Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Yumi Chiba | + 1933 word(s) | 1933 | 2021-11-01 07:13:44 | | | |
| 2 | Rita Xu | Meta information modification | 1933 | 2021-11-09 08:58:52 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Chiba, Y. Cortical Bone Stem Cells. Encyclopedia. Available online: https://encyclopedia.pub/entry/15820 (accessed on 29 March 2026).
Chiba Y. Cortical Bone Stem Cells. Encyclopedia. Available at: https://encyclopedia.pub/entry/15820. Accessed March 29, 2026.
Chiba, Yumi. "Cortical Bone Stem Cells" Encyclopedia, https://encyclopedia.pub/entry/15820 (accessed March 29, 2026).
Chiba, Y. (2021, November 09). Cortical Bone Stem Cells. In Encyclopedia. https://encyclopedia.pub/entry/15820
Chiba, Yumi. "Cortical Bone Stem Cells." Encyclopedia. Web. 09 November, 2021.
Copy Citation
The newly established mouse cortical-bone-derived stem cells (mCBSCs) are unique stem cells compared to mouse mesenchymal stem cells (mMSCs). The mCBSC-treated hearts after myocardial infarction have been reported to have greater improvement in myocardial structure and functions.
cortical bone stem cells
stemness
glycan profile
1. Introduction
Several cardiovascular diseases (CVDs) are the leading cause of death globally due to their high morbidity and mortality rates [1]. In the coming decades, the incidence of CVD caused by ischemic CVDs, such as myocardial infarction (MI), is expected to be in upward trend [2]. After MI, myocyte death and the reduction in the number of functional cardiac myocytes ultimately leads to heart failure. Until now, although there is scientific progress and advancements in surgical techniques, drugs and surgical treatments can only delay the progression of chronic heart disease, but not improve the function of infarcted myocardial cells [3]. Therefore, the use of stem cells has emerged as a promising treatment for heart disease [4]. Our research and others suggest that stem cells hold immense potential for cardiac repair and regeneration [5][6][7][8]. Clinical use of adult somatic stem cells (SSCs) is a reality today and many stem cell types, including bone-marrow-derived mesenchymal stem cells (MSCs) [9], bone marrow cells [10][11], cardiac-derived cardiac progenitor [12], and cardio-sphere-derived cells [13], have been tested. The beneficial effects of tested cell therapies on cardiac structure and function have been modest, and most studies to date have not been adequately powered to document efficacy. The emerging consensus from these studies suggests that the donated stem cell population falls short of fully restoring normal cardiac functional capacity because of a combination of issues, such as poor survival, lack of proliferation, engraftment, and differentiation. In addition, it seems that much of the benefit derived from cell therapy has come from the release of paracrine factors acting on the host myocardium rather than from differentiation of infused/injected stem cells into new cardiac tissue.
Recently, we have shown that a novel SSC, mouse cortical-bone-derived stem cells (mCBSCs), improve cardiac remodeling and functions. The mCBSC-treated hearts showed increased neovascularization, and newly formed cardiac myocytes were also observed [8][14]. mCBSCs produce a unique combination of immunomodulatory and angiogenic and proangiogenic factors, which may be the reason why mCBSCs were more effective in improving the post-MI hearts compared to cardiac-derived stem cells and MSCs [14]. Furthermore, mCBSCs possess enhanced proliferation capacity, better survival, and lineage commitment capacity than those in other stem cells [14]. Thus, it is suggested that CBSCs have greater potential to repair the damaged heart than other cells currently being tested clinically. However, there are still many unknown characteristics in mCBSCs, including stem-cell-like features, noncardiac therapeutic potentials, and cell surface markers. Such characterization is considered to be useful for enhancing the utility of CBSC.
