
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Asahiro Morishita | + 2223 word(s) | 2223 | 2021-08-03 06:21:24 | | | |
| 2 | Dean Liu | Meta information modification | 2223 | 2021-08-03 10:40:01 | | |
Video Upload Options
Several miRNAs are associated with organ-specific and systemic fibrosis in the liver. Individual expression of miRNAs in plasma or serum is useful for liver fibrosis detection. In addition, some miRNAs can distinguish between early and late fibrosis with high sensitivity and specificity equal to or greater than the APRI and Fib-4 index.
1. Introduction
Activation of hepatic stellate cells (HSCs) is a pivotal event in liver fibrosis [1], and various inflammatory and fibrotic pathways are involved [2]. Activated HSCs are precursors of myofibroblasts that produce extracellular matrix (ECM) in the liver. Therefore, the treatment of hepatic fibrosis requires the prevention of hepatocellular damage and control of activated HSCs [3]. If HSC inactivation can be induced by cell- or target-specific pharmacological interventions, then, more effective, less toxic, and more accurate anti-fibrotic treatments than the currently available ones can be developed.
HSCs, also known as Ito cells, are fat-storing cells containing vitamin A which are found in the space of Disse and account for approximately 10% of the total number of liver cells. HSCs correspond to pericytes that surround the sinusoidal endothelial cells with branch-like projections and contact hepatocytes [4]. HSCs have also been suggested to serve as antigen-presenting cells in the liver, presenting lipid antigens to CD1-restricted T lymphocytes such as natural killer T (NKT) cells and promoting NKT cell proliferation by presenting hepatic NKT cell lipid antigens via interleukin (IL)-15 [5].
EVs have attracted attention as delivery media for functional RNAs, including miRNAs. Exosomes are a subset of EVs which are secreted from the endoplasmic reticulum in cells with a lipid bilayer membrane with an average diameter of 30–100 nm [6][7]. They contain mRNAs and miRNAs which they transmit between cells [6]. The endoplasmic reticulum-derived EVs, including exosomes, contain RNA, proteins, and other information from the donor (EV–secreting) cell, and transmit this information to the recipient cell. miRNAs are mostly contained in EVs and are transported to recipient cells by intercellular transport, affecting signal transduction in recipient cells and cell phenotype. miRNAs in EVs extracted from patients are useful as disease biomarkers for various diseases [6]. In addition, a new drug delivery system that hijacks exosomes to deliver anti-miR oligonucleotides into the cells that receive them has been developed and is expected to have therapeutic applications [8].
Furthermore, it is expected to be applied to the diagnosis of liver fibrosis and to the development of novel therapeutic agents using miRNAs.
2. Epigenetic Changes in Liver Fibrosis/Cirrhosis
Several miRNAs are associated with organ-specific and systemic fibrosis in the liver [9]. Individual expression of miRNAs in plasma or serum is useful for liver fibrosis detection [10][11]. In addition, some miRNAs can distinguish between early and late fibrosis with high sensitivity and specificity equal to or greater than the APRI and Fib-4 index [12][13]. For example, patients with advanced cirrhosis showed significantly lower levels of miR-29a in their serum compared to healthy controls and patients with early fibrosis [14]. Additionally, serum levels of miR-138 and miR-143 are characteristic of the later stages of liver fibrosis and thus, miR-138 may be useful for detecting fibrosis in its early stages [15]. Furthermore, serum levels of miR-34a and miR-122 correlate with the progression of fibrosis, especially in patients with chronic hepatitis C or nonalcoholic fatty liver disease (NAFLD) [16]. In addition, miR-221 is upregulated in patients with liver cirrhosis [17]. Elucidation of the relationship between miRNAs and liver fibrosis may be useful for detection of fibrosis without invasive liver biopsy, early therapeutic intervention, and identification of high-risk patients. There are various microRNAs that are expected to be biomarkers for liver diseases ( Table 1 ).
| miRNA | Expression Level | References | |
|---|---|---|---|
| Liver fibrosis | miR-29a | Down | [14] |
| Liver fibrosis | miR-138 | Up | [15] |
| Liver fibrosis | miR-143 | Up | [15] |
| Liver fibrosis | miR-34a | Up | [16] |
| Liver fibrosis | miR-122 | Up | [16] |
| Liver fibrosis | miR-221 | Up | [17] |
| Portal vein thrombus | miR-19a | Up | [18] |
| Portal vein thrombus | miR-34a | Up | [18] |
| Portal vein thrombus | miR-21 | Up | [19] |
| SBP | miR-122 | Down | [20] |
| SBP | miR-155 | Up | [21][22] |
| SBP | miR-223 | Up | [23] |
| HBV | miR-21-5p | Up | [24] |
| HBV | miR-125a-5p | Up | [25] |
| HBV | miR-27a | Up | [26] |
