Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Giuseppe Mannino | + 2396 word(s) | 2396 | 2021-11-18 04:53:11 | | | |
| 2 | Jessie Wu | Meta information modification | 2396 | 2021-11-30 02:30:22 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Mannino, G. Chemistry and Biosynthesis of Melatonin. Encyclopedia. Available online: https://encyclopedia.pub/entry/16502 (accessed on 25 May 2026).
Mannino G. Chemistry and Biosynthesis of Melatonin. Encyclopedia. Available at: https://encyclopedia.pub/entry/16502. Accessed May 25, 2026.
Mannino, Giuseppe. "Chemistry and Biosynthesis of Melatonin" Encyclopedia, https://encyclopedia.pub/entry/16502 (accessed May 25, 2026).
Mannino, G. (2021, November 29). Chemistry and Biosynthesis of Melatonin. In Encyclopedia. https://encyclopedia.pub/entry/16502
Mannino, Giuseppe. "Chemistry and Biosynthesis of Melatonin." Encyclopedia. Web. 29 November, 2021.
Copy Citation
Melatonin is a ubiquitous indolamine, largely investigated for its key role in the regulation of several physiological processes in both animals and plants. In the last century, it was reported that this molecule may be produced in high concentrations by several species belonging to the plant kingdom and stored in specialized tissues.
Melatonin
Chemistry
biosynthesis
indolamine
1. Chemistry of Melatonin
From a physio-chemical point of view, pure melatonin resembles an off-white powder, having 232.28 g/mol as molecular weight and a density of 1.175 g/cm3. The melting point ranges between 116.5 °C and 118 °C; the boiling point is 512.8 °C [1]. From a chemical point of view, melatonin is identified by the chemical formula C13H16N2O2. The indole chemical scaffold is functionalized with a 3-amide group and a 5-alkoxygroup (Figure 1). Moreover, since it is originated starting from a molecule of tryptophan, is classified as an indolamine compound [2]. This particular chemical structure confers great stability by high resonance mesomerism.
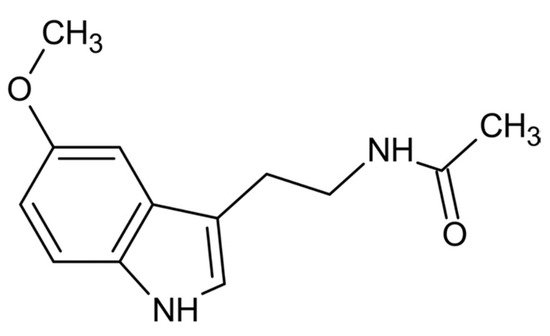
Figure 1. Chemical structure of melatonin.
Moreover, the 3-amide group and 5-alkoxygroup are also the main responsible of the amphiphilicity of this molecule. This property makes melatonin able to cross biological membranes and enter any cellular and subcellular compartments [2][3], allowing not only its easy distribution but also a high protection against oxidative stress in various cell compartments [2][4]. The antioxidant protection of melatonin is correlated both to its own redox active properties and to metabolites originated during its metabolism. Indeed, a series of new compounds having noteworthy antioxidant properties may be further generated by melatonin oxidation in a set of reactions known as melatonin antioxidant cascade [5][6]. Among these metabolites, cyclic 3-hydroxymelatonin (C3-OHM), N1-acetyl-N2-formyl-5-methoxykynuramine (AFMK), N1-acetyl-5-methoxykynuramine (AMK), 6-hydroxymelatonin (6-OHM), 2-hydroxymelatonin (2-OHM) are the most known (Figure 2).

Figure 2. Melatonin metabolism and its related metabolites. ONOO− = peroxynitrite; ROS = reactive oxygen species; RNS = reactive nitrogen species; O2−: superoxide anion; OH−: hydroxyl radical; AMFK: N1-acetyl-N2-formyl-5-methoxykynuramine; AMK: N1-acetyl-5-methoxykynuramine; AMMC: 3-acetamidomethyl-6-methoxycinnolinone; AMNK: N1-acetyl-5-methoxy-3 nitrokynura-mine.
