Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Thimios Mitsiadis | + 9005 word(s) | 9005 | 2021-11-23 07:15:42 | | | |
| 2 | Vivi Li | -4962 word(s) | 4043 | 2021-12-02 10:31:03 | | | | |
| 3 | Vivi Li | Meta information modification | 4043 | 2021-12-02 10:31:32 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Mitsiadis, T.; Marconi, G.D.; Porcheri, C.; Trubiani, O. Current Regenerative Medicine Approaches: Three-Dimensional Culture Systems. Encyclopedia. Available online: https://encyclopedia.pub/entry/16674 (accessed on 12 May 2026).
Mitsiadis T, Marconi GD, Porcheri C, Trubiani O. Current Regenerative Medicine Approaches: Three-Dimensional Culture Systems. Encyclopedia. Available at: https://encyclopedia.pub/entry/16674. Accessed May 12, 2026.
Mitsiadis, Thimios, Guya Diletta Marconi, Cristina Porcheri, Oriana Trubiani. "Current Regenerative Medicine Approaches: Three-Dimensional Culture Systems" Encyclopedia, https://encyclopedia.pub/entry/16674 (accessed May 12, 2026).
Mitsiadis, T., Marconi, G.D., Porcheri, C., & Trubiani, O. (2021, December 02). Current Regenerative Medicine Approaches: Three-Dimensional Culture Systems. In Encyclopedia. https://encyclopedia.pub/entry/16674
Mitsiadis, Thimios, et al. "Current Regenerative Medicine Approaches: Three-Dimensional Culture Systems." Encyclopedia. Web. 02 December, 2021.
Copy Citation
Three-dimensional (3D) culture systems opened up new horizons in the emerging field of regenerative medicine and in studying the biology of tissues and organs, modelling various diseases, and screening drugs. Producing accurate in vitro models increases the possibilities for studying molecular control of cell–cell and cell–microenvironment interactions in detail. The Notch signalling is linked to cell fate determination, tissue definition, and maintenance in both physiological and pathological conditions. Hence, 3D cultures provide new accessible platforms for studying activation and modulation of the Notch pathway.
spheroids
organoids
organ-on-a-chip
microfluidics
3D culture systems
Notch signalling
cancer
drug screening
regeneration
1. Introduction
In vitro cultures have been established over a century ago and are largely used for studying physiological and pathological conditions, modulation of signalling pathways, and gene expression modifications. Cells derived from normal and/or diseased tissues can be grown either as adherent monolayers or in suspension. These two-dimensional (2D) tissue cultures only partially recapitulate the cellular behaviour of the tissue of origin, as they cannot reproduce complex cell–cell and cell–extracellular matrix interactions. This prompted the development of three-dimensional (3D) cell culture systems, where the existing interactions between the various cell populations in the tissue of origin are largely maintained [1][2][3][4]. Recently, 3D culture systems have been further improved, by incorporating structural tissue elements, including components of the extracellular matrix (ECM). Cells can be grown in 3D culture settings that support the formation of tissue-specific spheroids and organoids, as well as in more complex organotypic models (e.g., “organ-on-a-chip”). These models provide a unique platform to dissect the molecular communication between cells and, ultimately, the role of molecular pathways in the regulation of tissue homeostasis.
2. The Notch Signalling Pathway
Notch signalling is one of the major pathways controlling cell fate specification, intercellular communication, tissue organisation, and morphology [5][6][7][8]. The canonical Notch signalling is a cell–cell communication mechanism, where juxtaposed cells physically interact via transmembrane receptors and ligands exposed on opposite cell membranes [9][10][11]. Four type of Notch receptors (Notch1, Notch2, Notch3, and Notch4) and five Notch ligands (Jagged1 and Jagged2, Delta-like1, Delta-like3, and Delta-like4) have been described in mammals. These molecules share a high degree of sequence homology with their Drosophila counterpart, which is an indicator of a highly conserved pathway throughout evolution. The interaction between receptors and ligands can give rise to either a lateral induction or a lateral inhibition, with respective activation or inhibition of ligand expression in the neighbouring cells [12]. Notch–ligand interaction triggers a cascade of protein cleavages, leading to the release of the active Notch intracellular domain (NICD). NICD translocates to the nucleus where it forms a transcriptional complex with Suppressor of Hairless (or RBP-Jk or CSL), Mastermind and transcriptional co-activators to modulate the expression of Notch-downstream target genes (mainly Hes and Hey families), ultimately resulting in the regulation of the Notch ligand transcription (Figure 1a,b). This signalling cascade is relatively simple since only few proteins are involved in the Notch pathway. However, the combinations of interactions between the four different ligands and the five different receptors might generate downstream diversity with distinctive signalling outputs. Although this is not yet well-studied, several findings suggest differences among ligands and receptors, as well as diversity of post-translational modifications of Notch receptors [13]. Non-canonical, CSL-independent NICD activity and ligand-independent activation of Notch signalling have been also reported [13]. In the case of lateral induction, the expression of the ligand in the receiving cell increases, while a lateral inhibition reduces the expression of the same ligand [10][14]. Finally, receptors and ligands can be expressed on the same cell, reducing their availability for interactions with the neighbouring cell. This cis-inhibition is a fine tuning of the Notch signalling that results in a general inhibition of the pathway (Figure 1c) [15].
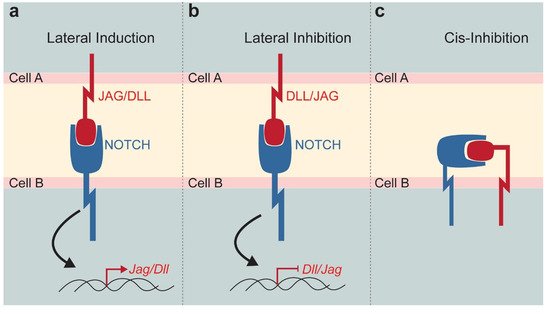
Figure 1. Schematic representation of the Notch signalling function. Notch pathway modulation strongly depends on the Notch receptor—DLL/JAG ligand interaction established between neighbouring cells. (a) Lateral induction triggers the expression of the Jag or Dll in the receiving cell. (b) Lateral inhibition reduces the expression of Dll or Jag in the neighbouring cell. (c) Cis-inhibition occurs when receptor and ligand expressed by the same cell, activates the pathway and sequesters active molecules on a single cell surface. Abbreviations: DLL, Delta-like; JAG, Jagged.
During organogenesis, the Notch pathway is involved in cell fate determination and the formation of tissue boundaries [16]. During adulthood, Notch signalling plays an important role in regulating the behaviour of stem cells, by maintaining their stemness and influencing their fate in most tissues [17][18]. Notch signalling is active in various adult stem cell niches [19]. For example, interaction between Notch1 and Delta-like4 (Dll4) is necessary for establishing hematopoietic clusters in the aorta-gonad-mesonephros, which is the primordial niche for hematopoietic stem cells [20]. Similarly, Notch signalling is primordial for neural stem cells maintenance during both the embryonic and adult life [21][22][23]. Neural stem cells expressing Notch keep their undifferentiated status, while absence of Notch induces their differentiation [23][24]. Indeed, adult stem cells of the dentate gyrus are maintained in a semi-quiescent state via the Notch1-Hes5 axis, and depletion of Notch1 results in a reduced number of neurons [25].
A number of diseases are linked to mutations in the Notch signalling pathway [13][26]. NOTCH3 mutations in humans cause the Cerebral Autosomal Dominant Anteriopathy with Subcortical Infracts and Leukoencephalopathy (CADASIL), a disease associated with recurring small brain infarcts and degeneration of vascular smooth muscle cells in the brain. Mutations in the JAGGED1 (JAG1) ligand and more rarely in the NOTCH2 receptor cause the Alagille syndrome that affects multiple organs, including heart, liver, kidney, and craniofacial organs. Furthermore, NOTCH1 mutations are linked to aortic valve diseases, and DELTA-LIKE3 (DLL3) mutations to spondylocostal dysostosis [13]. The Notch pathway is also involved in the generation of cancers [27]. Frequently, components of the Notch signalling are not mutated in most cancers. However, NOTCH1 mutations have been detected in patients with acute lymphoblastic leukaemia (T-ALL) tumours, as well as in patients with both, small cell lung cancer (SCLC) and non-small cell lung cancer (NSCLC). Furthermore, NOTCH1 mutations in the skin lead to the generation of skin cancer, while mutations in the hematopoietic compartment lead to myeloproliferative disorders [26][28]. In addition, upregulation of NOTCH1 and/or JAG1 can lead to various pathologies, such as breast cancer [29] and prostate cancer [30]. These findings suggest that Notch signalling disbalance plays a very important critical role in the generation and progression of a disease.
