| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Sherwin Reyes | + 2012 word(s) | 2012 | 2020-08-05 09:55:13 |
Video Upload Options
Urinary tract infection (UTI) is a common bacterial infection of the urethra, bladder, ureters, and the kidneys. A diagnosis of a positive UTI is any pathogen with a bacterial load of greater than 100,000 CFU/mL in the urine. A bacterial load under 100,000 CFU/mL is considered a negative UTI result. Escherichia coli (E.coli) is the most common pathogen that infects the urinary tract.
1. Abstract
Urinary tract infection (UTI) is one of the most common infections, accounting for a substantial portion of outpatient hospital and clinic visits. Standard diagnosis of UTI by culture and sensitivity can take at least 48 h, and improper diagnosis can lead to an increase in antibiotic resistance following therapy. To address these shortcomings, rapid bioluminescence assays were developed and evaluated for the detection of UTI using intact, viable cells of Photobacterium mandapamensis USTCMS 1132 or previously lyophilized cells of Photobacterium leiognathi ATCC 33981. Two platform technologies—tube bioluminescence extinction technology urine (TuBETUr) and cellphone-based UTI bioluminescence extinction technology (CUBET)—were developed and standardized using artificial urine to detect UTI. These assays could also provide information regarding pathogen concentration/level, helping guide treatment decisions. These technologies were able to detect microbes associated with UTI at less than 105 CFU/mL, which is usually the lower cut-off limit for a positive UTI diagnosis. These technologies could potentially address the need for point-of-care UTI detection while reducing the possibility of antibiotic resistance associated with misdiagnosed cases of urinary tract infections, especially in low-resource environments.
2. Introduction
Urinary tract infection (UTI) is a common infection that affects approximately 150 million people in the world[1][2][3]. Due to its commonality, a rapid and cost-efficient diagnostic test to identify UTI is important. It is needed especially for those in low-income countries because access to expensive technologies is difficult and on-site testing is preferable. Two common technologies used to identify UTI are urine culture and urinalysis. Urine culture is an accurate way to detect UTI because it can test for bacteria cell count, type of uro-pathogen, and antibiotic susceptibility[4][5]. The setback of urine culture is that it takes several days to get results. Urinalysis, on the other hand, is a rapid and inexpensive way to test UTI but the sensitivity and specificity is low therefore not reliable[6][7][8]. Thus, the need for rapid UTI detection with amenable on-site testing is a necessity[9].
We approached this need through the design and development of simple bioluminescence-based assays that required only living, unicellular bioluminescent organisms that could be kept for the long term in a dry form for the diagnosis of UTI. Based on our findings that exposure to infected urine resulted in the loss of organismal bioluminescence, we developed a bioluminescence-based cell suspension assay that could be clinically validated against currently accepted, conventional assays for UTI, such as the culture technique. With the use of standardized suspensions of luminous bacteria, infections in the urinary tract could be rapidly detected in less than 2 h compared to conventional urine culture that can take up to two days. Our whole-cell bioluminescence assay was adapted to two different platforms, a laboratory equipment-based detection called tube bioluminescence extinction technology urine (TuBETUr) and a point-of-care, cellphone-based detection termed cellphone-based UTI bioluminescence extinction technology (CUBET). The TuBETUr and CUBET assays worked by simply measuring the luminescence of Photobacterium mandapamensis (P. mandapamensis) or Photobacterium leiognathi (P. leiognathi), respectively. The loss of brightness indicated the detection of a bacterial UTI; conversely, a constant brightness measured beyond a certain cut-off time indicated a negative UTI. Both gave rapid results using the bioluminescent organism as the only reagent, making TuBETUr and CUBET extremely inexpensive and effective UTI diagnostic tools.