Glycans are expressed mainly on the surface of cells as components of glycoproteins, glycolipids, and proteoglycans, and are used as cell surface markers [15][16]. Glycoproteins contain N-glycans (including three major types: high mannose, hybrid, and complex glycans) synthesized on asparagine residues, part of the Asn-X-Ser/Thr consensus sequence (X: any amino acid except proline), and O-glycans synthesized on serine or threonine residues. It is well documented that glycan compositions of glycoproteins largely change during stem cell differentiation [17][18][19][20]. Furthermore, cell surface glycans contribute to fundamental biological functions, such as cell differentiation, cell adhesion, cell–cell interactions, pathogen–host recognition, toxin–receptor interactions, cancer metastasis, immune responses, and regulation of signaling pathways [16]. The glycans, including lactosamine and fucosylation, have been implicated in the self-renewal of stem cells by preventing differentiation [19][21][22]. In human MSCs (hMSCs), both N- and O-glycan processing functionally modulates early steps of osteogenic differentiation [23]. In adipose-derived hMSCs, the ability to differentiate is downregulated, with a decrease in α2–6-linked sialic acid associated with long-term cell culture in so-called “in vitro cellular aging” [24]. Thus, SSCs, including hMSCs, undergo tremendous changes, such as loss of differentiation potential, accompanying changes in glycans during long-term cell culture [25][26][27].
2. Isolation and Culture of mCBSCs
We isolated mCBSCs with modified protocols, as described in the Materials and Methods section. With modified protocols, mCBSCs were observed after approximately 2 months (passage 13) and cultured (Figure 1A). Fluorescence-activated cell sorting (FACS) analysis showed that the majority of mCBSCs (passage 17) expressed Sca-1 and β1-integrin (CD29), and around 50% of mCBSCs expressed c-kit (CD117) (Figure 1B). In contrast, later passaged mCBSCs (passage 22) showed weaker expression of c-kit, while Sca-1 and CD29 were expressed in the majority of the cells (Figure 1B). In both passaged mCBSCs (passage 17 and 22), CD45, Lin (hematopoietic lineage), and CD34 were not expressed (Figure 1B). We found that around 50% of mCBSCs (passages 16-18; defined early passage, EP) expressed c-kit, while c-kit was expressed in weak populations of mCBSCs (passages 22-24; defined late passage, LP) (Figure 1C). The growth curve showed the expected pattern of rapid growth for about 45 days, including EP and LP (Figure 1D). Furthermore, senescence-associated β-galactosidase (SA-β-Gal) activity, which is a well-known marker for senescent cells, was almost always negative in LP-mCBSCs compared to hydrogen peroxide (H2O2)-treated LP-mCBSCs, in which stress-induced premature senescence was considered to be induced (Figure 1E). These results indicate that populations of mCBSCs expressing c-kit change during long-term culture in so-called in vitro cellular aging, without cellular senescence. We compared characteristics of mCBSCs (EP and LP) with bone-marrow-derived mMSCs in further experiments.
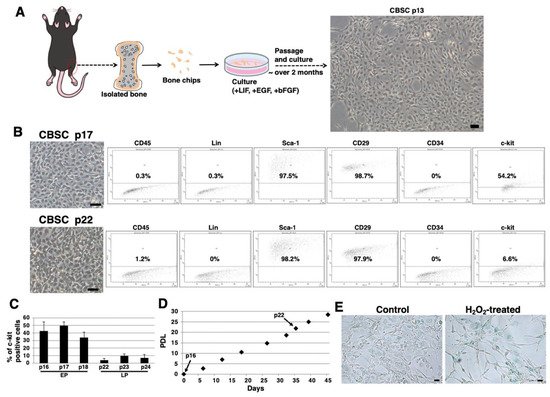
Figure 1. Isolation of mCBSCs. (A) Scheme of isolation and culture of mCBSCs. The bone biopsy was isolated from one limb. Then, bone chips were prepared and cultured with mCBSCs culture media. About 2 months after passage and culture, mCBSCs was observed. Scale bar in the phase contrast image: 2 μm. (B) FACS analysis of CD45, Lin, Sca-1, CD29, CD34, and c-kit in mCBSCs (p17 and p22). Three independent experiments were performed and representative results are shown. Scale bar in the phase contrast image: 2 µm. (C) Percent of c-kit positive cells in mCBSCs (p16-p18 and p22-p24) is shown. The values shown are the means ± SD from three independent experiments. (D) Growth curve showing the population doubling level (PDL) during culturing from p16 to p24. (E) Representative image of staining for SA-β-Gal in nontreated (control) or 400 µM hydrogen peroxide (H2O2)-treated LP-mCBSCs from two independent experiments is shown. Scale bar in the phase contrast image: 50 µm. Abbreviations: mCBSC, mouse cortical-bone-derived stem cell; EP, early passage; LP, late passage; PDL, population doubling level; SA-β-Gal, senescence-associated β-galactosidase; H2O2, hydrogen peroxide.
