Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Craig Vierra | + 1586 word(s) | 1586 | 2021-08-25 11:41:59 | | | |
| 2 | Camila Xu | -10 word(s) | 1576 | 2021-09-09 03:57:55 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Vierra, C. Black Widow Spider Tubuliform Silk. Encyclopedia. Available online: https://encyclopedia.pub/entry/14019 (accessed on 09 May 2026).
Vierra C. Black Widow Spider Tubuliform Silk. Encyclopedia. Available at: https://encyclopedia.pub/entry/14019. Accessed May 09, 2026.
Vierra, Craig. "Black Widow Spider Tubuliform Silk" Encyclopedia, https://encyclopedia.pub/entry/14019 (accessed May 09, 2026).
Vierra, C. (2021, September 08). Black Widow Spider Tubuliform Silk. In Encyclopedia. https://encyclopedia.pub/entry/14019
Vierra, Craig. "Black Widow Spider Tubuliform Silk." Encyclopedia. Web. 08 September, 2021.
Copy Citation
Spider silk is a high-performance fiber renowned for its extraordinary mechanical properties, rivaling steel, Kevlar, and a wide range of other natural and manmade materials.
spider silk
spidroins
tubuliform silk
black widow spider
1. Introduction
Spider silk is a high-performance fiber renowned for its extraordinary mechanical properties, rivaling steel, Kevlar, and a wide range of other natural and manmade materials [1][2]. The western black widow, Latrodectus hesperus, has become one of the most extensively studied spider species with respect to silk gene expression and fiber constituents, both at the level of transcriptomics and proteomics [3][4][5][6][7][8][9][10][11]. Black widow spiders, which are cobweavers, are members of the superfamily Araneoidea. Morphological studies of female black widow spiders reveal seven distinct silk-producing abdominal glands, including the major ampullate, minor ampullate, tubuliform, aciniform, pyriform, aggregate, and flagelliform glands [12][13]. Each gland produces fibers or glues with distinct mechanical and biomolecular properties that are highly tuned to specific biological functions [14]. Differences in the mechanical properties of the threads are largely due to altered protein compositions, which are strongly correlated with differential expression of structural proteins called spidroins, a contraction of the words spider and fibroin [15][16]. Spidroins, therefore, represent a family of spider-specific fibroins that have significant roles in fiber mechanics. Members of this family have common biochemical characteristics, including large molecular masses with internal block repeats and non-repetitive N- and C-terminal domains within their protein sequences [16][17][18][19]. Although the spidroin family protein architectures all share internal block repeats, their specific amino acid sequences and lengths differ, and these differences in primary amino acid sequences have been a focal point of the scientific community to establish relationships between protein sequence, structure, and fiber mechanics [20][21][22].
The major ampullate and tubuliform glands have different morphological features, but both lumens store a liquid spinning dope that is extruded into a dry fiber via external spigots on the spider’s spinnerets. Despite morphological distinctions, these two glands are reported to share similar spidroin gene expression profiles [3]. In black widow spiders, the major ampullate gland extrudes dragline silk (also called major ampullate silk or MA silk), a fiber type used for locomotion and web construction, whereas the tubuliform gland manufactures the main constituent of egg cases, tubuliform silk. Egg cases, which also contain aciniform silk, enclose eggs and protect them from predators and environmental fluctuations, such as changes in temperature and humidity [23]. Tubuliform silk has been shown to contain the spidroin TuSp1, along with two non-spidroin proteins called egg case protein 1 and 2 (ECP1 and ECP2) [6][24][25][26]. Additionally, a recent proteomic analysis of tubuliform glands, along with deep transcriptome sequencing studies to identify silk-gland specific transcripts (SSTs), has led to the discovery of 14 SST proteins in tubuliform glands of black widow spiders [5]. Two of these 14 SST proteins have been previously shown to be constituents of tubuliform silk, which include ECP1 and TuSp1 [6][24][25][26]. However, it is unclear whether the other 12 SST proteins are extruded from the tubuliform gland, or alternatively, whether these factors represent non-extruded proteins that participate in the spider silk assembly pathway. To further emphasize the importance of this work, the identification of novel constituents of spider silk, specifically tubuliform silk, has significant value for the spider silk community, as insight into the ingredients that comprise natural silk has direct implications on synthetic silk production and its potential use for military, industrial, and engineering applications.