3. Data, Model, Applications, and Influence
3.1 Relationship between the Cell Density of Selected Uropathogens and Time-to-Blackout in Artificial Urine
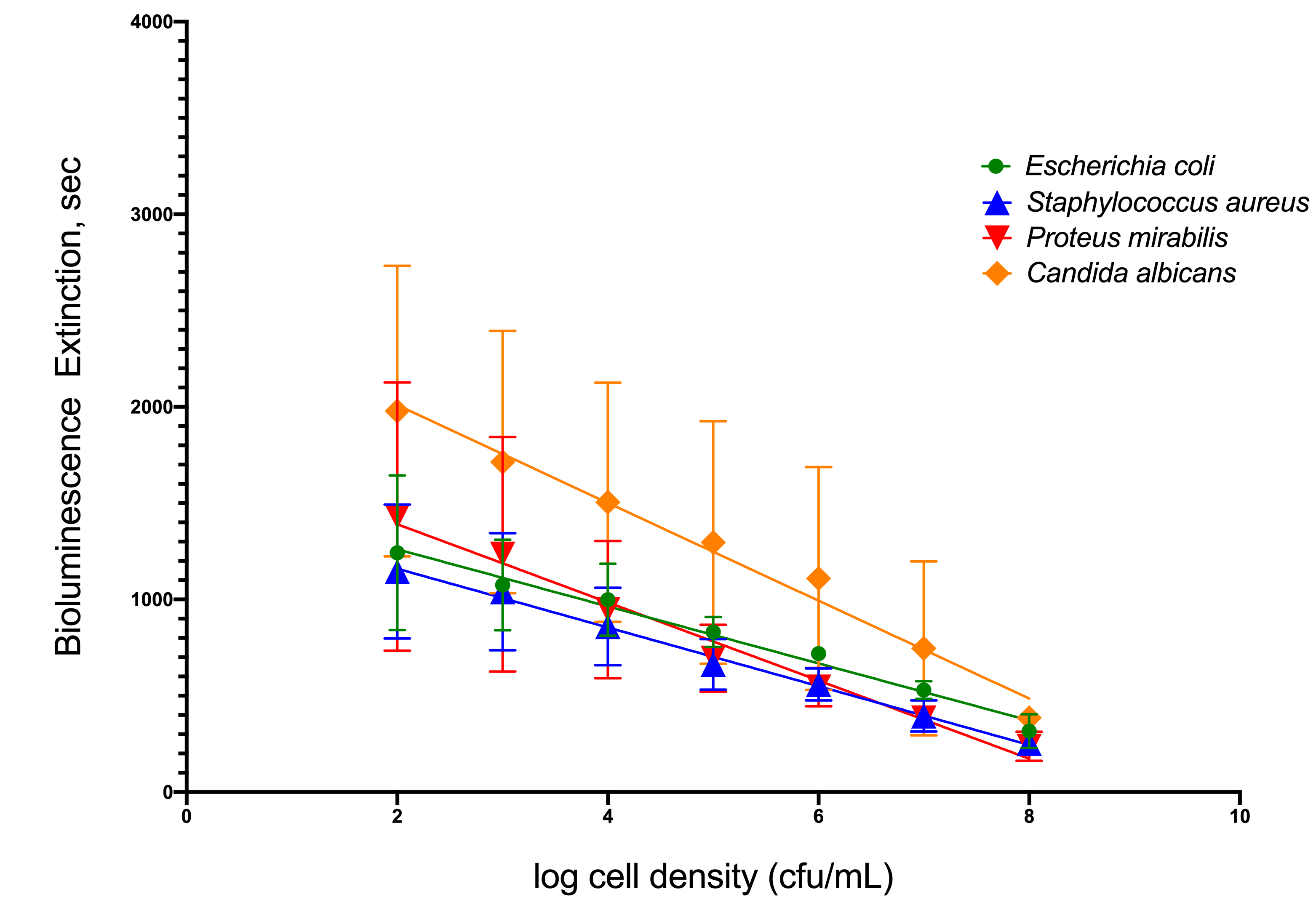
To simulate infection in patients’ urine, four common uropathogenic bacteria strains were utilized as standard organisms to determine the capability of TuBETUr technology in detecting bacteria spiked within a range of 102–108 CFU/mL in artificial urine. Each uropathogen showed a linear decrease in light intensity as the number of cells present in the artificial urine increased (Figure 1). Clinically, a 105 CFU/mL concentration[10][11][12], although indicating a significant number of uropathogens, is considered the lower limit of detection. Any bacterial concentration in the urine at or above this limit is considered serious enough to warrant treatment, whereas any bacterial cell count below this limit is considered negative. Thus, the lower limit of detection was determined by assigning a value, termed the cut-off time, that statistically defines an assay detection limit, as determined by the longest pathogen blackout time. The cut-off time, therefore, represents a point that is three standard deviations beyond the longest experimental mean blackout time for a culture containing ≥ 105 CFU/mL bacteria and defines an arbitrary “cut-off”, beyond which all samples are considered negative. Among the four uropathogens, Candida albicans, an opportunistic fungus associated with moniliasis in the urinary tract, showed the longest blackout time in this assay. Shown graphically in Figure 1, the mean blackout times for 105 CFU/mL of E. coli, Staphylococcus aureus, Proteus mirabilis, and C. albicans uropathogens in artificial urine were 831, 663, 694, and 1246 s, respectively. As a result, an assay cut-off time of 1257 s was utilized for TuBETUr in the diagnosis of UTI, as this mean blackout.