3. Stem-Cell-Like Features of mCBSCs
We examined stem-cell-like features of mCBSCs, specifically, Nanog and Sox2, as they are well known as stemness markers in the undifferentiated state of mouse ESCs [28]. In mCBSCs, Nanog was expressed, but lower than that in mMSCs (Figure 2A). In contrast, Sox2 was highly expressed in mCBSCs, comparable to that in mMSCs (Figure 2A). We further examined stemness by measuring the self-renewing ability of mCBSCs, using a colony-forming assay. The lower density of mCBSCs (both EP and LP) showed the formation of healthy colonies (Figure 2B), indicating the self-renewing ability in both EP- and LP-mCBSCs. Furthermore, the self-renewing ability in mCBSCs was higher than that in mMSCs (Figure 2B). We next examined the multilineage differentiation ability between mMSCs, EP-mCBSCs, and LP-mCBSCs. The mMSCs exhibited adipogenic, osteogenic, and chondrogenic differentiation, as shown in Figure 2C, while mCBSCs (EP and LP) exhibited only chondrogenic differentiation (Figure 2C). These results indicate that, among three lineages, mCBSCs have differentiation directly into the chondrogenic lineage only.
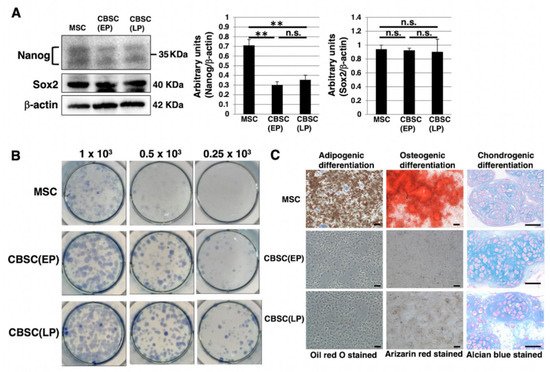
Figure 2. Stemness features of mCBSCs. (A) Western blot analysis of Nanog, Sox2, and β-actin (loading control) in mMSCs and mCBSCs (EP and LP). The blot images were cropped to highlight the Nanog, Sox2, and β-actin bands. The histogram shows the mean densitometric analysis ± SD of Nanog or Sox2 normalized to the loading control (β-actin). The values were obtained from three independent experiments. ** p < 0.01, n.s.: not significant. (B) Colony-forming assay in mMSCs and mCBSCs (EP and LP). Cells were plated at low density (0.25 × 103 cells, 0.5 × 103 cells, or 1 × 103 cells/well) and cultured. After 7 days, cells were stained and photos were captured. Two independent experiments were performed and representative images are shown. (C) Adipogenic, osteogenic, or chondrogenic differentiation was induced in mMSCs and mCBSCs (EP and LP). Two independent experiments were performed and representative images are shown. Scale bar in the phase contrast image of adipogenic and osteogenic: 2 µm. Scale bar in the phase contrast image of chondrogenic: 50 µm. Abbreviations: mMSC, mouse mesenchymal stem cell; mCBSC, mouse cortical-bone-derived stem cell; EP, early passage; LP, late passage.
