| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Malka Cohen-Armon | + 1624 word(s) | 1624 | 2020-09-09 09:49:09 | | | |
| 2 | Bruce Ren | + 13 word(s) | 1637 | 2020-09-14 03:34:25 | | | | |
| 3 | Bruce Ren | + 14 word(s) | 1638 | 2020-09-14 03:35:55 | | | | |
| 4 | Bruce Ren | + 14 word(s) | 1638 | 2020-09-14 03:37:30 | | | | |
| 5 | Bruce Ren | Meta information modification | 1638 | 2020-09-19 02:44:05 | | |
Video Upload Options
This entry summarizes recent data disclosing the efficacy of the modified phenanthridine PJ34 in exclusive eradication of a variety of human cancer cells without impairing healthy proliferating cells. Its cytotoxic activity in cancer cells is attributed to the insertion of specific un-repairable anomalies in the structure of their mitotic spindle, leading to mitotic catastrophe cell death.
1. Background
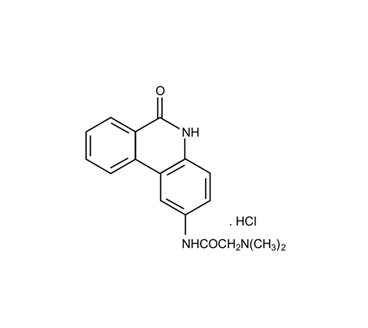
In the last twenty years the modified phenanthridine PJ34 has been known for its activity as a PARP (polyADP-ribose polymerase) inhibitor [1][2] (Figure 1). Recently, PARP inhibitors attract the attention of researchers and clinicians due to their FDA approval for cancer therapy [3][4][5][6], although some of them failed in clinical trials. Unexpectedly, it has been disclosed that PJ34 that has been originally invented to protect cells from cell death, imposed by pathological stress conditions, such as ischemia and inflammation [7][8][9], has an exceptional cytotoxicity in human cancer cells. Its cytotoxic activity does not involve PARP inhibition, and most importantly, it does not affect healthy cells [10][11][12][13][14][15][16][17].
Figure 1. The chemical structure of PJ34, N-(6-Oxo-5,6-dihydrophenanthridin-2-yl)-(N,N-dimethylamino)acetamide hydrochloride.
The modified phenanthridine PJ34 is a stable molecule, fairly soluble in water (22 mg/mL), and permeable in the cell membrane. Unlike other PARP inhibitors, PJ34 exclusively eradicates a variety of human cancer cells, including cancer cells resistant to current therapy, without affecting proliferating and non-proliferating healthy cells including human epithelial, mesenchymal and endothelial cells [10][11][12], and healthy cells of mouse origin, including mouse embryonic fibroblasts (MEF), fibroblasts, neurons in the central nervous system and neuronal progenitor stem cells. Its exclusive cytotoxic activity in a variety of human cancer cells, solid and hematological malignancies and in xenografts is summarized in a recent review [18].
Incubation with PJ34 completely eradicates within 48 -96 hours a variety of human cancer cells (18). In addition, PJ34 retardes the development of human tumors in mouse xenografts, and most surprisingly, PJ34 exclusively eradicates cancer cells in developing tumors. About 90% of human pancreas ductal adenocarcinoma, PANC1 cells in tumors developed in immunocompromised mice (xenografts ) were eradicated 30 days after 14 Intravenous treatments with PJ34 (60 mg/kg). Metastases were not detected. This and other treatments with PJ34 in xenografts did not impair the weight gain of the mice nor impact their activity and development, which were monitored during the experiments. Moreover, treatments with PJ34 in combination with other anti-cancer agents enabled reducing their cytotoxic doses, and achieved efficient treatment of some resistant cancer tumors [18].
Flow cytometry measurements revealed that PJ34 exclusively arrests mitosis in human cancer cells [10][11][14]. Therefore, the possibility that PJ34 induces mitotic arrest has been tested by testing its effect on the post-translational modification of all the currently known proteins implicated in mitosis. These proteins were screened in a group of human cancer cells versus healthy cells, in an attempt to identify different effects of PJ34 on the post-translational modifications of proteins in the cancer versus healthy cells. Changes induced by PJ34 in their post-translational modifications were measured by the shift in their isoelectric point (IP) in two-dimensional (2-D) gel electrophoresis [13]. Also, the effect of PJ34 on the isoelectric point of these proteins in the two groups of cells was compared to the effect of a potent PARP inhibitor, in order to examine possible involvement of PARP inhibition in the effect of PJ34 [13].
