
Video Upload Options
Coronaviruses cause diseases in humans and livestock. The SARS-CoV-2 is infecting millions of human beings, with high morbidity and mortality worldwide. The main protease (Mpro) of coronavirus plays a pivotal role in viral replication and transcription, which, in theory, is an attractive drug target for antiviral drug development. It has been extensively discussed whether Xanthohumol is able to help COVID-19 patients. Here, researchers report that Xanthohumol, a small molecule in clinical trials from hops (Humulus lupulus), was a potent pan-inhibitor for various coronaviruses by targeting Mpro, for example, betacoronavirus SARS-CoV-2 (IC50 value of 1.53 μM), and alphacoronavirus PEDV (IC50 value of 7.51 μM). Xanthohumol inhibited Mpro activities in the enzymatical assays, while pretreatment with Xanthohumol restricted the SARS-CoV-2 and PEDV replication in Vero-E6 cells. Therefore, Xanthohumol is a potent pan-inhibitor of coronaviruses and an excellent lead compound for further drug development.
1. Introduction
2. Coronaviral Mpro Is Highly Conserved
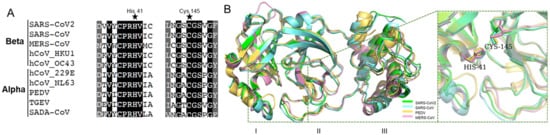
3. SARS-CoV-2 Mpro Inhibitor Screening
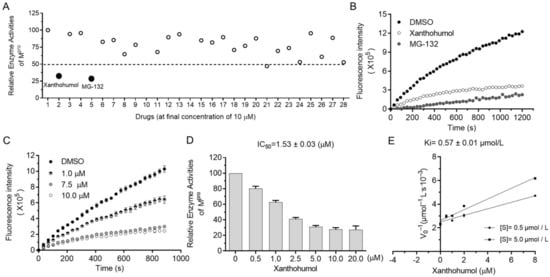
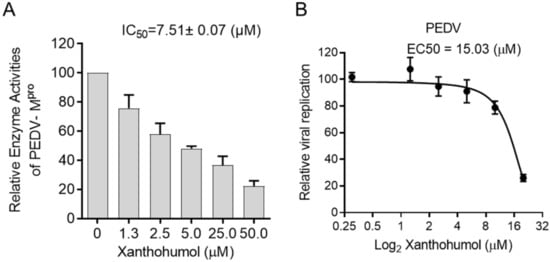
4. Xanthohumol Potentially Inhibits Various Coronaviral Mpro
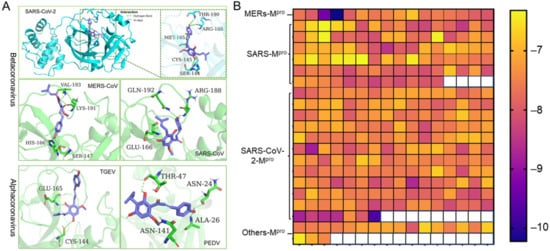
5. Xanthohumol Restricts SARS-CoV-2 and PEDV Replication in the Cell-Based Assay
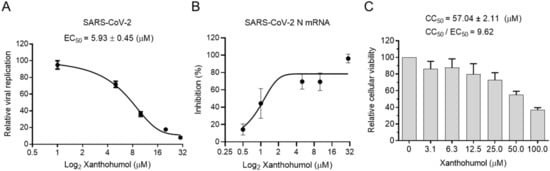
References
- Cui, J.; Li, F.; Shi, Z.L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 2019, 17, 181–192.
- Kirtipal, N.; Bharadwaj, S.; Kang, S.G. Evolution. From SARS to SARS-CoV-2, insights on structure, pathogenicity and immunity aspects of pandemic human coronaviruses. Infect. Genet. Evol. 2020, 85, 104502.
- Tizard, I.R. Vaccination against coronaviruses in domestic animals. Vaccine 2020, 38, 5123–5130.
- Edwards, C.E.; Yount, B.L.; Graham, R.L.; Leist, S.R.; Hou, Y.J.; Dinnon, K.H.; Sims, A.C.; Swanstrom, J.; Gully, K.; Scobey, T.D.; et al. Swine acute diarrhea syndrome coronavirus replication in primary human cells reveals potential susceptibility to infection. Proc. Natl. Acad. Sci. USA 2020, 117, 26915–26925.