| HBV | miR-181b | Up | [27] |
| HCV | miR-222 | Up | [28] |
| HCV | miR-221 | Up | [28][29] |
| HCV | miR-21 | Up | [30] |
| HCV | miR-99a | Down | [31] |
| HCV | miR-215-5p | Up | [32] |
| HCV | miR-483-5p | Up | [32] |
| HCV | miR-193b-3p | Up | [32] |
| HCV | miR-34a | Up | [29][32] |
| HCV | miR-885-5p | Up | [32] |
| HCV | miR-26b-5p | Down | [32] |
| HCV | miR-197-3p | Down | [32] |
| HCV | miR-122 | Down | [33] |
| HCV | miR-16 | Up | [29] |
| HCV | miR-200c | Up | [34] |
| HCV | miR-20a | Up | [35] |
| NASH/NAFLD | miR-34a | Up | [36][37] |
| NASH/NAFLD | miR-372-3p | Down | [38] |
| NASH/NAFLD | miR-373-3p | Down | [38] |
| AIH | miR-133a | Up | [39][40] |
| AIH | miR-122 | Down | [41] |
| AIH | miR-21 | Down | [41] |
| PBC | miR-21 | Up | [42] |
| PBC | miR-210 | Up | [43] |
| PBC | let-7b | Up | [44] |
| PBC | miR-520a-5p | Up | [44] |
| PBC | miR-125b | Down | [44] |
| PBC | miR-92a | Down | [45] |
| PBC | miR-223-3p | Down | [46] |
| PBC | miR-21-5p | Down | [46] |
| PBC | miR-139-5p | Down | [47] |
| PSC | miR-150-5p | Down | [48] |
| PSC | miR-122 | Down | [49] |
| PSC | miR-200c | Down | [50] |
Recently, the relationship between liver diseases and various miRNAs has been confirmed ( Table 2 ). Some miRNAs such as miR-21, miR-221/222, and miR-181b, promote liver fibrosis through the TGF-β and NF-κB pathways [51]. In addition, miR-221 regulates multiple targets, including cyclin-dependent kinase inhibitors (CDKN1C or CDKN1B), cytokine signaling 1, E-cadherin, phosphatase and tensin homolog (PTEN), and Bcl-2 modifying factor, which are involved in liver fibrosis [17]. miR-214 also plays an important role in liver fibrosis by regulating the expression of suppressor of fused homolog protein, and knocking down its expression alleviates liver fibrosis in carbon tetrachloride (CCL4)-treated mice [52]. Moreover, the knockdown of the miR-23b miRNA cluster promotes bile duct differentiation and suppresses or restores TGF-β-induced liver fibrosis depending on stellate cell activation [53].
| miRNA | Predicted Target | Involvement in Disease Progression | References | |
|---|---|---|---|---|
| Liver fibrosis | miR-21 | TGF-β pathway, NF-κB pathway | promote | [51] |
| Liver fibrosis | miR-221/222 | TGF-β pathway, NF-κB pathway | promote | [51] |
| Liver fibrosis | miR-181b | TGF-β pathway, NF-κB pathway | promote | [51] |
| Liver fibrosis | miR-221 | CDKN1C, CDKN1B, Socs1, E-cadherin, PTEN, BMF | promote | [17] |
| Liver fibrosis | miR-214 | Sufu | promote | [52] |
| Liver fibrosis | miR-23b | TGF-β pathway | inhibit | [53] |
| Liver fibrosis | miR-30a | Beclin1,α-SMA,TIMP-1,Collagen I | inhibit | [54] |
| Liver fibrosis | miR-29b | TGF-β pathway | inhibit | [51] |
| Liver fibrosis | miR-101 | TGF-β pathway | inhibit | [51] |
| Liver fibrosis | miR-122 | TGF-β pathway | inhibit | [51] |
| Liver fibrosis | miR-214-3p | TGF-β pathway | inhibit | [51] |
| Liver fibrosis | miR-29a | BRD4, CD36 | inhibit | [55][56][57] |
| Liver fibrosis | miR-34 | TGF-β1/Smad3 pathway | inhibit | [58] |
| Liver fibrosis | miR-223 | NLRP3 | inhibit | [59] |
| Liver fibrosis | miR-455-3p | HSF1 | inhibit | [60] |
| Liver fibrosis | miR-125b | Gli3 | inhibit | [61] |
| Liver fibrosis | miR-378 | Gli3 | inhibit | [62] |
| Liver fibrosis | miR-152 | Gli3 | inhibit | [63] |
| Portal hypertension | miR-29 | VEGF | inhibit | [64] |
| Portal hypertension | miR-126 family | SPRED1, PIK3R2/p85-beta | inhibit | [65] |
| HBV | miR-21-5p | TGF-β1 | promote | [24] |
| HBV | miR-27a | PPARγ, FOXO1, APC, P53, RXRα | promote | [26] |
| HBV | miR-181b | PTEN/Akt pathway | promote | [27] |
| HCV | miR-21 | Smad7 | promote | [30] |
| HCV | miR-99a | mTOR | inhibit | [31] |
| HCV | miR-200c | FAP-1 | promote | [34] |
| ALD | miR-122 | HIF1α | inhibit | [66] |
| ALD | miR-223 | IL-6-p47phox pathway | inhibit | [67] |
| ALD | miR-155 | PPARγ, PPRE | promote | [68] |
| ALD | miR181b-3p | importin α5 | inhibit | [69] |
| ALD | miR-217 | SIRT1 | promote | [70] |
| NASH/NAFLD | miR-21 | STAT3 signaling pathway, TGF-β/Smad3/Smad7 signaling pathway | promote | [71] |
| NASH/NAFLD | miR-29a | CD36 | inhibit | [72] |
| NASH/NAFLD | miR-122 | AGPAT1, DGAT1 | inhibit | [73] |
| NASH/NAFLD | miR-34a | PPARα signaling pathway | promote | [36][37] |
| NASH/NAFLD | miR-129-5p | PEG3 | inhibit | [74] |
| NASH/NAFLD | miR-188-5p | PTEN/PI3K/AKT pathway | promote | [75] |
| NASH/NAFLD | miR-223 | Cxcl10, Nlrp3, Taz | inhibit | [76] |
| NASH/NAFLD | miR-27 | insulin signaling pathway | promote | [77] |
| NASH/NAFLD | miR-372-3p | AEBP1 | inhibit | [38] |
| NASH/NAFLD | miR-373-3p | AEBP1 | inhibit | [38] |
| AIH | miR-375 | AEG-1 | promote | [78] |
| AIH | miR-138 | p53 | promote | [79] |
| AIH | miR-15a/miR-16-1 | aryl hydrocarbon receptor-IL-22 regulatory axis | promote | [80] |
| AIH | miR-674-5p | 5-LO | inhibit | [81] |
| AIH | miRNA-143 | phosphorylation of TAK1 | inhibit | [82] |