1.1. N1-Acetyl-N2-Formyl-5-Methoxykynuramine (AFMK)
Kynuramine compounds, such as AFMK and its de-formylated form (AMK), are molecules produced during tryptamine degradation. The redox activity and antioxidative properties of AFMK have been evaluated in several experimental models. Unlike antioxidants, such as vitamin C and vitamin E, AMFK can donate more than one electron [7]. In particular, Rosen and colleagues showed that AFMK can donate four electrons leading to the production of indolinone derivatives, such as Z-, E- isomers of N-(1-formyl-5-methoxy-3oxo-2,3-dihydro-1H-indol-2-ylidenemethyl)-acetamide and N-(1-formyl-2-hydroxy-5-methoxy-3-oxo-2,3-dihydro-1H-indol-2-ylmethyl)-acetamide [8]. However, AFMK was reported to be a less effective free radical scavenger than AMK and melatonin [9][10][11][12]. The antioxidant properties of AFMK were demonstrated also in biological models. In particular, Tan and colleagues showed that the addition of AFMK to calf thymus DNA in presence of a mixture of prooxidant agents strongly reduced in a dose-dependent way the levels of 8-OH-dG (an indicator of DNA damage) [7]. Moreover, in rat liver homogenates incubated with H2O2 and Fe2+, 100 µM AFMK inhibited lipid peroxidation (LPO) and improved cell viability, although Fe2+ chelation was not observed [7].
1.2. N1-Acetyl-5-Methoxykynuramine (AMK)
AFMK can be both enzymatically and non-enzymatically de-formylated resulting in the formation of AMK [13][14]. This compound showed a higher efficiency for scavenging ROS and preventing protein oxidation with respect to AFMK [15]. Radical scavenging action of AMK leads to the production of AMK oligomers, such as 3-acetamidomethyl-6-methoxycinnolinone and N1-acetyl-5-methoxy-3 nitrokynuramine. This property of AMK strongly depends on the environmental conditions [12]. Indeed, it has been observed that in aqueous solution, AMK is a radical scavenger stronger than melatonin, although it is a good scavenger also in nonpolar environment. In particular, AMK is a better OH•− and NO scavenger than both melatonin [16] and AFMK [17].
1.3. 3-Hydroxymelatonin (C3-OHM)
Melatonin oxidation by reactive oxygen species (ROS) and reactive nitrogen species (RNS) scavenging may produce also C3-OHM. Experimental data showed an antioxidant protection by radicals. In particular, as with AFMK, also C3-OHM prevented DNA oxidation induced by Fenton reaction [18]. The presence of C3-OHM was always coupled to AFMK formation, both in vitro and in vivo experimentations [19]. The ratio between oxidants and melatonin affected the amount of melatonin oxidation products. In particular, higher were the ROS levels, more AFMK was produced [20]. Indeed, in this condition, C3-OHM can also be oxidised to AFMK.
1.4. 6-Hydroxymelatonin (6-OHM)
6-Hydroxymelatonin (6-OHM), for the first time discovered in animal urine in the form of 6-hydroxymelatonin sulfate, is one of the major melatonin catabolites in animals. Experimental data showed that 6-OHM prevented lipid peroxidation [21] and DNA damage induced by environmental pollutants, chromium [22], and OH•− generated by Fenton reaction [23]. Duan et al. also showed neuronal protection by 6-OHM in a model of ischemia/reperfusion-mediated injury. In this model, the anti-apoptotic action involved the inhibition of cytochrome C, inhibition of caspase 3 activity, and stabilization of the mitochondrial membrane potential [24]. Although the known protective effect of 6-OHM, a slight prooxidant activity was also shown. In particular, it was reported that 6-OHM caused oxidative DNA damage with double-strand breaks via redox cycling [25].
1.5. 2-Hydroxymelatonin (2-OHM)
Melatonin oxidation also leads to the production of 2-OHM, especially after scavenging of HClO [26], oxoferryl haemoglobin [27] and OH•− [28]. Conversely to 3-OHM, 2-OHM is one the prevalent products of the hydroxylation of melatonin in plants. 2-OHM production is coupled to the formation of the keto tautomer melatonin 2-indolinone. In addition, in cytochrome C in vitro models the oxidation 2-OHM into AFMK was observed [29].
2. Biosynthesis of Melatonin
2.1. Biosynthetic Route in Plants
It has been shown that the cellular compartments with the highest melatonin levels in plants are mitochondria and chloroplasts [30]. This observation, together with the demonstrated localization of serotonin N-acetyltransferase (SNAT), one of the rate-limiting enzymes involved in melatonin biosynthesis, in chloroplasts [31][32] and in mitochondria [33], leads to hypothesize that these organelles are the major sites involved in the biosynthesis of this indolamine. The genes encoding for all the enzymes catalysing the whole melatonin biosynthetic pathway in plants have been discovered in several plant species, with the exception of one putative gene encoding for a tryptophan hydroxylase (TPH), which catalyses the conversion of tryptophan into 5-hydroxytryptophan [34]. In particular, this enzyme, already known in vertebrates, was only recently proposed in plants.
