
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Adewale Fadaka | + 3668 word(s) | 3668 | 2021-11-03 05:01:03 | | | |
| 2 | Peter Tang | Meta information modification | 3668 | 2021-11-04 02:50:01 | | |
Video Upload Options
Antimicrobial resistance (AMR) is a significant threat to global health. Antimicrobial peptides (AMPs) have shown potential as alternative diagnostic and therapeutic agents in biomedical applications. Their clinical applications are limited to topical application due to their systemic toxicity, susceptibility to protease degradation, short half-life, and rapid renal clearance. To circumvent these challenges and improve AMP’s efficacy, different approaches such as peptide chemical modifications and the development of AMP delivery systems have been employed. Nanomaterials have been shown to improve the activity of antimicrobial drugs by providing support and synergistic effect against pathogenic microbes.
1. Introduction
2. Antimicrobial Agents and their Activity
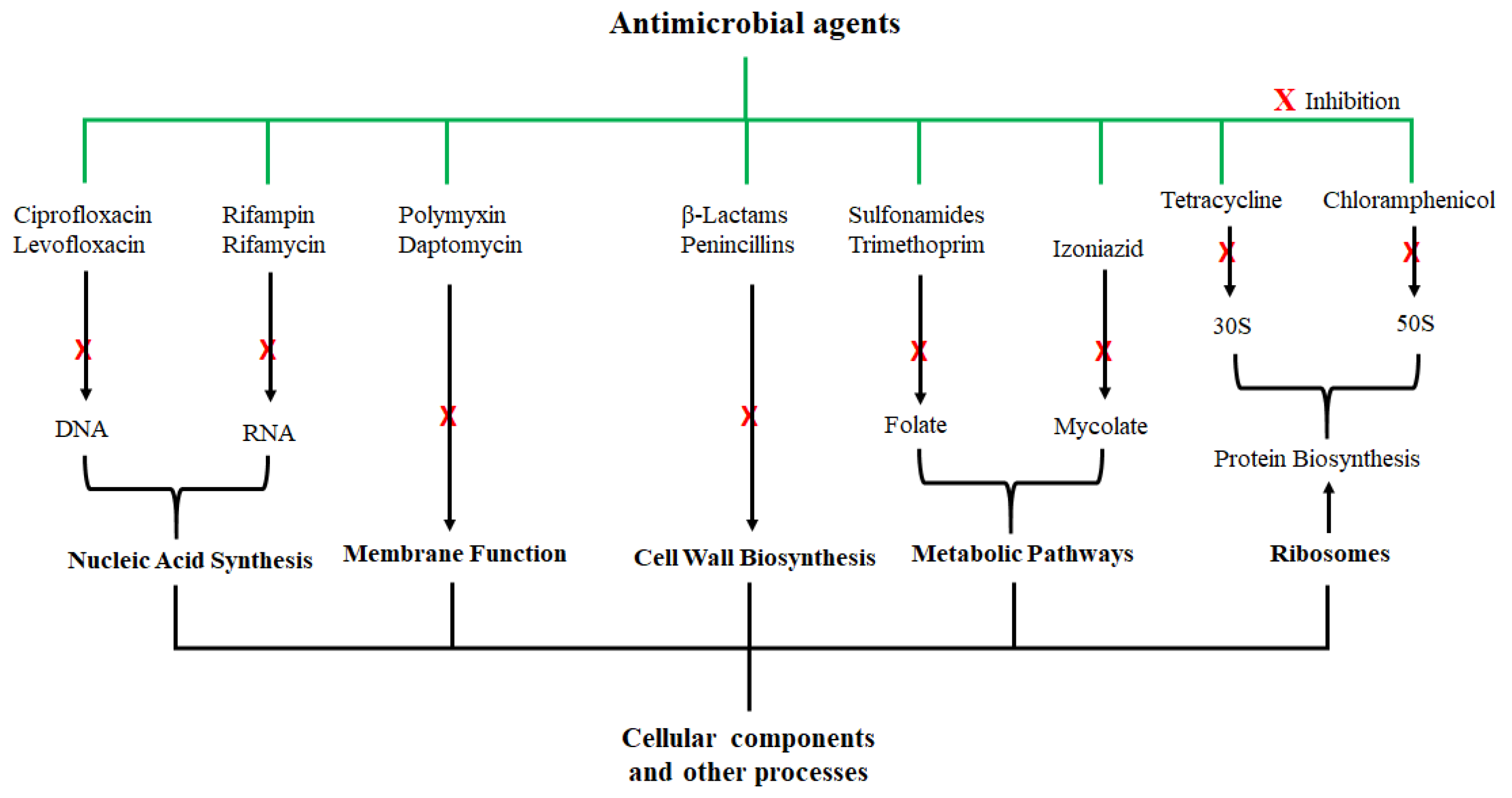
3. Overview and Properties of AMPs

|
Groups |
Characteristics |
Examples |
Mode of Action |
Refs |
|---|---|---|---|---|
|
α-helical peptides |
Amidated C-terminus, N-terminal signal peptides |
FALL-39 Magainins Cecropins |
Pore formation |
[39] |
|
[42] |
||||
|
β-sheet |
cationic with disulfide bridges |
β-defensins |
Membrane disruption |
|
|
plectasin |
[45] |
|||
|
protegrins |
[46] |
|||
|
Extended AMPs or Non-αβ peptides |
Contains proline, arginine, tryptophan, glycine or histidine rich amino acids |
Indolicidin |
Membrane disruption Disruption of intracellular function |
[47] |
|
Bactenecins |
[48] |
|||
|
Histatins |
[49] |