4. The Feature of Glycan Profiles in mCBSCs (EP and LP) and mMSCs
To identify the glycan profiles of the mCBSCs and mMSCs, lectin microarray analysis was performed. As a result, the signal intensities were observed in various types of lectins (Figure 3A and Table 1). Initially, the glycan profiles of both EP- and LP-mCBSCs were compared with mMSCs. The signal intensities of SNA, SSA, and TJA-I (α2-6sialic acid-binding lectins) in mMSCs were significantly higher than those in both EP- and LP-mCBSCs. The signal intensity of TxLC-I in mMSCs was slightly higher than that in both mCBSCs. In contrast, the signal intensities of RCA120 and DSA (lactosamine (Galβ1-4GlcNAc)-binding lectins), and LEL (poly lactosamine- or poly N-acetylglucosamine-binding lectin) in both EP- and LP-mCBSCs were higher than those in mMSCs. The signal intensity of WFA (LacdiNAc (GalNAcβ1-4GlcNAc)-binding lectin) in mCBSCs was slightly higher than that in mMSCs. Moreover, the signal intensities of MAL-I, ACG, and WGA (α2-3sialic acid-binding lectins) in mCBSCs were higher than those in mMSCs. These results suggest that there are differences in sialic acid structures and the number of lactosamine units between mMSCs and mCBSCs.
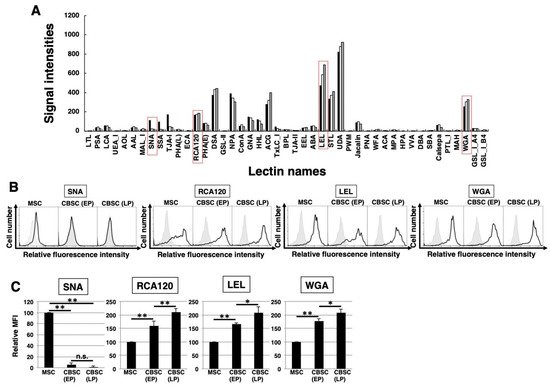
Figure 3. The feature of glycan profiles of mCBSCs and mMSCs. (A) Glycan profiles of the mMSCs, mouse cortical-bone-derived stem cells at early passage (EP-mCBSCs) and those at late passage (LP-mCBSCs) by averaged data (n = 10). Bar graph representation of signal intensities of 45 lectins in lectin microarray data. Closed, open, and gray bars show mMSCs, EP-mCBSCs, and LP-mCBSCs, respectively. The lectins enclosed in a red line are further experimented in (B) and (C). (B) FACS analysis of mMSCs, EP-mCBSCs, and LP-mCBSCs using lectins (SNA, RCA120, LEL, and WGA). Three independent experiments were performed and representative results are shown. Negative control is shown in gray. (C) MFIs relative to those of mMSCs (value = 100) are shown. Results are presented as means ± SD from three independent experiments. * p < 0.05, ** p < 0.01. Abbreviations: mMSC, mouse mesenchymal stem cell; mCBSC, mouse cortical-bone-derived stem cell; EP, early passage; LP, late passage; SNA, Sambucus nigra; RCA120, Ricinus communis agglutinin I; LEL, Lycopersicon esculentum; WGA, wheat germ agglutinin; MFIs, mean fluorescence intensities.
Table 1. The signal intensities of each cell type in lectin microarray data.
| Lectin Name | mMSC-1 | mMSC-2 | mMSC-3 | CBSC (EP)-1 | CBSC (EP)-2 | CBSC (EP)-3 | CBSC (LP)-1 | CBSC (LP)-2 | CBSC (LP)-3 |