Figure 1. Relationship of blackout time and approximate log10 cell density/mL of the four common
uropathogens, which included Escherichia coli ATCC 25922TM, Staphylococcus aureus ATCC 23235TM,
Proteus mirabilis ATCC 35659TM, and Candida albicans ATCC 14053TM in artificial urine.
3.2 Application of TuBETUr for UTI Diagnosis
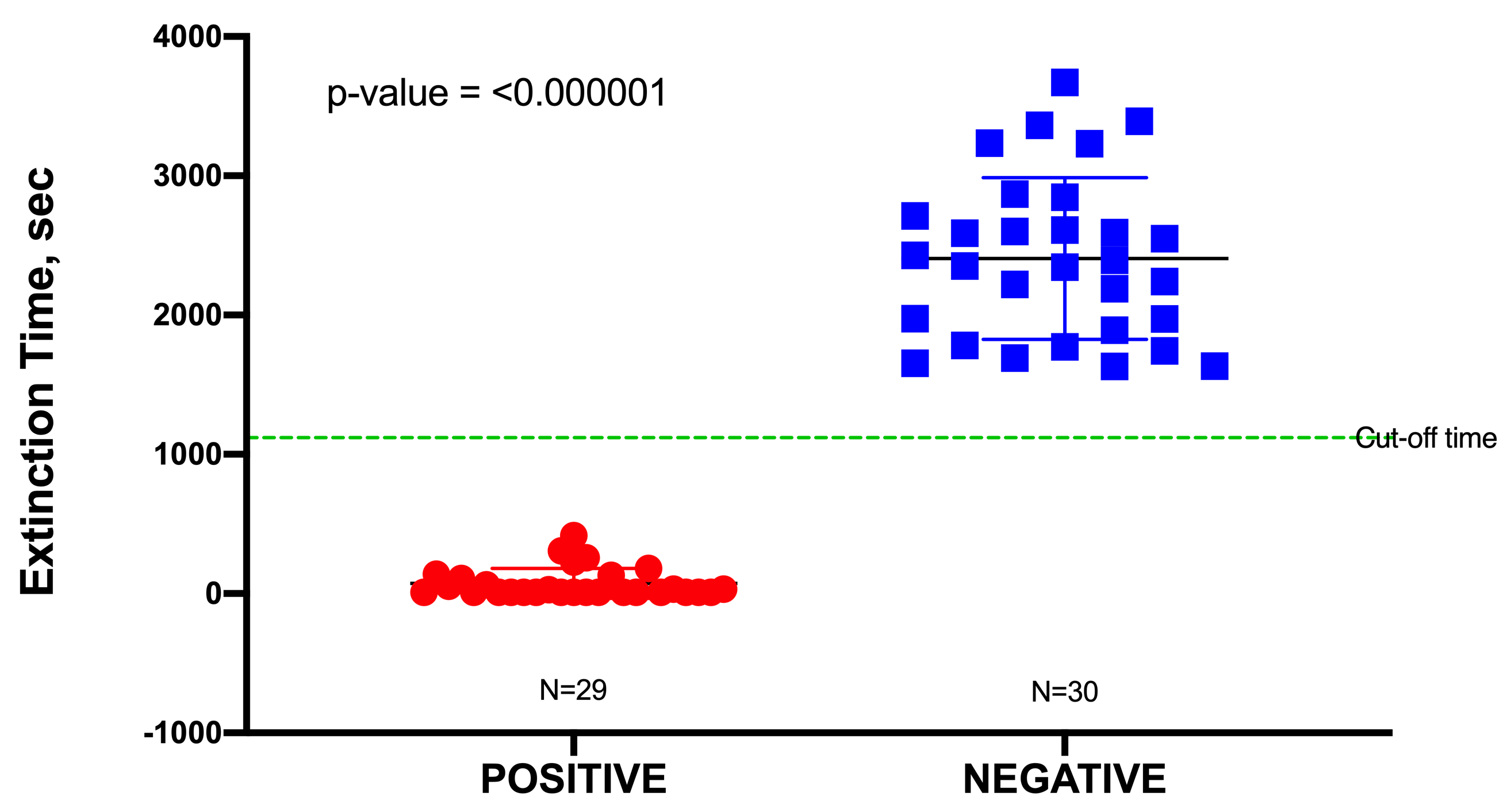
In this study, a total of 60 urine specimens were used, comprising 30 negative samples and 30 positive samples representing uncomplicated UTIs, although one outlier in the positive data was identified from a Dixon’s Q test and was subsequently removed. Uncomplicated urine specimens were used to minimize false positives due to potential interference (toxicity) from any unknown pharmaceuticals present. The pH of these urine samples was found to range from 6.0 to 7.5; this had no significant effect on the blackout times, as most bacteria would readily grow in a pH range of 6.0–8.0. All positive urine specimens yielded a culture of ≥105 CFU/mL, which was an indication of a clinically positive UTI. Blackout times ranged from as few as 10 to 1110 s, and the 29 urine specimens were correctly identified by TuBETUr. As the blackout times for the 29 validated positive samples fell below the cut-off time of 1257 s expected for 105 CFU/mL, the actual concentrations were likely much higher. The blackout times for negative specimens were all above the arbitrary 1257 s cut-off time for the ≥105 log10 CFU/mL detection limit and were correctly diagnosed by TuBETUr with a specificity of 100%. Thus, TuBETUr provided a sensitivity for positive UTI diagnosis of 29/29 or 100% (Figure 2). These results confirmed TuBETUr to be an exceptional diagnostic method for UTI. Further, the substantially longer blackout times for negative samples indicated that the upper time limit could potentially be adjusted to 1257 s in order to prevent further false negatives. Although TuBETUr was not designed to identify the species of microorganisms present, this was not a drawback as quantification of cell density in the urine is still considered the most important parameter in diagnosing UTI.