|||
|
Loop peptides |
Dodecapeptides Tachyplesins Protigrin-1 Bactenecin-1 Ranalexin Brevinin 1E Lactoferricin |
Disruption of bacterial membrane |
[50] [53] |
4. NPs with Antimicrobial Activity and Their Mode of Action
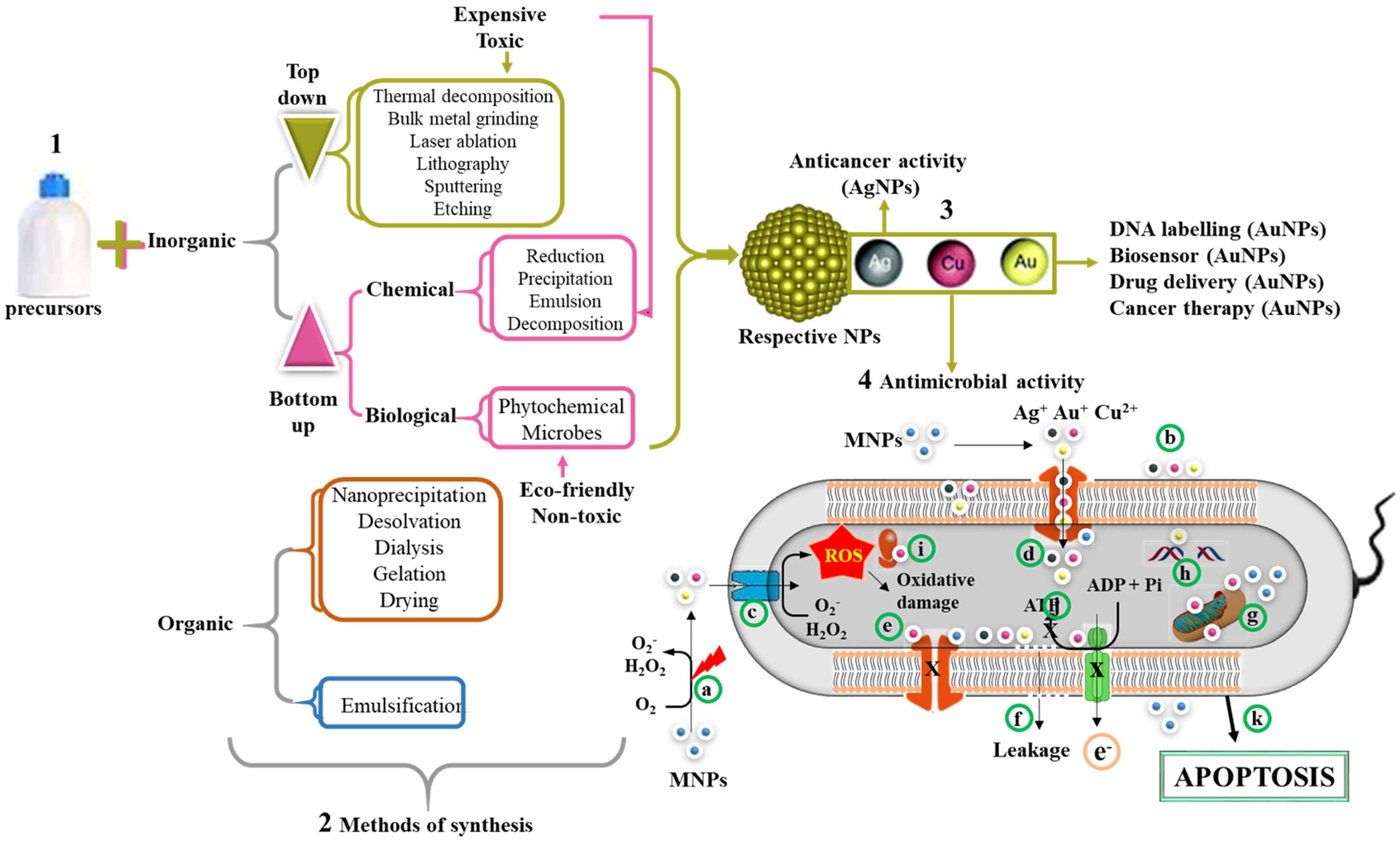
5. Nanocarriers of AMPs
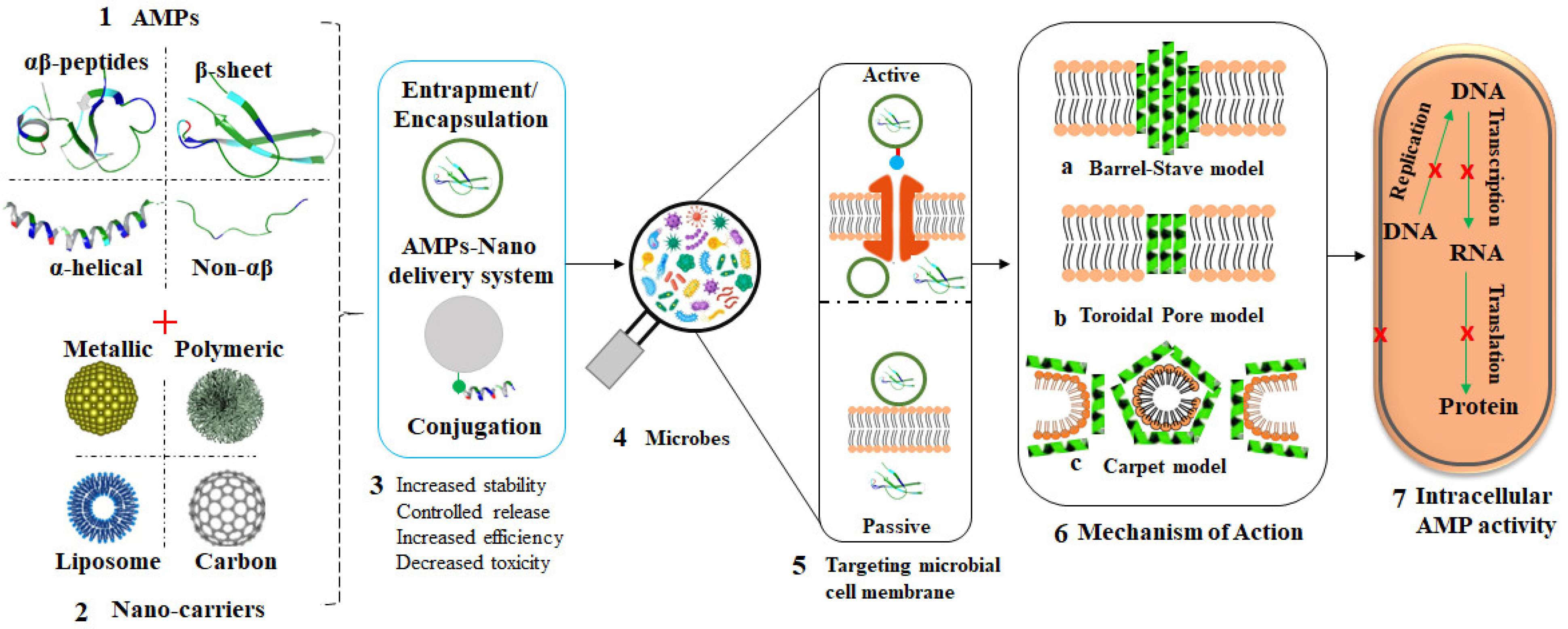
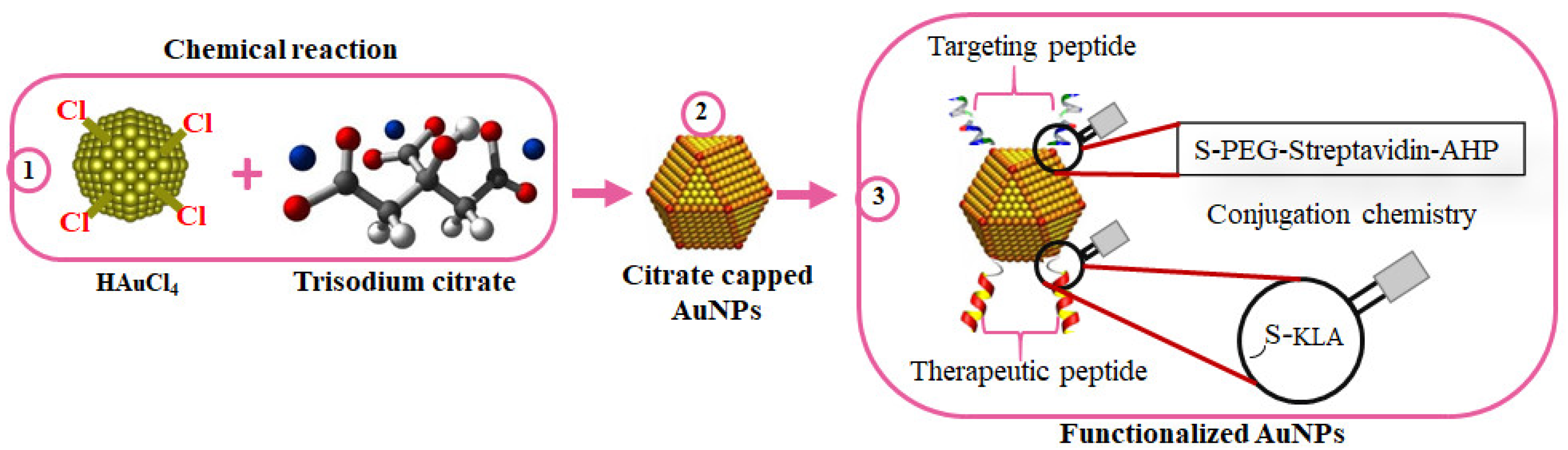
References
- Wall, S. Prevention of antibiotic resistance—An epidemiological scoping review to identify research categories and knowledge gaps. Glob. Health Action 2019, 12, 1756191.
- Sengupta, S.; Chattopadhyay, M.K.; Grossart, H.-P. The multifaceted roles of antibiotics and antibiotic resistance in nature. Front. Microbiol. 2013, 4, 47.
- Spellberg, B.; Gilbert, D.N. The future of antibiotics and resistance: A tribute to a career of leadership by John Bartlett. Clin. Infect. Dis. 2014, 59, S71–S75.
- Centers for Disease Control and Prevention. Office of Infectious Disease. In Antibiotic Resistance Threats in the United States; CDC: Atlanta, GA, USA, 2013; pp. 1–114.
- Ventola, C.L. The antibiotic resistance crisis: Part 1: Causes and threats. Pharm. Ther. 2015, 40, 277.
- Smith, R.A.; M’ikanatha, N.M.; Read, A.F. Antibiotic resistance: A primer and call to action. Health Commun. 2015, 30, 309–314.
- Jasovský, D.; Littmann, J.; Zorzet, A.; Cars, O. Antimicrobial resistance—A threat to the world’s sustainable development. Upsala J. Med. Sci. 2016, 121, 159–164.
- Prestinaci, F.; Pezzotti, P.; Pantosti, A. Antimicrobial resistance: A global multifaceted phenomenon. Pathog. Glob. Health 2015, 109, 309–318.
- Davies, J.; Davies, D. Origins and evolution of antibiotic resistance. Microbiol. Mol. Biol. Rev. 2010, 74, 417–433.