Melatonin biosynthesis begins with the amino acid tryptophan, a compound that plants are able to synthesize de novo via the shikimate pathway (Figure 3). This pathway consists of seven different steps that allow to the biosynthesis of all aromatic amino acids in plants, including tryptophan [35]. Briefly, 3-Deoxy-D-arabinoheptulosonate 7-phosphate (DAHP) synthase (EC 2.5.1.54) transforms phosphoenol pyruvate (PEP) and erythrose-4-phosphate in DAHP, that is then cyclized into 3-dehydroquinate (DHQ) by the action of the DHQ synthase (EC 4.2.3.4). Finally, shikimate is synthetized through the dehydration and dehydrogenation catalyzed by DHQ dehydratase (EC 4.2.1.10) and shikimate dehydrogenase (EC 1.1.1.25). Therefore, shikimate is phosphorylated by the shikimate kinase (EC 2.7.1.71) and converted in 5-enolpyruvylshikimate-3-phosphate (EPSP) by the EPSP synthase (EC 2.5.1.19). Finally, chorismate is formed through the activity of chorismate synthase (EC 4.2.3.5) that converts EPSP in chorismate, the essential intermediate in tryptophan biosynthesis (Figure 3). Chorismate is converted in anthranilate via anthranilate synthase (EC 4.1.3.27) that is consequently condensed with phosphoribosylpyrophosphate (PRPP), generating phosphoribosyl anthranilate (PRA). The ribose ring added in this last reaction is then opened by PRA isomerase (PRAI; EC 5.3.1.24), subjected to reductive decarboxylation in order to form indole-3-glycerol phosphate that is spontaneously converted into the indole scaffold. Finally, tryptophan is produced via the reaction of the indole with serine through the action of tryptophan synthase (TPS; EC 4.2.1.20) (Figure 3).

Figure 3. Biosynthetic pathway involved in the synthesis of tryptophan, the key compound for the formation of melatonin in plants. PEP: 2-phosphoenolpyruvate; DAHP: 3-deoxy-D-arabinoheptulosonate 7-phosphate; DHQ: 3-dehydroquinic acid; DHS: 3-dehydroshikimate; PEP: 2-phosphoenolpyruvate; EPSP: 5-enolpyruvylshikimate-3-phosphate; PRA: Phosporibosyl antranilate; PRAI: PRA isomerase; PRPP: phosphoribosylpyrophosphate; IGP: indole-3-glycerol phosphate; EC: enzyme commission number).
At least six enzymes are known to be involved in melatonin biosynthesis from tryptophan, indicating that multiple biosynthetic pathways may be present in this process. The six enzymes known to be involved in the synthesis of melatonin are: (i) L-tryptophan decarboxylase (TDC), (ii) tryptamine 5-hydroxylase (T5H), (iii) serotonin N-acetyltransferase (SNAT), (iv) acetylserotonin O-methyltransferase (ASMT), (v) caffeic acid 3-O-methyltransferase (COMT), and (vi) a putative tryptophan hydroxylase (TPH) not yet identified.
The first step of the melatonin biosynthetic process in plants is related to the production of serotonin from tryptophan. Two different pathways may be involved (Figure 4). The first way begins with the decarboxylation of tryptophan into tryptamine by TPH, and then tryptamine is hydroxylated to serotonin by TDC. On the other hand, another possibility first involves the hydroxylation of tryptophan into 5-hydroxytryptophan by TPH, and then the decarboxylation of 5-hydrotryptophan into serotonin by TDC. These routes are both possible, because TDC shows a good affinity for both tryptophan and 5-hydroxytriptophan in vitro [36]. However, it has been demonstrated that in plants is more frequent the decarboxylation than the hydroxylation as first step [36].
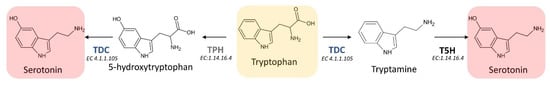
Figure 4. First two reactions of the melatonin biosynthetic pathway leading to the formation of the essential intermediate serotonin. TDC: L-tryptophan decarboxylase; TPH: tryptophan hydroxylase; T5H: tryptamine 5-hydroxylase; EC: enzyme commission number).
Melatonin synthesis from serotonin is a two-step reaction involving three different enzymes (SNATs, ASMTs, and COMT) that may have various isoforms. The first enzyme catalyzes an acetylation, whereas the other two enzymes are methyltransferases [36] (Figure 5). As the tree enzymes exhibit a substrate affinity for serotonin, N-acetylserotonin, and 5-methoxytryptamine, also in this case the order by which the different enzymes act can vary [37][38][39].