Figure 2. Sample distribution of urinary tract infection (UTI) negative and positive samples detected by tube
bioluminescence extinction technology urine (TuBETUr) technology compared to standard urine culture
3.3 Cellphone-Based Urinary Tract Infection Bioluminescence Extinction Technology (CUBET)
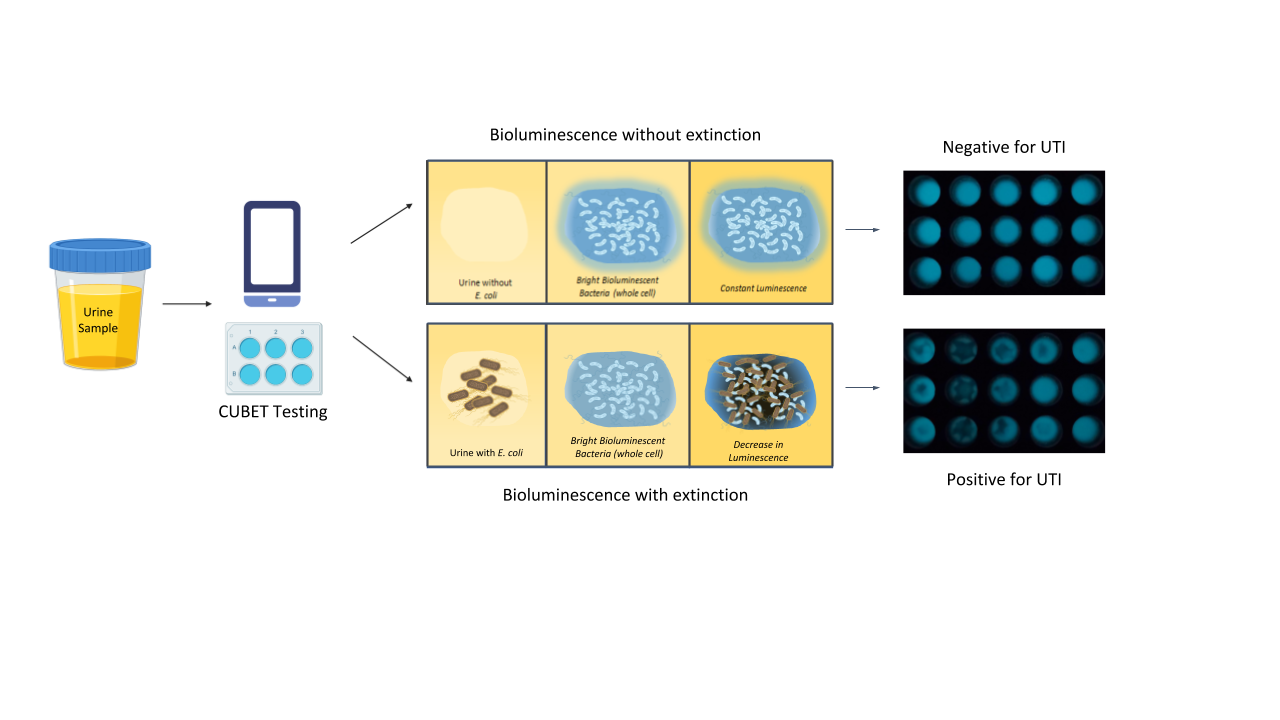
Upon demonstration that lyophilized P. leiognathi cells could be used as the bioluminescent signaling reagent for UTI detection, we developed a point-of-care device to aid in diagnosing urinary tract infections in remote areas or high-traffic clinics. The CUBET device was 3D printed based on a schematic rendered in Rhinoceros® (v. 6) software. As shown in Figure 3, the CUBET assay did not require sophisticated instrumentation beyond a simple camera-equipped cellphone to confirm suspected UTIs in patients. Furthermore, the results obtained from CUBET were comparable to that of our previous instrumental results with greater than 99% confidence. Figure 3 demonstrates the simplicity of the process, and the inclusion of lyophilized reporter enabled the long-term storage of the assay kit. Within minutes, CUBET could detect if a patient had a UTI by simply adding the urine sample to the chamber and waiting. If the P. leiognathi did not blackout after 5 min, then the sample was negative for UTI, and positive samples could be detected with high sensitivity within
the first 5 min of adding urine specimen.
CUBET is a UTI diagnostic tool that can provide early detection of UTI on the parameter that the bacterial load is ≥105 CFU/mL. Although it cannot detect the specific uro-pathogen causing the UTI, it can prevent antimicrobial resistance caused by misdiagnose. In addition, it is an inexpensive and affordable for low income countries. Furthermore, it is able to detect within minutes if a patient has UTI or not.
Figure 3. Flow of urinary tract infection detection using cellphone-based UTI bioluminescence
extinction technology (CUBET)
4. Materials and Methods
4.1 Urine Samples
Urine samples, representing 60 individuals, were obtained from Far Eastern University–Nicanor Reyes Medical Foundation Clinical Laboratory Bacteriology and Clinical Microscopy section, Quezon City Philippines. Samples were acquired as a gift following the laboratory assessment of biosafety and deidentification.