- Nuti, R.; Goud, N.S.; Saraswati, A.P.; Alvala, R.; Alvala, M. Antimicrobial peptides: A promising therapeutic strategy in tackling antimicrobial resistance. Curr. Med. Chem. 2017, 24, 4303–4314.
- Pirtskhalava, M.; Amstrong, A.A.; Grigolava, M.; Chubinidze, M.; Alimbarashvili, E.; Vishnepolsky, B.; Gabrielian, A.; Rosenthal, A.; Hurt, D.E.; Tartakovsky, M. DBAASP v3: Database of antimicrobial/cytotoxic activity and structure of peptides as a resource for development of new therapeutics. Nucleic Acids Res. 2021, 49, D288–D297.
- Wang, Z.; Wang, G. APD: The Antimicrobial Peptide Database. Nucleic Acids Res. 2004, 32, D590–D592.
- Jhong, J.-H.; Chi, Y.-H.; Li, W.-C.; Lin, T.-H.; Huang, K.-Y.; Lee, T.-Y. dbAMP: An integrated resource for exploring antimicrobial peptides with functional activities and physicochemical properties on transcriptome and proteome data. Nucleic Acids Res. 2018, 47, D285–D297.
- Wang, G.; Li, X.; Wang, Z. APD3: The antimicrobial peptide database as a tool for research and education. Nucleic Acids Res. 2016, 44, D1087–D1093.
- Lei, J.; Sun, L.; Huang, S.; Zhu, C.; Li, P.; He, J.; Mackey, V.; Coy, D.H.; He, Q. The antimicrobial peptides and their potential clinical applications. Am. J. Transl. Res. 2019, 11, 3919–3931.
- Mulder, K.; Lima, L.A.; Miranda, V.; Dias, S.C.; Franco, O.L. Current scenario of peptide-based drugs: The key roles of cationic antitumor and antiviral peptides. Front. Microbiol. 2013, 4, 321.
- Jiang, Y.; Yang, D.; Li, W.; Wang, B.; Jiang, Z.; Li, M. Antiviral activity of recombinant mouse β-defensin 3 against influenza A virus in vitro and in vivo. Antivir. Chem. Chemother. 2012, 22, 255–262.