2. Proteomic Analysis of Tubuliform Glands, Spinning Dope, and Egg Sacs
In order to perform a comprehensive, in-depth analysis of the constituents of tubuliform silk, we developed a generalized proteomic workflow designed to analyze proteins expressed in tubuliform glands, the spinning dope, which represents the liquid components within the lumen of the glands and the egg sacs of black widow spiders using mass spectrometry (Figure 1). Scanning electron microscope studies of black widow spider egg cases have previously revealed two distinct fiber types [25]. The diameter sizes are approximately 5 μm and 0.5 μm, corresponding to tubuliform and aciniform silk, respectively, with the majority of the volume comprising tubuliform silk (Figure 2A,B).
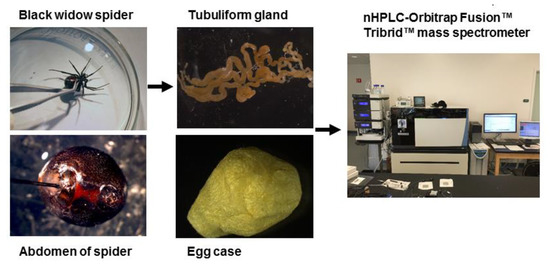
Figure 1. General workflow for isolation of the tubuliform gland, egg sac, and dope (liquid removed from the tubuliform glands), followed by analysis of the tryptic peptides by MS/MS analysis.
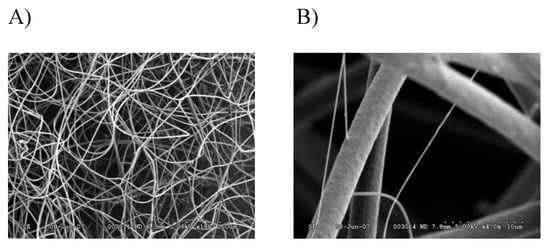

Figure 2. SEM images of black widow spider egg case silk reveal two different diameter fibers: (A) 180× magnification; (B) 4000× magnification.
We collected egg sacs from black widow spiders, dissolved the fibers in chaotropic solvents, and performed in solution tryptic digestion, followed by nLC-MS/MS analysis. Analysis of the egg cases led to the identification of 30 protein groups, 174 peptide groups, 523 peptide spectral matches (PSMs), and 30,985 MS2 spectra. After manual inspection, the number of protein groups was reduced from 30 to 25 due to protein duplications entered into the database with different accession codes. The number of unique peptides that mapped back to targets ranged from 1 to 34. Some of the proteins found within egg cases included ECP1, ECP2, TuSp1, AcSp1, MaSp1, MaSp2, Flagelliform spidroin variant 1, aggregate spidroin variant 1, PySp1, cysteine-rich protein 2 (CRP2), and CRP4. ECP1, ECP2, and TuSp1 have previously been shown to represent constituents of tubuliform silk, while AcSp1 has been shown to be components of dragline, wrapping, and egg case silk [5][6][8][24][26]. Moreover, both MaSp1 and MaSp2 mRNAs have been shown to be present within tubuliform glands, suggesting tubuliform silk represents a composite of spidroins. The detection of aggregate spidroin variant 1 in egg sacs also supports the presence of glue proteins extruded from the aggregate gland during the spinning of egg cases, while the presence of PySp1 may indicate attachment disc silk is used to help immobilize the egg cases to a solid support.