|---|---|---|---|---|---|---|---|---|---|
| LTL*2 | 0.1 | 0.1 | 1.3 | 0.2 | 0.1 | 0.6 | 0.0 | 0.0 | 0.1 |
| PSA*2 | 29.6 | 33.1 | 50.5 | 41.3 | 45.4 | 36.8 | 23.5 | 28.5 | 23.8 |
| LCA*2 | 46.2 | 49.9 | 74.4 | 56.6 | 59.2 | 50.1 | 35.7 | 41.8 | 34.2 |
| UEA_I | 0.1 | 0.0 | 3.5 | 0.0 | 0.1 | 0.5 | 0.0 | 0.1 | 0.3 |
| AOL*2 | 2.8 | 2.5 | 15.7 | 5.8 | 6.5 | 7.2 | 2.0 | 3.5 | 6.1 |
| AAL*2 | 28.6 | 28.9 | 54.6 | 48.2 | 52.4 | 38.6 | 36.4 | 40.4 | 31.4 |
| MAL_I*2 | 13.1 | 14.6 | 21.9 | 27.2 | 30.6 | 21.1 | 18.8 | 22.1 | 16.6 |
| SNA*1, *2 | 105.2 | 104.2 | 129.8 | 25.2 | 23.2 | 22.0 | 16.7 | 20.5 | 20.7 |
| SSA*1, *2 | 89.2 | 84.6 | 112.9 | 17.7 | 18.2 | 16.3 | 11.1 | 15.2 | 12.7 |
| TJA-I*1, *2 | 153.2 | 160.4 | 197.7 | 45.0 | 45.0 | 47.5 | 34.8 | 44.8 | 40.9 |
| PHA(L)*2 | 11.8 | 12.5 | 27.4 | 18.9 | 21.3 | 19.0 | 9.0 | 12.7 | 12.7 |
| ECA*2 | 1.1 | 1.4 | 5.7 | 5.3 | 5.6 | 5.2 | 2.9 | 5.3 | 4.4 |
| RCA120 | 148.3 | 151.8 | 194.2 | 162.8 | 167.2 | 187.2 | 164.3 | 191.9 | 192.9 |
| PHA(E)*2 | 66.1 | 70.0 | 107.9 | 74.1 | 80.4 | 94.3 | 54.3 | 60.7 | 74.6 |
| DSA*1 | 354.7 | 354.1 | 405.5 | 426.4 | 424.0 | 443.8 | 423.0 | 454.3 | 450.3 |
| GSL-II*2 | 0.5 | 1.1 | 4.0 | 1.7 | 2.4 | 3.7 | 0.1 | 0.8 | 2.0 |
| NPA*1 | 400.6 | 413.7 | 369.2 | 373.9 | 372.4 | 302.9 | 326.5 | 313.9 | 282.0 |
| ConA*1, *2 | 48.8 | 53.1 | 58.7 | 69.1 | 71.0 | 58.7 | 45.4 | 51.0 | 41.2 |
| GNA | 144.5 | 162.0 | 145.7 | 152.2 | 164.2 | 105.9 | 122.0 | 147.0 | 72.1 |
| HHL*2 | 90.0 | 89.5 | 165.5 | 111.7 | 102.1 | 103.8 | 68.3 | 67.8 | 81.0 |
| ACG*2 | 341.5 | 331.2 | 193.7 | 349.2 | 324.7 | 298.2 | 456.7 | 396.0 | 355.6 |
| TxLC_I*1, *2 | 33.2 | 36.7 | 62.0 | 24.1 | 26.8 | 26.1 | 16.1 | 18.2 | 17.8 |
| BPL | 12.6 | 11.9 | 23.9 | 14.0 | 17.0 | 18.1 | 14.9 | 17.8 | 17.0 |
| TJA-II | 5.9 | 5.5 | 17.4 | 4.2 | 4.3 | 6.1 | 1.0 | 9.5 | 5.3 |
| EEL | 33.0 | 35.1 | 39.7 | 34.3 | 38.5 | 32.3 | 35.8 | 37.0 | 37.3 |
| ABA*2 | 37.9 | 42.8 | 71.0 | 54.5 | 55.4 | 57.3 | 35.5 | 39.7 | 44.0 |
| LEL*1, *2 | 486.8 | 474.2 | 469.9 | 577.2 | 540.0 | 625.4 | 645.6 | 656.2 | 747.8 |
| STL*2 | 376.3 | 336.5 | 305.0 | 349.0 | 342.0 | 409.7 | 402.3 | 424.4 | 405.9 |
| UDA | 984.6 | 965.7 | 600.8 | 854.2 | 838.9 | 925.6 | 958.4 | 804.8 | 987.9 |
| PWM | 0.1 | 0.0 | 1.8 | 0.7 | 0.3 | 1.4 | 0.2 | 0.9 | 0.3 |
| Jacalin*2 | 85.1 | 85.3 | 99.3 | 99.7 | 103.6 | 93.8 | 78.2 | 81.8 | 63.5 |
| PNA | 0.0 | 0.0 | 0.3 | 0.1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.2 |
| WFA*1 | 7.1 | 6.7 | 15.0 | 16.0 | 18.0 | 16.1 | 15.3 | 23.7 | 15.6 |
| ACA*2 | 3.4 | 4.0 | 12.7 | 6.8 | 7.5 | 6.5 | 2.6 | 4.6 | 4.9 |
| MPA*2 | 11.4 | 12.0 | 20.0 | 15.6 | 16.1 | 12.1 | 7.1 | 9.4 | 6.8 |