- Pachón-Ibáñez, M.E.; Smani, Y.; Pachón, J.; Sánchez-Céspedes, J. Perspectives for clinical use of engineered human host defense antimicrobial peptides. FEMS Microbiol. Rev. 2017, 41, 323–342.
- Dürr, U.H.; Sudheendra, U.; Ramamoorthy, A. LL-37, the only human member of the cathelicidin family of antimicrobial peptides. Biochim. Biophys. Acta (BBA)—Biomembr. 2006, 1758, 1408–1425.
- Méndez-Samperio, P. The human cathelicidin hCAP18/LL-37: A multifunctional peptide involved in mycobacterial infections. Peptides 2010, 31, 1791–1798.
- Kanthawong, S.; Bolscher, J.G.; Veerman, E.C.; van Marle, J.; de Soet, H.J.; Nazmi, K.; Wongratanacheewin, S.; Taweechaisupapong, S. Antimicrobial and antibiofilm activity of LL-37 and its truncated variants against Burkholderia pseudomallei. Int. J. Antimicrob. Agents 2012, 39, 39–44.
- Vandamme, D.; Landuyt, B.; Luyten, W.; Schoofs, L. A comprehensive summary of LL-37, the factotum human cathelicidin peptide. Cell. Immunol. 2012, 280, 22–35.
- Sørensen, O.E.; Cowland, J.B.; Theilgaard-Mönch, K.; Liu, L.; Ganz, T.; Borregaard, N. Wound healing and expression of antimicrobial peptides/polypeptides in human keratinocytes, a consequence of common growth factors. J. Immunol. 2003, 170, 5583–5589.
- Grossman, P.; Tiefenthaler-Gilmer, U.; Raysz, A.; Kesper, U. Mindfulness training as an intervention for fibromyalgia: Evidence of postintervention and 3-year follow-up benefits in well-being. Psychother. Psychosom. 2007, 76, 226–233.
- Chamorro, C.I.; Weber, G.; Grönberg, A.; Pivarcsi, A.; Ståhle, M. The human antimicrobial peptide LL-37 suppresses apoptosis in keratinocytes. J. Investig. Dermatol. 2009, 129, 937–944.
- Tomasinsig, L.; Pizzirani, C.; Skerlavaj, B.; Pellegatti, P.; Gulinelli, S.; Tossi, A.; Di Virgilio, F.; Zanetti, M. The human cathelicidin LL-37 modulates the activities of the P2X7 receptor in a structure-dependent manner. J. Biol. Chem. 2008, 283, 30471–30481.
- Girnita, A.; Zheng, H.; Grönberg, A.; Girnita, L.; Ståhle, M. Identification of the cathelicidin peptide LL-37 as agonist for the type I insulin-like growth factor receptor. Oncogene 2012, 31, 352–365.
- Ramos, R.; Silva, J.P.; Rodrigues, A.C.; Costa, R.; Guardão, L.; Schmitt, F.; Soares, R.; Vilanova, M.; Domingues, L.; Gama, M. Wound healing activity of the human antimicrobial peptide LL37. Peptides 2011, 32, 1469–1476.
- Nemeth, J.; Oesch, G.; Kuster, S.P. Bacteriostatic versus bactericidal antibiotics for patients with serious bacterial infections: Systematic review and meta-analysis. J. Antimicrob. Chemother. 2014, 70, 382–395.
- Adzitey, F. Antibiotic Classes and Antibiotic Susceptibility of Bacterial Isolates from Selected Poultry; A Mini Review. World’s Vet. J. 2015, 5, 36–41.
- Grossman, T.H. Tetracycline Antibiotics and Resistance. Cold Spring Harb. Perspect. Med. 2016, 6, a025387.
- Mendes, R.E.; Farrell, D.J.; Sader, H.S.; Streit, J.M.; Jones, R.N. Update of the telavancin activity in vitro tested against a worldwide collection of Gram-positive clinical isolates (2013), when applying the revised susceptibility testing method. Diagn. Microbiol. Infect. Dis. 2015, 81, 275–279.
- Mallapragada, S.; Wadhwa, A.; Agrawal, P. Antimicrobial peptides: The miraculous biological molecules. J. Indian Soc. Periodontol. 2017, 21, 434.
- Hancock, R.E. Peptide antibiotics. Lancet 1997, 349, 418–422.
- Tennessen, J. Molecular evolution of animal antimicrobial peptides: Widespread moderate positive selection. J. Evol. Biol. 2005, 18, 1387–1394.
- Li, A.; Lee, P.; Ho, B.; Ding, J.; Lim, C. Atomic force microscopy study of the antimicrobial action of Sushi peptides on Gram negative bacteria. Biochim. Biophys. Acta (BBA)—Biomembr. 2007, 1768, 411–418.
- Meincken, M.; Holroyd, D.; Rautenbach, M. Atomic force microscopy study of the effect of antimicrobial peptides on the cell envelope of Escherichia coli. Antimicrob. Agents Chemother. 2005, 49, 4085–4092.
- Pushpanathan, M.; Gunasekaran, P.; Rajendhran, J. Antimicrobial Peptides: Versatile Biological Properties. Int. J. Pept. 2013, 2013, 675391.
- Agerberth, B.; Gunne, H.; Odeberg, J.; Kogner, P.; Boman, H.G.; Gudmundsson, G.H. FALL-39, a putative human peptide antibiotic, is cysteine-free and expressed in bone marrow and testis. Proc. Natl. Acad. Sci. USA 1995, 92, 195–199.
- Berkowitz, B.A.; Bevins, C.L.; Zasloff, M.A. Magainins: A new family of membrane-active host defense peptides. Biochem. Pharmacol. 1990, 39, 625–629.
- Zasloff, M. Magainins, a class of antimicrobial peptides from Xenopus skin: Isolation, characterization of two active forms, and partial cDNA sequence of a precursor. Proc. Natl. Acad. Sci. USA 1987, 84, 5449–5453.
- Andersson, M.; Boman, A.; Boman, H. Ascaris nematodes from pig and human make three anti-bacterial peptides: Isolation of cecropin P1 and two ASABF peptides. Cell. Mol. Life Sci. 2003, 60, 599–606.
- Nguyen, L.T.; Haney, E.F.; Vogel, H.J. The expanding scope of antimicrobial peptide structures and their modes of action. Trends Biotechnol. 2011, 29, 464–472.
- Ganz, T. Defensins: Antimicrobial peptides of innate immunity. Nat. Rev. Immunol. 2003, 3, 710–720.