2. PJ34 Efficiently Eradicates a Variety of Human Cancer Cells in Tissue Cultures
This analysis identified only three proteins in the tested cancer cells with isoelectric point significantly shifted by PJ34, while not affected in healthy cells [13]. These proteins included two motor proteins [19], human kinesins 14/HSET/kifC1 and kif18A, and the non-motor protein NuMA (nuclear mitotic apparatus protein) [20][21][22][23][24]. PARP inhibition did not exert any similar effect on these proteins.
HSET/kifC1 has an essential role in the spindle structure of human cancer cells [20][21][22][25][26][27][28]. Differences in the expression and function of HSET in cancer versus healthy cells have been reported [25][26][27]. HSET/kifC1 inhibition or silencing causes small aberrant spindles in human malignant cells [26].
The kinesin Kif18A is implicated in microtubules de-polymerization, necessary for the binding of the duplicated chromosomes to kinetochores in the spindle mid-zone [23].
The third identified protein, NuMA, is essential for mitosis in both malignant and benign cells [29][30].A clear-cut difference has been disclosed by the effect of PJ34 on the post-translational modification of NuMA in the human malignant cells [13] versus the effect of PJ34 on the post translational modification of NuMA in benign human cells (13). PJ34 did not affect the isoelectric point of NuMA in healthy epithelial cells [13], while prevented the post translational modification of NuMA in the tested human cancer cells [13]. Concomitantly, the ability of NuMA to bind proteins was lost in the PJ34-treated malignant cells [13]. Moreover, the lost ability of NuMA to bind proteins was accompanied by un-clustering of NuMA in the spindle poles of malignant cells treated with PJ34, as disclosed by confocal imaging [13] (Figure 2). In contrast, the bi-polar clustering of NuMA in the mitotic spindles of the healthy benign cells was not affected by the same treatment with PJ34 [13] (Figure 2).
A post-translational modification of NuMA by either polyADP-ribosylation or phosphorylation promotes the binding of NuMA to proteins [31][32][33][34][35][36]. NuMA is phosphorylated by serine-threonine kinase pim1, and NuMA phosphorylation by serine threonine kinases at a specific site in the protein is crucial for its ability to bind proteins [31][32]. Similarly, polyADP-ribosylation of NuMA by tankyrase1 in cancer cells promotes the ability of NuMA to bind other proteins [33].
Pim kinases and tankyrase1 are both inhibited by PJ34 at the same concentrations range of PJ34 that causes cell death in human cancer cells (measured IC50 = 3.7 μM for pim1 inhibition by PJ34, and IC50 = 1 μM for tankyrase1 inhibition by PJ34) [34][35][36][37]. Furthermore, tankyrase1 and pim kinases are hardly expressed in healthy somatic cells, while highly expressed in human cancer cells [36][37].
Clustered NuMA in the spindle poles and tethering of microtubules to the clustered proteins in the spindle poles are essential for the construction of stable poles, which are required for the alignment of the chromosomes in the spindle mid-zone [27][28][29][30]. Blocking the post-translational modification of NuMA by PJ34 can prevent the clustering of NuMA in the spindle poles [13].
Thus, blocking the post translational modification of kinesines HSET and kif18A and the post translational modification of NuMA can also prevent the construction of spindles in cancer cells by HSET, and the construction of stable spindle poles by NuMA clustering and the tethering of microtubules in the spindle poles. of chromosomes in the mid-zone to the kinetochores ([18], Figure 2).