| HPA*2 | 0.0 | 0.0 | 3.3 | 0.3 | 0.6 | 0.8 | 0.0 | 0.0 | 0.0 |
| VVA | 0.0 | 0.0 | 0.1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.1 |
| DBA | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| SBA | 2.5 | 3.8 | 8.8 | 6.1 | 6.4 | 5.4 | 4.2 | 7.1 | 4.4 |
| Calsepa*2 | 54.3 | 56.4 | 75.3 | 69.4 | 72.1 | 62.2 | 42.7 | 48.4 | 32.3 |
| PTL_I | 3.4 | 3.8 | 7.9 | 3.5 | 4.7 | 3.4 | 2.9 | 3.6 | 2.2 |
| MAH*2 | 5.1 | 5.6 | 11.9 | 13.6 | 13.9 | 11.0 | 4.6 | 7.7 | 6.3 |
| WGA*1 | 247.6 | 263.2 | 256.9 | 304.0 | 331.3 | 286.0 | 341.9 | 341.1 | 311.2 |
| GSL_I_A4 | 23.6 | 21.8 | 41.6 | 26.3 | 30.8 | 27.5 | 26.8 | 29.2 | 26.6 |
| GSL_I_B4 | 10.0 | 10.2 | 16.1 | 13.9 | 16.2 | 10.5 | 12.7 | 16.2 | 7.8 |
The data were averaged after normalization (sample 1 and 2, n = 3; sample 3, n = 4). T-test was performed with signal intensities before average (n = 10). *1 There was the significant difference between mCBSCs (both EP and LP) and mMSCs. *2 There was the significant difference between EP-mCBSCs and LP-mCBSCs.
Next, the glycan profiles were analyzed with regard to cellular aging of mCBSCs by comparing between EP and LP. The signal intensities of PSA, LCA, and AAL (α1-6fucose-binding lectins) decreased with LP-mCBSCs. The signal intensities of LEL and STL (poly N-acetylglucosamine-binding lectin), RCA120, and WGA increased with LP-mCBSCs. These results suggest that the glycan profile in CBSCs changes with cellular aging.
References
- Virani, S.S.; Alonso, A.; Aparicio, H.J.; Benjamin, E.J.; Bittencourt, M.S.; Callaway, C.W.; Carson, A.P.; Chamberlain, A.M.; Cheng, S.; Delling, F.N.; et al. Heart Disease and Stroke Statistics-2021 Update: A Report From the American Heart Association. Circulation 2021, 143, e254–e743.
- Sliwa, K.; Ntusi, N. Battling Cardiovascular Diseases in a Perfect Storm. Circulation 2019, 139, 1658–1660.
- Ji, S.T.; Kim, H.; Yun, J.; Chung, J.S.; Kwon, S.M. Promising Therapeutic Strategies for Mesenchymal Stem Cell-Based Cardiovascular Regeneration: From Cell Priming to Tissue Engineering. Stem Cells Int. 2017, 2017, 3945403.
- Karantalis, V.; Balkan, W.; Schulman, I.H.; Hatzistergos, K.E.; Hare, J.M. Cell-based therapy for prevention and reversal of myocardial remodeling. Am. J. Physiol. Heart Circ. Physiol. 2012, 303, H256–H270.
- Urbanek, K.; Torella, D.; Sheikh, F.; Angelis, A.D.; Nurzynska, D.; Silvestri, F.; Beltrami, C.A.; Bussani, R.; Beltrami, A.P.; Quaini, F.; et al. Myocardial regeneration by activation of multipotent cardiac stem cells in ischemic heart failure. Proc. Natl. Acad. Sci. USA 2005, 102, 8692–8697.