- Mygind, P.H.; Fischer, R.L.; Schnorr, K.M.; Hansen, M.T.; Sönksen, C.P.; Ludvigsen, S.; Raventós, D.; Buskov, S.; Christensen, B.; De Maria, L. Plectasin is a peptide antibiotic with therapeutic potential from a saprophytic fungus. Nature 2005, 437, 975–980.
- Kokryakov, V.N.; Harwig, S.S.; Panyutich, E.A.; Shevchenko, A.A.; Aleshina, G.M.; Shamova, O.V.; Korneva, H.A.; Lehrer, R.I. Protegrins: Leukocyte antimicrobial peptides that combine features of corticostatic defensins and tachyplesins. FEBS Lett. 1993, 327, 231–236.
- Falla, T.J.; Karunaratne, D.N.; Hancock, R.E. Mode of action of the antimicrobial peptide indolicidin. J. Biol. Chem. 1996, 271, 19298–19303.
- Frank, R.W.; Gennaro, R.; Schneider, K.; Przybylski, M.; Romeo, D. Amino acid sequences of two proline-rich bactenecins. Antimicrobial peptides of bovine neutrophils. J. Biol. Chem. 1990, 265, 18871–18874.
- Takahashi, D.; Shukla, S.K.; Prakash, O.; Zhang, G. Structural determinants of host defense peptides for antimicrobial activity and target cell selectivity. Biochimie 2010, 92, 1236–1241.
- Dings, R.P.; Haseman, J.R.; Leslie, D.B.; Luong, M.; Dunn, D.L.; Mayo, K.H. Bacterial membrane disrupting dodecapeptide SC4 improves survival of mice challenged with Pseudomonas aeruginosa. Biochim. Et Biophys. Acta (Bba)-Gen. Subj. 2013, 1830, 3454–3457.
- Nakamura, T.; Furunaka, H.; Miyata, T.; Tokunaga, F.; Muta, T.; Iwanaga, S.; Niwa, M.; Takao, T.; Shimonishi, Y. Tachyplesin, a class of antimicrobial peptide from the hemocytes of the horseshoe crab (Tachypleus tridentatus). Isolation and chemical structure. J. Biol. Chem. 1988, 263, 16709–16713.
- Xu, F.; Meng, K.; Wang, Y.-R.; Luo, H.-Y.; Yang, P.-L.; Wu, N.-F.; Fan, Y.-L.; Yao, B. Eukaryotic expression and antimicrobial spectrum determination of the peptide tachyplesin II. Protein Expr. Purif. 2008, 58, 175–183.
- Di Giulio, A.; Zhao, H. Antimicrobial peptides: Basic mechanisms of action and emerging pharmacological interest. Asian J. Biochem. 2006, 1, 28–40.
- Brogden, K.A. Antimicrobial peptides: Pore formers or metabolic inhibitors in bacteria? Nat. Rev. Microbiol. 2005, 3, 238–250.
- Roversi, D.; Luca, V.; Aureli, S.; Park, Y.; Mangoni, M.L.; Stella, L. How many antimicrobial peptide molecules kill a bacterium? The case of PMAP-23. ACS Chem. Biol. 2014, 9, 2003–2007.
- Marr, A.K.; Gooderham, W.J.; Hancock, R.E. Antibacterial peptides for therapeutic use: Obstacles and realistic outlook. Curr. Opin. Pharmacol. 2006, 6, 468–472.
- Yount, N.Y.; Yeaman, M.R. Emerging themes and therapeutic prospects for anti-infective peptides. Annu. Rev. Pharmacol. Toxicol. 2012, 52, 337–360.
- Fox, J.L. Antimicrobial peptides stage a comeback: Better understanding of the mechanisms of action, modification and synthesis of antimicrobial peptides is reigniting commercial development. Nat. Biotechnol. 2013, 31, 379–383.
- Riool, M.; de Breij, A.; de Boer, L.; Kwakman, P.H.; Cordfunke, R.A.; Cohen, O.; Malanovic, N.; Emanuel, N.; Lohner, K.; Drijfhout, J.W. Controlled Release of LL-37-Derived Synthetic Antimicrobial and Anti-Biofilm Peptides SAAP-145 and SAAP-276 Prevents Experimental Biomaterial-Associated Staphylococcus aureus Infection. Adv. Funct. Mater 2017, 27, 1606623.
- Correa, M.G.; Martínez, F.B.; Vidal, C.P.; Streitt, C.; Escrig, J.; de Dicastillo, C.L. Antimicrobial metal-based nanoparticles: A review on their synthesis, types and antimicrobial action. Beilstein J. Nanotechnol. 2020, 11, 1450–1469.
- Alves, M.M.; Bouchami, O.; Tavares, A.; Córdoba, L.; Santos, C.F.; Miragaia, M.; de Fátima Montemor, M. New insights into antibiofilm effect of a nanosized ZnO coating against the pathogenic methicillin resistant Staphylococcus aureus. ACS Appl. Mater. Interfaces 2017, 9, 28157–28167.
- Oun, A.A.; Rhim, J.-W. Carrageenan-based hydrogels and films: Effect of ZnO and CuO nanoparticles on the physical, mechanical, and antimicrobial properties. Food Hydrocoll. 2017, 67, 45–53.
- Shankar, S.; Rhim, J.-W. Facile approach for large-scale production of metal and metal oxide nanoparticles and preparation of antibacterial cotton pads. Carbohydr. Polym. 2017, 163, 137–145.
- Li, J.; Cha, R.; Zhao, X.; Guo, H.; Luo, H.; Wang, M.; Zhou, F.; Jiang, X.J.A.N. Gold nanoparticles cure bacterial infection with benefit to intestinal microflora. ACS Nano 2019, 13, 5002–5014.
- Akbar, A.; Sadiq, M.B.; Ali, I.; Muhammad, N.; Rehman, Z.; Khan, M.N.; Muhammad, J.; Khan, S.A.; Rehman, F.U.; Anal, A.K. Synthesis and antimicrobial activity of zinc oxide nanoparticles against foodborne pathogens Salmonella typhimurium and Staphylococcus aureus. Biocatal. Agric. Biotechnol. 2019, 17, 36–42.
- Chen, J.; Wu, L.; Lu, M.; Lu, S.; Li, Z.; Ding, W. Comparative Study on the Fungicidal Activity of Metallic MgO Nanoparticles and Macroscale MgO Against Soilborne Fungal Phytopathogens. Front. Microbiol. 2020, 11, 365.