Following the spinning of the egg sacs by female black widow spiders, we removed their tubuliform glands by microdissection, dissolved the glands in HFIP, and then performed in-solution tryptic digestion, followed by MS/MS analysis (Figure 1). Prior to lysis, we removed the spinning dope from the tubuliform glands, which was performed by the collection of internal liquid contents from the tubuliform glands. Analysis of the tubuliform glands led to the identification of 409 protein groups (collapsed to 341 protein groups; Figure 3A), 1118 peptide groups, 1832 PSMs, and 15,777 MS2 spectra. Consistent with previous studies, ECP1, ECP2, and TuSp1 were expressed in tubuliform glands. CRP4, a CRP family member, was also demonstrated to exist for the first time at the protein level within tubuliform glands. Although CRP4 was expressed in tubuliform glands, this observation differs from a previous transcriptomic analysis of L. hesperus, in which CRP1 and CRP3 mRNAs were detected in tubuliform glands [27]. Other proteins involved in a wide range of molecular and cellular functions were identified, including translation, protein folding and processing, cellular communication (signaling), metabolism, cytoskeletal activities, cell adhesion, and gene regulation (Figure 3B).
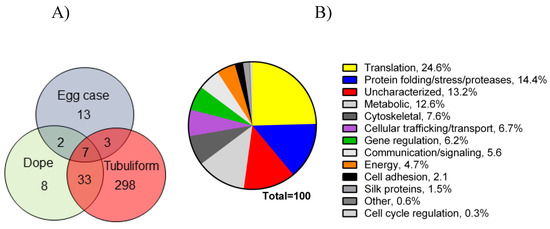
Figure 3. Venn diagram representing number of proteins expressed in tubuliform glands, liquid spinning dope, and egg sacs, along with a pie chart of percentages of proteins expressed in the tubuliform gland categorized into several different molecular and cellular functions: (A) Venn diagram; (B) pie chart.
Translation, protein folding and processing, and enzymes involved in metabolic reactions constituted approximately 25%, 14%, and 13%, respectively, of the proteome (Figure 3B). Some of the proteins expressed in tubuliform glands represented uncharacterized factors, while others fell into a broad range of cellular activities (Figure 3B). Overall, these data support that a large number of tubuliform gland proteins participate in silk protein biosynthesis.
Analysis of the spinning dope revealed 50 different protein groups. ECP1, ECP2, MaSp1, MaSp2, TuSp1, along with CRP4, were identified in the dope. Comparison of our proteomic data from egg cases, tubuliform glands, and spinning dope revealed seven proteins common to all three locations, which included TuSp1 (egg case fibroin), ECP1, ECP2, fasciclin, CRP4, and two uncharacterized proteins (Figure 3A). Representative MS/MS spectra corresponding to CRP4 and one of the uncharacterized proteins are shown (Figure 4A,B). The discovery of a common group of seven proteins in the liquid dope and egg cases further substantiates the assertion that these molecules are constituents of tubuliform silk. Moreover, it implies CRP4, which has been shown to be a constituent of dragline silk, may have a generalized functional role in both fiber types [7].
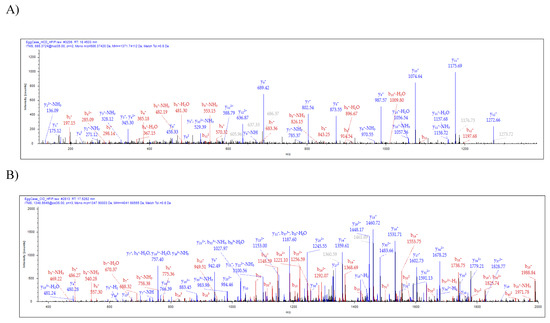
Figure 4. Tandem MS analysis of tryptic fragments generated from in-solution digestion of tubuliform spinning dope identifies the presence of a CRP4 peptide sequence, VVGPFPICDYGLR, and peptide sequence of an uncharacterized protein, SQGNVAMSSSQAGYG QGQSYSSNYAATGDSGTGQGGYSSMR. (A) HCD (MS/MS) spectrum of CRP4 precursor ion with MH+ m/z 1271.74 supports the presence of CRP4 in spinning dope; (B) CID (MS/MS) spectrum of the uncharacterized protein’s precursor ion with MH+ m/z 4041.68 supports the presence of ECP3 in spinning dope. Y- and b-ions and their corresponding masses are shown above the peaks.
