
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Marina Francis | + 1235 word(s) | 1235 | 2020-06-29 05:03:56 | | | |
| 2 | Nicole Yin | Meta information modification | 1235 | 2020-07-16 11:14:57 | | |
Video Upload Options
Sphingolipids are essential structural components of biological membranes that mediate a wide array of physiological functions such as inflammation, cell proliferation, survival, senescence, and death. An emerging body of evidence suggests that bioactive sphingolipids modulate the DNA damage response (DDR) induced by genotoxic stress and therein determine cell fate.
1. Definition
Sphingolipids are characterized by an amino-alcohol, sphingosine, backbone. They constitute major components of cell membranes. Sphingosine is synthesized in the endoplasmic reticulum from non-sphingolipid precursors [1]. Alterations in this basic structure give rise to different sphingolipid metabolites (sphingomyelin, ceramide (Cer), ceramide-1-phosphate (C1P), sphingosine-1-phosphate (S1P), glycosphingolipids…) that mediate various physiological and pathophysiological processes.
2. Introduction
Sphingolipids constitute a class of ubiquitous lipids that regulate the cell membranes’ fluidity and subdomain structure [2][3]. Besides their structural role, research over the past few decades highlighted the importance of sphingolipids in modulating several cellular processes such as proliferation, differentiation, inflammation, senescence, and death [4]. These bioactive molecules mainly include sphingomyelin, ceramide, ceramide-1-phosphate, sphingosine, and sphingosine-1-phosphate. Several studies were conducted to unravel sphingolipids functions, along with the implications of their altered metabolic pathways in several pathological states such as cancer, atherosclerosis, angiogenesis, inflammation, and diabetes [4].
3. Sphingolipids Metabolic Pathway
Ceramide (Cer) is the central metabolite generated within the sphingolipid metabolic pathway through three different pathways. (i) Ceramide de novo synthesis: Palmitoyl-CoA is condensed with serine by the action of serine palmitoyl transferase, followed by a set of reduction and acetylation reactions to generate ceramide; (ii) Sphingomyelin (SM) catabolism: SM is catabolized by sphingomyelinases to generate ceramide; (iii) Salvage pathway: N-acylation of fatty acids with a sphingosine backbone produces ceramide through the action of ceramide synthases [5]. The generated pro-apoptotic Cer can be phosphorylated by ceramide kinase into ceramide-1-phosphate (C1P) in trans-Golgi or plasma membranes. C1P plays an important role in inflammatory responses, cell survival and proliferation [1][4]. Afterwards, C1P can be dephosphorylated by C1P phosphatases or other unspecific lipid phosphate phosphatases (LPP family) [1][6]. Cer is also utilized to generate two major groups of complex glycosphingolipids. Glucosylceramide synthase generates glucosphingolipids by adding glucose as the first residue to Cer at C1 hydroxyl position, whereas galactosylceramide synthase generates galactosphingolipids by adding galactose to Cer [1]. Moreover, Cer can be catabolized by ceramidases into sphingosine which promotes cell cycle arrest and apoptosis. In its turn, sphingosine can be phosphorylated by sphingosine kinases into the pro-survival lipid sphingosine-1-phosphate (S1P) [7]. S1P can be dephosphorylated by S1P phosphatases [8][9] or unspecific LPP [10]. The generated sphingosine can be further used to produce Cer or S1P [9]. S1P lyase (SPL) is considered as the last enzyme in the sphingolipid catabolic pathway because it can irreversibly break down S1P into phosphoethanolamine and hexadecenal [11] (Figure 1).
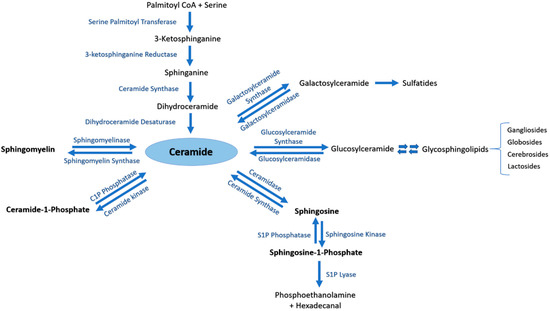
Figure 1. The sphingolipid metabolic pathway. Ceramide is the central metabolite generated in the sphingolipid metabolism by three distinct pathways. Ceramide de novo synthesis consists of Palmitoyl-CoA condensation with serine by the action of serine palmitoyl transferase, followed by a set of reduction and acetylation reactions to generate ceramide. Sphingomyelin (SM) catabolism generates ceramide through the action of sphingomyelinases. The salvage pathway involves N-acylation of fatty acids with a sphingosine backbone to produce ceramide by ceramide synthases. Ceramide can be further phosphorylated to ceramide-1-phosphate (C1P) by ceramide kinase, or converted into complex glycosphingolipids by glucosylceramide or galactosylceramide synthases. Ceramidase is responsible of catabolizing ceramide into sphingosine, which may be further metabolized by sphingosine kinases to generate sphingosine-1-phosphate (S1P).