- Guilger-Casagrande, M.; Lima, R.D. Synthesis of Silver Nanoparticles Mediated by Fungi: A Review. Front. Bioeng. Biotechnol. 2019, 7, 287.
- Swain, P.; Nayak, S.K.; Sasmal, A.; Behera, T.; Barik, S.K.; Swain, S.K.; Mishra, S.S.; Sen, A.K.; Das, J.K.; Jayasankar, P. Antimicrobial activity of metal based nanoparticles against microbes associated with diseases in aquaculture. World J. Microbiol. Biotechnol. 2014, 30, 2491–2502.
- Mallmann EJ, J.; Cunha, F.A.; Castro, B.N.; Maciel, A.M.; Menezes, E.A.; Fechine, P.B.A. Antifungal activity of silver nanoparticles obtained by green synthesis. Rev. Inst. Med. Trop. São Paulo 2015, 57, 165–167.
- Galdiero, S.; Falanga, A.; Vitiello, M.; Cantisani, M.; Marra, V.; Galdiero, M. Silver nanoparticles as potential antiviral agents. Molecules 2011, 16, 8894–8918.
- Sharmin, S.; Rahaman, M.M.; Sarkar, C.; Atolani, O.; Islam, M.T.; Adeyemi, O.S. Nanoparticles as antimicrobial and antiviral agents: A literature-based perspective study. Heliyon 2021, 7, e06456.
- Erkoc, P.; Ulucan-Karnak, F.J.P. Nanotechnology-Based Antimicrobial and Antiviral Surface Coating Strategies. Prosthesis 2021, 3, 25–52.
- Deepika, M.S.; Thangam, R.; Sundarraj, S.; Sheena, T.S.; Sivasubramanian, S.; Kulandaivel, J.; Thirumurugan, R. Co-delivery of diverse therapeutic compounds using PEG–PLGA nanoparticle cargo against drug-resistant bacteria: An improved anti-biofilm strategy. ACS Appl. Bio Mater. 2019, 3, 385–399.
- Durak, S.; Arasoglu, T.; Ates, S.C.; Derman, S. Enhanced antibacterial and antiparasitic activity of multifunctional polymeric nanoparticles. Nanotechnology 2020, 31, 175705.
- Alamdaran, M.; Movahedi, B.; Mohabatkar, H.; Behbahani, M. In-vitro study of the novel nanocarrier of chitosan-based nanoparticles conjugated HIV-1 P24 protein-derived peptides. J. Mol. Liq. 2018, 265, 243–250.
- Real, D.; Hoffmann, S.; Leonardi, D.; Salomon, C.; Goycoolea, F.M. Chitosan-based nanodelivery systems applied to the development of novel triclabendazole formulations. PLoS ONE 2018, 13, e0207625.
- Zafar, N.; Shamaila, S.; Nazir, J.; Sharif, R.; Rafique, M.S.; Ul-Hasan, J.; Ammara, S.; Khalid, H. Antibacterial action of chemically synthesized and laser generated silver nanoparticles against human pathogenic bacteria. J. Mater. Sci. Technol. 2016, 32, 721–728.
- Khalid, H.; Shamaila, S.; Zafar, N.; Shahzadi, S. Synthesis of copper nanoparticles by chemical reduction method. Sci. Int. 2015, 27, 3085–3088.
- Adewale, O.B.; Anadozie, S.O.; Potts-Johnson, S.S.; Onwuelu, J.O.; Obafemi, T.O.; Osukoya, O.A.; Fadaka, A.O.; Davids, H.; Roux, S. Investigation of bioactive compounds in Crassocephalum rubens leaf and in vitro anticancer activity of its biosynthesized gold nanoparticles. Biotechnol. Rep. 2020, 28, e00560.
- Pelgrift, R.Y.; Friedman, A.J. Nanotechnology as a therapeutic tool to combat microbial resistance. Adv. Drug Deliv. Rev. 2013, 65, 1803–1815.
- Ivask, A.; Kurvet, I.; Kasemets, K.; Blinova, I.; Aruoja, V.; Suppi, S.; Vija, H.; Käkinen, A.; Titma, T.; Heinlaan, M. Size-dependent toxicity of silver nanoparticles to bacteria, yeast, algae, crustaceans and mammalian cells in vitro. PLoS ONE 2014, 9, e102108.
- Azam, A.; Ahmed, A.S.; Oves, M.; Khan, M.; Memic, A. Size-dependent antimicrobial properties of CuO nanoparticles against Gram-positive and-negative bacterial strains. Int. J. Nanomed. 2012, 7, 3527.
- Natan, M.; Banin, E. From Nano to Micro: Using nanotechnology to combat microorganisms and their multidrug resistance. Fems Microbiol. Rev. 2017, 41, 302–322.
- Popa, M.; Pradell, T.; Crespo, D.; Calderón-Moreno, J.M. Stable silver colloidal dispersions using short chain polyethylene glycol. Colloids Surf. A Physicochem. Eng. Asp. 2007, 303, 184–190.
- Tolaymat, T.M.; El Badawy, A.M.; Genaidy, A.; Scheckel, K.G.; Luxton, T.P.; Suidan, M. An evidence-based environmental perspective of manufactured silver nanoparticle in syntheses and applications: A systematic review and critical appraisal of peer-reviewed scientific papers. Sci. Total Environ. 2010, 408, 999–1006.
- Aboyewa, J.A.; Sibuyi, N.R.; Meyer, M.; Oguntibeju, O.O. Green Synthesis of Metallic Nanoparticles Using Some Selected Medicinal Plants from Southern Africa and Their Biological Applications. Plants 2021, 10, 1929.
- Sibuyi, N.R.S.; Thipe, V.C.; Panjtan-Amiri, K.; Meyer, M.; Katti, K.V. Green synthesis of gold nanoparticles using Acai berry and Elderberry extracts and investigation of their effect on prostate and pancreatic cancer cells. Nanobiomedicine 2021, 8, 1849543521995310.
- Adewale, O.B.; Egbeyemi, K.A.; Onwuelu, J.O.; Potts-Johnson, S.S.; Anadozie, S.O.; Fadaka, A.O.; Osukoya, O.A.; Aluko, B.T.; Johnson, J.; Obafemi, T.O. Biological synthesis of gold and silver nanoparticles using leaf extracts of Crassocephalum rubens and their comparative in vitro antioxidant activities. Heliyon 2020, 6, e05501.
