated Macrophages Control Metabolic Homeostasis in a Trem2-Dependent Manner. Cell 2019, 178, 686–698.e14.
- Sárvári, A.K.; Van Hauwaert, E.L.; Markussen, L.K.; Gammelmark, E.; Marcher, A.-B.; Ebbesen, M.F.; Nielsen, R.; Brewer, J.R.; Madsen, J.G.S.; Mandrup, S. Plasticity of Epididymal Adipose Tissue in Response to Diet-Induced Obesity at Single-Nucleus Resolution. Cell Metab. 2020, 33, 437–453.e5.
- Magalhaes, M.S.; Smith, P.; Portman, J.R.; Jackson-Jones, L.H.; Bain, C.C.; Ramachandran, P.; Michailidou, Z.; Stimson, R.H.; Dweck, M.R.; Denby, L.; et al. Role of Tim4 in the regulation of ABCA1+ adipose tissue macrophages and post-prandial cholesterol levels. Nat. Commun. 2021, 12, 4434.
- Schober, A.; Maleki, S.S.; Nazari-Jahantigh, M. Regulatory Non-coding RNAs in Atherosclerosis. Handb. Exp. Pharmacol. 2020, 270, 463.
- Fernández-Hernando, C.; Ackah, E.; Yu, J.; Suárez, Y.; Murata, T.; Iwakiri, Y.; Prendergast, J.; Miao, R.Q.; Birnbaum, M.; Sessa, W.C. Loss of Akt1 Leads to Severe Atherosclerosis and Occlusive Coronary Artery Disease. Cell Metab. 2007, 6, 446–457.
- Yan, L.L.; Daviglus, M.L.; Liu, K.; Stamler, J.; Wang, R.; Pirzada, A.; Garside, D.B.; Dyer, A.R.; Van Horn, L.; Liao, Y.; et al. Midlife Body Mass Index and Hospitalization and Mortality in Older Age. JAMA: J. Am. Med. Assoc. 2006, 295, 190–198.
- Dykstra, B.; Olthof, S.; Schreuder, J.; Ritsema, M.; de Haan, G. Clonal analysis reveals multiple functional defects of aged murine hematopoietic stem cells. J. Exp. Med. 2011, 208, 2691–2703.
- Pang, W.W.; Price, E.A.; Sahoo, D.; Beerman, I.; Maloney, W.J.; Rossi, D.J.; Schrier, S.L.; Weissman, I.L. Human bone marrow hematopoietic stem cells are increased in frequency and myeloid-biased with age. Proc. Natl. Acad. Sci. USA 2011, 108, 20012–20017.
- Lumeng, C.; Liu, J.; Geletka, L.; Delaney, C.E.; DelProposto, J.; Desai, A.; Oatmen, K.; Martinez-Santibanez, G.; Julius, A.; Garg, S.; et al. Aging Is Associated with an Increase in T Cells and Inflammatory Macrophages in Visceral Adipose Tissue. J. Immunol. 2011, 187, 6208–6216.
- Sene, A.; Khan, A.A.; Cox, D.; Nakamura, R.E.; Santeford, A.; Kim, B.M.; Sidhu, R.; Onken, M.D.; Harbour, J.W.; Hagbi-Levi, S.; et al. Impaired Cholesterol Efflux in Senescent Macrophages Promotes Age-Related Macular Degeneration. Cell Metab. 2013, 17, 549–561.
- Hunt, E.A.; Broyles, D.; Head, T.; Deo, S.K. MicroRNA Detection: Current Technology and Research Strategies. Annu. Rev. Anal. Chem. 2015, 8, 217–237.
- Kratz, M.; Coats, B.R.; Hisert, K.B.; Hagman, D.; Mutskov, V.; Peris, E.; Schoenfelt, K.Q.; Kuzma, J.N.; Larson, I.; Billing, P.S.; et al. Metabolic Dysfunction Drives a Mechanistically Distinct Proinflammatory Phenotype in Adipose Tissue Macrophages. Cell Metab. 2014, 20, 614–625.
- Xue, J.; Schmidt, S.V.; Sander, J.; Draffehn, A.; Krebs, W.; Quester, I.; De Nardo, D.; Gohel, T.D.; Emde, M.; Schmidleithner, L.; et al. Transcriptome-based network analysis reveals a spectrum model of human macrophage activation. Immunity 2014, 40, 274–288.
- Sander, J.; Schmidt, S.V.; Cirovic, B.; McGovern, N.; Papantonopoulou, O.; Hardt, A.-L.; Aschenbrenner, A.; Kreer, C.; Quast, T.; Xu, A.; et al. Cellular Differentiation of Human Monocytes Is Regulated by Time-Dependent Interleukin-4 Signaling and the Transcriptional Regulator NCOR2. Immunity 2017, 47, 1051–1066.e12.
- Li, C.; Menoret, A.; Farragher, C.; Ouyang, Z.; Bonin, C.; Holvoet, P.; Vella, A.T.; Zhou, B. Single cell transcriptomics based-MacSpectrum reveals novel macrophage activation signatures in diseases. JCI Insight 2019, 5, 126453.
- Li, C.; Qu, L.; Matz, A.J.; Murphy, P.A.; Liu, Y.; Manichaikul, A.W.; Aguiar, D.; Rich, S.S.; Herrington, D.M.; Vu, D.; et al. AtheroSpectrum Reveals Novel Macrophage Foam Cell Gene Signatures Associated With Atherosclerotic Cardiovascular Disease Risk. Circulation 2022, 145, 206–218.





