
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Dar-Bin Shieh | + 2642 word(s) | 2642 | 2021-08-10 06:35:44 | | | |
| 2 | Beatrix Zheng | + 40 word(s) | 2682 | 2021-08-19 10:39:47 | | |
Video Upload Options
Clostridioides difficile, a spore-forming bacterium, is a nosocomial infectious pathogen which can be found in animals as well. Although various antibiotics and disinfectants were developed, C. difficile infection (CDI) remains a serious health problem. C. difficile spores have complex structures and dormant characteristics that contribute to their resistance to harsh environments, successful transmission and recurrence.
1. Introduction
Clostridioides (formerly Clostridium ) difficile , a Gram-positive bacterium that causes severe antibiotic-associated diarrheas and colitis, was first isolated from new-born infants in 1935 [1]. It produces oval terminal endospores [1] and is commonly acquired through community and hospital (nosocomial) infections [2]. The disease is associated with inappropriate antibiotic treatment, which causes an imbalance of the host’s intestine microbial flora, in turn activating the dormant C. difficile [3]. Clindamycin, carbapenems and fluoroquinolones are the antibiotics most commonly associated with increasing the risk of C. difficile infection (CDI) [4]. Additionally, gastric acid-suppressant and older age (>65 years) are also important risk factors [5]. The symptoms of CDI include watery diarrhea, fever, abdominal pain and toxic megacolon [6]. From a recent epidemiological analysis reported by NHS trusts in England, there were a total of 13,177 CDI cases diagnosed between 2019 and 2020 [7], a small increase of 7.4% compared to the previous year ( n = 12,274). The incidence of hospital-onset CDI cases mirrors the trends in the incidence of all cases, with a decline between 2007 and 2014 followed by a relatively stable state till 2018. The rate (hospital-onset cases/100,000 bed-days) of hospital-onset CDI cases increased from 12.2 to 13.6 between 2018 and 2020. In the USA, a statistical analysis revealed that CDI cases in 10 hospitals increased between 2011 and 2017, while the adjusted estimate of the burden of hospitalizations for CDI decreased by 24% [8]. Whereas the adjusted estimates of the burden of first recurrences and in-hospital deaths did not change significantly, suggesting the effectiveness of infection-prevention practices, and new more refined diagnostic techniques, it is important to eliminate false positives and to improve infection prevention. However, reducing the burden of CDI remains one of the imperative health care priorities in western countries. Different ribotypes occur according to geographic localization and time of the episodes, associated with evolutionary sophistication. Ribotype 027 and 078 strains have spread worldwide since the millennium, and this was attributed to their ability to metabolize disaccharide trehalose approved by the USA Food and Drug Administration (FDA) since 2000 [9]. Epidemic C. difficile ribotypes yield more toxins and have higher sporulation compared to non-epidemic ones [10][11], in spite of some controversial results [11].
The C. difficile spores play an important role in its pathogenesis and are well known to resist gastric acid, harsh environments and antibiotics treatment, or even survive in dry inorganic surfaces for months [12]. Hand washing has been recommended as a good practice to reduce risk of pathogen transmission, not only among health care workers but also visitors [13][14]. Hand washing with soap and water is significantly more effective at removing C. difficile spores than alcohol-based hand rubs [15][16]. The unique structure of the spores helps the pathogen to overcome UV-A and UV-B irradiation, heat (up to 71 °C), extreme freezing, biocides, chemical disinfectants, desiccation and nutrient deficiency [17][18]. The exosporium is the outermost layer of the spore, containing cysteine (CdeC)-rich proteins that enhance their surface adhesion and spore-host interactions [19][20] and were demonstrated to assist in resistance to heat, enzymes and macrophage-inactivation [19][21]. The next layer is the coat, which blocks oxidizing agents, hypochlorite and enzymes from damaging the microorganism [18][22]. Inside the coat is the outer membrane, and the cortex which keeps the spore in a dehydrated state [23]. The low permeability of the inner membrane prevents the core from invasion from water and other small molecules [24]. The innermost part of the spore is the dehydrated core, containing DNA, RNA, ribosomes, small acid-soluble spore proteins (SASPs) and large amounts of calcium dipicolinic acid (Ca-DPA) [18][25]. The high level of the Ca-DPA increases their resistance to environmental stressors, such as disinfectants and ultraviolet radiation [26].
2. Molecular Pathogenesis of C. difficile Infection
Transmission of C. difficile spores or vegetative cells occurs through the fecal—oral route, and from direct contact with contaminated items [27]. However, only spore-form C. difficile can pass through the gastric acid and achieve residence in the large intestine [6]. As the spores are exposed to an appropriate environment containing bile acid and co-germinant, they can be reactivated into the vegetative state [28][29]. C. difficile spores germinate mostly within the ileum due to the higher environment pH (around 7.4) [29]. At a molecular level, CspC serves as the bile acid germinant receptor, while CspA acts as the co-germinant receptor. Upon activation, the signal is transmitted to CspB [30][31][32], which converts pro-SleC into its active form to degrade the cortex [33][34]. This leads to expansion of the germ cell wall and rehydration of the core, together with the release of dipicolinic acid (DPA) [35][36]. The outgrowth of C. difficile spores into a vegetative cell is the result. The vegetative form of C. difficile proliferates and produces toxin A (TcdA) and toxin B (TcdB), which contribute to the major pathogenesis process. TcdE protein contributes to the secretion of TcdA and TcdB whcih promotes C. difficile growth by obtaining nutrients from toxin-mediated collagen degradation and suppression of competitors in the gut [37][38][39]. Secreted TcdA binds to carbohydrates on the apical surface of colonic epithelial cells, while TcdB recognizes their Wnt receptor frizzled family (FZDs) proteins [40]. Both TcdA and TcdB toxins enter cells via endocytosis, followed by fusion with the lysosome [41][42]. Upon acidification of the organelle, protonation triggers conformation changes in TcdA and TcdB to form hairpin pores; this is followed by release of the glucosyltransferase domains of the toxins from the organelles via autoproteolysis [43][44]. These domains further glycosylate Rho and Rac in the cytosol, preventing them from being activated by guanine nucleotide exchange factors (GEFs), thereby triggering apoptosis and loss of tight junction integrity in mucosal epithelial cells [41][45].