Since Notch signalling plays a major role in organogenesis, pathology, and regeneration, different 3D culture platforms such as spheroids, organoids, and “organ-on-a-chip” devices can be used to dissect the roles and the regulation of this pathway in various healthy and pathological cell populations. The fine tuning of the levels of Notch signalling may provide new therapeutic scenarios for the various Notch-linked diseases, and this possibility could be fully investigated and explored in the above-mentioned 3D culture systems.
3. 3D Culture Systems
3D culture systems provide the new means of generating tissues in a more physiological manner. However, these systems never fully recapitulate heterogeneity and the complexity of the organs and tissues, since they lack the fluctuations that exist in vivo. Furthermore, the major components of 3D systems, such as cell input, ECM parameters, protein concentrations, are combined in a manner that represents the simplified version of the in vivo unit. Here, we present the most used 3D culture systems, namely the spheroids, organoids, and “organ-on-a-chip” devices.
3.1. Spheroids
Isolated cells cultured in low-attachment wells naturally aggregate to form 3D microtissues, known as multi-cellular spheroids. This mechanism of cell self-assembly has been initially observed in sponges, where upon their dissociation, cells were able to aggregate and recreate a new sponge body [31][32]. More recent studies have shown that pluripotent stem cells (PSCs) derived from murine blastocyst were also able to aggregate, thus forming the embryoid bodies (EBs), which can be maintained as 3D structures in vitro [33]. This simplified 3D culture system allowed the investigation of complex cellular and molecular mechanisms occurring during embryo growth. Intercellular communication and differentiation inputs were preserved in the EBs, thus modelling the development of the three germ cell layers and their reciprocal interaction (i.e., ectoderm, mesoderm, and endoderm) [34][35]. Similarly, Chinese hamster V79 lung cells grown as single cell suspension were able to aggregate and form spheroids that are characterised by a hierarchical structure, where proliferating cells occupy the most peripheral parts and quiescent cells the central part (core) of the spheroids [36]. This spheroidal organisation broadly mirrors the structural complexity of living tissues, where different cell types position themselves according to their needs for oxygen, nutritional gradients, and interactions with other subgroups of cells. Therefore, multicellular spheroids preserve the important biological properties of the tissue of origin, providing higher cell viability, stable morphology, and physiological metabolic activity (Figure 2a) [2][37][38].
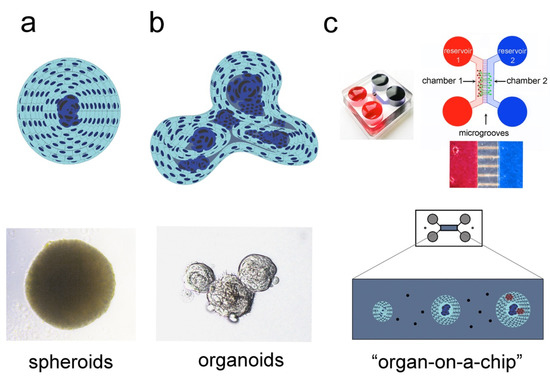
Figure 2. 3D culture systems used to recapitulate the in vivo tissue organisation and complexity. Simplified overview of three different culture models: spheroids (a), organoids (b), and “organ-on-a-chip” (c). (a) Spheroids represent the simplest 3D culture system. Spheroids constitute efficient model systems for studying cell–cell communication, cellular metabolism, oxygen gradients, and nutrients distribution. Their complexity can be scaled up by integrating into the spheroids different cell types, such as stem cells and endothelial cells, thus providing the ideal platform for biochemical and molecular testing, as well as rapid drug screening. (b) Organoids reproduce the complexity of the native tissue, where cells at various cytodifferentiation states, ranging from stem cells and progenitors to fully differentiated cells, coexist. (c) “Organ-on-a-chip” closely mimics several aspects of the native organ. Integrated into a microfluidic system, 2D cultures, spheroids and organoids can be exposed to a dynamic microenvironment that allows controlled fluid exchange and interactions between different tissues, such as vessels and nerves (adapted from [39][40]).
The 3D structure of spheroids provides the ideal setting for studying the fine regulation of Notch signalling and the precise Notch ligand–receptor interactions. Endothelial cells have been successfully co-cultured with specific mesenchymal stem cell (MSCs) populations for the formation of mixed spheroids. These spheroids demonstrate an increased survival rate that correlates with upregulation of Notch expression [41]. More particularly, the vasculogenic potential in mixed spheroids appears to be Notch3-dependent, as Notch3 deletion blocks the sprouting of new forming vessels [42]. Although nutrients and oxygen are easily distributed in the external layers of the spheroids, these elements are less accessible in their core. Cells situated in the core of the spheroids react to these hypoxic conditions by activating the Notch pathway, which in turn correlates with the upregulation of hypoxia-inducible factor-1 (HIF-1), hence triggering angiogenesis [42][43].
3.2. Organoids
Organoids are generated from adult stem cells (ASCs) and/or PSCs of various tissues and organs. In these 3D cultures, the differentiation program of the tissue of origin is maintained, and a simplified miniature version of the organ develops in vitro (Figure 2b). Organoids can acquire a spherical shape in the initial phase, but their further growth leads to the establishment of more complex 3D structures that closely resemble the organ of origin [37][44][45]. Organoids have been successfully generated from brain, intestine, heart, thymus, liver, lung, and pancreas [31][37][45][46][47][48][49][50]. Therefore, organoids are well-accepted model systems for studying stem cell behaviour in pathophysiological conditions, and further analyse cell–cell communication and the cells–ECM interactions. Unlike spheroids, which develop in the absence of scaffold, the development and survival of organoids often requires the presence of various biomimetic materials (e.g., Hydrogel, Matrigel) [51][52]. Organoids are more complex than spheroids, but largely maintain basic features of the tissue of origin by recapitulating its essential function and spatially-restricted cell lineage commitment [52][53].
Modulation of Notch signalling in organoids might influence the fate and behaviour of cells, as well as the crosstalk between stem cells and stroma cells, thus recapitulating the in vivo situation. Indeed, the oscillatory expression patterns of the Notch effector Hes1, reported in embryonic stem cells (ESCs) and during neurogenesis, is maintained in cortical organoids generated from induced PSCs (iPSCs) [54]. Cell differentiation is disrupted when Hes1 oscillations cease, which suggests that tissue homeostasis and stem cell maintenance are strictly controlled by the Notch signalling [55][56][57].
3.3. “Organ-on-a-chip” Technology
Spheroids and organoids represent an important step forward in the reproduction of the in vivo physiology, although major elements of complexity are still lacking (e.g., lack of innervation, absence of fluid perfusion to create a dynamic environment, inefficient nutrient and waste transport, etc.) [44][45][46][57][58][59]. Organoids can be placed in microfluidic chambers that work as an irrigation system allowing continuous flow of nutrients, thus mimicking the vascular system by carrying nutrients and oxygen [58][60]. Spheroids/organoids grown in a microfluidic chamber face physical constraints, including shearing forces, compression-induced stretches and stiffness variations in the ECM [61][62][63]. These biomechanical cues reflect the in vivo stimuli and contribute to create more accurate copies of the native organs [63][64][65]. Drugs, chemical products, and signalling molecules can be evenly distributed in the chamber where cells are grown, thus emulating the biological tissue reactions upon the various pharmaceutical treatments in vivo (Figure 2c) [39]. Different cell types from a given tissue/organ can be co-cultured in these microfluidic devices, thus establishing the inter-cellular crosstalk that guarantees tissue functionality. “Organ-on-a-chip” devices are already available for various organs and tissues, including kidney, bone, cartilage, skin, and ovary [40].