| AIH | miR-223 | AIM2 | inhibit | [83] |
| PBC | miR-21 | CDK2AP1 | promote | [42] |
| PBC | miR-210 | MLL4 | promote | [43] |
| PBC | miR-506 | AE 2, InsP3R3 | promote | [41] |
| PBC | miR-92a | IL-17A | inhibit | [45] |
| PBC | miR-223-3p | TGFBR2,MEF2C,FOXP1,RBPJ | promote | [46] |
| PBC | miR-21-5p | TGFBR2,MEF2C,FOXP2,RBPJ | promote | [24] |
| PBC | miR-139-5p | c-FOS | promote | [47] |
| PSC | miR-7a | Ngn-3 | promote | [84] |
| PSC | miR-873-5p | GNMT | promote | [85] |
| PSC | miR-24 | menin | inhibit | [86] |
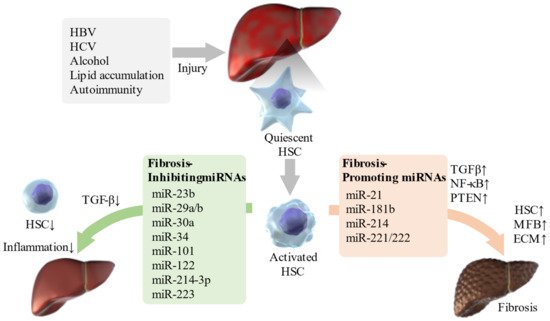
Thus, various miRNAs are involved in the regulation of liver fibrosis ( Figure 1 ). The identification of miRNAs involved in the pathogenesis of liver fibrosis will enable miRNA-based therapies, and even if this is difficult, miRNA-associated mechanisms can be targeted for therapy.
In contrast to EVs in serum extracted from mice with induced liver fibrosis, miR-34c, miR-151-3p, miR-483-5p, miR-532-5p, and miR-687 were upregulated in serum EVs from mice without fibrosis. When these EVs were administered to mice with CCL4-induced liver injury, hepatocellular damage and liver fibrosis were suppressed in the healthy mouse-derived EV group, and inflammatory cytokines and transaminases in the blood were reduced. Furthermore, in serum EVs from patients with F3/4 hepatic fibrosis and healthy subjects, levels of miR-34c, miR-151-3p, miR-483-5p, and miR-532-5p were upregulated in healthy subjects compared to levels in patients with F3/4 hepatic fibrosis. When EVs from healthy subjects were administered to human-derived HSCs, HSC activation was suppressed. These EV miRNAs suppress HSC activation and contribute to the suppression of liver fibrosis [87].
3. miRNAs Associated with Complications of Liver Cirrhosis
In addition to HCC, gastrointestinal bleeding due to portal hypertension, ascites, portal vein thrombosis, and HE are some of the most common complications of cirrhosis.
Portal hypertension is caused by an increase in portal venous inflow and intrahepatic vascular resistance, leading to esophageal varices, ascites, HE, and hypersplenism, and it is a major cause of death in patients with cirrhosis [88]. TGF-β-mediated HSC activation is involved in ECM production and is a mechanistic factor in regulating vascular resistance and pressure in the liver [89]. Animal studies have shown that inhibition of TGF-β1 synthesis and blockade of TGF-β receptors can significantly reduce portal hypertensive pressure [90]. Intrahepatic angiogenesis by vascular endothelial growth factor (VEGF), an angiogenic growth factor, is also a cause of sinusoidal systemic circulation and portal hypertension [91]. miR-29 acts as an anti-fibrotic mediator by inhibiting angiogenic factors such as VEGF [64]. The miR-126 family is associated with angiogenesis and directly inhibits negative regulators of the VEGF pathway, such as the Sprouty-related, equine herpesvirus-1 domain-containing protein 1 (SPRED1) and phosphoinositol-3 kinase regulatory subunit 2 (PIK3R2) [65]. Thus, these miRNAs, which affect TGF-β and VEGF, may be key to the treatment of portal hypertension.
A portal vein thrombus in cirrhosis causes worsening of the liver reserve, gastroesophageal varices, and ascites. The pathophysiology of portal vein thrombosis encompasses one or more of the following features: decreased portal blood flow, hypercoagulable state, and damage to the vascular endothelium. As liver function declines, platelets and hepatic-derived coagulation factors are reduced, while hepatic-derived anticoagulation factors are also reduced, and when this balance is disrupted, portal vein thrombi form [92]. Notably, miR-19a and miR-34a levels are correlated with portal vein thrombosis [18], and miR-21 is considered an independent predictor of portal vein thrombosis in patients with HCC [19].
Patients with ascites, spontaneous bacterial peritonitis (SBP), and hepatorenal syndrome had significantly lower levels of miR-122 than those without these complications. In addition, serum miR-122 levels were associated with survival of patients with cirrhosis, independent of the Model for End-Stage Liver Disease (MELD) score or patient ’s age [20]. One study indicated that miR-155 is an outstanding diagnostic marker for SBP, and detection of both serum CD64 and calprotectin levels also provide a more useful diagnosis when using blood samples from patients with cirrhosis and ascites [21]. Furthermore, miR-155 [22] and miR-223 [23] were elevated in ascites from patients with cirrhosis and SBP, indicating that these miRNAs may be involved in the immune response in ascites upon SBP. Animal models of HE also show changes in miRNAs in the blood and cerebral cortex [93][94]. In the HE model, ammonia-induced changes in miRNA expression regulate the expression of heme oxygenase 1 (HO-1) and induce astrocyte senescence [95].