4. Nuclear Sphingolipids
Many studies have identified sphingolipids as important modulators of key nuclear processes. So far, various subnuclear compartments including the nuclear envelope, nuclear matrix, nucleolus, and chromatin have been described to host various sphingolipid species [12][13][14][15][16][17][18][19][20]. Although nuclear pores should permit the nucleo-cytoplasmic exchange of sphingolipids, many of these nuclear metabolites are in a dynamic state and undergo turnover. Nuclear localization of sphingolipid metabolizing enzymes has been demonstrated. To this end, the utilization of several analytical, biochemical, and microscopic techniques led to the identification and quantification of various nuclear sphingolipid species along with their metabolizing enzymes [21]. In fact, sphingomyelin (SM) is the dominant nuclear sphingolipid variant [22]. Through its metabolism, SM gives rise to ceramides, sphingosine, and S1P, which in turn give other metabolites (table 1).
Table 1. Nuclear sphingolipid metabolites and metabolizing enzymes. This table recapitulates the various nuclear sphingolipid metabolites and enzymes detected in the nuclear compartment with a brief description of their important nuclear functions. NE: nuclear envelop, dsRNA: double stranded RNA.
|
Nuclear Sphingolipid Metabolites |
Nuclear Sphingolipid Producing Enzymes |
Nuclear Sphingolipid Degrading or Converting Enzymes |
Main Nuclear Functions |
|
Sphingomyelin |
Sphingomyelin synthase |
Reverse sphingomyelin synthase Neutral sphingomyelinase |
Maintenance of NE and nucleoplasm structure Regulation of NE permeability and Fluidity Stabilization of DNA and dsRNA |
|
Ceramide |
Ceramide synthase Ceramide desaturase Neutral sphingomyelinase |
Ceramidase Ceramide kinase |
Regulation of Cell cycle arrest, Senescence, and Apoptosis |
|
Ceramide-1-phosphate |
Ceramide kinase |
C1P phosphatase |
Regulation of cell growth and survival |
|
Sphingosine |
Ceramidase |
Ceramide synthase Sphingosine kinase 2 |
Regulation of gene transcription and apoptosis |
|
Sphingosine-1-phosphate |
Sphingosine kinase 2 |
S1P lyase S1P phosphatase |
Epigenetic modulation of gene transcription Regulation of cell cycle progression and apoptosis Stabilization of human telomerase |
5. Role of Sphingolipids in DNA Damage Response
Various chemotherapeutic drugs and DNA damaging agents target sphingolipid metabolizing enzymes. Strong evidence suggests that lipids are involved in DDR and determining cell fate [5]. Most cancer treatments lead to Cer generation which is implicated in cell death response [23]. However, cancer cells tend to develop survival strategies like generating the pro-survival sphingolipid metabolite S1P after the phosphorylation of sphingosine generated by Cer hydrolysis [24]. Hence, the regulation of these metabolites production is of significant importance in determining the cells’ fate in response to DNA damage [5].
All non-surgical cancer therapies aim to eradicate tumor cells while sparing normal tissues through complex cell signaling pathways. Research over the past few decades confirmed that the stress induced by these therapies involves the accumulation of ceramide. However, any dysregulation in this process, due to either decreased generation or increased metabolism of ceramide, confers resistance against these therapies [25]. From this perspective, emerging therapeutic and clinical interventions are under investigation to maximize the positive outcomes of these therapies by a combinatorial approach. For instance, as recombinant human acid sphingomyelinase (rhASM) was previously evaluated in patients with Niemann-Pick disease, the idea of its administration in cancer therapies flourishes. rhASM might be used to induce pro-apoptotic ceramide levels beyond the tolerance of cells. This treatment is more likely to affect tumors than normal tissues [26]. Moreover, a recent study reported that gentamicin, a commonly used anti-microbial drug, can potentially play a role in cancer therapies. The administration of gentamicin highly upregulated acid sphingomyelinase and induced apoptosis in human gastric cancer cells [27]. As cancer cells can develop survival strategies like generating the pro-survival S1P, inhibitors for both SK1 and SK2 were developed. However, sphingosine kinase inhibitors exhibited some downstream off-target effects such as inhibiting ERK and Akt pathways [28]. Hence, further studies should address the development of more specific sphingosine kinase targets for possible clinical trials. Interestingly, the total plasma levels of ceramide, measured in early days after the combined treatment of radio-chemotherapy, can predict tumor responses in patients with liver and lung metastases of colorectal cancer. It allows the identification of patients with high risks of metastases [29]. Therefore, successful discoveries of sphingolipid therapeutic targets and biomarkers will potentially enhance the standard of care therapies by overcoming tumor resistance and developing new effective diagnostic and prognostic sphingolipidomic-based tests.