Innervation in microfluidics can be simulated upon co-culture of ganglia or single neurons with spheroids/organoids, where the establishment of neuronal connections with specific cell populations can be studied in detail [66]. The interactions between neurons and cells might define the fate of cells and affect their cytoskeleton restructuring [57]. Microelectrode arrays have been also used in microfluidic constructions for the simulation of the electrical activity of the neuronal network [65][67].
The infusion of components of the immune system in the microfluidic systems could be useful for studying tissue reactions under pathological conditions [68][69][70]. The Notch pathway is central in the regulation of the immune system [71][72]. Notch signalling, which is directly involved in the maturation of lymphoid organs, is activated during pro-inflammatory response by cytokines such as Tumour Necrosis Factor (TNF) and Interleukin-1 (IL1), and regulates the myelomonocytic differentiation via inhibition of Hes1 [73][74][75][76][77].
4. 3D Systems in Regenerative Medicine
The recent developments in 3D culture systems open new horizons in medical fields relying on the therapeutic potential of stem cells. Stem cells require specific microenvironmental cues to preserve their self-renewal capacities and undifferentiation status. 3D cultures have been established with a variety of stem cell types, using ESCs, ASCs and iPSCs, among others [45][46]. Stem cells grown in organoids preserve their self-renewal characteristics, the expression of molecules linked to stemness, and their ability to progress into various specific cell fates [78][79]. Adult bone marrow was the initially identified source of MSCs [80]. Over the last decades, MSCs populations were isolated from many other tissues and organs, such as the adipose tissue, periosteum, trabecular bone, synovium, skeletal muscle, and teeth [81][82][83]. MSCs isolated from human teeth can be grown in 3D spheroids, and are able to differentiate into many cell types, such as osteocytes, chondrocytes, adipocytes, and neuronal cells [21][81][82][83][84][85]. Both in vitro and in vivo studies have demonstrated that the osteogenic potential of dental stem cells (DSCs) cultured as spheroids has improved, when compared to that of the 2D cultured DSCs [17][86][87][88]. Similarly, in vitro studies have shown a higher cell differentiation potential in spheroids than in 2D culture systems [89]. To analyse the in vivo regenerative/healing potential of spheroids and organoids, these 3D structures can be implanted in pathological and defective tissues of various animal models. Upon implantation into the malfunctioning tissue, stem cells from the 3D structure could initiate the healing process and contribute to the complete tissue restoration, as this was the case for the intestinal tissue [41][50][90]. 3D structures generated by patient-derived stem cells, can also be used for analysing and testing various pharmaceutical products (i.e., drug screening), thus establishing the basis for future regenerative personalised treatments [53].
These novel tissue engineering techniques, combined with modern genetic and pharmacological tools provide unprecedented powerful solutions in the field of regenerative medicine for the repair of damaged tissues. However, important limitations still exist for transferring these techniques in the clinics. Implantation of any sort of undifferentiated material into the live tissues, for therapeutic purposes, is not yet free of side-effects and carries an enormous risk of uncontrolled cell proliferation and cancer initiation [91][92][93]. Therefore, a deep knowledge on the signalling molecules involved in the establishment of the stem cells niches in the 3D structures is necessary in order to overcome these limitations. The maintenance of the stem cell niches is complex and requires the interaction between molecules of different signalling pathways [18][94]. Accessible in vitro systems mimicking the characteristics and structure of stem cell compartments of various organs have been already used for studying the role of Notch signalling in stem cell populations [15]. The interaction between Wnt and Notch signalling has been described in human iPSC-derived cortical spheroids, where inhibition of either one or both pathways can affect brain tissue identity [95].
5. 3D Cultures in Disease Modelling
Pathological conditions arise from structural and functional alterations of the organ. Cells from pathological tissues can be used in 3D systems for analysing the molecular and cellular mechanisms involved in these pathological processes. Additionally, bioengineered 3D models can be used for testing drug toxicity and efficacy, a first step in the development of novel pharmacological products. Albeit partially, several pathophysiological conditions such as wound healing processes, inflammatory diseases and cancers are currently reproduced in 3D cultures [96][97][98][99][100][101]. We here present three systems as a proof of principle for the usage of 3D culture in disease modelling.
Skin wound healing is a complex process involving vascularisation, stem cell migration, immunoreaction, growth factors release, and synthesis of ECM [102]. All these processes have to be strictly synchronised. Spheroids containing specific cell populations can be transplanted into wounded tissues, thus greatly improving the healing process, due to the release of growth factors, induced immunomodulation, and targeted integration to the damaged tissue [103][104]. In vitro skin models have been developed from keratinocytes and fibroblasts co-cultures, grown in specific ECM scaffolds [96]. Air–liquid interface adds to the complexity of these 3D structures that represent accurate models for skin wound healing therapies [96]. Notch signalling is activated during wound healing, while its blockage results in important delay of tissue regeneration. On the other hand, induction of its activity enhances tissue repair [105]. Similarly, modulation of Notch affects keratinocyte behaviour in scratch assays in vitro [105][106]. 3D cultures can therefore be used to analyse the roles of Notch molecules during the various stages of skin regeneration and wound healing [105].
Chronic liver diseases, where the inflammatory processes derive from infections or continuous exposure of chemicals, are often reproduced in vitro using patient-derived cells or cell lines [107][108]. However, primary cells are viable for only a limited number of passages, while immortalised cell lines may not completely mirror the physiological conditions [109]. Liver-derived organoids bypass these limitations, allowing for the maintenance of hepatoblasts in culture for longer time [110]. In contrast to classical hepatocyte monolayer models, the 3D culture system does not result in cell dedifferentiation and changes in their metabolic activities [109][111][112]. Hence, 3D spheroids from hepatic cells constitute interesting tools for studying chronic and acute conditions in liver diseases [113]. Genetic defects of Notch pathway components result in severe liver malformations. For example, mutations of the Jagged1 gene result in bile ducts reduction and consequent dysfunction of the biliary tree [114][115]. Hence, the generation of organoids from healthy and pathological liver tissues provides an additional tool for further exploring the potential therapeutic roles of Notch signalling [116].
Cancer is one of the most common disease models that can be reproduced in 3D cultures. Cancer establishes a self-protective environment that allows aberrant cells to access nutrients and resources, to ultimately expand and self-renew while escaping the immune surveillance of the host [117]. This specialised microenvironment can be partially reproduced in cancer-derived spheroids [118]. Most cancers have a component of undifferentiated cells, known as cancer stem cells, which maintain the tumour core and allow seeding of metastasis in distant locations [118][119][120]. Organoids and spheroids derived from cancer tissues represent excellent platforms for drug screening and personalised medicine. These 3D systems allow for expansion of the initially limited pool of cancer cells, thus recapitulating essential tumour features [121]. Cancer-derived spheroids or organoids can be also studied in microfluidic “organ-on-a-chip” devices, allowing a more accurate and complete investigation [3][122][123]. The Notch pathway can play both an oncogenic and anti-tumorigenic role, depending on the type of cancer and the tissue involved. Notch signalling regulation during neo-angiogenesis plays a crucial role in restructuring the tumorigenic microenvironment, influencing oxygen and nutrients income [124]. Additionally, the role of Notch in immunomodulation has a great impact in the tumorigenic growth [72]. The generation of tumours strongly depends on specific cell–cell communications and cell–ECM interactions. 3D culture models are of great help in order to understand tissue dysfunction and cancer formation due to aberrations in Notch signalling [120][125][126]. Neuroblastoma and breast cancer-derived spheroids showed upregulation of Notch expression, concomitant to the increase of cells that are positive to cancer stem cell markers [127][128]. Notch1 and Notch2 deletions lead to structural alterations in cutaneous squamous cell carcinoma [129][130]. Intestinal tumoroids are also used to study the effects of aberrant Notch signalling in the generation of colorectal cancers [131][132][133][134][135].