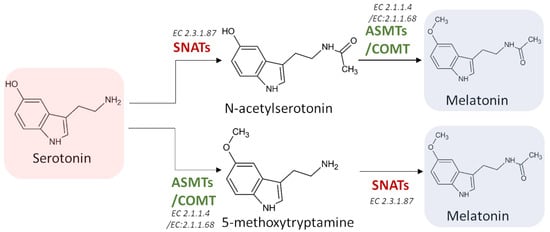
Figure 5. The last two potential reactions leading to the formation of melatonin. SNATs: serotonin N-acetyltransferase; ASMTs: acetylserotonin O-methyltransferase; COMT: caffeic acid 3-O-methyltransferase; EC: enzyme commission number.
The enzymes involved in melatonin biosynthesis from tryptophan have a different distribution in plant cells. TDC is localized in the cytoplasm [40], T5H in the endoplasmic reticulum [5], SNAT is expressed in chloroplasts [5], whereas ASMT and COMT are in the cytoplasm [41]. Among the four possible melatonin biosynthetic pathways reported in Figure 6, the first and the second pathways result in serotonin synthesis in the endoplasmic reticulum, whereas the third and fourth in cytoplasmic environment [36]. ASMTs/COMT are exclusively located in the cytoplasm and SNATs in the chloroplast, the final subcellular sites of melatonin synthesis and accumulation may vary. For example, in the cytoplasm serotonin is rapidly metabolized into phenylpropanoid amides, such as feruloylserotonin, by the serotonin N-hydroxycinnamoyl transferase (SHT) [42] and melatonin is rapidly converted into cyclic 3-hydroxymelatonin (3-OHM) by the melatonin 3-hydroxylase (M3H), whereas in chloroplasts melatonin can be metabolized into 2-hydroxymelatonin (2-OHM) by the melatonin 2-hydroxylase (M2H) [43][44].
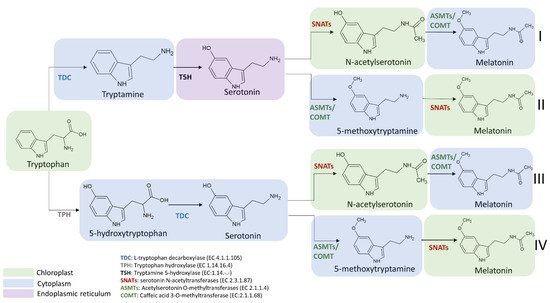
Figure 6. Subcellular localization of melatonin intermediates and enzymes involved in the transformation of tryptophan into melatonin in the different melatonin biosynthetic routes. TDC: L-tryptophan decarboxylase; TPH: tryptophan hydroxylase; T5H: tryptamine 5-hydroxylase; SNATs: serotonin N-acetyltransferase; ASMTs: acetylserotonin O-methyltransferase; COMT: caffeic acid 3-O-methyltransferase.
The selection of the pathway for melatonin biosynthesis depends on plant growth conditions. Indeed, under standard or stress conditions that do not cause a high accumulation of serotonin, the melatonin biosynthetic pathway proceeds from tryptophan via the intermediate tryptamine/serotonin/N-acetylserotonin up to melatonin [45] (Figure 6). In this pathway, serotonin levels are relatively low and this molecule is preferentially acetylated to N-acetylserotonin by SNAT due to its higher affinity (Km = 0.385 mmol/L) for serotonin in comparison to ASMT (Km = 1.035 mmol/L) and COMT (Km = 3.396 mmol/L). The produced N-acetylserotonin is rapidly O-methylated into melatonin by either ASMT or COMT with a 30-fold higher catalytic efficiency than SNAT, leading to low levels of N-acetylserotonin. Based on previously published data, it was observed that COMT exhibits higher catalytic efficiency than ASMT at 37 °C but in vivo experiments, and it was observed that the activity of COMT to methylate N-acetylserotonin into melatonin was markedly inhibited due to the fact that COMT preferred methylate other substrates such as caffeic acid and 5-hydroxyconiferaldehyde [46][47]. These phenomena resulted in the functional loss of COMT activity for melatonin synthesis and a dominant role of ASMT in methylating N-acetylserotonin into melatonin (Figure 6).