4.2. Bioluminescence Assay Using Reference Organisms in Artificial Urine for the Tube Bioluminescence Extinction Assay Urine (TuBETUr)
Ten-fold serial dilutions ranging from 102–108 CFU/mL of Escherichia coli ATCC 25922TM, Staphylococcus aureus ATCC 23235TM, Proteus mirabilis ATCC 35659TM, and Candida albicans ATCC 14053TM cultures were individually grown in artificial urine (AU-Siriraj) for several hours, salinized in 2.5% (w/v) sodium chloride (EMD Chemicals; Gibbstown, NJ, USA), and mixed with a 2 McFarland standard bioluminescent cell suspension of P. mandapamensis. In practice, this translated to 9 mL of each cell suspension prepared as above, mixed with a 1 mL aliquot of 1010 CFU/mL saline suspension of P. mandapamensis. The tube was capped and inverted once, and the time-to-blackout in min was recorded per dilution level for each culture. The result was graphed against the approximate cell density determined for the uropathogens and expressed as CFU/mL.
4.3. Tube Bioluminescence Extinction Technology for Urine (TuBETUr) Samples
The bioluminescence extinction (blackout time) for the 60 urine samples was examined. These samples were initially collected from patients with uncomplicated UTI. The urine samples were coded and measured for their pH and turbidity. The estimated viable plate count of each urine sample was determined side by side as the blackout time was being determined. Freshly collected urine samples were prepared identically to the artificial urine samples. Blackout was observed visually in a dark room, and photographs were obtained. From the blackout period of the 59 remaining uncomplicated urine samples (following outlier removal), a cut-off time was determined from the 3-sigma rule applied to the longest positive blackout time that would distinguish normal (non-infected) urine from the UTI-diagnosed samples. Infected urine samples were those that yielded an estimated viable plate count of 105 CFU/mL. Non-infected urine had an estimated viable plate count of less than 105 CFU/mL. From this sample size, the mean time-to-blackout for the normal and UTI-infected urine samples was determined and compared statistically to the results obtained for the 105 CFU/mL standards in artificial urine.
4.4. Detection of UTI in Artificial Urine Using the CUBET Assay
The CUBET platform was prepared and set-up in the laboratory. The 200 μL of the negative control (DSMZ (Deutsche Sammlung von Mikroorganismen und Zellkulturen), Artificial Seawater) [37], artificial urine spiked with 105 CFU/mL of uropathogenic E. coli (test sample), and artificial urine without uropathogenic E. coli were added to the wells (in triplicates), mixed with the previously lyophilized cells of P. leiognathi in the CUBET attachment cartridge, and incubated at room temperature for 5 min. After incubation, the cartridge was placed in the area allotted within the CUBET platform. A high-resolution smartphone camera was placed on top of the platform, and, with the use of long exposure photography, a picture was taken and analyzed using ImageJ. 1.x software [46]. The same procedure was performed using a luminometer (CLARIOstar, BMG Labtech; Cary, NC, USA). The two results were compared using one-way ANOVA with a significance cutoff of p ≤ 0.0001.
References
- Thomas, S.T.; Heneghan, C.; Price, C.P.; Van den Bruel, A.; Plüddemann, A. Point-Of-Care Testing for Urinary Tract Infections; Horizon Scan Report 0045; NIHR: Woodstock Road Oxford, UK, 2016. Available online: https://www.community.healthcare.mic.nihr.ac.uk/reports-and-resources/horizonscanning-reports/point-of-care-testing-for-urinary-tract-infections (accessed on 14 July 2020).
- Ivnitski, D.; Abdel-Hamid, I.; Atanasov, P.; Wilkins, E. Biosensors for detection of pathogenic bacteria. Biosens. Bioelectron. 1999, 14, 599–624.
- Davenport, M.; Mach, K.E.; Shortli e, L.M.D.; Banaei, N.; Wang, T.-H.; Liao, J.C. New and developing diagnostic technologies for urinary tract infections. Nat. Rev. Urol. 2017, 14, 296.