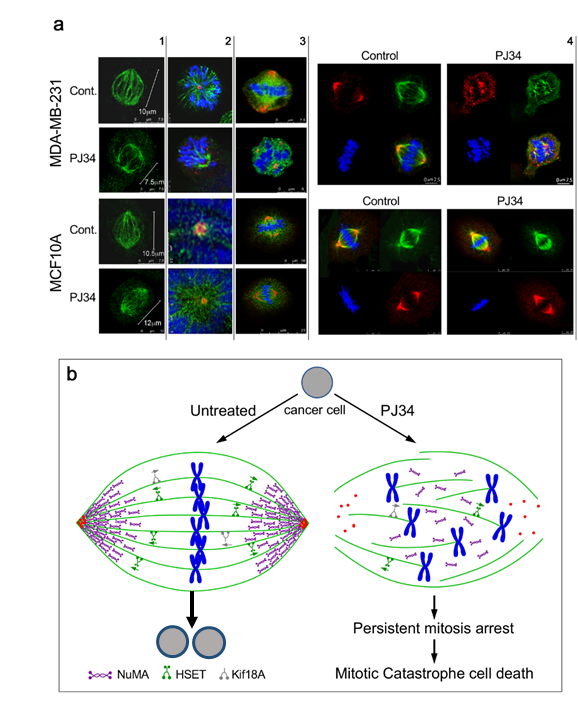
Figure 2. (a) Confocal images of mitotic spindles in human triple negative breast cancer cells (MDA-MB-231) and in human healthy breast epithelial cells (MCF10A), untreated or incubated with PJ34. Incubation of human cancer cells with PJ34 (20 μM, 27 h) impaired spindle poles (labeled by immunolabelling, -tubulin in the centrosomes—red), microtubules (labeled by immunolabelling—kinesin HSET or by immunolabelling -tubulin—green), segregation and alignment of chromosomes (labeled by DAPI—blue), and NuMA clustering in the spindle poles (Immunolabeled NuMA—red). Column 1: Microtubules in spindles of healthy and cancer cells immunolabeled by the kinesin HSET in cancer and healthy cells, untreated and treated with PJ34. Column 2: Spindle poles labeled by γ-tubulin in healthy and cancer cells, untreated and treated with PJ34. HSET is immunolabeled in the microtubules. Column 3: Clustered NuMA in bipolar spindles of healthy cells either treated or not with PJ34, and in untreated cancer cells. Un-clustered NuMA in spindles of cancer cells treated with PJ34. Column 4: upper frame: In cancer cells—clustered NuMA in spindle poles and aligned chromosomes in the midzone of untreated cancer cells. Aberrant spindles, un-clustered NuMA and scattered chromosomes in cancer cells treated with PJ34. Lower frame: In healthy cells—clustered NuMA in the spindle poles and segregated chromosomes aligned in the mid-zone of the mitotic spindle of healthy cells either untreated or treated with PJ34. (b) A schematic presentation indicating the effect of PJ34 on the spindle structure in human cancer cell. In the untreated cancer cell, normal bipolar spindles with clustered NuMA, clustered multi-centrosomes, and aligned chromosomes in the spindle mid-zone. In the PJ-34 treated cancer cell, aberrant microtubules (green), aberrant spindle poles, un-clustered NuMA (as indicated), dispersed chromosomes (blue) and un-clustered multi-centrosome (red), From: Visochek et al., 2017, Oncotarget.
The unstable structure of the spindle poles preventing chromosomes arrangement in the midzone activates the spindle assembly control (SAC) proteins, which leads to mitosis arrest followed by mitotic catastrophe cell death when the structural anomaly is not amended [38][39][40][41]. This is exactly the phenomenon observed by confocal imaging in a variety of human cancer cells treated with PJ34 [13]. De-clustering of centrosomes observed in multi-centrosomal cancer cells treated with PJ34 could result from the un-stable aberrant spindle poles [13]. Thus, by preventing the post translation modification of HSET, kif18A and NuMA in human cancer cells, specific anomalies in their mitotic spindle structure are inserted ([13] and Figure 2). These results are in consistence with previous findings [42][43].
The potency of PJ34 to exclusively eradicate human cancer cells without impairing healthy cells can be attributed to the anomalies exclusively inserted in the structure of the mitotic spindle of human cancer cells, which arrest mitosis and kills cancer cell in the pre-anaphase stage by mitotic catastrophe cell death. Therefore, the more frequently cancer cells enter mitosis, the more efficiently they are eradicated. Thus, despite the permeability of PJ34 in the cell membrane, and despite its rapid distribution in the animal’s tissues, treatment with PJ34 did not impair healthy tissues in the tested animals, nor their development and weight-gain.
3. Conclusion
In conclusion, the modified phenanthridine PJ34, which has been invented for PARP inhibition, efficiently eradicates a variety of human cancer cells by mitotic catastrophe cell death caused by faults in their mitotic spindle.