- Lee, P.I.; Hsueh, P.R. Emerging threats from zoonotic coronaviruses-from SARS and MERS to 2019-nCoV. J. Microbiol. Immunol. Infect. 2020, 53, 365–367.
- Chen, Y.; Liu, Q.; Guo, D. Emerging coronaviruses: Genome structure, replication, and pathogenesis. J. Med. Virol. 2020, 92, 418–423.
- Wit, E.D.; Feldmann, F.; Cronin, J.; Jordan, R.; Feldmann, H. Prophylactic and therapeutic remdesivir (GS-5734) treatment in the rhesus macaque model of MERS-CoV infection. Proc. Natl. Acad. Sci. USA 2020, 117, 6771–6776.
- Amirian, E.S.; Levy, J.K. Current knowledge about the antivirals remdesivir (GS-5734) and GS-441524 as therapeutic options for coronaviruses. One Health 2020, 9, 100128.
- Wang, Y.; Zhang, D.; Du, G.; Du, R.; Zhao, J.; Jin, Y.; Fu, S.; Gao, L.; Cheng, Z.; Lu, Q.; et al. Remdesivir in adults with severe COVID-19: A randomised, double-blind, placebo-controlled, multicentre trial. Lancet 2020, 395, 1569–1578.
- Jin, Z.; Du, X.; Xu, Y.; Deng, Y.; Liu, M.; Zhao, Y.; Zhang, B.; Li, X.; Zhang, L.; Peng, C.; et al. Structure of Mpro from SARS-CoV-2 and discovery of its inhibitors. Nature 2020, 582, 289–293.
- Xue, X.; Yang, H.; Shen, W.; Zhao, Q.; Li, J.; Yang, K.; Chen, C.; Jin, Y.; Bartlam, M.; Rao, Z. Production of authentic SARS-CoV M(pro) with enhanced activity: Application as a novel tag-cleavage endopeptidase for protein overproduction. J. Mol. Biol. 2007, 366, 965–975.
- Li, Q.; Kang, C.B. Progress in Developing Inhibitors of SARS-CoV-2 3C-Like Protease. Microorganisms 2020, 8, 1250.
- Dai, W.; Zhang, B.; Jiang, X.M.; Su, H.; Liu, H. Structure-Based Design, Synthesis and Biological Evaluation of Peptidomimetic Aldehydes as a Novel Series of Antiviral Drug Candidates Targeting the SARS-CoV-2 Main Protease. BioRxiv 2020.
- Ma, C.; Hu, Y.; Townsend, J.A.; Lagarias, P.I.; Marty, M.T.; Kolocouris, A.; Wang, J. Ebselen, disulfiram, carmofur, PX-12, tideglusib, and shikonin are non-specific promiscuous SARS-CoV-2 main protease inhibitors. ACS Pharm. Transl. Sci. 2020, 3, 1265–1277.
- Vandyck, K.; Deval, J. Considerations for the discovery and development of 3-chymotrypsin-like cysteine protease inhibitors targeting SARS-CoV-2 infection. Curr. Opin. Virol. 2021, 49, 36–40.
- Kowalczyk, M.; Golonko, A.; Świsłocka, R.; Kalinowska, M.; Parcheta, M.; Swiergiel, A.; Lewandowski, W.J. Drug Design Strategies for the Treatment of Viral Disease. Plant Phenolic Compounds and Their Derivatives. Front. Pharm. 2021, 12, 709104.
- Liu, X.; Song, Z.; Bai, J.; Nauwynck, H.; Zhao, Y.; Jiang, P. Xanthohumol inhibits PRRSV proliferation and alleviates oxidative stress induced by PRRSV via the Nrf2–HMOX1 axis. Vet. Res. 2019, 50, 61.
- Liu, W.; Li, W.; Liu, H.; Yu, X. Xanthohumol inhibits colorectal cancer cells via downregulation of Hexokinases II-mediated glycolysis. Int. J. Biol. Sci. 2019, 15, 2497–2508.
- Liu, H.; Zhang, L.; Li, G.; Gao, Z. Xanthohumol protects against Azoxymethane-induced colorectal cancer in Sprague-Dawley rats. Environ. Toxicol. 2020, 35, 136–144.
- Yong, W.K.; Malek, S.N.A. Xanthohumol induces growth inhibition and apoptosis in ca ski human cervical cancer cells. Evid. Based Complement. Altern. Med. 2015, 2015, 921306.