C. difficile cells start to sporulate when nutrients are scarce in the environment, although the quorum-sensing signals to relay the environmental state are yet to be identified [25]. Stage 0 sporulation protein A (Spo0A) is a transcription factor critical for the C. difficile life cycle that regulates genes associated with biofilm formation, metabolism, toxin production and sporulation [46]. Phosphorylation of Spo0A is an early triggering factor for sporulation, followed by the activation of sigma factor F (σF) to control the downstream effectors σG in the forespore compartment. Activation of σG is required for spore cell wall synthesis and cortex formation [47]. Spo0A also activates σE and further activates σk to modulate coat protein expression and DPA synthesis, as well as their structural assembly after asymmetric division. At this stage, the replicated DNA is already packed into the forespore [48]. After completing the development of the membrane, spore coat and cortex proteins, the mother cell will be lysed, and the mature spore will be released ( Figure 1 ).
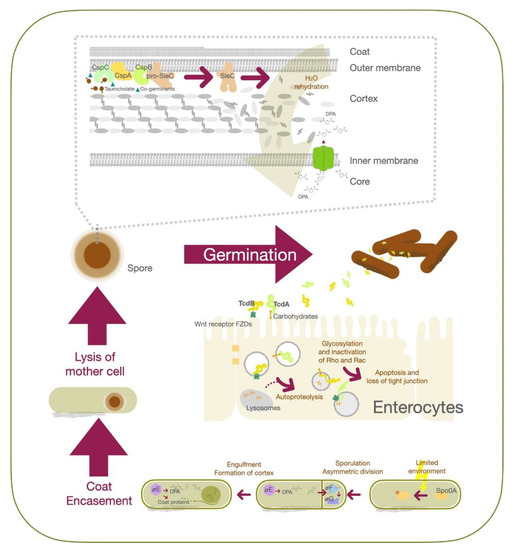
Symptomatic C. difficile patients shed out the vegetative cells and spores leading to contamination of their environment [49]. The vegetative cells can survive 6 h in room air, while the spores may remain alive for as long as 5 months [50][51]. As the health-care workers and patients’ family members contact the spore-contaminated surface, the spores tightly attach to the skin [52]. Noticeably, asymptomatic C. difficile carriers can shed out spores and cause another CDI outbreak [53]. Prevention of C. difficile spore formation is, therefore, an important strategy for CDI management and could reduce the threat of relapse [54].
Biofilm formation by vegetative cells and spores has been identified in several Clostridium species, including Clostridium perfringens , C. thermocellum and C. acetobutylicum [55]. Biofilm formation could play major roles in all phases of CDI, especially in their recurrence [56][57], since it helps to enhance microorganism retention, enabling them to resist the flow of luminal material in the gastrointestinal (GI) tract and to prevent the host immune system’s attack. Biofilms provide a powerful shield against antibiotics and create a comfort zone for the microorganism to survive and prosper [58]. Intriguingly, biofilms could also reduce germination efficiency in C. difficile . Such controversy complicates the development of treatment strategies.
3. Emergent Roles for Nanotechnology in Infectious Diseases
The use of nanotechnology to defeat multidrug resistance gained significant global attention as new effective antibiotic development is extremely challenging and costly. Modifications in nanoparticles (NPs) could enable multifunctional purposes and bring about advanced applications in medicine. Nanomaterials could be modified with specific targeting moieties such as antibodies or aptamers, to enhance the therapeutic specificity and minimize collateral damage to healthy tissues [59]. Various types of nanomaterials have been shown to deliver drugs with good releasing profile and improve efficacy. These nanomedicines have been extensively explored for applications in the infectious disease area as well [60][61].
Organic nanoparticles (e.g., liposomes, polymeric, micelles and ferritin) have been used to enhance the bioavailability of therapeutic compounds and to increase their delivery and efficacy [62]. They were developed as drug delivery systems, offering a controlled-release profile and targeting the desired tissues or cells. These nanocarrier systems may control drug release by an excipient that enabled slow dissolution of poorly soluble drug crystals, from the core compartment to the interstitial space. Sustained release can also be obtained by encapsulating drugs in nanocarriers capable of loading both hydrophobic and hydrophilic drugs. Organic nanocarrier systems have been evaluated for the treatment of local infections of the female reproductive tract, lungs and skin. Injectable nanocarriers have also been explored for the systemic delivery of drugs [63]. Regarding the types of pathogens targeted, nanoparticles have also been extensively explored for treating fungal [64], bacterial [63] and viral infections, including by Candida albicans and severe acute respiratory syndrome 2 (SARS-CoV-2) [65].