Previous studies have identified 14 SST proteins in tubuliform glands [5]. Of the 14 SST proteins, 8 were shown to be expressed exclusively in tubuliform glands; these eight SST proteins were not detected in the major or minor ampullate glands. Of the eight SST tubuliform gland proteins, two were known constituents of tubuliform silk (ECP1 and TuSp1), while the other six SST proteins were classified as uncharacterized proteins [5]. In order to determine whether these six SSTs were present in tubuliform spinning dope and egg cases, we analyzed a customized protein database containing only these six translated SSTs. Two of the SST proteins were detected in both locations (dope and egg cases), which corresponded to transcriptome scaffold ID numbers 129 (silk_comp16922_c0_seq1) and 503 (Contig242). One of the SST proteins, transcriptome scaffold ID #717 (silk_Contig894), was found exclusively in tubuliform glands, while the remaining three SST proteins were not found in either tubuliform glands, dope, or egg case fibers.
References
- Gosline, J.M.; Guerette, P.A.; Ortlepp, C.S.; Savage, K.N. The mechanical design of spider silks: From fibroin sequence to mechanical function. J. Exp. Biol. 1999, 202, 3295–3303.
- Moore, A.M.; Tran, K. Material properties of cobweb silk from the black widow spider Latrodectus hesperus. Int. J. Biol. Macromol. 1999, 24, 277–282.
- Lane, A.K.; Hayashi, C.Y.; Whitworth, G.B.; Ayoub, N.A. Complex gene expression in the dragline silk producing glands of the western black widow (Latrodectus hesperus). BMC Genomics 2013, 14, 846.
- Clarke, T.H.; Garb, J.E.; Hayashi, C.Y.; Haney, R.A.; Lancaster, A.K.; Corbett, S.; Ayoub, N.A. Multi-tissue transcriptomics of the black widow spider reveals expansions, co-options, and functional processes of the silk gland gene toolkit. BMC Genomics 2014, 15, 365.
- Chaw, R.C.; Correa-Garhwal, S.M.; Clarke, T.H.; Ayoub, N.A.; Hayashi, C.Y. Proteomic evidence for components of spider silk synthesis from black widow silk glands and fibers. J. Proteome Res. 2015, 14, 4223–4231.
- Hu, X.; Kohler, K.; Falick, A.M.; Moore, A.M.; Jones, P.R.; Vierra, C. Spider egg case core fibers: Trimeric complexes assembled from TuSp1, ECP-1, and ECP-2. Biochemistry 2006, 45, 3506–3516.
- Larracas, C.; Hekman, R.; Dyrness, S.; Arata, A.; Williams, C.; Crawford, T.; Vierra, C.A. Comprehensive proteomic analysis of spider dragline silk from black widows: A recipe to build synthetic silk fibers. Int. J. Mol. Sci. 2016, 17, 1537.
- Vasanthavada, K.; Hu, X.; Falick, A.M.; La Mattina, C.; Moore, A.M.; Jones, P.R.; Yee, R.; Reza, R.; Tuton, T.; Vierra, C. Aciniform spidroin, a constituent of egg case sacs and wrapping silk fibers from the black widow spider Latrodectus hesperus. J. Biol. Chem. 2007, 282, 35088–35097.
- La Mattina, C.; Reza, R.; Hu, X.; Falick, A.M.; Vasanthavada, K.; McNary, S.; Yee, R.; Vierra, C.A. Spider minor ampullate silk proteins are constituents of prey wrapping silk in the cob weaver Latrodectus hesperus. Biochemistry 2008, 47, 4692–4700.
- He, Q.; Duan, Z.; Yu, Y.; Liu, Z.; Liang, S. The venom gland transcriptome of Latrodectus tredecimguttatus revealed by deep sequencing and cdna library analysis. PLoS ONE 2013, 8, e81357.
- Casem, M.L.; Collin, M.A.; Ayoub, N.A.; Hayashi, C.Y. Silk gene transcripts in the developing tubuliform glands of the western black widow, Latrodectus hesperus. J. Arachnol. 2010, 38, 99–103.