- Gieroba, B.; Arczewska, M.; Sławińska-Brych, A.; Rzeski, W.; Stepulak, A.; Gagoś, M. Prostate and breast cancer cells death induced by xanthohumol investigated with Fourier transform infrared spectroscopy. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2020, 231, 118112.
- Zhang, W.; Pan, Y.; Gou, P.; Zhou, C.; Ma, L.; Liu, Q.; Du, Y.; Yang, J.; Wang, Q. Effect of xanthohumol on Th1/Th2 balance in a breast cancer mouse model. Oncol. Rep. 2018, 39, 280–288.
- Buckwold, V.E.; Wilson, R.J.; Nalca, A.; Beer, B.B.; Voss, T.G.; Turpin, J.A.; Buckheit, R.W., III; Wei, J.; Wenzel-Mathers, M.; Walton, E.M.; et al. Antiviral activity of hop constituents against a series of DNA and RNA viruses. Antivir. Res. 2004, 61, 57–62.
- Wang, Q.; Ding, Z.H.; Liu, J.K.; Zheng, Y.T. Xanthohumol, a novel anti-HIV-1 agent purified from Hops Humulus lupulus. Antivir. Res. 2004, 64, 189–194.
- Bradley, R.; Langley, B.O.; Ryan, J.J.; Phipps, J.; Hanes, D.A.; Stack, E.; Jansson, J.K.; Metz, T.O.; Stevens, J.F. Xanthohumol microbiome and signature in healthy adults (the XMaS trial): A phase I triple-masked, placebo-controlled clinical trial. Trials 2020, 21, 835.
- Yang, H.; Yang, M.; Ding, Y.; Liu, Y.; Lou, Z.; Zhou, Z.; Sun, L.; Mo, L.; Ye, S.; Pang, H.; et al. The crystal structures of severe acute respiratory syndrome virus main protease and its complex with an inhibitor. Proc. Natl. Acad. Sci. USA 2003, 100, 13190–13195.
- Fu, L.; Ye, F.; Feng, Y.; Yu, F.; Wang, Q.; Wu, Y.; Zhao, C.; Sun, H.; Huang, B.; Niu, P.; et al. Both Boceprevir and GC376 efficaciously inhibit SARS-CoV-2 by targeting its main protease. Nat. Commun. 2020, 11, 4417.
- Vuong, W.; Khan, M.B.; Fischer, C.; Arutyunova, E.; Lamer, T.; Shields, J.; Saffran, H.A.; McKay, R.T.; van Belkum, M.J.; Joyce, M.A.; et al. Feline coronavirus drug inhibits the main protease of SARS-CoV-2 and blocks virus replication. Nat. Commun. 2020, 11, 4282.
- Qiao, J.; Li, Y.S.; Zeng, R.; Liu, F.L.; Luo, R.-H.; Huang, C.; Wang, Y.-F.; Zhang, J.; Quan, B.; Shen, C.; et al. SARS-CoV-2 M pro inhibitors with antiviral activity in a transgenic mouse model. Science 2021, 371, 1374–1378.
- Zhu, L.; George, S.; Schmidt, M.F.; Al-Gharabli, S.I.; Rademann, J.; Hilgenfeld, R. Peptide aldehyde inhibitors challenge the substrate specificity of the SARS-coronavirus main protease. Antivir. Res. 2011, 92, 204–212.
- Tomar, S.; Johnston, M.L.; John, S.; Osswald, H.L.; Nyalapatla, P.R.; Paul, L.N.; Ghosh, A.K.; Denison, M.; Mesecar, A.D. Ligand-induced Dimerization of Middle East Respiratory Syndrome (MERS) Coronavirus nsp5 Protease (3CLpro): Implications for Nsp5 Regulation and the Development of Antivirals. J. Biol. Chem. 2015, 290, 19403–19422.
- Wang, F.; Chen, C.; Yang, K.; Xu, Y.; Liu, X.; Gao, F.; Liu, H.; Chen, X.; Zhao, Q.; Liu, X.J.; et al. Michael Acceptor-Based Peptidomimetic Inhibitor of Main Protease from Porcine Epidemic Diarrhea Virus. J. Med. Chem. 2017, 60, 3212–3216.
- Anand, K.; Ziebuhr, J.; Wadhwani, P.; Mesters, J.R.; Hilgenfeld, R. Coronavirus Main Proteinase (3CLpro) Structure: Basis for Design of Anti-SARS Drugs. Science 2003, 300, 1763–1767.