On the other hand, during plant senescence or under abiotic stress conditions, plants tend to accumulate large amounts of melatonin intermediates (such as tryptophan, tryptamine and serotonin) [48]. Consequently, the biosynthetic route preferably proceeds from tryptophan via the intermediates tryptamine/serotonin/5-MT up to melatonin [36], where serotonin is O-methylated into 5-MT by COMT, and then it is acylated by SNAT leading to the formation of melatonin (Figure 6). However, it was observed that a serotonin boost was not proportionally correlated to a significant increase in melatonin level. Indeed, in several experimental conditions, despite the content of tryptophan and serotonin was slightly enhanced during the senescence, an equal increment of melatonin level was not observed. For example, data from senescent detached rice leaves have shown a difference of more than threefold in metabolic capacity between serotonin and melatonin synthesis [49][50]. This huge difference between the two compounds could be explained by the relatively low catalytic efficiencies of COMT and SNAT in senescence compared to those under normal growth conditions. However, despite SNATs auto-inhibition by serotonin was not currently observed and other regulatory roles of serotonin on the melatonin synthetic pathway are still unknown, low levels of melatonin and relatively high levels of 5-MT are obtained compared to N-acetylserotonin [51][52].
Despite the melatonin biosynthetic pathway under normal conditions producing more melatonin that in senescence and serotonin boost conditions, the melatonin levels are not related anyway to the levels of tryptophan and serotonin present in the cells. Consequently, the limiting step could be attributed to the production of N-acetylserotonin by SNAT [52]. Indeed, N-acetylserotonin must first cross the chloroplast membrane into the cytoplasm where ASMT or COMT can now transform it into melatonin [39][52].
Another potential route based on the studies conducted on T5H-deficient and T5H-suppression rice plants seems to be related to 5-hydroxytryptophan-mediated serotonin synthesis. In the investigated plants, the serotonin levels were much lower compared to control plants, but melatonin levels were higher. This interesting result was incompatible with the previously described melatonin biosynthetic pathways [53][54]. In this case, it was supposed that 5-HT is produced from tryptophan by the action of a putative TPH and then converted into serotonin by TDC. Given the low levels of serotonin and the high levels of melatonin found in T5H-deficient plants, the 5-HT pathway does not result in a serotonin boost but plays a key role in inducing melatonin levels.
References
- Mannino, G.; Caradonna, F.; Cruciata, I.; Lauria, A.; Perrone, A.; Gentile, C. Melatonin reduces inflammatory response in human intestinal epithelial cells stimulated by interleukin-1β. J. Pineal Res. 2019, 67, e12598.
- Omer, R.A.; Koparir, P.; Ahmed, L.; Koparir, M. Computational and spectroscopy study of melatonin. Indian J. Chem. B 2021, 60, 732–741.
- Tan, D.; Reiter, R.; Manchester, L.; Yan, M.; El-Sawi, M.; Sainz, R.; Mayo, J.; Kohen, R.; Allegra, M.; Hardelan, R. Chemical and Physical Properties and Potential Mechanisms: Melatonin as a Broad Spectrum Antioxidant and Free Radical Scavenger. Curr. Top. Med. Chem. 2005, 2, 181–197.
- Simopoulos, A.P.; Tan, D.X.; Manchester, L.C.; Reiter, R.J. Purslane: A plant source of omega-3 fatty acids and melatonin. J. Pineal Res. 2005, 39, 331–332.
- Back, K. Melatonin metabolism, signaling and possible roles in plants. Plant J. 2021, 105, 376–391.
- Reina, M.; Martínez, A. A new free radical scavenging cascade involving melatonin and three of its metabolites (3OHM, AFMK and AMK). Comput. Theor. Chem. 2018, 1123, 111–118.
- Tan, D.X.; Manchester, L.C.; Burkhardt, S.; Sainz, R.M.; Mayo, J.C.; Kohen, R.; Shohami, E.; Huo, Y.S.; Hardeland, R.; Reiter, R.J. N1-acetyl-N2-formyl-5-methoxykynuramine, a biogenic amine and melatonin metabolite, functions as a potent antioxidant. FASEB J. 2001, 15, 2294–2296.
- Rosen, J.; Than, N.N.; Koch, D.; Poeggeler, B.; Laatsch, H.; Hardeland, R. Interactions of melatonin and its metabolites with the ABTS cation radical: Extension of the radical scavenger cascade and formation of a novel class of oxidation products, C2-substituted 3-indolinones. J. Pineal Res. 2006, 41, 374–381.
- Hardeland, R. Antioxidative protection by melatonin: Multiplicity of mechanisms from radical detoxification to radical avoidance. Endocrine 2005, 27, 119–130.
- Iwashita, H.; Matsumoto, Y.; Maruyama, Y.; Watanabe, K.; Chiba, A.; Hattori, A. The melatonin metabolite N1-acetyl-5-methoxykynuramine facilitates long-term object memory in young and aging mice. J. Pineal Res. 2021, 70, e12703.