6. Drug Screening and Therapy Testing in 3D Culture Systems
Drug discovery is a multifaceted, time-consuming, risky, and expensive process, which aims at the identification of novel molecules with therapeutic potential. Newly developed molecules might present unexpected and unwanted effects during the clinical trial phases, that could be associated with the limitations of current preclinical models [136]. The conventional 2D cell culture systems do not accurately simulate the in vivo tissue reactions in both physiological and pathological conditions, thus limiting their value to study pharmacokinetics and pharmacodynamics of new drugs [4]. Instead, 3D culture models have gained attention in recent years in the pharmacological field, including the drug discovery field, mainly due to their capability to emulate faithfully the in vivo testing conditions [136]. The advantages of 3D models to simulate the characteristics of the native tissue, promoted their applicability in clinical testing, drug screening, disease modelling, and in predicting the acquisition of drug resistance [1][137][138]. Spheroids and/or organoids allow accurate evaluation of cell alterations due to oxygen, growth factors, nutrients, and drug modifications [139]. Spheroids and organoids can be used in personalised medicine for high-throughput screening of pharmacological responses. Microfluidic devices assess more faithfully drug concentrations, but several caveats still remain in translating the results to in vivo settings [138][140][141]. It is therefore obvious that new important information is collected in recent years by combining tissue, genetic and molecular engineering technologies and “organ-on-a-chip” platforms [142][143]. For example, main features of the “organ-on-a-chip” technology, such as the vascular support, the tissue–tissue communication and the use of different cell types, contributed to more detailed and valid studies on drug effects (in tissues or single cells) [143][144]. These 3D microfluidic devices are also used to predict the pharmacological reactions in humans, ultimately reducing the usage of animal models and clinical trials [144][145][146] (Figure 3).
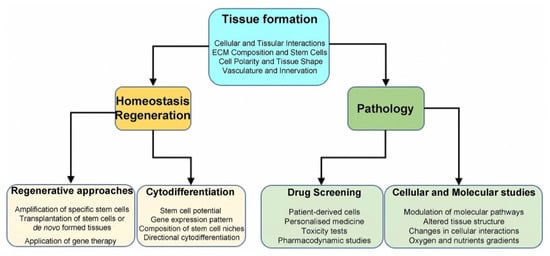
Figure 3. Summary of 3D systems and their applications. Several aspects of tissue organisation can be modelled in 3D cultures. For all applications, spheroids, organoids, and “organ-on-a-chip” represent more faithfully the main features of native tissues when compared to 2D cultures. 3D systems allow characterisation of specific stem cell populations and could be used for regenerative purposes. Furthermore, 3D structures can reproduce a disease environment and serve for drug testing.
Notch pathway deregulation creates severe pathological conditions, including cancer and cardiovascular diseases, and therefore constitutes a promising target for novel pharmacological treatments [126]. Blockage of the Notch activity can be achieved by interference between receptor–ligand interaction (e.g., via monoclonal antibodies), or by preventing receptor cleavages (GSI-DAPT molecules) [126][147]. The efficiency of these approaches is strongly limited by severe side effects, mainly due to systemic off-target effects [147][148][149]. Thus, developing 3D in vitro models that preserve the tissue complexity might constitute a safe way for molecular analysis and understanding the big potential of Notch signalling modulation in disease treatment.
7. Conclusions
3D culture systems that faithfully mimic the in vivo environment and cell behaviour are excellent simulation models for physiological and pathological conditions. These technologically advanced tools are extremely useful for studying specific molecular cues and analysing the role of specific signalling pathways, including the Notch pathway, in the context of specific disease models, drug responses and tissue regeneration.
Abbreviations
| NICD | Notch Intracellular Domain |
| Dll4 | Delta-like4 |
| 3D | Three-dimensional |
| EBs | Embryoid Bodies |
| PSCs | Pluripotent Stem Cells |
| MSCs | Mesenchymal Stem Cells |
| HIF1 | Hypoxia inducible factor 1 |
| ASCs | Adult Stem Cells |
| ECM | Extracellular Matrix |
| ESCs | Embryonic Stem Cells |
| iPSC | Induced-Pluripotent Stem Cells |
| TNF | Tumour Necrosis Factor |
| IL1 | Interleukin-1 |
| NGS | Next Generation Sequencing |
| DSCs | Dental Stem Cells |
References
- Jensen, C.; Teng, Y. Is It Time to Start Transitioning From 2D to 3D Cell Culture? Front. Mol. Biosci. 2020, 7, 33.
- Chaicharoenaudomrung, N.; Kunhorm, P.; Noisa, P. Three-dimensional cell culture systems as an in vitro platform for cancer and stem cell modeling. World J. Stem Cells 2019, 11, 1065–1083.
- Doctor, A.; Seifert, V.; Ullrich, M.; Hauser, S.; Pietzsch, J. Three-Dimensional Cell Culture Systems in Radiopharmaceutical Cancer Research. Cancers 2020, 12, 2765.
- Fang, Y.; Eglen, R.M. Three-Dimensional Cell Cultures in Drug Discovery and Development. SLAS Discov. Adv. Life Sci. R D 2017, 22, 456–472.
- Koch, U.; Lehal, R.; Radtke, F. Stem cells living with a Notch. Development 2013, 140, 689–704.
- Mitsiadis, T.A.; Graf, D.; Luder, H.; Gridley, T.; Bluteau, G. BMPs and FGFs target Notch signalling via jagged 2 to regulate tooth morphogenesis and cytodifferentiation. Development 2010, 137, 3025–3035.
- Mitsiadis, T.A.; Henrique, D.; Thesleff, I.; Lendahl, U. Mouse Serrate-1 (Jagged-1): Expression in the developing tooth is regulated by epithelial-mesenchymal interactions and fibroblast growth factor-4. Development 1997, 124, 1473–1483.
- Mitsiadis, T.A.; Lardelli, M.; Lendahl, U.; Thesleff, I. Expression of Notch 1, 2 and 3 is regulated by epithelial-mesenchymal interactions and retinoic acid in the developing mouse tooth and associated with determination of ameloblast cell fate. J. Cell Biol. 1995, 130, 407–418.
- Simpson, P. Lateral inhibition and the development of the sensory bristles of the adult peripheral nervous system of Drosophila. Dev. Camb. Engl. 1990, 109, 509–519.
- Louvi, A.; Artavanis-Tsakonas, S. Notch signalling in vertebrate neural development. Nat. Rev. Neurosci. 2006, 7, 93–102.
- Artavanis-Tsakonas, S. Notch Signaling: Cell Fate Control and Signal Integration in Development. Science 1999, 284, 770–776.
- Lewis, J. Notch signalling and the control of cell fate choices in vertebrates. Semin. Cell Dev. Biol. 1998, 9, 583–589.
- Braune, E.-B.; Lendahl, U. Notch—A goldilocks signaling pathway in disease and cancer therapy. Discov. Med. 2016, 21, 189–196.
- Artavanis-Tsakonas, S.; Delidakis, C.; Fehon, R.G. The Notch locus and the cell biology of neuroblast segregation. Annu. Rev. Cell Biol. 1991, 7, 427–452.
- Bigas, A.; Porcheri, C. Notch and Stem Cells. Adv. Exp. Med. Biol. 2018, 1066, 235–263.
- Irvine, K.D.; Vogt, T.F. Dorsal-ventral signaling in limb development. Curr. Opin. Cell Biol. 1997, 9, 867–876.
- Monterubbianesi, R.; Bencun, M.; Pagella, P.; Woloszyk, A.; Orsini, G.; Mitsiadis, T.A. A comparative in vitro study of the osteogenic and adipogenic potential of human dental pulp stem cells, gingival fibroblasts and foreskin fibroblasts. Sci. Rep. 2019, 9, 1761.
- Mitsiadis, T.A.; Barrandon, O.; Rochat, A.; Barrandon, Y.; De Bari, C. Stem cell niches in mammals. Exp. Cell Res. 2007, 313, 3377–3385.
- Pagella, P.; Vargas Roditi, L.; Stadlinger, B.; Moor, A.E.; Mitsiadis, T.A. Notch signaling in the dynamics of perivascular stem cells and their niches. Stem Cells Transl. Med. 2021, 10, 1433–1445.
- Porcheri, C.; Golan, O.; Calero-Nieto, F.J.; Thambyrajah, R.; Ruiz-Herguido, C.; Wang, X.; Catto, F.; Guillén, Y.; Sinha, R.; González, J.; et al. Notch ligand Dll4 impairs cell recruitment to aortic clusters and limits blood stem cell generation. EMBO J. 2020, 39, e104270.