4. Association of the Causes of Liver Cirrhosis and miRNAs
Direct-acting antivirals are highly effective, but HCV elimination is not a cure for liver disease, especially in patients with advanced fibrosis or cirrhosis. However, miRNAs show promise as a treatment for residual liver fibrosis after antiviral treatment.
Since the incidence of NAFLD/NASH is expected to continue to increase, there is an urgent need to develop early diagnosis and treatment methods using miRNAs.
Autoimmune liver diseases include autoimmune hepatitis (AIH), primary biliary cholangitis (PBC), and primary sclerosing cholangitis (PSC), which are characterized by chronic liver and biliary inflammation. These diseases require persistent treatment, but the underlying causes of these diseases are still unknown. To date, numerous susceptible loci for autoimmune liver diseases in the human leukocyte antigen (HLA) and non-HLA regions have been identified by a genome-wide association studies (GWAS) [96][97]. However, genetic analysis alone is inadequate to identify the cause of autoimmune liver diseases, and environmental factors may be involved in the development of these diseases.
In recent years, the pathological and diagnostic relevance of miRNAs in autoimmune liver diseases have been reported.
References
- Koyama, Y.; Brenner, D.A. Liver inflammation and fibrosis. J. Clin. Investig. 2017, 127, 55–64.
- Seki, E.; Brenner, D.A. Recent advancement of molecular mechanisms of liver fibrosis. J. Hepato Biliary Pancreat. Sci. 2015, 22, 512–518.
- Higashi, T.; Friedman, S.L.; Hoshida, Y. Hepatic stellate cells as key target in liver fibrosis. Adv. Drug Deliv. Rev. 2017, 121, 27–42.
- Blomhoff, R.; Wake, K. Perisinusoidal stellate cells of the liver: Important roles in retinol metabolism and fibrosis. FASEB J. 1991, 5, 271–277.
- Winau, F.; Hegasy, G.; Weiskirchen, R.; Weber, S.; Cassan, C.; Sieling, P.A.; Modlin, R.L.; Liblau, R.S.; Gressner, A.M.; Kaufmann, S.H. Ito cells are liver-resident antigen-presenting cells for activating T cell responses. Immunity 2007, 26, 117–129.
- Valadi, H.; Ekstrom, K.; Bossios, A.; Sjostrand, M.; Lee, J.J.; Lotvall, J.O. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654–659.
- Kalluri, R.; LeBleu, V.S. The biology, function, and biomedical applications of exosomes. Science 2020, 367.
- Yamayoshi, A.; Oyama, S.; Kishimoto, Y.; Konishi, R.; Yamamoto, T.; Kobori, A.; Harada, H.; Ashihara, E.; Sugiyama, H.; Murakami, A. Development of antibody-oligonucleotide complexes for targeting exosomal microRNA. Pharmaceutics 2020, 12, 545.
- O’Reilly, S. MicroRNAs in fibrosis: Opportunities and challenges. Arthritis Res. Ther. 2016, 18, 11.
- Iacob, D.G.; Rosca, A.; Ruta, S.M. Circulating microRNAs as non-invasive biomarkers for hepatitis B virus liver fibrosis. World J. Gastroenterol. 2020, 26, 1113–1127.
- Wang, J.; Chu, E.S.; Chen, H.Y.; Man, K.; Go, M.Y.; Huang, X.R.; Lan, H.Y.; Sung, J.J.; Yu, J. MicroRNA-29b prevents liver fibrosis by attenuating hepatic stellate cell activation and inducing apoptosis through targeting PI3K/AKT pathway. Oncotarget 2015, 6, 7325–7338.
- Wang, T.Z.; Lin, D.D.; Jin, B.X.; Sun, X.Y.; Li, N. Plasma microRNA: A novel non-invasive biomarker for HBV-associated liver fibrosis staging. Exp. Ther. Med. 2019, 17, 1919–1929.
- Appourchaux, K.; Dokmak, S.; Resche-Rigon, M.; Treton, X.; Lapalus, M.; Gattolliat, C.H.; Porchet, E.; Martinot-Peignoux, M.; Boyer, N.; Vidaud, M.; et al. MicroRNA-based diagnostic tools for advanced fibrosis and cirrhosis in patients with chronic hepatitis B and C. Sci. Rep. 2016, 6, 34935.
- Roderburg, C.; Urban, G.W.; Bettermann, K.; Vucur, M.; Zimmermann, H.; Schmidt, S.; Janssen, J.; Koppe, C.; Knolle, P.; Castoldi, M.; et al. Micro-RNA profiling reveals a role for miR-29 in human and murine liver fibrosis. Hepatology 2011, 53, 209–218.
- El-Ahwany, E.; Nagy, F.; Zoheiry, M.; Shemis, M.; Nosseir, M.; Taleb, H.A.; El Ghannam, M.; Atta, R.; Zada, S. Circulating miRNAs as predictor markers for activation of hepatic stellate cells and progression of HCV-induced liver fibrosis. Electron. Physician 2016, 8, 1804–1810.
- Cermelli, S.; Ruggieri, A.; Marrero, J.A.; Ioannou, G.N.; Beretta, L. Circulating microRNAs in patients with chronic hepatitis C and non-alcoholic fatty liver disease. PLoS ONE 2011, 6, e23937.
- Markovic, J.; Sharma, A.D.; Balakrishnan, A. MicroRNA-221: A fine tuner and potential biomarker of chronic liver injury. Cells 2020, 9, 1767.