- Monowar, T.; Rahman, M.S.; Bhore, S.J.; Raju, G.; Sathasivam, K.V. Silver Nanoparticles Synthesized by Using the Endophytic Bacterium Pantoea ananatis are Promising Antimicrobial Agents against Multidrug Resistant Bacteria. Molecules 2018, 23, 3220.
- Escárcega-González, C.E.; Garza-Cervantes, J.A.; Vazquez-Rodríguez, A.; Montelongo-Peralta, L.Z.; Treviño-Gonzalez, M.T.; Castro, E.D.B.; Saucedo-Salazar, E.M.; Morales, R.C.; Soto, D.R.; González, F.T. In vivo antimicrobial activity of silver nanoparticles produced via a green chemistry synthesis using Acacia rigidula as a reducing and capping agent. Int. J. Nanomed. 2018, 13, 2349.
- Lopez-Abarrategui, C.; Figueroa-Espi, V.; Lugo-Alvarez, M.B.; Pereira, C.D.; Garay, H.; Barbosa, J.A.; Falcão, R.; Jiménez-Hernández, L.; Estévez-Hernández, O.; Reguera, E.; et al. The intrinsic antimicrobial activity of citric acid-coated manganese ferrite nanoparticles is enhanced after conjugation with the antifungal peptide Cm-p5. Int. J. Nanomed. 2016, 11, 3849–3857.
- Brancolini, G.; Kokh, D.B.; Calzolai, L.; Wade, R.C.; Corni, S. Docking of ubiquitin to gold nanoparticles. ACS Nano 2012, 6, 9863–9878.
- Kamar, R.; Réjasse, A.; Jéhanno, I.; Attieh, Z.; Courtin, P.; Chapot-Chartier, M.-P.; Nielsen-Leroux, C.; Lereclus, D.; El Chamy, L.; Kallassy, M. DltX of Bacillus thuringiensis is essential for D-alanylation of teichoic acids and resistance to antimicrobial response in insects. Front. Microbiol. 2017, 8, 1437.
- Meireles, D.; Pombinho, R.; Carvalho, F.; Sousa, S.; Cabanes, D. Listeria monocytogenes wall teichoic acid glycosylation promotes surface anchoring of virulence factors, resistance to antimicrobial peptides, and decreased susceptibility to antibiotics. Pathogens 2020, 9, 290.
- Joo, H.-S.; Otto, M. Mechanisms of resistance to antimicrobial peptides in staphylococci. Biochim. Biophys. Acta (BBA)—Biomembr. 2015, 1848, 3055–3061.
- Joo, H.-S.; Fu, C.-I.; Otto, M. Bacterial strategies of resistance to antimicrobial peptides. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2016, 371, 20150292.
- Spohn, R.; Daruka, L.; Lázár, V.; Martins, A.; Vidovics, F.; Grézal, G.; Méhi, O.; Kintses, B.; Számel, M.; Jangir, P.K.; et al. Integrated evolutionary analysis reveals antimicrobial peptides with limited resistance. Nat. Commun. 2019, 10, 4538.
- Rodríguez-Rojas, A.; Baeder, D.Y.; Johnston, P.; Regoes, R.R.; Rolff, J. Bacteria primed by antimicrobial peptides develop tolerance and persist. PLoS Pathog. 2021, 17, e1009443.
- Baindara, P.; Ghosh, A.K.; Mandal, S.M. Coevolution of resistance against antimicrobial peptides. Microb. Drug Resist. 2020, 26, 880–899.
- Andersson, D.I.; Hughes, D.; Kubicek-Sutherland, J.Z. Mechanisms and consequences of bacterial resistance to antimicrobial peptides. Drug Resist. Updates 2016, 26, 43–57.
- El Shazely, B.; Yu, G.; Johnston, P.R.; Rolff, J. Resistance Evolution Against Antimicrobial Peptides in Staphylococcus aureus Alters Pharmacodynamics Beyond the MIC. Front. Microbiol. 2020, 11.
- Mahlapuu, M.; Håkansson, J.; Ringstad, L.; Björn, C. Antimicrobial peptides: An emerging category of therapeutic agents. Front. Cell. Infect. Microbiol. 2016, 6, 194.
- Moravej, H.; Moravej, Z.; Yazdanparast, M.; Heiat, M.; Mirhosseini, A.; Moosazadeh Moghaddam, M.; Mirnejad, R. Antimicrobial peptides: Features, action, and their resistance mechanisms in bacteria. Microb. Drug Resist. 2018, 24, 747–767.
- Abo-zeid, Y.; Williams, G.R. The potential anti-infective applications of metal oxide nanoparticles: A systematic review. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2020, 12, e1592.
- Beyth, N.; Houri-Haddad, Y.; Domb, A.; Khan, W.; Hazan, R. Alternative antimicrobial approach: Nano-antimicrobial materials. Evid.-Based Complement. Altern. Med. 2015, 2015, 246012.
- Caster, J.M.; Patel, A.N.; Zhang, T.; Wang, A. Investigational nanomedicines in 2016: A review of nanotherapeutics currently undergoing clinical trials. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2017, 9, e1416.
- Smerkova, K.; Dolezelikova, K.; Bozdechova, L.; Heger, Z.; Zurek, L.; Adam, V. Nanomaterials with active targeting as advanced antimicrobials. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2020, 12, e1636.
- Jelinkova, P.; Mazumdar, A.; Sur, V.P.; Kociova, S.; Dolezelikova, K.; Jimenez, A.M.J.; Koudelkova, Z.; Mishra, P.K.; Smerkova, K.; Heger, Z. Nanoparticle-drug conjugates treating bacterial infections. J. Control. Release 2019, 307, 166–185.
- Baier, G.; Cavallaro, A.; Vasilev, K.; Mailänder, V.; Musyanovych, A.; Landfester, K. Enzyme responsive hyaluronic acid nanocapsules containing polyhexanide and their exposure to bacteria to prevent infection. Biomacromolecules 2013, 14, 1103–1112.
- Boas, U.; Heegaard, P.M. Dendrimers in drug research. Chem. Soc. Rev. 2004, 33, 43–63.
- Esmaeili, F.; Hosseini-Nasr, M.; Rad-Malekshahi, M.; Samadi, N.; Atyabi, F.; Dinarvand, R. Preparation and antibacterial activity evaluation of rifampicin-loaded poly lactide-co-glycolide nanoparticles. Nanomed. Nanotechnol. Biol. Med. 2007, 3, 161–167.