- Pagella, P.; Miran, S.; Neto, E.; Martin, I.; Lamghari, M.; Mitsiadis, T.A. Human dental pulp stem cells exhibit enhanced properties in comparison to human bone marrow stem cells on neurites outgrowth. FASEB J. 2020, 34, 5499–5511.
- Knoblich, J.A.; Jan, L.Y.; Jan, Y.N. Asymmetric segregation of Numb and Prospero during cell division. Nature 1995, 377, 624–627.
- Gaiano, N.; Nye, J.S.; Fishell, G. Radial glial identity is promoted by Notch1 signaling in the murine forebrain. Neuron 2000, 26, 395–404.
- Knoblich, J.A. Asymmetric cell division during animal development. Nat. Rev. Mol. Cell Biol. 2001, 2, 11–20.
- Ables, J.L.; DeCarolis, N.A.; Johnson, M.A.; Rivera, P.D.; Gao, Z.; Cooper, D.C.; Radtke, F.; Hsieh, J.; Eisch, A.J. Notch1 Is Required for Maintenance of the Reservoir of Adult Hippocampal Stem Cells. J. Neurosci. 2010, 30, 10484–10492.
- Louvi, A.; Artavanis-Tsakonas, S. Notch and disease: A growing field. Semin. Cell Dev. Biol. 2012, 23, 473–480.
- Ho, D.M.; Artavanis-Tsakonas, S. The Notch-Mediated Proliferation Circuitry. Curr. Top. Dev. Biol. 2016, 116, 17–33.
- Reichrath, J.; Reichrath, S. The Impact of Notch Signaling for Carcinogenesis and Progression of Nonmelanoma Skin Cancer: Lessons Learned from Cancer Stem Cells, Tumor Angiogenesis, and Beyond. Adv. Exp. Med. Biol. 2021, 1287, 123–154.
- Stylianou, S.; Clarke, R.B.; Brennan, K. Aberrant activation of notch signaling in human breast cancer. Cancer Res. 2006, 66, 1517–1525.
- Deng, G.; Ma, L.; Meng, Q.; Ju, X.; Jiang, K.; Jiang, P.; Yu, Z. Notch signaling in the prostate: Critical roles during development and in the hallmarks of prostate cancer biology. J. Cancer Res. Clin. Oncol. 2016, 142, 531–547.
- Wilson, H.V. On some phenomena of coalescence and regeneration in sponges. J. Exp. Zool. 1907, 5, 245–258.
- Wilson, H.V. A new method by which sponges may be artificially reared. Science 1907, 25, 912–915.
- Martin, G.R. Isolation of a pluripotent cell line from early mouse embryos cultured in medium conditioned by teratocarcinoma stem cells. Proc. Natl. Acad. Sci. USA 1981, 78, 7634–7638.
- Desai, N.; Rambhia, P.; Gishto, A. Human embryonic stem cell cultivation: Historical perspective and evolution of xeno-free culture systems. Reprod. Biol. Endocrinol. 2015, 13, 9.
- Morgani, S.M.; Canham, M.A.; Nichols, J.; Sharov, A.A.; Migueles, R.P.; Ko, M.S.H.; Brickman, J.M. Totipotent embryonic stem cells arise in ground-state culture conditions. Cell Rep. 2013, 3, 1945–1957.
- Achilli, T.-M.; Meyer, J.; Morgan, J.R. Advances in the formation, use and understanding of multi-cellular spheroids. Expert Opin. Biol. Ther. 2012, 12, 1347–1360.
- Clevers, H. Modeling Development and Disease with Organoids. Cell 2016, 165, 1586–1597.
- Natsiou, D.; Granchi, Z.; Mitsiadis, T.A.; Jimenez-Rojo, L. Generation of Spheres from Dental Epithelial Stem Cells. Front. Physiol. 2017, 8, 7.
- Pagella, P.; Neto, E.; Lamghari, M.; Mitsiadis, T.A. Investigation of orofacial stem cell niches and their innervation through microfluidic devices. Eur. Cell. Mater. 2015, 29, 213–223.
- Pagella, P.; Cordiale, A.; Marconi, G.; Trubiani, O.; Rasponi, M.; Mitsiadis, T. Bioengineered tooth emulation systems for regenerative and pharmacological purposes. Eur. Cell. Mater. 2021, 41, 502–516.
- Vorwald, C.E.; Joshee, S.; Leach, J.K. Spatial localization of endothelial cells in heterotypic spheroids influences Notch signaling. J. Mol. Med. Berl. Ger. 2020, 98, 425–435.
- De Francesco, E.; Maggiolini, M.; Musti, A. Crosstalk between Notch, HIF-1α and GPER in Breast Cancer EMT. Int. J. Mol. Sci. 2018, 19, 2011.
- Wang, W.-M.; Zhao, Z.-L.; Ma, S.-R.; Yu, G.-T.; Liu, B.; Zhang, L.; Zhang, W.-F.; Kulkarni, A.B.; Sun, Z.-J.; Zhao, Y.-F. Epidermal Growth Factor Receptor Inhibition Reduces Angiogenesis via Hypoxia-Inducible Factor-1α and Notch1 in Head Neck Squamous Cell Carcinoma. PLoS ONE 2015, 10, e0119723.
- Drost, J.; Clevers, H. Organoids in cancer research. Nat. Rev. Cancer 2018, 18, 407–418.
- Lancaster, M.A.; Knoblich, J.A. Organogenesis in a dish: Modeling development and disease using organoid technologies. Science 2014, 345, 1247125.
- Hofer, M.; Lutolf, M.P. Engineering organoids. Nat. Rev. Mater. 2021, 6, 402–420.
- Kim, J.; Koo, B.-K.; Knoblich, J.A. Human organoids: Model systems for human biology and medicine. Nat. Rev. Mol. Cell Biol. 2020, 21, 571–584.
- Simian, M.; Bissell, M.J. Organoids: A historical perspective of thinking in three dimensions. J. Cell Biol. 2017, 216, 31–40.
- Mariani, J.; Vaccarino, F.M. Breakthrough Moments: Yoshiki Sasai’s Discoveries in the Third Dimension. Cell Stem Cell 2019, 24, 837–838.
- Corrò, C.; Novellasdemunt, L.; Li, V.S.W. A brief history of organoids. Am. J. Physiol. Cell Physiol. 2020, 319, C151–C165.
- De Lau, W.; Barker, N.; Low, T.Y.; Koo, B.-K.; Li, V.S.W.; Teunissen, H.; Kujala, P.; Haegebarth, A.; Peters, P.J.; van de Wetering, M.; et al. Lgr5 homologues associate with Wnt receptors and mediate R-spondin signalling. Nature 2011, 476, 293–297.
- Sato, T.; Vries, R.G.; Snippert, H.J.; van de Wetering, M.; Barker, N.; Stange, D.E.; van Es, J.H.; Abo, A.; Kujala, P.; Peters, P.J.; et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 2009, 459, 262–265.
- Sahu, S.; Sharan, S.K. Translating Embryogenesis to Generate Organoids: Novel Approaches to Personalized Medicine. iScience 2020, 23, 101485.
- Trujillo, C.A.; Gao, R.; Negraes, P.D.; Gu, J.; Buchanan, J.; Preissl, S.; Wang, A.; Wu, W.; Haddad, G.G.; Chaim, I.A.; et al. Complex Oscillatory Waves Emerging from Cortical Organoids Model Early Human Brain Network Development. Cell Stem Cell 2019, 25, 558–569.e7.
- Shimojo, H.; Ohtsuka, T.; Kageyama, R. Oscillations in notch signaling regulate maintenance of neural progenitors. Neuron 2008, 58, 52–64.
- Shimojo, H.; Isomura, A.; Ohtsuka, T.; Kori, H.; Miyachi, H.; Kageyama, R. Oscillatory control of Delta-like1 in cell interactions regulates dynamic gene expression and tissue morphogenesis. Genes Dev. 2016, 30, 102–116.
- Kobayashi, T.; Mizuno, H.; Imayoshi, I.; Furusawa, C.; Shirahige, K.; Kageyama, R. The cyclic gene Hes1 contributes to diverse differentiation responses of embryonic stem cells. Genes Dev. 2009, 23, 1870–1875.