Inorganic nanoparticles (e.g., metals or metal oxides) have also been investigated for the prevention and treatment of infectious pathogens. Some inorganic nanomaterials were discovered to exhibit diverse activities against multi-drug-resistant pathogens [66]. These include silver (Ag), zinc oxide (ZnO), iron-containing nanoparticles and more. The antibacterial properties of the metallic nanoparticles may be attributed to the generation of reactive oxygen species (ROS), disruption of cell membranes, ability to bind thiol groups (SH-)/disulfide bonds (R-S-S-R) in biomolecules and the release of soluble metal ions [67][68]. The most widely studied metallic materials in infection control are silver (Ag) nanoparticles. The antimicrobial mechanisms of Ag nanoparticles are associated with ROS generation and silver ion release from the nanoparticles. Ag nanoparticles and ions interact with the thiol group, sulfur and phosphorus in the microbial cells subsequently bringing about DNA damage and protein dysfunction [69]. In addition, Ag nanoparticles could also anchor to the bacterial cell wall and cause structural changes in the cell membrane, thus radically affecting cell membrane permeability and inducing cell death. Free radicals generated by Ag nanoparticles upon contact with the bacterial cell membrane are another important mechanism for their anti-microbial activity, as confirmed by electron spin resonance analysis [69]. Compared to their bulk state, Ag nanoparticles also display efficient antimicrobial properties due to their large surface-to-volume ratio, providing better contact interface with the microorganisms [70].
Nanotechnology has also been applied to disease prevention in vaccine formulation, the so called nanovaccinology. Over the past decade, nanoscale size materials such as virus-like particles (VLPs), liposomes, polymeric, inorganic nanoparticles and emulsions have gained attention as potential delivery vehicles for vaccine antigens, which can both stabilize vaccine antigens and act as adjuvants [71]. These advantages are attributable to the nanoscale particle size, which facilitates uptake by antigen-presenting cells (APCs), leading to efficient antigen recognition and presentation. Modifying the surfaces of nanoparticles with different targeting moieties permits the delivery of antigens to specific receptors on the cell surface, thereby stimulating selective and specific immune responses [72]. The most well-known example is the liposome applied for encapsulating modified mRNAs and stimulating host immune response in SARS -CoV-2 vaccines developed by Moderna and Pfizer-BioNTech [73]. Nanoparticles in the vaccine formulations allow for enhanced immunogenicity and stability of the payload (antigen or nucleic acids encoding the expression of antigen), but also targeted delivery and slow release for longer immunostimulation [74].
4. Nanomaterials for CDI Therapeutics
Organic nanoparticles for the delivery of anti-sense anti-microbial oligonucleotides have been reported recently for anti-CDI therapy [75]. The modified anti-sense oligonucleotides can specifically target five essential C. difficile genes simultaneously. They used three (APDE-8, CODE-9, CYDE-21) novel cationic amphiphilic bolaamphiphiles (CABs) to form nano-sized vesicles or vesicle-like aggregates (CABVs) and encapulate 25-mer antisense oligonucleotides (ASO). The empty CABVs had little effect on C. difficile growth and could deliver an effective amount of ASO against C. difficile. Through encapsulation by bolaamphiphile-based nanocomplex, the oligonucleotides could be effectively transported into C. difficile to modulate the translation of specific mRNA which achieved inhibitory concentrations in C. difficile without affecting normal microbiota [76][77].
Polyurethane containing crystal violet (CV) and 3–4 nm ZnO nanoparticles have been reported to present bactericidal activity against hospital-acquired pathogens including multidrug-resistant E. coli , Pseudomonas aeruginosa , methicillin-resistant S. aureus (MRSA) and even highly resistant endospores of C. difficile [78]. However, recent studies showed that ZnO nanoparticles may affect other microorganisms and impact normal intestinal microflora despite their widespread use in biomedicine [79]. DNA damage induced by ZnO also limits its biomedical application in clinical settings [80]. Other types of nanomaterials responsive to photonic energy (e.g., photodynamic or photothermal therapy) have also been developed for anti-microbial therapy, to enhance drug delivery and local activation. However, the gastrointestinal tract is not the ideal organ for light illumination.