On the basis of these findings, we hope that cell death evoked by structural faults in the mitotic spindle of human cancer cells will pave the way to a new concept in cancer therapy.
References
- Abdelkarim, G.E.; Gertz, K.; Harm, C.; Katchanov, J.; Dirnagi, U.; Szabo, C.; Enders, M. Protective effects of PJ34, a novel potent inhibitor of poly(ADP-ribose)polymerase (PARP) in in-vitro and in-vivo models of stroke. Int. J. Mol. Med. 2001, 7, 255–260.
- Jagtap, P.; Szabo, C. Poly(ADP-ribose)polymerase and the therapeutic effects of its inhibitors. Nat. Rev. Drug Discov. 2005, 4, 421–440.
- Slade, D. PARP and PARG inhibitors in cancer treatment. Gene Dev. 2020, 34, 1–35.
- Carden, C.P.; Yap, T.A.; Kaye, S.B. PARP inhibition: Targeting the Achilles heelof DNA repair to treat germline and sporadic ovarian cancers. Curr. Opin. Oncol. 2010, 22, 473–480.
- Plummer, R. Perspective on the pipeline of drugs being developed with modulationof DNA damage as a target. Clin. Cancer Res. 2010, 16, 4527–4531.
- Mangerich, A.; Burkle, A. How to kill tumor cells with inhibitors of poly(ADP-ribosyl)ation. Int. J. Cancer 2011, 128, 251–265.
- Hassa, P.O.; Hottiger, M.O. The diverse biological roles of mammalian PARPs, a small but powerful family of poly-ADP-ribose polymerases. Front. Biosci. 2008, 13, 3046–3082.
- 8. Wahlberg, E.; Karlberg, T.; Kouznetsova, E.; Markova, N.; Macchiarulo, A.; Torsell, A.-G.; Pol, E.; Frostell, A.; Ekblad, T.; Öncü,D.; Kull, B.; et al. Family-wide chemical profiling and structural analysis of PARP and tankyrase inhibitors. Nat. Biotech. 2012, 30, 283–289.
- Citarelli, M.; Teotia, S.; Lamb, R.S. Evolutionary history of the poly(ADP-ribose)polymerase gene family in eukaryotes. BMC Evol. Biol. 2010, 10, 308, doi:10.1186/1471-2148-10-308.
- Krishnakumar, R.; Kraus, W.L. The PARP side of the nucleus: Molecular actions, physiological outcomes, and clinical targets. Mol. Cell 2010, 39, 8–24.
- Gibson, B.A.; Kraus, W.L. New insights into the molecular and cellular functions of poly(ADP-ribose) and PARPs. Nat. Rev. Mol. Cell Biol. 2012, 13, 411–424.
- Kauppinen, T.M.; Swanson, R.A. The role of poly(ADP-ribose) polymerase-1 in CNS disease. Neuroscience 2007, 145, 1267–1272.
- Strosznajder, R.P.; Czubowicz, K.; Jesko, H.; Strosznajder, J.B. Poly(ADP-ribose) metabolism in brain and its role in ischemia pathology. Mol. Neurobiol. 2010, 41, 187–196.
- Ba, X.; Garg, N.J. Signaling mechanism of poly(ADP-ribose)polymerase-1 (PARP-1) in inflammatory diseases. Am. J. Pathol. 2011, 178, 946–955.
- Gregersen, L.H.; Jesper, J.Q. The cellular response to transcription blocking DNA damage. Trend Biochem. Sci. 2018, 43, 327–341.
- Lesueur, P.; Chevalier, F.; Austry, J.-B.; Waissi, W.; Burckel, H.; Noel, G.; Habrand, J.-L.; Saintigny, Y.; Joly, F. PolyADP-ribose) polymerase inhibitors as radiosensitizers: A systematic review of preclinical human studies. Oncotarget 2017, 8, 69105–69124.
- Bryant, H.E.; Schultz, N.; Thomas, H.D.; Parker, K.M.; Flower, D.; Lopez, E.; Kyle, S.; Meuth, M.; Curtin, N.J.; Helleday, T. Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature 2005, 434, 913–917.