- Motawi, T.K.; Shaker, O.G.; El-Maraghy, S.A.; Senousy, M.A. Serum microRNAs as potential biomarkers for early diagnosis of hepatitis C virus-related hepatocellular carcinoma in Egyptian patients. PLoS ONE 2015, 10, e0137706.
- Yoon, J.S.; Kim, G.; Lee, Y.R.; Park, S.Y.; Tak, W.Y.; Kweon, Y.O.; Park, J.G.; Lee, H.W.; Han, Y.S.; Ha, H.T.; et al. Clinical significance of microRNA-21 expression in disease progression of patients with hepatocellular carcinoma. Biomark. Med. 2018, 12, 1105–1114.
- Waidmann, O.; Koberle, V.; Brunner, F.; Zeuzem, S.; Piiper, A.; Kronenberger, B. Serum microRNA-122 predicts survival in patients with liver cirrhosis. PLoS ONE 2012, 7, e45652.
- Nabiel, Y.; Barakat, G.; Abed, S. Serum CD64 and ascitic fluid calprotectin and microRNA-155 as potential biomarkers of spontaneous bacterial peritonitis. Eur. J. Gastroenterol. Hepatol. 2019, 31, 1064–1069.
- Lutz, P.; M’haimid, M.; Pohlmann, A.; Lehmann, J.; Jansen, C.; Schierwagen, R.; Klein, S.; Strassburg, C.P.; Spengler, U.; Trebicka, J. MicroRNA-155 is upregulated in ascites in patients with spontaneous bacterial peritonitis. Sci. Rep. 2017, 7, 40556.
- Schindler, P.; Kupcinskas, J.; Juzenas, S.; Skieceviciene, J.; Salteniene, V.; Schulz, C.; Weigt, J.; Malfertheiner, P.; Link, A. Expression of microRNAs in the ascites of patients with peritoneal carcinomatosis and peritonitis. Cancer Cytopathol. 2018, 126, 353–363.
- Wang, W.; Liu, R.; Su, Y.; Li, H.; Xie, W.; Ning, B. MicroRNA-21-5p mediates TGF-beta-regulated fibrogenic activation of spinal fibroblasts and the formation of fibrotic scars after spinal cord injury. Int. J. Biol. Sci. 2018, 14, 178–188.
- Zheng, J.; Zhou, Z.; Xu, Z.; Li, G.; Dong, P.; Chen, Z.; Lin, D.; Chen, B.; Yu, F. Serum microRNA-125a-5p, a useful biomarker in liver diseases, correlates with disease progression. Mol. Med. Rep. 2015, 12, 1584–1590.
- Zhang, H.; Yan, X.L.; Guo, X.X.; Shi, M.J.; Lu, Y.Y.; Zhou, Q.M.; Chen, Q.L.; Hu, Y.Y.; Xu, L.M.; Huang, S.; et al. MiR-27a as a predictor for the activation of hepatic stellate cells and hepatitis B virus-induced liver cirrhosis. Oncotarget 2018, 9, 1075–1090.
- Yu, F.; Zhou, G.; Li, G.; Chen, B.; Dong, P.; Zheng, J. Serum miR-181b is correlated with hepatitis B virus replication and disease progression in chronic hepatitis B patients. Dig. Dis. Sci. 2015, 60, 2346–2352.
- Abdel-Al, A.; El-Ahwany, E.; Zoheiry, M.; Hassan, M.; Ouf, A.; Abu-Taleb, H.; Abdel Rahim, A.; El-Talkawy, M.D.; Zada, S. MiRNA-221 and miRNA-222 are promising biomarkers for progression of liver fibrosis in HCV Egyptian patients. Virus Res. 2018, 253, 135–139.
- Mourad, L.; El-Ahwany, E.; Zoheiry, M.; Abu-Taleb, H.; Hassan, M.; Ouf, A.; Rahim, A.A.; Hassanien, M.; Zada, S. Expression analysis of liver-specific circulating microRNAs in HCV-induced hepatocellular carcinoma in Egyptian patients. Cancer Biol. Ther. 2018, 19, 400–406.
- Marquez, R.T.; Bandyopadhyay, S.; Wendlandt, E.B.; Keck, K.; Hoffer, B.A.; Icardi, M.S.; Christensen, R.N.; Schmidt, W.N.; McCaffrey, A.P. Correlation between microRNA expression levels and clinical parameters associated with chronic hepatitis C viral infection in humans. Lab. Investig. 2010, 90, 1727–1736.
- Lee, E.B.; Sung, P.S.; Kim, J.H.; Park, D.J.; Hur, W.; Yoon, S.K. MicroRNA-99a restricts replication of hepatitis C virus by targeting mTOR and de novo Lipogenesis. Viruses 2020, 12, 696.
- Cabral, B.C.A.; Hoffmann, L.; Bottaro, T.; Costa, P.F.; Ramos, A.L.A.; Coelho, H.S.M.; Villela-Nogueira, C.A.; Urmenyi, T.P.; Faffe, D.S.; Silva, R. Circulating microRNAs associated with liver fibrosis in chronic hepatitis C patients. Biochem. Biophys. Rep. 2020, 24, 100814.
- Halasz, T.; Horvath, G.; Par, G.; Werling, K.; Kiss, A.; Schaff, Z.; Lendvai, G. MiR-122 negatively correlates with liver fibrosis as detected by histology and FibroScan. World J. Gastroenterol. 2015, 21, 7814–7823.
- Ramachandran, S.; Ilias Basha, H.; Sarma, N.J.; Lin, Y.; Crippin, J.S.; Chapman, W.C.; Mohanakumar, T. Hepatitis C virus induced miR200c down modulates FAP-1, a negative regulator of Src signaling and promotes hepatic fibrosis. PLoS ONE 2013, 8, e70744.