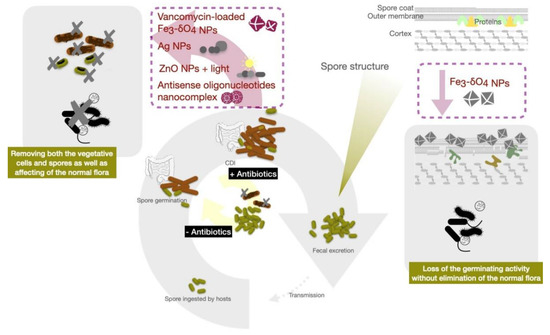
Iron is the most abundant transition metal in the human body and participates in important physiological functions such as oxygen transport and electron transfer. The human body has developed sophisticated systems for the uptake, transport, storage and metabolism of iron. Iron-containing nanoparticles usually exhibited magnetic properties and are among the pioneer nanomaterials used in a wide variety of biomedical and bioengineering applications. Many iron-containing nanoparticles have been evaluated in preclinical and clinical trials, and some of those have reached the market. Zero-valent-iron (ZVI) nanoparticles exhibited excellent biocompatibility while harboring prominently bactericidal efficacy against E. coli . Iron oxide nanoparticles also have been widely used for biomedical applications, including hyperthermia therapy and magnetic resonance imagining [81]. There are reports describing that iron oxide nanoparticles have reduced bacterial biofilm formation and viability via an increase in oxidative stress [82][83]. The anti-bacterial mechanisms of these metallic nanoparticles are attributed to their ability to generate reactive oxygen species, disrupt cell membranes, bind thiol groups and release toxic ions. Iron oxide nanoparticles, widely used as T2 weighed MRI imaging contrast agents, were recently used in CDI treatment. The Fe 3-δ O 4 magnetite nanoparticles (500 µg/mL) were reported to display sporicidal activity against C. difficile spores without adversely affecting the gut microbiota of experimental mice [84]. Fe 3-δ O 4 magnetite nanoparticles bind to the surface of C. difficile spores, decreasing Ca-DPA release from the spores. The nanoparticles eventually inhibited spore viability in vitro and attenuated C. difficile -induced colitis in this mouse model. However, Fe 3-δ O 4 nanoparticles did not kill the vegetative cells. A ZVI signal, detected in the Fe 3-δ O 4 nanoparticles by X-ray diffraction, has been reported to involve the induction of intracellular oxidative stress and depleting mitochondrial membrane potential in malignant cells [85][86]. Generation of ROS may contribute to the inhibition of C. difficile spore germination. As the Fe 3-δ O 4 nanoparticles only showed efficacy in anti-spore germination without killing the vegetative form of C. difficile , vancomycin-loaded Fe 3-δ O 4 nanoparticles (van-IONPs) were synthesized to further enhance CDI control. These nanoparticles demonstrated the ability to inhibit both vegetative cell growth and spore germination [84][87] through direct binding to C. difficile spores and blocking their germination, while inhibiting vegetative cells by releasing antibiotics in a synchronized manner ( Figure 2 ). Moreover, van-IONPs protected the intestinal mucosa from C. difficile spore adhesion and significantly decreased the level of C. difficile -induced inflammation in mice. These nanoparticles outperform both Fe 3-δ O 4 nanoparticles and free vancomycin in overall anti-CDI efficacy, due to the dual-function activities of targeting both spores and vegetative cells [87]. ROS production has also been reported as the cause for the anti-bacterial properties of another type of Fe 3-δ O 4 nanoparticles [88]. One important advantage of these iron-based nanoparticles in CDI control is their selectivity for C. difficile spores and biocompatibility to intestinal mucosa cells. Such properties preserved the normal intestinal flora critical for preventing invasion by other pathogens and protecting the host from recurrence [89].
References
- Hall, I.C.; O’toole, E. Intestinal flora in new-born infants: With a description of a new pathogenic anaerobe, Bacillus difficilis. Am. J. Dis. Child. 1935, 49, 390–402.
- Bloomfield, L.E.; Riley, T.V. Epidemiology and risk factors for community-associated Clostridium difficile infection: A narrative review. Infect. Dis. Ther. 2016, 5, 231–251.
- Kochan, T.J.; Foley, M.H.; Shoshiev, M.S.; Somers, M.J.; Carlson, P.E.; Hanna, P.C. Updates to Clostridium difficile spore germination. J. Bacteriol. 2018, 200, e00218-18.
- Brown, K.A.; Khanafer, N.; Daneman, N.; Fisman, D.N. Meta-analysis of antibiotics and the risk of community-associated Clostridium difficile infection. Antimicrob. Agents Chemother. 2013, 57, 7.
- Jump, R.L. Clostridium difficile infection in older adults. Aging Health 2013, 9, 403–414.
- Poutanen, S.M.; Simor, A.E. Clostridium difficile—Associated diarrhea in adults. CMAJ 2004, 171, 51–58.
- Public Health England. Annual Epidemiological Commentary: Gram-Negative Bacteraemia, MRSA Bacteraemia, MSSA Bacteraemia and C. difficile Infections, up to and Including Financial Year April 2019 to March 2020; Public Health England: London, UK, 2020.
- Guh, A.Y.; Mu, Y.; Winston, L.G.; Johnston, H.; Olson, D.; Farley, M.M.; Wilson, L.E.; Holzbauer, S.M.; Phipps, E.C.; Dumyati, G.K. Trends in US burden of Clostridioides difficile infection and outcomes. N. Engl. J. Med. 2020, 382, 1320–1330.
- Collins, J.; Robinson, C.; Danhof, H.; Knetsch, C.; Van Leeuwen, H.; Lawley, T.; Auchtung, J.; Britton, R. Dietary trehalose enhances virulence of epidemic Clostridium difficile. Nature 2018, 553, 291–294.
- Akerlund, T.; Persson, I.; Unemo, M.; Norén, T.R.; Svenungsson, B.; Wullt, M.; Burman, L.G. Increased sporulation rate of epidemic Clostridium difficile type 027/NAP1. J. Clin. Microbiol. 2008, 46, 1530–1533.
- Vitucci, J.C.; Pulse, M.; Tabor-Simecka, L.; Simecka, J. Epidemic ribotypes of Clostridium (now Clostridioides) difficile are likely to be more virulent than non-epidemic ribotypes in animal models. BMC Microbiol. 2020, 20, 27.
- Weaver, L.; Michels, H.; Keevil, C. Survival of Clostridium difficile on copper and steel: Futuristic options for hospital hygiene. J. Hosp. Infect. 2008, 68, 145–151.