- Jeffery, F.; La Mattina, C.; Tuton-Blasingame, T.; Hsia, Y.; Gnesa, E.; Zhao, L.; Franz, A.; Vierra, C. Microdissection of black widow spider silk-producing glands. J. Vis. Exp. 2011, 47, e2382.
- Nentwig, W. Ecophysiology of Spiders; Springer: Berlin, Germany, 1987.
- Foelix, R. Biology of Spiders; Oxford University Press: New York, NY, USA, 1996.
- Guerette, P.A.; Ginzinger, D.G.; Weber, B.H.; Gosline, J.M. Silk properties determined by gland-specific expression of a spider fibroin gene family. Science 1996, 272, 112–115.
- Gatesy, J.; Hayashi, C.; Motriuk, D.; Woods, J.; Lewis, R. Extreme diversity, conservation, and convergence of spider silk fibroin sequences. Science 2001, 291, 2603–2605.
- Ayoub, N.A.; Garb, J.E.; Tinghitella, R.M.; Collin, M.A.; Hayashi, C.Y. Blueprint for a high-performance biomaterial: Full-length spider dragline silk genes. PLoS ONE 2007, 2, e514.
- Beckwitt, R.; Arcidiacono, S. Sequence conservation in the C-terminal region of spider silk proteins (spidroin) from Nephila clavipes (tetragnathidae) and Araneus bicentenarius (araneidae). J. Biol. Chem. 1994, 269, 6661–6663.
- Hayashi, C.Y.; Blackledge, T.A.; Lewis, R.V. Molecular and mechanical characterization of aciniform silk: Uniformity of iterated sequence modules in a novel member of the spider silk fibroin gene family. Mol. Biol. Evol. 2004, 21, 1950–1959.
- Trancik, J.E.; Czernuszka, J.T.; Bell, F.I.; Viney, C. Nanostructural features of a spider dragline silk as revealed by electron and X-ray diffraction studies. Polymer 2006, 47, 5633–5642.
- Jenkins, J.E.; Holland, G.P.; Yarger, J.L. High resolution magic angle spinning NMR investigation of silk protein structure within major ampullate glands of orb weaving spiders. Soft Matter 2012, 8, 1947–1954.
- Jenkins, J.E.; Creager, M.S.; Butler, E.B.; Lewis, R.V.; Yarger, J.L.; Holland, G.P. Solid-state NMR evidence for elastin-like beta-turn structure in spider dragline silk. Chem. Commun. 2010, 46, 6714–6716.
- Hieber, C.S. Spider cocoons and their suspension systems as barriers to generalist and specialist predators. Oecologia 1992, 91, 530–535.
- Tian, M.; Lewis, R.V. Molecular characterization and evolutionary study of spider tubuliform (eggcase) silk protein. Biochemistry 2005, 44, 8006–8012.
- Hu, X.; Kohler, K.; Falick, A.M.; Moore, A.M.; Jones, P.R.; Sparkman, O.D.; Vierra, C. Egg case protein-1. A new class of silk proteins with fibroin-like properties from the spider Latrodectus hesperus. J. Biol. Chem. 2005, 280, 21220–21230.
- Hu, X.; Lawrence, B.; Kohler, K.; Falick, A.M.; Moore, A.M.; McMullen, E.; Jones, P.R.; Vierra, C. Araneoid egg case silk: A fibroin with novel ensemble repeat units from the black widow spider, Latrodectus hesperus. Biochemistry 2005, 44, 10020–10027.
- Clarke, T.H.; Garb, J.E.; Haney, R.A.; Chaw, R.C.; Hayashi, C.Y.; Ayoub, N.A. Evolutionary shifts in gene expression decoupled from gene duplication across functionally distinct spider silk glands. Sci. Rep. 2017, 7, 8393.
More
Information
Subjects:
Biology
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.3K
Revisions:
2 times
(View History)
Update Date:
09 Sep 2021
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