- Shrivastava, S.; Petrone, J.; Steele, R.; Lauer, G.M.; Di Bisceglie, A.M.; Ray, R.B. Up-regulation of circulating miR-20a is correlated with hepatitis C virus-mediated liver disease progression. Hepatology 2013, 58, 863–871.
- Ding, J.; Li, M.; Wan, X.; Jin, X.; Chen, S.; Yu, C.; Li, Y. Effect of miR-34a in regulating steatosis by targeting PPARalpha expression in nonalcoholic fatty liver disease. Sci. Rep. 2015, 5, 13729.
- Xin, S.; Zhan, Q.; Chen, X.; Xu, J.; Yu, Y. Efficacy of serum miRNA test as a non-invasive method to diagnose nonalcoholic steatohepatitis: A systematic review and meta-analysis. BMC Gastroenterol. 2020, 20, 186.
- Gerhard, G.S.; Hanson, A.; Wilhelmsen, D.; Piras, I.S.; Still, C.D.; Chu, X.; Petrick, A.T.; DiStefano, J.K. AEBP1 expression increases with severity of fibrosis in NASH and is regulated by glucose, palmitate, and miR-372-3p. PLoS ONE 2019, 14, e0219764.
- Jia, H.Y.; Chen, F.; Chen, J.Z.; Wu, S.S.; Wang, J.; Cao, Q.Y.; Chen, Z.; Zhu, H.H. MicroRNA expression profiles related to early stage murine concanavalin A-induced hepatitis. Cell Physiol. Biochem. 2014, 33, 1933–1944.
- Tadokoro, T.; Morishita, A.; Sakamoto, T.; Fujihara, S.; Fujita, K.; Mimura, S.; Oura, K.; Nomura, T.; Tani, J.; Yoneyama, H.; et al. Galectin9 ameliorates fulminant liver injury. Mol. Med. Rep. 2017, 16, 36–42.
- Huang, C.; Xing, X.; Xiang, X.; Fan, X.; Men, R.; Ye, T.; Yang, L. MicroRNAs in autoimmune liver diseases: From diagnosis to potential therapeutic targets. Biomed. Pharmacother. 2020, 130, 110558.
- Afonso, M.B.; Rodrigues, P.M.; Simao, A.L.; Gaspar, M.M.; Carvalho, T.; Borralho, P.; Banales, J.M.; Castro, R.E.; Rodrigues, C.M.P. MiRNA-21 ablation protects against liver injury and necroptosis in cholestasis. Cell Death Differ. 2018, 25, 857–872.
- Kim, Y.C.; Jung, H.; Seok, S.; Zhang, Y.; Ma, J.; Li, T.; Kemper, B.; Kemper, J.K. MicroRNA-210 promotes bile acid-induced cholestatic liver injury by targeting mixed-lineage leukemia-4 methyltransferase in mice. Hepatology 2020, 71, 2118–2134.
- Sakamoto, T.; Morishita, A.; Nomura, T.; Tani, J.; Miyoshi, H.; Yoneyama, H.; Iwama, H.; Himoto, T.; Masaki, T. Identification of microRNA profiles associated with refractory primary biliary cirrhosis. Mol. Med. Rep. 2016, 14, 3350–3356.
- Liang, D.Y.; Hou, Y.Q.; Luo, L.J.; Ao, L. Altered expression of miR-92a correlates with Th17 cell frequency in patients with primary biliary cirrhosis. Int. J. Mol. Med. 2016, 38, 131–138.
- Wang, X.; Wen, X.; Zhou, J.; Qi, Y.; Wu, R.; Wang, Y.; Kui, Y.; Hua, R.; Jin, Q. MicroRNA-223 and microRNA-21 in peripheral blood B cells associated with progression of primary biliary cholangitis patients. PLoS ONE 2017, 12, e0184292.
- Katsumi, T.; Ninomiya, M.; Nishina, T.; Mizuno, K.; Tomita, K.; Haga, H.; Okumoto, K.; Saito, T.; Shimosegawa, T.; Ueno, Y. MiR-139-5p is associated with inflammatory regulation through c-FOS suppression, and contributes to the progression of primary biliary cholangitis. Lab. Investig. 2016, 96, 1165–1177.
- Wu, X.; Xia, M.; Chen, D.; Wu, F.; Lv, Z.; Zhan, Q.; Jiao, Y.; Wang, W.; Chen, G.; An, F. Profiling of downregulated blood-circulating miR-150-5p as a novel tumor marker for cholangiocarcinoma. Tumor Biol. 2016, 37, 15019–15029.
- Friedrich, K.; Baumann, C.; Wannhoff, A.; Rupp, C.; Mehrabi, A.; Weiss, K.H.; Gotthardt, D.N. Serum miRNA-122 is an independent biomarker of survival in patients with primary sclerosing cholangitis. J. Gastrointest. Liver Dis. 2018, 27, 145–150.
- Bernuzzi, F.; Marabita, F.; Lleo, A.; Carbone, M.; Mirolo, M.; Marzioni, M.; Alpini, G.; Alvaro, D.; Boberg, K.M.; Locati, M.; et al. Serum microRNAs as novel biomarkers for primary sclerosing cholangitis and cholangiocarcinoma. Clin. Exp. Immunol. 2016, 185, 61–71.
- Hayes, C.N.; Chayama, K. MicroRNAs as biomarkers for liver disease and hepatocellular carcinoma. Int. J. Mol. Sci. 2016, 17, 280.
- Ma, L.; Yang, X.; Wei, R.; Ye, T.; Zhou, J.K.; Wen, M.; Men, R.; Li, P.; Dong, B.; Liu, L.; et al. MicroRNA-214 promotes hepatic stellate cell activation and liver fibrosis by suppressing Sufu expression. Cell Death Dis. 2018, 9, 718.