- Ragusa, R.; Giorgianni, G.; Lupo, L.; Sciacca, A.; Rametta, S.; La Verde, M.; Mulè, S.; Marranzano, M. Healthcare-associated Clostridium difficile infection: Role of correct hand hygiene in cross-infection control. J. Prev. Med. Hyg. 2018, 59, E145.
- Scaria, E.; Barker, A.K.; Alagoz, O.; Safdar, N. Association of Visitor Contact Precautions with Estimated Hospital-Onset Clostridioides difficile Infection Rates in Acute Care Hospitals. JAMA Netw. Open 2021, 4, e210361.
- Jabbar, U.; Leischner, J.; Kasper, D.; Gerber, R.; Sambol, S.P.; Parada, J.P.; Johnson, S.; Gerding, D.N. Effectiveness of alcohol-based hand rubs for removal of Clostridium difficile spores from hands. Infect. Control Hosp. Epidemiol. 2010, 31, 565.
- Boyce, J.M.; Pittet, D. Guideline for hand hygiene in health-care settings: Recommendations of the Healthcare Infection Control Practices Advisory Committee and the HICPAC/SHEA/APIC/IDSA Hand Hygiene Task Force. Am. J. Infect. Control. 2002, 30, S1–S46.
- Barbut, F. Comparison of the efficacy of a hydrogen peroxide dry-mist disinfection system and sodium hypochlorite solution for eradication of Clostridium difficile spores. Infect. Control Hosp. Epidemiol. 2009, 30, 507–514.
- Leggett, M.J.; McDonnell, G.; Denyer, S.P.; Setlow, P.; Maillard, J.Y. Bacterial spore structures and their protective role in biocide resistance. J. Appl. Microbiol. 2012, 113, 485–498.
- Barra-Carrasco, J.; Olguín-Araneda, V.; Plaza-Garrido, Á.; Miranda-Cárdenas, C.; Cofré-Araneda, G.; Pizarro-Guajardo, M.; Sarker, M.R.; Paredes-Sabja, D. The Clostridium difficile exosporium cysteine (CdeC)-rich protein is required for exosporium morphogenesis and coat assembly. J. Bacteriol. 2013, 195, 3863.
- Mora-Uribe, P.; Miranda-Cárdenas, C.; Castro-Córdova, P.; Gil, F.; Calderón, I.; Fuentes, J.A.; Rodas, P.I.; Banawas, S.; Sarker, M.R.; Paredes-Sabja, D. Characterization of the adherence of Clostridium difficile spores: The integrity of the outermost layer affects adherence properties of spores of the epidemic strain R20291 to components of the intestinal mucosa. Front. Cell. Infect. Microbiol. 2016, 6, 99.
- Calderón-Romero, P.; Castro-Córdova, P.; Reyes-Ramírez, R.; Milano-Céspedes, M.; Guerrero-Araya, E.; Pizarro-Guajardo, M.; Olguín-Araneda, V.; Gil, F.; Paredes-Sabja, D. Clostridium difficile exosporium cysteine-rich proteins are essential for the morphogenesis of the exosporium layer, spore resistance, and affect C. difficile pathogenesis. PLoS Pathog. 2018, 14, e1007199.
- Permpoonpattana, P.; Phetcharaburanin, J.; Mikelsone, A.; Dembek, M.; Tan, S.; Brisson, M.-C.; La Ragione, R.; Brisson, A.R.; Fairweather, N.; Hong, H.A. Functional characterization of Clostridium difficile spore coat proteins. J. Bacteriol. 2013, 195, 1492.
- Sunde, E.P.; Setlow, P.; Hederstedt, L.; Halle, B. The physical state of water in bacterial spores. Proc. Natl. Acad. Sci. USA 2009, 106, 19334–19339.
- Bressuire-Isoard, C.; Broussolle, V.; Carlin, F. Sporulation environment influences spore properties in Bacillus: Evidence and insights on underlying molecular and physiological mechanisms. FEMS Microbiol. Rev. 2018, 42, 614–626.
- Paredes-Sabja, D.; Shen, A.; Sorg, J.A. Clostridium difficile spore biology: Sporulation, germination, and spore structural proteins. Trends Microbiol. 2014, 22, 406–416.
- Jamroskovic, J.; Chromikova, Z.; List, C.; Bartova, B.; Barak, I.; Bernier-Latmani, R. Variability in DPA and calcium content in the spores of Clostridium species. Front. Microbiol. 2016, 7, 1791.
- Rupnik, M.; Wilcox, M.H.; Gerding, D.N. Clostridium difficile infection: New developments in epidemiology and pathogenesis. Nat. Rev. Microbiol. 2009, 7, 526–536.
- Sorg, J.A.; Sonenshein, A.L. Bile salts and glycine as cogerminants for Clostridium difficile spores. J. Bacteriol. 2008, 190, 2505.
- Kochan, T.J.; Shoshiev, M.S.; Hastie, J.L.; Somers, M.J.; Plotnick, Y.M.; Gutierrez-Munoz, D.F.; Foss, E.D.; Schubert, A.M.; Smith, A.D.; Zimmerman, S.K. Germinant synergy facilitates Clostridium difficile spore germination under physiological conditions. mSphere 2018, 3.
- Francis, M.B.; Allen, C.A.; Shrestha, R.; Sorg, J.A. Bile acid recognition by the Clostridium difficile germinant receptor, CspC, is important for establishing infection. PLoS Pathog. 2013, 9, e1003356.