- Andrabi, S.A.; Kim, N.-S.; Yu, S.W., Wang, H.; Koh, D.W.; Sassaki, M.; Klaus, J.A.; Otsuka, T.; Zhang, Z.; Koehler, C.K.; et al. Poly(ADP-ribose) (PAR) polymer is a death signal. Proc. Natl. Acad. Sci. USA 2006, 103, 18308–18313.
- Chang, H.H.Y.; Pannunzio, N.R.; Adachi, N.; Lieber, M.R. Non-homologous DNA end joining and alternative pathways to double-strand break repair Nat. Rev. Mol. Cell Biol. 2017, 18, 495–506.
- Paddock, M.N.; Bauman, A.T.; Higdon, R.; Kolker, E.; Takeda, S.; Scharenberg, A.M. Competition between PARP-1 and Ku70 control the decision between high-fidelity and mutagenic DNA repair. DNA Repair 2011, 10, 338–343.
- Farmer, H.; McCabe, N.; Lord, C.J.; Tutt, A.N.; Johnson, D.A.; Richardson, T.B.; Santarosa, M.; Dillon, K.J.; Hickson, I.; Knights, C.; et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 2005, 434, 917–921.
- Mak, J.P.Y.; Ma, H.T.; Poon, R.Y.C. Synergism between ATM and PARP1 inhibition involves DNA damage and abrogating the G2 DNA damage checkpoint. Mol. Cancer Ther. 2020, 19, 123–134.
- Zhang, R.; Hong, J.-J.; Yang, Q.; Ong, C.-T.; Li, B.-A.; Liou, Y-C. Poly(ADP- ribosyl)ation of OVOL2 regulates aneuploidy and cell death in cancer cells. Oncogene 2019, 38, 2750–2766.
- Zhou, J.X.; Feng, L.J.; Zhang, X. Risk of severe hematologic toxicities in cancer patients treated with PARP inhibitors: A meta-analysis of randomized controlled trials. Drug Des. Dev. Ther. 2017, 11, 3009–3017.
- Lee, Y.R.; Yu, D.-S.; Liang, Y.-C.; Huang, K.-F.; Chou, S.-J.; Chen, T.-C.; Lee, C.-C.; Chen, C.-L.; Chiou, S.-H.; Huang, H.-S. New approaches of PARP1 inhibitors in human lung cancer cells and cancer stem-like cells by some selected anthraquinone-derived small molecules. PLoS ONE 2013, 8, e56284, doi:10.1371/journal.pone.0056284.
- Kishi, Y.; Fujibara, H.; Kawaguchi, K.; Yamada, H.; Nakayaima, R.; Yamamoto, N.; Fujihara, Y.; Hamada, Y.; Satomura, K.; Masutani, M. PARP inhibitor PJ34 supresses osteogenic differentiation in mouse mesenchymal stem cells by modulating BMP-2 signaling pathway. Int. J. Mol. Sci. 2015, 16, 24820–24838.
- Antolin, A.A.; Ameratunga, M.; Banerji, U.; Clarke, P.A.; Workman, P.; Al-Lazikani, B. The kinase polypharmacology landscape of clinical PARP inhibitors. Sci. Rep. 2020, 10, 2585, doi:10.1038/s41598-020-59074-41.
- Inbar-Rozensal, D.; Visochek, L.; Castel, D.; Castiel, A.; Izraeli, S.; Dantzer, F.; Cohen-Armon, M. A selective eradication of human nonhereditary breast cancer cells by phenanthridine-derived polyADP-ribose polymerase inhibitors. Breast Cancer Res. 2009, 11, R78.
- Castiel, A.; Visochek, L.; Mittelman, L.; Dantzer, F.; Izraeli, S.; Cohen-Armon, M. A phenanthrene derived PARP inhibitor is an extra-centrosomes de-clustering agent exclusively eradicating human cancer cells. BMC Cancer 2011, 11, 412.
- Castiel, A.; Visochek, L.; Mittelman, L.; Zilberstein, Y.; Dantzer, F.; Izraeli, S.; Cohen-Armon, M. Cell death associated with abnormal mitosis observed by confocal imaging in live cancer cells. J. Vis. Exp. 2013, 78, e50568.
- Visochek, L.; Castiel, A.; Mittelman, L.; Elkin, M.; Atias, D.; Golan, T.; Izraeli, S.; Peretz, T.; Cohen-Armon, M. Exclusive destruction of mitotic spindles in human cancer cells. Oncotarget 2017, 8, 20813–20824.