- Das, S.; Gordián-Vélez, W.J.; Ledebur, H.C.; Mourkioti, F.; Rompolas, P.; Chen, H.I.; Serruya, M.D.; Cullen, D.K. Innervation: The missing link for biofabricated tissues and organs. NPJ Regen. Med. 2020, 5, 11.
- Zhang, Y.S.; Aleman, J.; Shin, S.R.; Kilic, T.; Kim, D.; Mousavi Shaegh, S.A.; Massa, S.; Riahi, R.; Chae, S.; Hu, N.; et al. Multisensor-integrated organs-on-chips platform for automated and continual in situ monitoring of organoid behaviors. Proc. Natl. Acad. Sci. USA 2017, 114, E2293–E2302.
- Wörsdörfer, P.; Takashi, I.; Asahina, I.; Sumita, Y.; Ergün, S. Do not keep it simple: Recent advances in the generation of complex organoids. J. Neural Transm. 2020, 127, 1569–1577.
- Zhang, C.; Zhao, Z.; Abdul Rahim, N.A.; van Noort, D.; Yu, H. Towards a human-on-chip: Culturing multiple cell types on a chip with compartmentalized microenvironments. Lab. Chip 2009, 9, 3185–3192.
- Cavo, M.; Fato, M.; Peñuela, L.; Beltrame, F.; Raiteri, R.; Scaglione, S. Microenvironment complexity and matrix stiffness regulate breast cancer cell activity in a 3D in vitro model. Sci. Rep. 2016, 6, 35367.
- Giobbe, G.G.; Crowley, C.; Luni, C.; Campinoti, S.; Khedr, M.; Kretzschmar, K.; De Santis, M.M.; Zambaiti, E.; Michielin, F.; Meran, L.; et al. Extracellular matrix hydrogel derived from decellularized tissues enables endodermal organoid culture. Nat. Commun. 2019, 10, 5658.
- Ng, S.; Tan, W.J.; Pek, M.M.X.; Tan, M.-H.; Kurisawa, M. Mechanically and chemically defined hydrogel matrices for patient-derived colorectal tumor organoid culture. Biomaterials 2019, 219, 119400.
- Bhatia, S.N.; Ingber, D.E. Microfluidic organs-on-chips. Nat. Biotechnol. 2014, 32, 760–772.
- Low, L.A.; Mummery, C.; Berridge, B.R.; Austin, C.P.; Tagle, D.A. Organs-on-chips: Into the next decade. Nat. Rev. Drug Discov. 2020, 20, 345–361.
- Pagella, P.; Neto, E.; Jiménez-Rojo, L.; Lamghari, M.; Mitsiadis, T.A. Microfluidics co-culture systems for studying tooth innervation. Front. Physiol. 2014, 5, 326.
- Pagella, P.; Mitsiadis, T.A. Analysis of Tooth Innervation in Microfluidic Coculture Devices. Methods Mol. Biol. Clifton NJ 2020, 2155, 99–106.
- Ye, W.; Luo, C.; Li, C.; Huang, J.; Liu, F. Organoids to study immune functions, immunological diseases and immunotherapy. Cancer Lett. 2020, 477, 31–40.
- Votanopoulos, K.I.; Forsythe, S.; Sivakumar, H.; Mazzocchi, A.; Aleman, J.; Miller, L.; Levine, E.; Triozzi, P.; Skardal, A. Model of Patient-Specific Immune-Enhanced Organoids for Immunotherapy Screening: Feasibility Study. Ann. Surg. Oncol. 2020, 27, 1956–1967.
- Yuki, K.; Cheng, N.; Nakano, M.; Kuo, C.J. Organoid Models of Tumor Immunology. Trends Immunol. 2020, 41, 652–664.
- Radtke, F.; Fasnacht, N.; Macdonald, H.R. Notch signaling in the immune system. Immunity 2010, 32, 14–27.
- Polini, A.; del Mercato, L.L.; Barra, A.; Zhang, Y.S.; Calabi, F.; Gigli, G. Towards the development of human immune-system-on-a-chip platforms. Drug Discov. Today 2019, 24, 517–525.
- Ando, K.; Kanazawa, S.; Tetsuka, T.; Ohta, S.; Jiang, X.; Tada, T.; Kobayashi, M.; Matsui, N.; Okamoto, T. Induction of Notch signaling by tumor necrosis factor in rheumatoid synovial fibroblasts. Oncogene 2003, 22, 7796–7803.
- Palaga, T.; Buranaruk, C.; Rengpipat, S.; Fauq, A.H.; Golde, T.E.; Kaufmann, S.H.E.; Osborne, B.A. Notch signaling is activated by TLR stimulation and regulates macrophage functions. Eur. J. Immunol. 2008, 38, 174–183.
- Klinakis, A.; Lobry, C.; Abdel-Wahab, O.; Oh, P.; Haeno, H.; Buonamici, S.; van De Walle, I.; Cathelin, S.; Trimarchi, T.; Araldi, E.; et al. A novel tumour-suppressor function for the Notch pathway in myeloid leukaemia. Nature 2011, 473, 230–233.
- Franklin, R.A.; Liao, W.; Sarkar, A.; Kim, M.V.; Bivona, M.R.; Liu, K.; Pamer, E.G.; Li, M.O. The cellular and molecular origin of tumor-associated macrophages. Science 2014, 344, 921–925.
- Jimenez-Rojo, L.; Pagella, P.; Harada, H.; Mitsiadis, T. Dental Epithelial Stem Cells as a Source for Mammary Gland Regeneration and Milk Producing Cells In Vivo. Cells 2019, 8, 1302.
- Han, J.; Menicanin, D.; Gronthos, S.; Bartold, P.M. Stem cells, tissue engineering and periodontal regeneration. Aust. Dent. J. 2014, 59 (Suppl. S1), 117–130.
- Charbord, P. Bone marrow mesenchymal stem cells: Historical overview and concepts. Hum. Gene Ther. 2010, 21, 1045–1056.
- Mitsiadis, T.A.; Orsini, G.; Jimenez-Rojo, L. Stem cell-based approaches in dentistry. Eur. Cell. Mater. 2015, 30, 248–257.
- Orsini, G.; Pagella, P.; Mitsiadis, T.A. Modern Trends in Dental Medicine: An Update for Internists. Am. J. Med. 2018, 131, 1425–1430.
- Trubiani, O.; Marconi, G.D.; Pierdomenico, S.D.; Piattelli, A.; Diomede, F.; Pizzicannella, J. Human Oral Stem Cells, Biomaterials and Extracellular Vesicles: A Promising Tool in Bone Tissue Repair. Int. J. Mol. Sci. 2019, 20, 4987.
- Diomede, F.; Marconi, G.D.; Fonticoli, L.; Pizzicanella, J.; Merciaro, I.; Bramanti, P.; Mazzon, E.; Trubiani, O. Functional Relationship between Osteogenesis and Angiogenesis in Tissue Regeneration. Int. J. Mol. Sci. 2020, 21, 3242.
- Park, Y.-J.; Cha, S.; Park, Y.-S. Regenerative Applications Using Tooth Derived Stem Cells in Other Than Tooth Regeneration: A Literature Review. Stem Cells Int. 2016, 2016, 9305986.
- Xiao, L.; Kumazawa, Y.; Okamura, H. Cell death, cavitation and spontaneous multi-differentiation of dental pulp stem cells-derived spheroids in vitro: A journey to survival and organogenesis. Biol. Cell 2014, 106, 405–419.
- Moritani, Y.; Usui, M.; Sano, K.; Nakazawa, K.; Hanatani, T.; Nakatomi, M.; Iwata, T.; Sato, T.; Ariyoshi, W.; Nishihara, T.; et al. Spheroid culture enhances osteogenic potential of periodontal ligament mesenchymal stem cells. J. Periodontal Res. 2018, 53, 870–882.
- Kim, H.-J.; Sung, I.-Y.; Cho, Y.-C.; Kang, M.-S.; Rho, G.-J.; Byun, J.-H.; Park, W.-U.; Son, M.-G.; Park, B.-W.; Lee, H.-J.; et al. Three-Dimensional Spheroid Formation of Cryopreserved Human Dental Follicle-Derived Stem Cells Enhances Pluripotency and Osteogenic Induction Properties. Tissue Eng. Regen. Med. 2019, 16, 513–523.