- Lawler, A.J.; Lambert, P.A.; Worthington, T. A Revised Understanding of Clostridioides difficile Spore Germination. Trends Microbiol. 2020, 28, 744–752.
- Bhattacharjee, D.; Francis, M.B.; Ding, X.; McAllister, K.N.; Shrestha, R.; Sorg, J.A. Reexamining the germination phenotypes of several Clostridium difficile strains suggests another role for the CspC germinant receptor. J. Bacteriol. 2015, 198, 777–786.
- Fimlaid, K.A.; Jensen, O.; Donnelly, M.L.; Francis, M.B.; Sorg, J.A.; Shen, A. Identification of a novel lipoprotein regulator of Clostridium difficile spore germination. PLoS Pathog. 2015, 11, e1005239.
- Francis, M.B.; Allen, C.A.; Sorg, J.A. Spore cortex hydrolysis precedes dipicolinic acid release during Clostridium difficile spore germination. J. Bacteriol. 2015, 197, 2276–2283.
- Paredes-Sabja, D.; Bond, C.; Carman, R.J.; Setlow, P.; Sarker, M.R. Germination of spores of Clostridium difficile strains, including isolates from a hospital outbreak of Clostridium difficile-associated disease (CDAD). Microbiology 2008, 154, 2241–2250.
- Wang, S.; Shen, A.; Setlow, P.; Li, Y.-Q. Characterization of the dynamic germination of individual Clostridium difficile spores using Raman spectroscopy and differential interference contrast microscopy. J. Bacteriol. 2015, 197, 2361.
- Govind, R.; Dupuy, B. Secretion of Clostridium difficile toxins A and B requires the holin-like protein TcdE. PLoS Pathog. 2012, 8, e1002727.
- Abt, M.C.; McKenney, P.T.; Pamer, E.G. Clostridium difficile colitis: Pathogenesis and host defence. Nat. Rev. Microbiol. 2016, 14, 609–620.
- Fletcher, J.R.; Pike, C.M.; Parsons, R.J.; Rivera, A.J.; Foley, M.H.; McLaren, M.R.; Montgomery, S.A.; Theriot, C.M. Clostridioides difficile exploits toxin-mediated inflammation to alter the host nutritional landscape and exclude competitors from the gut microbiota. Nat. Commun. 2021, 12, 462.
- Tao, L.; Zhang, J.; Meraner, P.; Tovaglieri, A.; Wu, X.; Gerhard, R.; Zhang, X.; Stallcup, W.B.; Miao, J.; He, X. Frizzled proteins are colonic epithelial receptors for C. difficile toxin B. Nature 2016, 538, 350–355.
- Papatheodorou, P.; Zamboglou, C.; Genisyuerek, S.; Guttenberg, G.; Aktories, K. Clostridial glucosylating toxins enter cells via clathrin-mediated endocytosis. PLoS ONE 2010, 5, e10673.
- Hirota, S.A.; Iablokov, V.; Tulk, S.E.; Schenck, L.P.; Becker, H.; Nguyen, J.; Al Bashir, S.; Dingle, T.C.; Laing, A.; Liu, J. Intrarectal instillation of Clostridium difficile toxin A triggers colonic inflammation and tissue damage: Development of a novel and efficient mouse model of Clostridium difficile toxin exposure. Infect. Immun. 2012, 80, 4474–4484.
- Chen, P.; Lam, K.-h.; Liu, Z.; Mindlin, F.A.; Chen, B.; Gutierrez, C.B.; Huang, L.; Zhang, Y.; Hamza, T.; Feng, H. Structure of the full-length Clostridium difficile toxin B. Nat. Struct. Mol. Biol. 2019, 26, 712–719.
- Orrell, K.E.; Zhang, Z.; Sugiman-Marangos, S.N.; Melnyk, R.A. Clostridium difficile toxins A and B: Receptors, pores, and translocation into cells. Crit. Rev. Biochem. Mol. Biol. 2017, 52, 461–473.
- Pfeifer, G.; Schirmer, J.; Leemhuis, J.; Busch, C.; Meyer, D.K.; Aktories, K.; Barth, H. Cellular uptake of Clostridium difficile toxin B translocation of the N-terminal catalytic domain into the cytosol of eukaryotic cells. J. Biol. Chem. 2003, 278, 44535–44541.
- Pettit, L.J.; Browne, H.P.; Yu, L.; Smits, W.K.; Fagan, R.P.; Barquist, L.; Martin, M.J.; Goulding, D.; Duncan, S.H.; Flint, H.J. Functional genomics reveals that Clostridium difficile Spo0A coordinates sporulation, virulence and metabolism. BMC Genomics 2014, 15, 160.
- Pereira, F.C.; Saujet, L.; Tomé, A.R.; Serrano, M.; Monot, M.; Couture-Tosi, E.; Martin-Verstraete, I.; Dupuy, B.; Henriques, A.O. The spore differentiation pathway in the enteric pathogen Clostridium difficile. PLoS Genet. 2013, 9, e1003782.
- Fimlaid, K.A.; Bond, J.P.; Schutz, K.C.; Putnam, E.E.; Leung, J.M.; Lawley, T.D.; Shen, A. Global analysis of the sporulation pathway of Clostridium difficile. PLoS Genet. 2013, 9, e1003660.
- Aithinne, K.A.; Cooper, C.W.; Lynch, R.A.; Johnson, D.L. Toilet plume aerosol generation rate and environmental contamination following bowl water inoculation with Clostridium difficile spores. Am. J. Infect. Control. 2019, 47, 515–520.