- Visochek, L.; Atias, D.; Spektor, I.; Castiel, A.; Golan, T.; Cohen-Armon, M. The phenanthrene derivative PJ34 exclusively eradicates human pancreatic cancer cells in xenografts. Oncotarget 2019, 10, 6269–6282, doi:10.18632/oncotarget.27268.
- Okuda, A.; Kurokawa, S.; Takehashi, M.; Maeda,A.; Fukuda, K.; Kubo, Y.; Nogusa, H.; Takatani‑Nakase,T.; Okuda, S.; Ueda, K.; Tanaka, S. Poly(ADP‑ribose) polymerase inhibitors activate the p53 signaling pathway in neural stem/progenitor cells. BMC Neurosci. 2017, 18, 2–18.
- Visochek, L.; Grigoryan, G.; Kalal, A.; Milshtein-Parush, H.; Gazit, N.; Slutsky, I.; Yeheskel, A.; Shainberg, A.; Castiel, A.; Seger, R.; et al. A PARP1-ERK2 synergism is required for the induction of LTP. Sci. Rep. 2016, 6, 24950.
- Madison, D.L.; Stauffer, D.; Lundblad, J.R. The PARP inhibitor PJ34 causes a PARP1-independent, p21 dependent mitotic arrest. DNA Repair 2011, 10, 1003–1013.
- Lamoral-Theys, D.; Andolfi, A.; Van Goietsenoven, G.; Cimmino, A.; Le Calv, B.; Wauthoz, N.; Mégalizzi, V.; Gras, T.; Bruyère, C.; Dubois, J.; et al. Lycorine, the Main Phenanthridine Amaryllidaceae Alkaloid, Exhibits Significant Antitumor Activity in Cancer Cells That Display Resistance to Proapoptotic Stimuli: An Investigation of Structure-Activity Relationship and Mechanistic Insight. J. Med. Chem. 2009, 52, 6244–6256.
- Wang, Y.-Y.; Taniguchi, T., Baba, T., Li, Y.-Y.; Isibashi, H.; Nukaida, N. Identification of a phenanthrene derivative as a potent anticancer drug with pim kinase inhibitory activity. Cancer Sci. 2011, 103, 107–115, doi:10.1111/j.1349-7006.2011.02117.x.
- Mariappan, A.; Soni, K.; Schorpp, K.; Zhao, F.; Minakar, A.; Zheng, X.; Mandad, S.; Macheleidt, I.; Ramani, A.; Kubelka, T.; et al. Inhibition of CPAP-tubulin interaction prevents proliferation of centrosome-amplified cancer cells. EMBO J. 2019, 38, e99876, doi:10.15252/emboj.201889876.
- Huang, S.H.; Xiong, M.; Chen, X.P.; Xiao, Z.Y.; Zhao, Y.F.; Huang, Z.Y. PJ34 an inhibitor of PARP1, suppresses cell growth and enhances the suppressive effects of cisplatin in liver cancer cells. Oncol. Rep. 2008, 20, 567–572.
- Pyriochou, A.; Olah, G.; Deitch, O.; Szabo, C.; Papaetropoulos, A. Inhibition of angiogenesis by the poly(ADP-ribose) polymerase inhibitor. J. Mol. Med. 2008, 22, 113–118.
- Gangopadhyay, N.N.; Luketich, J.D.; Opest, A.; Meyer, E.M.; Landreneau, R.; Schuchert, M.J. Inhibition of Poly(ADP-Ribose) Polymerase (PARP) Induces Apoptosis in Lung Cancer Cell Lines. Cancer Invest. 2011, 29, 608–616.
- Liang, B.; Xiong, M.; Ji, G.; Zhang, E.; Zhang, Z.; Dong, K.; Chen, X.; Huang, Z-Y. Synergistic suppressive effect of PARP-1 inhibitor PJ34 and HDAC inhibitor SAHA on proliferation of liver cancer cells. J. Huazhong Univ. Sci. Technol. 2015, 35, 535–540, doi:10.1007/s11596-015-1466-6.
- Xiong, T.; Chen, X.; Wei, H.; Xiao, H. Influence of PJ34 on the genotoxicity induced by melphalan in human multiple myeloma cells. Arch. Med. Sci. 2015, 11, 301–306.