- Ryu, N.-E.; Lee, S.-H.; Park, H. Spheroid Culture System Methods and Applications for Mesenchymal Stem Cells. Cells 2019, 8, 1620.
- Sugimoto, S.; Sato, T. Establishment of 3D Intestinal Organoid Cultures from Intestinal Stem Cells. Methods Mol. Biol. 2017, 1612, 97–105.
- Jeon, S.; Lee, H.-S.; Lee, G.-Y.; Park, G.; Kim, T.-M.; Shin, J.; Lee, C.; Oh, I.-H. Shift of EMT gradient in 3D spheroid MSCs for activation of mesenchymal niche function. Sci. Rep. 2017, 7, 6859.
- Lukomska, B.; Stanaszek, L.; Zuba-Surma, E.; Legosz, P.; Sarzynska, S.; Drela, K. Challenges and Controversies in Human Mesenchymal Stem Cell Therapy. Stem Cells Int. 2019, 2019, 9628536.
- Petrenko, Y.; Syková, E.; Kubinová, Š. The therapeutic potential of three-dimensional multipotent mesenchymal stromal cell spheroids. Stem Cell Res. Ther. 2017, 8, 94.
- Ferraro, F.; Celso, C.L.; Scadden, D. Adult stem cells and their niches. Adv. Exp. Med. Biol. 2010, 695, 155–168.
- Bejoy, J.; Bijonowski, B.; Marzano, M.; Jeske, R.; Ma, T.; Li, Y. Wnt-Notch Signaling Interactions during Neural and Astroglial Patterning of Human Stem Cells. Tissue Eng. Part A 2020, 26, 419–431.
- Iyer, K.; Chen, Z.; Ganapa, T.; Wu, B.M.; Tawil, B.; Linsley, C.S. Keratinocyte Migration in a Three-Dimensional In Vitro Wound Healing Model Co-Cultured with Fibroblasts. Tissue Eng. Regen. Med. 2018, 15, 721–733.
- Dosh, R.H.; Jordan-Mahy, N.; Sammon, C.; Le Maitre, C.L. Long-term in vitro 3D hydrogel co-culture model of inflammatory bowel disease. Sci. Rep. 2019, 9, 1812.
- Sun, L.; Wang, X.; Kaplan, D.L. A 3D cartilage—Inflammatory cell culture system for the modeling of human osteoarthritis. Biomaterials 2011, 32, 5581–5589.
- Damerau, A.; Gaber, T. Modeling Rheumatoid Arthritis In Vitro: From Experimental Feasibility to Physiological Proximity. Int. J. Mol. Sci. 2020, 21, 7916.
- Lv, D.; Hu, Z.; Lu, L.; Lu, H.; Xu, X. Three-dimensional cell culture: A powerful tool in tumor research and drug discovery. Oncol. Lett. 2017, 14, 6999–7010.
- Zhang, C.; Yang, Z.; Dong, D.-L.; Jang, T.-S.; Knowles, J.C.; Kim, H.-W.; Jin, G.-Z.; Xuan, Y. 3D culture technologies of cancer stem cells: Promising ex vivo tumor models. J. Tissue Eng. 2020, 11, 2041731420933407.
- Sorg, H.; Tilkorn, D.J.; Hager, S.; Hauser, J.; Mirastschijski, U. Skin Wound Healing: An Update on the Current Knowledge and Concepts. Eur. Surg. Res. 2017, 58, 81–94.
- Park, I.-S.; Chung, P.-S.; Ahn, J.C. Enhancement of Ischemic Wound Healing by Spheroid Grafting of Human Adipose-Derived Stem Cells Treated with Low-Level Light Irradiation. PLoS ONE 2015, 10, e0122776.
- Kim, S.; Kim, E.M.; Yamamoto, M.; Park, H.; Shin, H. Engineering Multi-Cellular Spheroids for Tissue Engineering and Regenerative Medicine. Adv. Healthc. Mater. 2020, 9, 2000608.
- Chigurupati, S.; Arumugam, T.V.; Son, T.G.; Lathia, J.D.; Jameel, S.; Mughal, M.R.; Tang, S.-C.; Jo, D.-G.; Camandola, S.; Giunta, M.; et al. Involvement of Notch Signaling in Wound Healing. PLoS ONE 2007, 2, e1167.
- Okuyama, R.; Nguyen, B.-C.; Talora, C.; Ogawa, E.; di Vignano, A.T.; Lioumi, M.; Chiorino, G.; Tagami, H.; Woo, M.; Dotto, G.P. High Commitment of Embryonic Keratinocytes to Terminal Differentiation through a Notch1-caspase 3 Regulatory Mechanism. Dev. Cell 2004, 6, 551–562.
- Del Campo, J.A.; Gallego, P.; Grande, L. Role of inflammatory response in liver diseases: Therapeutic strategies. World J. Hepatol. 2018, 10, 1–7.
- Tilg, H.; Diehl, A.M. Cytokines in Alcoholic and Nonalcoholic Steatohepatitis. N. Engl. J. Med. 2000, 343, 1467–1476.
- Ruoß, M.; Vosough, M.; Königsrainer, A.; Nadalin, S.; Wagner, S.; Sajadian, S.; Huber, D.; Heydari, Z.; Ehnert, S.; Hengstler, J.G.; et al. Towards improved hepatocyte cultures: Progress and limitations. Food Chem. Toxicol. 2020, 138, 111188.
- Nuciforo, S.; Heim, M.H. Organoids to model liver disease. JHEP Rep. 2021, 3, 100198.
- Huch, M.; Dorrell, C.; Boj, S.F.; van Es, J.H.; Li, V.S.W.; van de Wetering, M.; Sato, T.; Hamer, K.; Sasaki, N.; Finegold, M.J.; et al. In vitro expansion of single Lgr5+ liver stem cells induced by Wnt-driven regeneration. Nature 2013, 494, 247–250.
- Hu, H.; Gehart, H.; Artegiani, B.; LÖpez-Iglesias, C.; Dekkers, F.; Basak, O.; van Es, J.; Chuva de Sousa Lopes, S.M.; Begthel, H.; Korving, J.; et al. Long-Term Expansion of Functional Mouse and Human Hepatocytes as 3D Organoids. Cell 2018, 175, 1591–1606.e19.
- Kozyra, M.; Johansson, I.; Nordling, Å.; Ullah, S.; Lauschke, V.M.; Ingelman-Sundberg, M. Human hepatic 3D spheroids as a model for steatosis and insulin resistance. Sci. Rep. 2018, 8, 14297.
- Li, L.; Krantz, I.D.; Deng, Y.; Genin, A.; Banta, A.B.; Collins, C.C.; Qi, M.; Trask, B.J.; Kuo, W.L.; Cochran, J.; et al. Alagille syndrome is caused by mutations in human Jagged1, which encodes a ligand for Notch1. Nat. Genet. 1997, 16, 243–251.
- Oda, T.; Elkahloun, A.G.; Pike, B.L.; Okajima, K.; Krantz, I.D.; Genin, A.; Piccoli, D.A.; Meltzer, P.S.; Spinner, N.B.; Collins, F.S.; et al. Mutations in the human Jagged1 gene are responsible for Alagille syndrome. Nat. Genet. 1997, 16, 235–242.
- Guan, Y.; Xu, D.; Garfin, P.M.; Ehmer, U.; Hurwitz, M.; Enns, G.; Michie, S.; Wu, M.; Zheng, M.; Nishimura, T.; et al. Human hepatic organoids for the analysis of human genetic diseases. JCI Insight 2017, 2, e94954.
- Spill, F.; Reynolds, D.S.; Kamm, R.D.; Zaman, M.H. Impact of the physical microenvironment on tumor progression and metastasis. Curr. Opin. Biotechnol. 2016, 40, 41–48.
- Tevis, K.M.; Colson, Y.L.; Grinstaff, M.W. Embedded Spheroids as Models of the Cancer Microenvironment. Adv. Biosyst. 2017, 1, 1700083.
- Tamaki, M.; McDonald, W.; Amberger, V.R.; Moore, E.; Del Maestro, R.F. Implantation of C6 astrocytoma spheroid into collagen type I gels: Invasive, proliferative, and enzymatic characterizations. J. Neurosurg. 1997, 87, 602–609.