- Fekety, R.; Kim, K.-H.; Brown, D.; Batts, D.H.; Cudmore, M.; Silva, J., Jr. Epidemiology of antibiotic-associated colitis: Isolation of Clostridium difficile from the hospital environment. Am. J. Med. 1981, 70, 906–908.
- Jump, R.L.; Pultz, M.J.; Donskey, C.J. Vegetative Clostridium difficile survives in room air on moist surfaces and in gastric contents with reduced acidity: A potential mechanism to explain the association between proton pump inhibitors and C. difficile-associated diarrhea? Antimicrob. Agents Chemother. 2007, 51, 2883–2887.
- Pokrywka, M.; Feigel, J.; Douglas, B.; Grossberger, S.; Hensler, A.; Weber, D. A bundle strategy including patient hand hygiene to decrease Clostridium difficile infections. Medsurg Nurs. 2014, 23, 145.
- Furuya-Kanamori, L.; Marquess, J.; Yakob, L.; Riley, T.V.; Paterson, D.L.; Foster, N.F.; Huber, C.A.; Clements, A.C. Asymptomatic Clostridium difficile colonization: Epidemiology and clinical implications. BMC Infect. Dis. 2015, 15, 516.
- Garneau, J.R.; Valiquette, L.; Fortier, L.-C. Prevention of Clostridium difficile spore formation by sub-inhibitory concentrations of tigecycline and piperacillin/tazobactam. BMC Infect. Dis. 2014, 14, 29.
- Dapa, T.; Unnikrishnan, M. Biofilm formation by Clostridium difficile. Gut Microbes 2013, 4, 397–402.
- Normington, C.; Moura, I.B.; Bryant, J.A.; Ewin, D.J.; Clark, E.V.; Kettle, M.J.; Harris, H.C.; Spittal, W.; Davis, G.; Henn, M.R. Biofilms harbour Clostridioides difficile, serving as a reservoir for recurrent infection. NPJ Biofilms Microbiomes 2021, 7, 16.
- Frost, L.R.; Cheng, J.K.; Unnikrishnan, M. Clostridioides difficile biofilms: A mechanism of persistence in the gut? PLoS Pathog. 2021, 17, e1009348.
- Uruén, C.; Chopo-Escuin, G.; Tommassen, J.; Mainar-Jaime, R.C.; Arenas, J. Biofilms as Promoters of Bacterial Antibiotic Resistance and Tolerance. Antibiotics 2021, 10, 3.
- Amin, M.L.; Joo, J.Y.; Yi, D.K.; An, S.S.A. Surface modification and local orientations of surface molecules in nanotherapeutics. J. Control Release 2015, 207, 131–142.
- De Jong, W.H.; Borm, P.J. Drug delivery and nanoparticles: Applications and hazards. Int. J. Nanomed. 2008, 3, 133–149.
- Gao, W.; Chan, J.M.; Farokhzad, O.C. pH-responsive nanoparticles for drug delivery. Mol. Pharm. 2010, 7, 1913–1920.
- Yetisgin, A.A.; Cetinel, S.; Zuvin, M.; Kosar, A.; Kutlu, O. Therapeutic nanoparticles and their targeted delivery applications. Molecules 2020, 25, 2193.
- Kirtane, A.R.; Verma, M.; Karandikar, P.; Furin, J.; Langer, R.; Traverso, G. Nanotechnology approaches for global infectious diseases. Nat. Nanotechnol. 2021, 16, 369–384.
- Mba, I.E.; Nweze, E.I. The use of nanoparticles as alternative therapeutic agents against Candida infections: An up-to-date overview and future perspectives. World J. Microbiol. Biotechnol. 2020, 36, 163.
- Mba, I.E.; Sharndama, H.C.; Osondu-Chuka, G.O.; Okeke, O.P. Immunobiology and nanotherapeutics of severe acute respiratory syndrome 2 (SARS-CoV-2): A current update. Infect. Dis. 2021, 53, 559–580.
- Turner, R.J. Metal-based antimicrobial strategies. Microb. Biotechnol. 2017, 10, 1062–1065.
- Slavin, Y.N.; Asnis, J.; Häfeli, U.O.; Bach, H. Metal nanoparticles: Understanding the mechanisms behind antibacterial activity. J. Nanobiotechnol. 2017, 15, 65.
- Shaikh, S.; Nazam, N.; Rizvi, S.M.D.; Ahmad, K.; Baig, M.H.; Lee, E.J.; Choi, I. Mechanistic insights into the antimicrobial actions of metallic nanoparticles and their implications for multidrug resistance. Int. J. Mol. Sci. 2019, 20, 2468.
- Prabhu, S.; Poulose, E.K. Silver nanoparticles: Mechanism of antimicrobial action, synthesis, medical applications, and tox-icity effects. Int. Nano Lett. 2012, 2, 32.
- Lin, Z.; Monteiro-Riviere, N.A.; Riviere, J.E. Pharmacokinetics of metallic nanoparticles. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2015, 7, 189–217.
- Kreuter, J. Nanoparticles as adjuvants for vaccines. Vaccine Design. 1995, 463–472.
- Mehrabi, M.; Dounighi, N.M.; Mohammadi, M.; Masoudi, A. Nanoparticles and vaccine development. Pharm. Nanotechnol. 2020, 8, 6–21.