- Schumacher, I.; Margalit, R. Liposome-encapsulated ampicillin: Physicochemical and antibacterial properties. J. Pharm. Sci. 1997, 86, 635–641.
- Torchilin, V.P. Micellar nanocarriers: Pharmaceutical perspectives. Pharm. Res. 2007, 24, 1–16.
- Amina, S.J.; Guo, B. A Review on the Synthesis and Functionalization of Gold Nanoparticles as a Drug Delivery Vehicle. Int. J. Nanomed. 2020, 15, 9823–9857.
- Murphy, C.J.; Gole, A.M.; Stone, J.W.; Sisco, P.N.; Alkilany, A.M.; Goldsmith, E.C.; Baxter, S.C. Gold nanoparticles in biology: Beyond toxicity to cellular imaging. Acc. Chem. Res. 2008, 41, 1721–1730.
- Lee, K.-S.; El-Sayed, M.A. Gold and silver nanoparticles in sensing and imaging: Sensitivity of plasmon response to size, shape, and metal composition. J. Phys. Chem. B 2006, 110, 19220–19225.
- Toderas, F.; Baia, M.; Maniu, D.; Astilean, S. Tuning the plasmon resonances of gold nanoparticles by controlling their size and shape. J. Optoelectron. Adv. Mater. 2008, 10, 2282–2284.
- Boisselier, E.; Astruc, D. Gold nanoparticles in nanomedicine: Preparations, imaging, diagnostics, therapies and toxicity. Chem. Soc. Rev. 2009, 38, 1759–1782.
- Tran, Q.H.; Le, A.-T. Silver nanoparticles: Synthesis, properties, toxicology, applications and perspectives. Adv. Nat. Sci. Nanosci. Nanotechnol. 2013, 4, 033001.
- Shen, J.; Yu, M.; Meng, Q.; Li, J.; Lv, Y.; Lu, W. Fatty acid-based strategy for efficient brain targeted gene delivery. Pharm. Res. 2013, 30, 2573–2583.
- Nikanjam, M.; Gibbs, A.R.; Hunt, C.A.; Budinger, T.F.; Forte, T.M. Synthetic nano-LDL with paclitaxel oleate as a targeted drug delivery vehicle for glioblastoma multiforme. J. Control. Release 2007, 124, 163–171.
- Dhar, S.; Gu, F.X.; Langer, R.; Farokhzad, O.C.; Lippard, S.J. Targeted delivery of cisplatin to prostate cancer cells by aptamer functionalized Pt(IV) prodrug-PLGA-PEG nanoparticles. Proc. Natl. Acad. Sci. USA 2008, 105, 17356–17361.
- Górski, A.; Wasik, M.; Nowaczyk, M.; Korczak-Kowalska, G. Immunomodulating activity of heparin. FASEB J. 1991, 5, 2287–2291.
- Zhang, H.; Luo, J.; Li, Y.; Henderson, P.T.; Wang, Y.; Wachsmann-Hogiu, S.; Zhao, W.; Lam, K.S.; Pan, C.X. Characterization of high-affinity peptides and their feasibility for use in nanotherapeutics targeting leukemia stem cells. Nanomedicine 2012, 8, 1116–1124.
- Liao, D.; Liu, Z.; Wrasidlo, W.; Chen, T.; Luo, Y.; Xiang, R.; Reisfeld, R.A. Synthetic enzyme inhibitor: A novel targeting ligand for nanotherapeutic drug delivery inhibiting tumor growth without systemic toxicity. Nanomed. Nanotechnol. Biol. Med. 2011, 7, 665–673.
- Chikkaveeraiah, B.V.; Soldà, A.; Choudhary, D.; Maran, F.; Rusling, J.F. Ultrasensitive nanostructured immunosensor for stem and carcinoma cell pluripotency gatekeeper protein NANOG. Nanomedicine 2012, 7, 957–965.
- Lee, G.Y.; Kim, J.H.; Oh, G.T.; Lee, B.H.; Kwon, I.C.; Kim, I.S. Molecular targeting of atherosclerotic plaques by a stabilin-2-specific peptide ligand. J. Control. Release 2011, 155, 211–217.
- Almer, G.; Wernig, K.; Saba-Lepek, M.; Haj-Yahya, S.; Rattenberger, J.; Wagner, J.; Gradauer, K.; Frascione, D.; Pabst, G.; Leitinger, G.; et al. Adiponectin-coated nanoparticles for enhanced imaging of atherosclerotic plaques. Int. J. Nanomed. 2011, 6, 1279–1290.
- Li, N.; Larson, T.; Nguyen, H.H.; Sokolov, K.V.; Ellington, A.D. Directed evolution of gold nanoparticle delivery to cells. Chem. Commun. 2010, 46, 392–394.
- Iqbal, M.Z.; Ali, I.; Khan, W.S.; Kong, X.; Dempsey, E. Reversible self-assembly of gold nanoparticles in response to external stimuli. Mater. Des. 2021, 205, 109694.
- Park, S.; Lee, W.J.; Park, S.; Choi, D.; Kim, S.; Park, N. Reversibly pH-responsive gold nanoparticles and their applications for photothermal cancer therapy. Sci. Rep. 2019, 9, 1–9.
- Bernardim, B.; Matos, M.J.; Ferhati, X.; Compañón, I.; Guerreiro, A.; Akkapeddi, P.; Burtoloso, A.C.; Jiménez-Osés, G.; Corzana, F.; Bernardes, G.J. Efficient and irreversible antibody–cysteine bioconjugation using carbonylacrylic reagents. Nat. Protoc. 2019, 14, 86–99.
- S Sibuyi, N.R.; Thovhogi, N.; Gabuza, K.B.; Meyer, M.D.; Drah, M.; Onani, M.O.; Skepu, A.; Madiehe, A.M.; Meyer, M. Peptide-functionalized nanoparticles for the selective induction of apoptosis in target cells. Nanomedicine 2017, 12, 1631–1645.
- Hossen, M.N.; Kajimoto, K.; Akita, H.; Hyodo, M.; Ishitsuka, T.; Harashima, H. Ligand-based targeted delivery of a peptide modified nanocarrier to endothelial cells in adipose tissue. J. Control. Release 2010, 147, 261–268.