- Silva, S.O.; Rodrigues, M.R.; Carvalho, S.R.Q.; Catalani, L.H.; Campa, A.; Ximenes, V.F. Oxidation of melatonin and its catabolites, N1-acetyl-N 2-formyl-5-methoxykynuramine and N1-acetyl-5-methoxykynuramine, by activated leukocytes. J. Pineal Res. 2004, 37, 171–175.
- Galano, A.; Tan, D.X.; Reiter, R.J. On the free radical scavenging activities of melatonin’s metabolites, AFMK and AMK. J. Pineal Res. 2013, 54, 245–257.
- Gitto, E.; Tan, D.-X.; Reiter, R.J.; Karbownik, M.; Manchester, L.C.; Cuzzocrea, S.; Fulia, F.; Barberi, I. Individual and synergistic antioxidative actions of melatonin: Studies with vitamin E, vitamin C, glutathione and desferrrioxamine (desferoxamine) in rat liver homogenates. J. Pharm. Pharmacol. 2010, 53, 1393–1401.
- Hardeland, R.; Tan, D.X.; Reiter, R.J. Kynuramines, metabolites of melatonin and other indoles: The resurrection of an almost forgotten class of biogenic amines. J. Pineal Res. 2009, 47, 109–126.
- Ressmeyer, A.R.; Mayo, J.C.; Zelosko, V.; Sáinz, R.M.; Tan, D.X.; Poeggeler, B.; Antolín, I.; Zsizsik, B.K.; Reiter, R.J.; Hardeland, R. Antioxidant properties of the melatonin metabolite N1-acetyl-5-methoxykynuramine (AMK): Scavenging of free radicals and prevention of protein destruction. Redox Rep. 2003, 8, 205–213.
- León, J.; Escames, G.; Rodríguez, M.I.; López, L.C.; Tapias, V.; Entrena, A.; Camacho, E.; Carrión, M.D.; Gallo, M.A.; Espinosa, A.; et al. Inhibition of neuronal nitric oxide synthase activity by N 1-acetyl-5-methoxykynuramine, a brain metabolite of melatonin. J. Neurochem. 2006, 98, 2023–2033.
- Hardeland, R.; Backhaus, C.; Fadavi, A.; Hess, M. N1-acetyl-5-methoxykynuramine contrasts with other tryptophan metabolites by a peculiar type of NO scavenging: Cyclization to a cinnolinone prevents formation of unstable nitrosamines. J. Pineal Res. 2007, 43, 104–105.
- Hacışevki, A.; Baba, B. An overview of melatonin as an antioxidant molecule: A biochemical approach. Melatonin Mol. Biol. Clin. Pharm. Approaches 2018, 59–85.
- Tan, D.X.; Manchester, L.C.; Terron, M.P.; Flores, L.J.; Reiter, R.J. One molecule, many derivatives: A never-ending interaction of melatonin with reactive oxygen and nitrogen species? J. Pineal Res. 2007, 42, 28–42.
- Tan, D.X.; Hardeland, R.; Manchester, L.C.; Poeggeler, B.; Lopez-Burillo, S.; Mayo, J.C.; Sainz, R.M.; Reiter, R.J. Mechanistic and comparative studies of melatonin and classic antioxidants in terms of their interactions with the ABTS cation radical. J. Pineal Res. 2003, 34, 249–259.
- Hara, M.; Iigo, M.; Ohtani-Kaneko, R.; Nakamura, N.; Suzuki, T.; Reiter, R.J.; Hirata, K. Administration of melatonin and related indoles prevents exercise-induced cellular oxidative changes in rats. Neurosignals 1997, 6, 90–100.
- Qi, W.; Reiter, R.J.; Tan, D.X.; Manchester, L.C.; Siu, A.W.; Garcia, J.J. Increased levels of oxidatively damaged DNA induced by chromium(III) and H2O2: Protection by melatonin and related molecules. J. Pineal Res. 2000, 29, 54–61.
- López-Burillo, S.; Tan, D.X.; Rodriguez-Gallego, V.; Manchester, L.C.; Mayo, J.C.; Sainz, R.M.; Reiter, R.J. Melatonin and its derivatives cyclic 3-hydroxymelatonin, N1-acetyl-N2-formyl-5-methoxykynuramine and 6-methoxymelatonin reduce oxidative DNA damage induced by Fenton reagents. J. Pineal Res. 2003, 34, 178–184.