- Shin, M.D.; Shukla, S.; Chung, Y.H.; Beiss, V.; Chan, S.K.; Ortega-Rivera, O.A.; Wirth, D.M.; Chen, A.; Sack, M.; Pokorski, J.K.; et al. COVID-19 vaccine development and a potential nanomaterial path forward. Nat. Nanotechnol. 2020, 15, 646–655.
- Demento, S.L.; Cui, W.; Criscione, J.M.; Stern, E.; Tulipan, J.; Kaech, S.M.; Fahmy, T.M. Role of sustained antigen release from nanoparticle vaccines in shaping the T cell memory phenotype. Biomaterials 2012, 33, 4957–4964.
- Stewart, D.B. Anti-sense antibiotic agents as treatment for bacterial infections. Surg. Infect. 2018, 19, 831–835.
- Sharma, A.K.; Krzeminski, J.; Weissig, V.; Hegarty, J.P.; Stewart, D.B. Cationic amphiphilic bolaamphiphile-based delivery of antisense oligonucleotides provides a potentially microbiome sparing treatment for C. difficile. J. Antibiot. 2018, 71, 713–721.
- Hegarty, J.P.; Krzeminski, J.; Sharma, A.K.; Guzman-Villanueva, D.; Weissig, V.; Stewart Sr, D.B. Bolaamphiphile-based nanocomplex delivery of phosphorothioate gapmer antisense oligonucleotides as a treatment for Clostridium difficile. Int. J. Nanomed. 2016, 11, 3607.
- Sehmi, S.K.; Lourenco, C.; Alkhuder, K.; Pike, S.D.; Noimark, S.; Williams, C.K.; Shaffer, M.S.; Parkin, I.P.; MacRobert, A.J.; Allan, E. Antibacterial surfaces with activity against antimicrobial resistant bacterial pathogens and endospores. ACS Infect. Dis. 2020, 6, 939–946.
- Elshama, S.S.; Abdallah, M.E.; Abdel-Karim, R.I. Zinc oxide nanoparticles: Therapeutic benefits and toxicological hazards. Open Nanomed. J. 2018, 5, 16–22.
- Hemeg, H.A. Nanomaterials for alternative antibacterial therapy. Int. J. Nanomed. 2017, 12, 8211.
- Arias, L.S.; Pessan, J.P.; Vieira, A.P.M.; Lima, T.M.T.d.; Delbem, A.C.B.; Monteiro, D.R. Iron oxide nanoparticles for biomedical applications: A perspective on synthesis, drugs, antimicrobial activity, and toxicity. Antibiotics 2018, 7, 46.
- Jamzad, M.; Bidkorpeh, M.K. Green synthesis of iron oxide nanoparticles by the aqueous extract of Laurus nobilis L. leaves and evaluation of the antimicrobial activity. J. Nanostruct. Chem. 2020, 10, 193–201.
- Thukkaram, M.; Sitaram, S.; Subbiahdoss, G. Antibacterial efficacy of iron-oxide nanoparticles against biofilms on different biomaterial surfaces. Int. J. Biomater. 2014, 2014, 716080.
- Lee, W.-T.; Wu, Y.-N.; Chen, Y.-H.; Wu, S.-R.; Shih, T.-M.; Li, T.-J.; Yang, L.-X.; Yeh, C.-S.; Tsai, P.-J.; Shieh, D.-B. Octahedron iron oxide nanocrystals prohibited Clostridium difficile spore germination and attenuated local and systemic inflammation. Sci. Rep. 2017, 7, 8124.
- Huang, C.-C.; Chuang, K.-Y.; Chou, C.-P.; Wu, M.-T.; Sheu, H.-S.; Shieh, D.-B.; Tsai, C.-Y.; Su, C.-H.; Lei, H.-Y.; Yeh, C.-S. Size-control synthesis of structure deficient truncated octahedral Fe3− δO4 nanoparticles: High magnetization magnetites as effective hepatic contrast agents. J. Mater. Chem. 2011, 21, 7472–7479.
- Wu, Y.-N.; Chen, D.-H.; Shi, X.-Y.; Lian, C.-C.; Wang, T.-Y.; Yeh, C.-S.; Ratinac, K.R.; Thordarson, P.; Braet, F.; Shieh, D.-B. Cancer-cell-specific cytotoxicity of non-oxidized iron elements in iron core-gold shell NPs. Nanomedicine 2011, 7, 420–427.
- Chen, Y.-H.; Li, T.-J.; Tsai, B.-Y.; Chen, L.-K.; Lai, Y.-H.; Li, M.-J.; Tsai, C.-Y.; Tsai, P.-J.; Shieh, D.-B. Vancomycin-loaded nanoparticles enhance sporicidal and antibacterial efficacy for Clostridium difficile infection. Front. Microbiol. 2019, 10, 1141.
- Arakha, M.; Pal, S.; Samantarrai, D.; Panigrahi, T.K.; Mallick, B.C.; Pramanik, K.; Mallick, B.; Jha, S. Antimicrobial activity of iron oxide nanoparticle upon modulation of nanoparticle-bacteria interface. Sci. Rep. 2015, 5, 14813.
- Jandhyala, S.M.; Talukdar, R.; Subramanyam, C.; Vuyyuru, H.; Sasikala, M.; Reddy, D.N. Role of the normal gut microbiota. World J. Gastroenterol. 2015, 21, 8787–8803.