- Ho, H.J.; Tan, M.X.; Chen, M.I.; Tan, T.Y.; Koo, S.H.; Koong, A.Y.; Ng, L.P.; Hu, P.L.; Tan, K.T.; Moey, P.K. Interaction between antibiotic resistance, resistance genes, and treatment response for urinary tract infections in primary care. J. Clin. Microbiol. 2019, 57, e00143-19.
- Klein, R.D.; Hultgren, S.J. Urinary tract infections: Microbial pathogenesis, host–pathogen interactions and new treatment strategies. Nat. Rev. Microbiol. 2020, 18, 1–16. [CrossRef] [PubMed]
- Flenker, K.S.; Burghardt, E.L.; Dutta, N.; Burns, W.J.; Grover, J.M.; Kenkel, E.J.; Weaver, T.M.; Mills, J.; Kim, H.; Huang, L. Rapid detection of urinary tract infections via bacterial nuclease activity. Mol. Ther. 2017, 25, 1353–1362.
- Thomas, S.T.; Heneghan, C.; Price, C.P.; Van den Bruel, A.; Plüddemann, A. Point-Of-Care Testing for Urinary Tract Infections; Horizon Scan Report 0045; NIHR: Woodstock Road Oxford, UK, 2016. Available online: https://www.community.healthcare.mic.nihr.ac.uk/reports-and-resources/horizonscanning-reports/point-of-care-testing-for-urinary-tract-infections (accessed on 14 July 2020).
- Ivnitski, D.; Abdel-Hamid, I.; Atanasov, P.; Wilkins, E. Biosensors for detection of pathogenic bacteria. Biosens. Bioelectron. 1999, 14, 599–624.
- Davenport, M.; Mach, K.E.; Shortli e, L.M.D.; Banaei, N.; Wang, T.-H.; Liao, J.C. New and developing diagnostic technologies for urinary tract infections. Nat. Rev. Urol. 2017, 14, 296.
- Rowe, T.A.; Juthani-Mehta, M. Diagnosis and management of urinary tract infection in older adults. Infect. Dis. Clin. N. Am. 2014, 28, 75.
- Rowe, T.A.; Juthani-Mehta, M. Diagnosis and management of urinary tract infection in older adults. Infect. Dis. Clin. N. Am. 2014, 28, 75.
- Grabe, M.; Bjerklund-Johansen, T.; Botto, H.; Çek, M.; Naber, K.; Tenke, P.;Wagenlehner, F. Guidelines on urological infections. Eur. Assoc. Urol. 2015, 182.
- Flenker, K.S.; Burghardt, E.L.; Dutta, N.; Burns, W.J.; Grover, J.M.; Kenkel, E.J.; Weaver, T.M.; Mills, J.; Kim, H.; Huang, L. Rapid detection of urinary tract infections via bacterial nuclease activity. Mol. Ther. 2017, 25, 1353–1362.
- Thomas, S.T.; Heneghan, C.; Price, C.P.; Van den Bruel, A.; Plüddemann, A. Point-Of-Care Testing for Urinary Tract Infections; Horizon Scan Report 0045; NIHR: Woodstock Road Oxford, UK, 2016. Available online: https://www.community.healthcare.mic.nihr.ac.uk/reports-and-resources/horizonscanning-reports/point-of-care-testing-for-urinary-tract-infections (accessed on 14 July 2020).
- Ivnitski, D.; Abdel-Hamid, I.; Atanasov, P.; Wilkins, E. Biosensors for detection of pathogenic bacteria. Biosens. Bioelectron. 1999, 14, 599–624.
- Leigh, D.; Williams, J. Method for the detection of significant bacteriuria in large groups of patients. J. Clin. Pathol. 1964, 17, 498–503. [PubMed]
- Grabe, M.; Bjerklund-Johansen, T.; Botto, H.; Çek, M.; Naber, K.; Tenke, P.;Wagenlehner, F. Guidelines on urological infections. Eur. Assoc. Urol. 2015, 182.