- Porcheri, C.; Meisel, C.T.; Mitsiadis, T.A. Molecular and Cellular Modelling of Salivary Gland Tumors Open New Landscapes in Diagnosis and Treatment. Cancers 2020, 12, E3107.
- Zanoni, M.; Cortesi, M.; Zamagni, A.; Arienti, C.; Pignatta, S.; Tesei, A. Modeling neoplastic disease with spheroids and organoids. J. Hematol. Oncol. 2020, 13, 97.
- Weiswald, L.-B.; Bellet, D.; Dangles-Marie, V. Spherical cancer models in tumor biology. Neoplasia 2015, 17, 1–15.
- Hellström, M.; Phng, L.-K.; Hofmann, J.J.; Wallgard, E.; Coultas, L.; Lindblom, P.; Alva, J.; Nilsson, A.-K.; Karlsson, L.; Gaiano, N.; et al. Dll4 signalling through Notch1 regulates formation of tip cells during angiogenesis. Nature 2007, 445, 776–780.
- Siebel, C.; Lendahl, U. Notch Signaling in Development, Tissue Homeostasis, and Disease. Physiol. Rev. 2017, 97, 1235–1294.
- Lobry, C.; Oh, P.; Aifantis, I. Oncogenic and tumor suppressor functions of Notch in cancer: It’s NOTCH what you think. J. Exp. Med. 2011, 208, 1931–1935.
- Ayla, S.; Bilir, A.; Soner, B.C.; Yilmaz-Dilsiz, O.; Ergüven, M.; Oktem, G. Notch signaling-related therapeutic strategies with novel drugs in neuroblastoma spheroids. J. Pediatr. Hematol. Oncol. 2014, 36, 37–44.
- Xiao, Y.; Ye, Y.; Zou, X.; Jones, S.; Yearsley, K.; Shetuni, B.; Tellez, J.; Barsky, S.H. The lymphovascular embolus of inflammatory breast cancer exhibits a Notch 3 addiction. Oncogene 2011, 30, 287–300.
- Rebay, I.; Fleming, R.J.; Fehon, R.G.; Cherbas, L.; Cherbas, P.; Artavanis-Tsakonas, S. Specific EGF repeats of Notch mediate interactions with Delta and serrate: Implications for notch as a multifunctional receptor. Cell 1991, 67, 687–699.
- Rebay, I.; Fehon, R.G.; Artavanis-Tsakonas, S. Specific truncations of Drosophila Notch define dominant activated and dominant negative forms of the receptor. Cell 1993, 74, 319–329.
- Miyaki, M.; Yamaguchi, T.; Iijima, T.; Takahashi, K.; Matsumoto, H.; Mori, T. Somatic Mutations of the CDC4 (FBXW7) Gene in Hereditary Colorectal Tumors. Oncology 2009, 76, 430–434.
- Babaei-Jadidi, R.; Li, N.; Saadeddin, A.; Spencer-Dene, B.; Jandke, A.; Muhammad, B.; Ibrahim, E.E.; Muraleedharan, R.; Abuzinadah, M.; Davis, H.; et al. FBXW7 influences murine intestinal homeostasis and cancer, targeting Notch, Jun, and DEK for degradation. J. Exp. Med. 2011, 208, 295–312.
- Camps, J.; Pitt, J.J.; Emons, G.; Hummon, A.B.; Case, C.M.; Grade, M.; Jones, T.L.; Nguyen, Q.T.; Ghadimi, B.M.; Beissbarth, T.; et al. Genetic Amplification of the NOTCH Modulator LNX2 Upregulates the WNT/β-Catenin Pathway in Colorectal Cancer. Cancer Res. 2013, 73, 2003–2013.
- Sancho, R.; Jandke, A.; Davis, H.; Diefenbacher, M.E.; Tomlinson, I.; Behrens, A. F-box and WD Repeat Domain-Containing 7 Regulates Intestinal Cell Lineage Commitment and Is a Haploinsufficient Tumor Suppressor. Gastroenterology 2010, 139, 929–941.
- Zhu, J.; Wang, J.; Shi, Z.; Franklin, J.L.; Deane, N.G.; Coffey, R.J.; Beauchamp, R.D.; Zhang, B. Deciphering Genomic Alterations in Colorectal Cancer through Transcriptional Subtype-Based Network Analysis. PLoS ONE 2013, 8, e79282.
- Lee, S.H.; Choi, N.; Sung, J.H. Pharmacokinetic and pharmacodynamic insights from microfluidic intestine-on-a-chip models. Expert Opin. Drug Metab. Toxicol. 2019, 15, 1005–1019.
- Candini, O.; Grisendi, G.; Foppiani, E.M.; Brogli, M.; Aramini, B.; Masciale, V.; Spano, C.; Petrachi, T.; Veronesi, E.; Conte, P.; et al. Author Correction: A Novel 3D In Vitro Platform for Pre-Clinical Investigations in Drug Testing, Gene Therapy, and Immuno-oncology. Sci. Rep. 2020, 10, 1845.
- Fatehullah, A.; Tan, S.H.; Barker, N. Organoids as an in vitro model of human development and disease. Nat. Cell Biol. 2016, 18, 246–254.
- Sung, J.H. Multi-organ-on-a-chip for pharmacokinetics and toxicokinetic study of drugs. Expert Opin. Drug Metab. Toxicol. 2021, 17, 969–986.
- Langhans, S.A. Three-Dimensional in Vitro Cell Culture Models in Drug Discovery and Drug Repositioning. Front. Pharmacol. 2018, 9, 6.
- Polini, A.; Prodanov, L.; Bhise, N.S.; Manoharan, V.; Dokmeci, M.R.; Khademhosseini, A. Organs-on-a-chip: A new tool for drug discovery. Expert Opin. Drug Discov. 2014, 9, 335–352.
- Gupta, N.; Liu, J.R.; Patel, B.; Solomon, D.E.; Vaidya, B.; Gupta, V. Microfluidics-based 3D cell culture models: Utility in novel drug discovery and delivery research. Bioeng. Transl. Med. 2016, 1, 63–81.
- Esch, E.W.; Bahinski, A.; Huh, D. Organs-on-chips at the frontiers of drug discovery. Nat. Rev. Drug Discov. 2015, 14, 248–260.
- Van Zundert, I.; Fortuni, B.; Rocha, S. From 2D to 3D Cancer Cell Models-The Enigmas of Drug Delivery Research. Nanomaterials 2020, 10, 2236.
- Lim, J.; Ching, H.; Yoon, J.-K.; Jeon, N.L.; Kim, Y. Microvascularized tumor organoids-on-chips: Advancing preclinical drug screening with pathophysiological relevance. Nano Converg. 2021, 8, 12.
- Prantil-Baun, R.; Novak, R.; Das, D.; Somayaji, M.R.; Przekwas, A.; Ingber, D.E. Physiologically Based Pharmacokinetic and Pharmacodynamic Analysis Enabled by Microfluidically Linked Organs-on-Chips. Annu. Rev. Pharmacol. Toxicol. 2018, 58, 37–64.
- Kimura, H.; Sakai, Y.; Fujii, T. Organ/body-on-a-chip based on microfluidic technology for drug discovery. Drug Metab. Pharmacokinet. 2018, 33, 43–48.
- Golde, T.E.; Koo, E.H.; Felsenstein, K.M.; Osborne, B.A.; Miele, L. γ-Secretase inhibitors and modulators. Biochim. Biophys. Acta BBA—Biomembr. 2013, 1828, 2898–2907.
- Andersson, E.R.; Lendahl, U. Therapeutic modulation of Notch signaling—Are we there yet? Nat. Rev. Drug Discov. 2014, 13, 357–378.
- Real, P.J.; Tosello, V.; Palomero, T.; Castillo, M.; Hernando, E.; de Stanchina, E.; Sulis, M.L.; Barnes, K.; Sawai, C.; Homminga, I.; et al. γ-secretase inhibitors reverse glucocorticoid resistance in T cell acute lymphoblastic leukemia. Nat. Med. 2009, 15, 50–58.
More
Information
Subjects:
Cell & Tissue Engineering
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.1K
Entry Collection:
Biopharmaceuticals Technology
Revisions:
3 times
(View History)
Update Date:
02 Dec 2021
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