- Rogler, C.E.; Matarlo, J.S.; Kosmyna, B.; Fulop, D.; Rogler, L.E. Knockdown of miR-23, miR-27, and miR-24 alters fetal liver development and blocks fibrosis in mice. Gene Expr. 2017, 17, 99–114.
- Chen, J.; Yu, Y.; Li, S.; Liu, Y.; Zhou, S.; Cao, S.; Yin, J.; Li, G. MicroRNA-30a ameliorates hepatic fibrosis by inhibiting Beclin1-mediated autophagy. J. Cell. Mol. Med. 2017, 21, 3679–3692.
- Matsumoto, Y.; Itami, S.; Kuroda, M.; Yoshizato, K.; Kawada, N.; Murakami, Y. MiR-29a assists in preventing the activation of human stellate cells and promotes recovery from liver fibrosis in mice. Mol. Ther. 2016, 24, 1848–1859.
- Lin, H.Y.; Wang, F.S.; Yang, Y.L.; Huang, Y.H. MicroRNA-29a suppresses CD36 to ameliorate high fat diet-induced steatohepatitis and liver fibrosis in mice. Cells 2019, 8, 1298.
- Huang, Y.H.; Kuo, H.C.; Yang, Y.L.; Wang, F.S. MicroRNA-29a is a key regulon that regulates BRD4 and mitigates liver fibrosis in mice by inhibiting hepatic stellate cell activation. Int. J. Med. Sci. 2019, 16, 212–220.
- Feili, X.; Wu, S.; Ye, W.; Tu, J.; Lou, L. MicroRNA-34a-5p inhibits liver fibrosis by regulating TGF-beta1/Smad3 pathway in hepatic stellate cells. Cell Biol. Int. 2018, 42, 1370–1376.
- Calvente, C.J.; Tameda, M.; Johnson, C.D.; Del Pilar, H.; Lin, Y.C.; Adronikou, N.; De Mollerat Du Jeu, X.; Llorente, C.; Boyer, J.; Feldstein, A.E. Neutrophils contribute to spontaneous resolution of liver inflammation and fibrosis via microRNA-223. J. Clin. Investig. 2019, 129, 4091–4109.
- Wei, S.; Wang, Q.; Zhou, H.; Qiu, J.; Li, C.; Shi, C.; Zhou, S.; Liu, R.; Lu, L. MiR-455-3p alleviates hepatic stellate cell activation and liver fibrosis by suppressing HSF1 expression. Mol. Ther. Nucleic Acids 2019, 16, 758–769.
- Hu, Z.; Li, L.; Ran, J.; Chu, G.; Gao, H.; Guo, L.; Chen, J. MiR-125b acts as anti-fibrotic therapeutic target through regulating Gli3 In Vivo and In Vitro. Ann. Hepatol. 2019, 18, 825–832.
- Hyun, J.; Wang, S.; Kim, J.; Rao, K.M.; Park, S.Y.; Chung, I.; Ha, C.S.; Kim, S.W.; Yun, Y.H.; Jung, Y. MicroRNA-378 limits activation of hepatic stellate cells and liver fibrosis by suppressing Gli3 expression. Nat. Commun. 2016, 7, 10993.
- Li, L.; Zhang, L.; Zhao, X.; Cao, J.; Li, J.; Chu, G. Downregulation of miR-152 contributes to the progression of liver fibrosis via targeting Gli3 In Vivo and In Vitro. Exp. Ther. Med. 2019, 18, 425–434.
- Zhu, H.; Fan, G.C. Role of microRNAs in the reperfused myocardium towards post-infarct remodelling. Cardiovasc. Res. 2012, 94, 284–292.
- Fish, J.E.; Santoro, M.M.; Morton, S.U.; Yu, S.; Yeh, R.F.; Wythe, J.D.; Ivey, K.N.; Bruneau, B.G.; Stainier, D.Y.; Srivastava, D. MiR-126 regulates angiogenic signaling and vascular integrity. Dev. Cell 2008, 15, 272–284.
- Satishchandran, A.; Ambade, A.; Rao, S.; Hsueh, Y.C.; Iracheta-Vellve, A.; Tornai, D.; Lowe, P.; Gyongyosi, B.; Li, J.; Catalano, D.; et al. MicroRNA 122, regulated by GRLH2, protects livers of mice and patients from ethanol-induced liver disease. Gastroenterology 2018, 154, 238–252.e7.
- Li, M.; He, Y.; Zhou, Z.; Ramirez, T.; Gao, Y.; Gao, Y.; Ross, R.A.; Cao, H.; Cai, Y.; Xu, M.; et al. MicroRNA-223 ameliorates alcoholic liver injury by inhibiting the IL-6-p47(phox)-oxidative stress pathway in neutrophils. Gut 2017, 66, 705–715.
- Bala, S.; Csak, T.; Saha, B.; Zatsiorsky, J.; Kodys, K.; Catalano, D.; Satishchandran, A.; Szabo, G. The pro-inflammatory effects of miR-155 promote liver fibrosis and alcohol-induced steatohepatitis. J. Hepatol. 2016, 64, 1378–1387.
- Saikia, P.; Bellos, D.; McMullen, M.R.; Pollard, K.A.; de la Motte, C.; Nagy, L.E. MicroRNA 181b-3p and its target importin alpha5 regulate toll-like receptor 4 signaling in Kupffer cells and liver injury in mice in response to ethanol. Hepatology 2017, 66, 602–615.
- Yin, H.; Hu, M.; Zhang, R.; Shen, Z.; Flatow, L.; You, M. MicroRNA-217 promotes ethanol-induced fat accumulation in hepatocytes by down-regulating SIRT1. J. Biol. Chem. 2012, 287, 9817–9826.