- Duan, Q.; Wang, Z.; Lu, T.; Chen, J.; Wang, X. Comparison of 6-hydroxylmelatonin or melatonin in protecting neurons against ischemia/reperfusion-mediated injury. J. Pineal Res. 2006, 41, 351–357.
- Sakano, K.; Oikawa, S.; Hiraku, Y.; Kawanishi, S. Oxidative DNA damage induced by a melatonin metabolite, 6-hydroxymelatonin, via a unique non-o-quinone type of redox cycle. Biochem. Pharmacol. 2004, 68, 1869–1878.
- Dellegar, S.M.; Murphy, S.A.; Bourne, A.E.; Dicesare, J.C.; Purser, G.H. Identification of the factors affecting the rate of deactivation of hypochlorous acid by melatonin. Biochem. Biophys. Res. Commun. 1999, 257, 431–439.
- Agozzino, P.; Avellone, G.; Bongiorno, D.; Ceraulo, L.; Filizzola, F.; Natoli, M.C.; Livrea, M.A.; Tesoriere, L. Melatonin: Structural characterization of its non-enzymatic mono-oxygenate metabolite. J. Pineal Res. 2003, 35, 269–275.
- Horstman, J.A.; Wrona, M.Z.; Dryhurst, G. Further insights into the reaction of melatonin with hydroxyl radical. Bioorg. Chem. 2002, 30, 371–382.
- Semak, I.; Naumova, M.; Korik, E.; Terekhovich, V.; Wortsman, J.; Slominski, A. A novel metabolic pathway of melatonin: Oxidation by cytochrome c. Biochemistry 2005, 44, 9300–9307.
- Kanwar, M.K.; Yu, J.; Zhou, J. Phytomelatonin: Recent advances and future prospects. J. Pineal Res. 2018, 65, e12526.
- Yu, Y.; Bian, L.; Jiao, Z.; Yu, K.; Wan, Y.; Zhang, G.; Guo, D. Molecular cloning and characterization of a grapevine (Vitis vinifera L.) serotonin N-acetyltransferase (VvSNAT2) gene involved in plant defense. BMC Genom. 2019, 20, 880.
- Wang, X.; Zhang, H.; Xie, Q.; Liu, Y.; Lv, H.; Bai, R.; Ma, R.; Li, X.; Zhang, X.; Guo, Y.D.; et al. SlSNAT Interacts with HSP40, a Molecular Chaperone, to Regulate Melatonin Biosynthesis and Promote Thermotolerance in Tomato. Plant Cell Physiol. 2020, 61, 909–921.
- Wang, L.; Feng, C.; Zheng, X.; Guo, Y.; Zhou, F.; Shan, D.; Liu, X.; Kong, J. Plant mitochondria synthesize melatonin and enhance the tolerance of plants to drought stress. J. Pineal Res. 2017, 63, e12429.
- Vigliante, I.; Mannino, G.; Maffei, M.E. Chemical Characterization and DNA Fingerprinting of Griffonia simplicifolia Baill. Molecules 2019, 24, 1032.
- Mannino, G.; Gentile, C.; Ertani, A.; Serio, G.; Bertea, C.M. Anthocyanins: Biosynthesis, Distribution, Ecological Role, and Use of Biostimulants to Increase Their Content in Plant Foods—A Review. Agriculture 2021, 11, 212.
- Back, K.; Tan, D.X.; Reiter, R.J. Melatonin biosynthesis in plants: Multiple pathways catalyze tryptophan to melatonin in the cytoplasm or chloroplasts. J. Pineal Res. 2016, 61, 426–437.
- Lee, H.Y.; Byeon, Y.; Lee, K.; Lee, H.J.; Back, K. Cloning of Arabidopsis serotonin N-acetyltransferase and its role with caffeic acid O-methyltransferase in the biosynthesis of melatonin in vitro despite their different subcellular localizations. J. Pineal Res. 2014, 57, 418–426.
- Park, S.; Byeon, Y.; Back, K. Functional analyses of three ASMT gene family members in rice plants. J. Pineal Res. 2013, 55, 409–415.
- Byeon, Y.; Lee, H.Y.; Lee, K.; Back, K. Caffeic acid O-methyltransferase is involved in the synthesis of melatonin by methylating N-acetylserotonin in Arabidopsis. J. Pineal Res. 2014, 57, 219–227.
- Zhou, Y.; Liao, L.; Liu, X.; Liu, B.; Chen, X.; Guo, Y.; Huang, C.; Zhao, Y.; Zeng, Z. Crystal structure of Oryza sativa TDC reveals the substrate specificity for TDC-mediated melatonin biosynthesis. J. Adv. Res. 2020, 24, 501–511.