References
- Christopher R. Gault; Lina M. Obeid; Yusuf A Hannun; An overview of sphingolipid metabolism: from synthesis to breakdown.. Single Molecule and Single Cell Sequencing 2010, 688, 1-23, 10.1007/978-1-4419-6741-1_1.
- Anthony H. Futerman; Yusuf A. Hannun; The complex life of simple sphingolipids. EMBO reports 2004, 5, 777-782, 10.1038/sj.embor.7400208.
- Wenjing Zheng; Jessica Kollmeyer; Holly Symolon; Amin Momin; Elizabeth Munter; Elaine Wang; Samuel Kelly; Jeremy C. Allegood; Ying Liu; Qiong Peng; et al.Harsha RamarajuM. Cameron SullardsMyles CabotAlfred H. Merrill Ceramides and other bioactive sphingolipid backbones in health and disease: Lipidomic analysis, metabolism and roles in membrane structure, dynamics, signaling and autophagy. Biochimica et Biophysica Acta (BBA) - Biomembranes 2006, 1758, 1864-1884, 10.1016/j.bbamem.2006.08.009.
- Alaa Abou Daher; Tatiana El Jalkh; Assaad A. Eid; Alessia Fornoni; Brian Marples; Youssef H. Zeidan; Translational Aspects of Sphingolipid Metabolism in Renal Disorders. International Journal of Molecular Sciences 2017, 18, 2528, 10.3390/ijms18122528.
- Brittany Carroll; Jane Catalina Donaldson; Lina M. Obeid; Sphingolipids in the DNA damage response.. Advances in Biological Regulation 2014, 58, 38-52, 10.1016/j.jbior.2014.11.001.
- Alistair Boath; Christine Graf; Emilie Lidome; Thomas Ullrich; Peter Nussbaumer; Frédéric Bornancin; Regulation and Traffic of Ceramide 1-Phosphate Produced by Ceramide Kinase. Journal of Biological Chemistry 2007, 283, 8517-8526, 10.1074/jbc.m707107200.
- Yusuf A. Hannun; Lina M. Obeid; Principles of bioactive lipid signalling: lessons from sphingolipids. Nature Reviews Molecular Cell Biology 2008, 9, 139-150, 10.1038/nrm2329.
- Chie Ogawa; Akio Kihara; Maiko Gokoh; Yasuyuki Igarashi; Identification and Characterization of a Novel Human Sphingosine-1-phosphate Phosphohydrolase, hSPP2. Journal of Biological Chemistry 2002, 278, 1268-1272, 10.1074/jbc.m209514200.
- Suzanne M. Mandala; Rosemary Thornton; Ismael Galve-Roperh; Samantha Poulton; Courtney Peterson; Ana Olivera; James Bergstrom; Myra B. Kurtz; Sarah Spiegel; Molecular cloning and characterization of a lipid phosphohydrolase that degrades sphingosine-1- phosphate and induces cell death. Proceedings of the National Academy of Sciences 2000, 97, 7859-7864, 10.1073/pnas.120146897.
- S. Pyne; J.S. Long; N.T. Ktistakis; N.J. Pyne; Lipid phosphate phosphatases and lipid phosphate signalling. Biochemical Society Transactions 2005, 33, 1370-1374, 10.1042/bst0331370.
- Mika Ikeda; Akio Kihara; Yasuyuki Igarashi; Sphingosine-1-phosphate lyase SPL is an endoplasmic reticulum-resident, integral membrane protein with the pyridoxal 5′-phosphate binding domain exposed to the cytosol. Biochemical and Biophysical Research Communications 2004, 325, 338-343, 10.1016/j.bbrc.2004.10.036.
- Robert W. Ledeen; Gusheng Wu; Thematic Review Series: Sphingolipids.Nuclear sphingolipids: metabolism and signaling. Journal of Lipid Research 2008, 49, 1176-1186, 10.1194/jlr.r800009-jlr200.
- Alice Alessenko; Subroto Chatterjee; Neutral sphingomyelinase: Localization in rat liver nuclei and involvement in regeneration/proliferation. Molecular and Cellular Biochemistry 1995, 143, 169-174, 10.1007/bf01816950.
- Gusheng Wu; Zi-Hua Lu; Robert W. Ledeen; GM1 Ganglioside in the Nuclear Membrane Modulates Nuclear Calcium Homeostasis During Neurite Outgrowth. Journal of Neurochemistry 2002, 65, 1419-1422, 10.1046/j.1471-4159.1995.65031419.x.
- Marta Micheli; Elisabetta Albi; Claude Leray; Mariapia Viola Magni; Nuclear sphingomyelin protects RNA from RNase action.. FEBS Letters 1998, 431, 443-447, 10.1016/s0014-5793(98)00810-2.