- Mohsin, S.; Khan, M.; Toko, H.; Bailey, B.; Cottage, C.T.; Wallach, K.; Nag, D.; Lee, A.; Siddiqi, S.; Lan, F.; et al. Human cardiac progenitor cells engineered with Pim-I kinase enhance myocardial repair. J. Am. Coll. Cardiol. 2012, 60, 1278–1287.
- Oskouei, B.N.; Lamirault, G.; Joseph, C.; Treuer, A.V.; Landa, S.; Silva, J.D.; Hatzistergos, K.; Dauer, M.; Balkan, W.; McNiece, I.; et al. Increased potency of cardiac stem cells compared with bone marrow mesenchymal stem cells in cardiac repair. Stem Cells Transl. Med. 2012, 1, 116–124.
- Duran, J.M.; Makarewich, C.A.; Sharp, T.E.; Starosta, T.; Zhu, F.; Hoffman, N.E.; Chiba, Y.; Madesh, M.; Berretta, R.M.; Kubo, H.; et al. Bone-derived stem cells repair the heart after myocardial infarction through transdifferentiation and paracrine signaling mechanisms. Circ. Res. 2013, 113, 539–552.
- Karantalis, V.; DiFede, D.L.; Gerstenblith, G.; Pham, S.; Symes, J.; Zambrano, J.P.; Fishman, J.; Pattany, P.; McNiece, I.; Conte, J.; et al. Autologous mesenchymal stem cells produce concordant improvements in regional function, tissue perfusion, and fibrotic burden when administered to patients undergoing coronary artery bypass grafting: The Prospective Randomized Study of Mesenchymal Stem Cell Therapy in Patients Undergoing Cardiac Surgery (PROMETHEUS) trial. Circ. Res. 2014, 114, 1302–1310.
- Pätilä, T.; Lehtinen, M.; Vento, A.; Schildt, J.; Sinisalo, J.; Laine, M.; Hämmäinen, P.; Nihtinen, A.; Alitalo, R.; Nikkinen, P.; et al. Autologous bone marrow mononuclear cell transplantation in ischemic heart failure: A prospective, controlled, randomized, double-blind study of cell transplantation combined with coronary bypass. J. Heart Lung Transplant. 2014, 33, 567–574.
- Delewi, R.; van der Laan, A.M.; Robbers, L.F.; Hirsch, A.; Nijveldt, R.; van der Vleuten, P.A.; Tijssen, J.G.; Tio, R.A.; Waltenberger, J.; Ten Berg, J.M.; et al. Long term outcome after mononuclear bone marrow or peripheral blood cells infusion after myocardial infarction. Heart 2015, 101, 363–368.
- Bolli, R.; Chugh, A.R.; D’Amario, D.; Loughran, J.H.; Stoddard, M.F.; Ikram, S.; Beache, G.M.; Wagner, S.G.; Leri, A.; Hosoda, T.; et al. Cardiac stem cells in patients with ischaemic cardiomyopathy (SCIPIO): Initial results of a randomised phase 1 trial. Lancet 2011, 378, 1847–1857.
- Makkar, R.R.; Smith, R.R.; Cheng, K.; Malliaras, K.; Thomson, L.E.; Berman, D.; Czer, L.S.; Marbán, L.; Mendizabal, A.; Johnston, P.V.; et al. Intracoronary cardiosphere-derived cells for heart regeneration after myocardial infarction (CADUCEUS): A prospective, randomised phase 1 trial. Lancet 2012, 379, 895–904.
- Mohsin, S.; Troupes, C.D.; Starosta, T.; Sharp, T.E.; Agra, E.J.; Smith, S.; Duran, J.M.; Zalavadia, N.; Zhou, Y.; Kubo, H.; et al. Unique Features of Cortical Bone Stem Cells Associated With Repair of the Injured Heart. Circ. Res. 2015, 117, 1024–1033.
- Varki, A.; Cummings, R.D.; Esko, J.D.; Stanley, P.; Hart, G.W.; Aebi, M.; Darvill, A.G.; Kinoshita, T.; Packer, N.H.; Prestegard, J.H.; et al. Essentials of Glycobiology, 3rd ed.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2017.