- Zhao, D.; Yao, Z.; Zhang, J.; Zhang, R.; Mou, Z.; Zhang, X.; Li, Z.; Feng, X.; Chen, S.; Reiter, R.J. Melatonin synthesis genes N-acetylserotonin methyltransferases evolved into caffeic acid O-methyltransferases and both assisted in plant terrestrialization. J. Pineal Res. 2021, e12737.
- Kang, K.; Park, S.; Kim, Y.S.; Lee, S.; Back, K. Biosynthesis and biotechnological production of serotonin derivatives. Appl. Microbiol. Biotechnol. 2009, 83, 27–34.
- Byeon, Y.; Back, K. Molecular cloning of melatonin 2-hydroxylase responsible for 2-hydroxymelatonin production in rice (Oryza sativa). J. Pineal Res. 2015, 58, 343–351.
- Lee, K.; Zawadzka, A.; Czarnocki, Z.; Reiter, R.J.; Back, K. Molecular cloning of melatonin 3-hydroxylase and its production of cyclic 3-hydroxymelatonin in rice (Oryza sativa). J. Pineal Res. 2016, 61, 470–478.
- Ye, T.; Yin, X.; Yu, L.; Zheng, S.J.; Cai, W.J.; Wu, Y.; Feng, Y.Q. Metabolic analysis of the melatonin biosynthesis pathway using chemical labeling coupled with liquid chromatography-mass spectrometry. J. Pineal Res. 2019, 66, e12531.
- Byeon, Y.; Choi, G.H.; Lee, H.Y.; Back, K. Melatonin biosynthesis requires N-acetylserotonin methyltransferase activity of caffeic acid O-methyltransferase in rice. J. Exp. Bot. 2015, 66, 6917–6925.
- Koshiba, T.; Hirose, N.; Mukai, M.; Yamamura, M.; Hattori, T.; Suzuki, S.; Sakamoto, M.; Umezawa, T. Characterization of 5-hydroxyconiferaldehyde O-methyltransferase in Oryza sativa. Plant Biotechnol. 2013, 30, 157–167.
- Arnao, M.B. Phytomelatonin: Discovery, Content, and Role in Plants. Adv. Bot. 2014, 2014, 815769.
- Byeon, Y.; Park, S.; Kim, Y.S.; Park, D.H.; Lee, S.; Back, K. Light-regulated melatonin biosynthesis in rice during the senescence process in detached leaves. J. Pineal Res. 2012, 53, 107–111.
- Hardeland, R. Melatonin in plants and other phototrophs: Advances and gaps concerning the diversity of functions. J. Exp. Bot. 2015, 66, 627–646.
- Byeon, Y.; Lee, H.J.; Lee, H.Y.; Back, K. Cloning and functional characterization of the Arabidopsis N-acetylserotonin O-methyltransferase responsible for melatonin synthesis. J. Pineal Res. 2016, 60, 65–73.
- Byeon, Y.; Lee, H.Y.; Back, K. Chloroplastic and cytoplasmic overexpression of sheep serotonin N-acetyltransferase in transgenic rice plants is associated with low melatonin production despite high enzyme activity. J. Pineal Res. 2015, 58, 461–469.
- Park, S.; Byeon, Y.; Back, K. Transcriptional suppression of tryptamine 5-hydroxylase, a terminal serotonin biosynthetic gene, induces melatonin biosynthesis in rice (Oryza sativa L.). J. Pineal Res. 2013, 55, 131–137.
- Park, S.; Lee, K.; Kim, Y.S.; Back, K. Tryptamine 5-hydroxylase-deficient Sekiguchi rice induces synthesis of 5-hydroxytryptophan and N-acetyltryptamine but decreases melatonin biosynthesis during senescence process of detached leaves. J. Pineal Res. 2012, 52, 211–216.
- Balemans, M.G.M.; Ebels, I.; Hendriks, H.G.; van Berlo, M.; de Moreé, A. The influence of some pteridines on pineal 5-methoxyindole synthesis in male Wistar rats periodically exposed to either white or green light. J. Neural Transm. 1983, 58, 121–134.
- Tan, D.X.; Hardeland, R.; Back, K.; Manchester, L.C.; Alatorre-Jimenez, M.A.; Reiter, R.J. On the significance of an alternate pathway of melatonin synthesis via 5-methoxytryptamine: Comparisons across species. J. Pineal Res. 2016, 61, 27–40.
More
Information
Subjects:
Plant Sciences
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
4.8K
Revisions:
2 times
(View History)
Update Date:
29 Mar 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