- Kyoji Tsugane; Keiko Tamiya-Koizumi; M. Nagino; Y. Nimura; S Yoshida; A possible role of nuclear ceramide and sphingosine in hepatocyte apoptosis in rat liver.. Journal of Hepatology 1999, 31, 8-17, 10.1016/s0168-8278(99)80158-5.
- Graziella Rossi; Mariapia Viola Magni; Elisabetta Albi; Sphingomyelin-cholesterol and double stranded RNA relationship in the intranuclear complex. Archives of Biochemistry and Biophysics 2007, 459, 27-32, 10.1016/j.abb.2006.11.020.
- Elisabetta Albi; S. Cataldi; G. Rossi; M. Viola Magni; M. Toller; S. Casani; G. Perrella; The nuclear ceramide/diacylglycerol balance depends on the physiological state of thyroid cells and changes during UV-C radiation-induced apoptosis. Archives of Biochemistry and Biophysics 2008, 478, 52-58, 10.1016/j.abb.2008.07.018.
- Nitai C. Hait; Jeremy Allegood; Michael Maceyka; Graham M. Strub; Kuzhuvelil B. Harikumar; Sandeep K. Singh; Cheng Luo; Ronen Marmorstein; Tomasz Kordula; Sheldon Milstien; et al.Sarah Spiegel Regulation of Histone Acetylation in the Nucleus by Sphingosine-1-Phosphate. Science 2009, 325, 1254-1257, 10.1126/science.1176709.
- Celia F. Cave; P. B. Gahan; A Cytochemical and Autoradiographic Investigation of Nucleolar Phospholipids. Caryologia 1970, 23, 303-312, 10.1080/00087114.1970.10796371.
- Shakti Gupta; Mano Ram Maurya; Alfred H Merrill Jr; Christopher K. Glass; Shankar Subramaniam; Integration of lipidomics and transcriptomics data towards a systems biology model of sphingolipid metabolism. BMC Systems Biology 2011, 5, 26-26, 10.1186/1752-0509-5-26.
- Robert W. Ledeen; Gusheng Wu; Sphingolipids of the nucleus and their role in nuclear signaling. Biochimica et Biophysica Acta (BBA) - Molecular and Cell Biology of Lipids 2006, 1761, 588-598, 10.1016/j.bbalip.2006.04.010.
- C. Patrick Reynolds; Barry J. Maurer; Richard N. Kolesnick; Ceramide synthesis and metabolism as a target for cancer therapy. Cancer Letters 2004, 206, 169-180, 10.1016/j.canlet.2003.08.034.
- Christopher R. Gault; Lina M. Obeid; Still benched on its way to the bedside: sphingosine kinase 1 as an emerging target in cancer chemotherapy.. Critical Reviews in Biochemistry and Molecular Biology 2011, 46, 342-51, 10.3109/10409238.2011.597737.
- Thomas H. Beckham; Joseph C. Cheng; S. Tucker Marrison; James S. Norris; Xiang Liu; Interdiction of sphingolipid metabolism to improve standard cancer therapies.. Advances in Breast Cancer Research 2013, 117, 1-36, 10.1016/B978-0-12-394274-6.00001-7.
- Radoslav Savic; Edward Schuchman; Use of Acid Sphingomyelinase for Cancer Therapy. Advances in Cancer Research 2013, 117, 91-115, 10.1016/b978-0-12-394274-6.00004-2.
- Elisabetta Albi; SamuelA Cataldi; Maria Rachele Ceccarini; Carmela Conte; Ivana Ferri; Katia Fettucciari; Federica Filomena Patria; Tommaso Beccari; Michela Codini; Gentamicin Targets Acid Sphingomyelinase in Cancer: The Case of the Human Gastric Cancer NCI-N87 Cells.. International Journal of Molecular Sciences 2019, 20, 4375, 10.3390/ijms20184375.
- Mengda Cao; Chunmei Ji; Yanjun Zhou; Wen Huang; Weiwei Ni; Xunliang Tong; Ji-Fu Wei; Sphingosine kinase inhibitors: A patent review. International Journal of Molecular Medicine 2018, 41, 2450-2460, 10.3892/ijmm.2018.3505.
- Nolwenn Dubois; Emmanuel Rio; Natacha Ripoche; Véronique Ferchaud-Roucher; Marie-Hélène Gaugler; Loïc Campion; Michel Krempf; Christian Carrie; Marc Mahé; Xavier Mirabel; et al.Francois Paris Plasma ceramide, a real-time predictive marker of pulmonary and hepatic metastases response to stereotactic body radiation therapy combined with irinotecan. Radiotherapy and Oncology 2016, 119, 229-235, 10.1016/j.radonc.2016.03.014.