- Varki, A. Biological roles of glycans. Glycobiology 2017, 27, 3–49.
- An, H.J.; Gip, P.; Kim, J.; Wu, S.; Park, K.W.; McVaugh, C.T.; Schaffer, D.V.; Bertozzi, C.R.; Lebrilla, C.B. Extensive determination of glycan heterogeneity reveals an unusual abundance of high mannose glycans in enriched plasma membranes of human embryonic stem cells. Mol. Cell. Proteom. 2012, 11, M111.010660.
- Hasehira, K.; Tateno, H.; Onuma, Y.; Ito, Y.; Asashima, M.; Hirabayashi, J. Structural and quantitative evidence for dynamic glycome shift on production of induced pluripotent stem cells. Mol. Cell Proteomics 2012, 11, 1913–1923.
- Hamouda, H.; Ullah, M.; Berger, M.; Tauber, R.; Ringe, J.; Blanchard, V. N-glycosylation profile of undifferentiated and adipogenically differentiated human bone marrow mesenchymal stem cells: Towards a next generation of stem cell markers. Stem Cells Dev. 2013, 22, 3100–3113.
- Wilson, K.M.; Thomas-Oates, J.E.; Genever, P.G.; Ungar, D. Glycan profiling shows unvaried n-glycomes in MSC clones with distinct differentiation potentials. Front. Cell Dev. Biol. 2016, 4, 52.
- Kumar, A.; Torii, T.; Ishino, Y.; Muraoka, D.; Yoshimura, T.; Togayachi, A.; Narimatsu, H.; Ikenaka, K.; Hitoshi, S. The Lewis X-related alpha1,3-fucosyltransferase, Fut10, is required for the maintenance of stem cell populations. J. Biol. Chem. 2013, 288, 28859–28868.
- Sasaki, N.; Shinomi, M.; Hirano, K.; Ui-Tei, K.; Nishihara, S. LacdiNAc (GalNAc beta 1-4GlcNAc) contributes to self-renewal ofmouse embryonic stemcells by regulating leukemia inhibitory factor/STAT3 signaling. Stem Cells. 2011, 29, 641–650.
- Wilson, K.M.; Jagger, A.M.; Walker, M.; Seinkmane, E.; Fox, J.M.; Kröger, R.; Genever, P.; Ungar, D. Glycans modify mesenchymal stem cell differentiation to impact on the function of resulting osteoblasts. J. Cell Sci. 2018, 131, jcs209452.
- Tateno, H.; Saito, S.; Hiemori, K.; Kiyoi, K.; Hasehira, K.; Toyoda, M.; Onuma, Y.; Ito, Y.; Akutsu, H.; Hirabayashi, J. α2-6 sialylation is a marker of the differentiation potential of human mesenchymal stem cells. Glycobiology 2016, 26, 1328–1337.
- Banfi, A.; Muraglia, A.; Dozin, B.; Mastrogiacomo, M.; Cancedda, R.; Quarto, R. Proliferation kinetics and differentiation potential of ex vivo expanded human bone marrow stromal cells: Implications for their use in cell therapy. Exp. Hematol. 2000, 28, 707–715.
- Bonab, M.M.; Alimoghaddam, K.; Talebian, F.; Ghaffari, S.H.; Ghavamzadeh, A.; Nikbin, B. Aging of mesenchymal stem cell in vitro. BMC Cell Biol. 2006, 7, 14.
- Schellenberg, A.; Stiehl, T.; Horn, P.; Joussen, S.; Pallua, N.; Ho, A.D.; Wagner, W. Population dynamics of mesenchymal stromal cells during culture expansion. Cytotherapy 2012, 14, 401–411.
- Parfitt, D.E.; Shen, M.M. From blastocyst to gastrula: Gene regulatory networks of embryonic stem cells and early mouse embryogenesis. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2014, 369, 20130542.
More
Information
Subjects:
Cell & Tissue Engineering
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
749
Revisions:
2 times
(View History)
Update Date:
09 Nov 2021
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