- Lai, C.Y.; Yeh, K.Y.; Lin, C.Y.; Hsieh, Y.W.; Lai, H.H.; Chen, J.R.; Hsu, C.C.; Her, G.M. MicroRNA-21 plays multiple oncometabolic roles in the process of NAFLD-related hepatocellular carcinoma via PI3K/AKT, TGF-beta, and STAT3 signaling. Cancers 2021, 13, 940.
- Yang, Y.L.; Kuo, H.C.; Wang, F.S.; Huang, Y.H. MicroRNA-29a disrupts DNMT3b to ameliorate diet-induced non-alcoholic steatohepatitis in mice. Int. J. Mol. Sci. 2019, 20, 1499.
- Chai, C.; Cox, B.; Yaish, D.; Gross, D.; Rosenberg, N.; Amblard, F.; Shemuelian, Z.; Gefen, M.; Korach, A.; Tirosh, O.; et al. Agonist of RORA attenuates nonalcoholic fatty liver progression in mice via up-regulation of MicroRNA 122. Gastroenterology 2020, 159, 999–1014.e9.
- Zhang, Z.; Wen, H.; Peng, B.; Weng, J.; Zeng, F. Downregulated microRNA-129-5p by long non-coding RNA NEAT1 upregulates PEG3 expression to aggravate non-alcoholic steatohepatitis. Front. Genet. 2020, 11, 563265.
- Riaz, F.; Chen, Q.; Lu, K.; Osoro, E.K.; Wu, L.; Feng, L.; Zhao, R.; Yang, L.; Zhou, Y.; He, Y.; et al. Inhibition of miR-188-5p alleviates hepatic fibrosis by significantly reducing the activation and proliferation of HSCs through PTEN/PI3K/AKT pathway. J. Cell. Mol. Med. 2021, 25, 4073–4087.
- He, Y.; Rodrigues, R.M.; Wang, X.; Seo, W.; Ma, J.; Hwang, S.; Fu, Y.; Trojnar, E.; Matyas, C.; Zhao, S.; et al. Neutrophil-to-hepatocyte communication via LDLR-dependent miR-223-enriched extracellular vesicle transfer ameliorates nonalcoholic steatohepatitis. J. Clin. Investig. 2021, 131.
- Benito-Vicente, A.; Uribe, K.B.; Rotllan, N.; Ramirez, C.M.; Jebari-Benslaiman, S.; Goedeke, L.; Canfran-Duque, A.; Galicia-Garcia, U.; Saenz De Urturi, D.; Aspichueta, P.; et al. MiR-27b modulates insulin signaling in hepatocytes by regulating insulin receptor expression. Int. J. Mol. Sci. 2020, 21, 8675.
- Ke, Q.H.; Chen, H.Y.; He, Z.L.; Lv, Z.; Xu, X.F.; Qian, Y.G.; Zheng, S.S. Silencing of microRNA-375 affects immune function in mice with liver failure by upregulating astrocyte elevated gene-1 through reducing apoptosis of Kupffer cells. J. Cell Biochem. 2019, 120, 253–263.
- Wang, Y.Q.; Lan, Y.Y.; Guo, Y.C.; Yuan, Q.W.; Liu, P. Down-regulation of microRNA-138 improves immunologic function via negatively targeting p53 by regulating liver macrophage in mice with acute liver failure. Biosci. Rep. 2019, 39.
- Lu, Z.; Liu, J.; Liu, X.; Huang, E.; Yang, J.; Qian, J.; Zhang, D.; Liu, R.; Chu, Y. MicroRNA 15a/16-1 suppresses aryl hydrocarbon receptor-dependent interleukin-22 secretion in CD4(+) T cells and contributes to immune-mediated organ injury. Hepatology 2018, 67, 1027–1040.
- Su, K.; Wang, Q.; Qi, L.; Hua, D.; Tao, J.; Mangan, C.J.; Lou, Y.; Li, L. MicroRNA-674-5p/5-LO axis involved in autoimmune reaction of Concanavalin A-induced acute mouse liver injury. Toxicol. Lett. 2016, 258, 101–107.
- Tu, H.; Chen, D.; Cai, C.; Du, Q.; Lin, H.; Pan, T.; Sheng, L.; Xu, Y.; Teng, T.; Tu, J.; et al. MicroRNA-143-3p attenuated development of hepatic fibrosis in autoimmune hepatitis through regulation of TAK1 phosphorylation. J. Cell. Mol. Med. 2020, 24, 1256–1267.
- Yang, F.; Lou, G.; Zhou, X.; Zheng, M.; He, J.; Chen, Z. MicroRNA-223 acts as an important regulator to Kupffer cells activation at the early stage of Con A-induced acute liver failure via AIM2 signaling pathway. Cell. Physiol. Biochem. 2014, 34, 2137–2152.
- Marzioni, M.; Agostinelli, L.; Candelaresi, C.; Saccomanno, S.; De Minicis, S.; Maroni, L.; Mingarelli, E.; Rychlicki, C.; Trozzi, L.; Banales, J.M.; et al. Activation of the developmental pathway neurogenin-3/microRNA-7a regulates cholangiocyte proliferation in response to injury. Hepatology 2014, 60, 1324–1335.
- Fernandez-Ramos, D.; Fernandez-Tussy, P.; Lopitz-Otsoa, F.; Gutierrez-de-Juan, V.; Navasa, N.; Barbier-Torres, L.; Zubiete-Franco, I.; Simon, J.; Fernandez, A.F.; Arbelaiz, A.; et al. MiR-873-5p acts as an epigenetic regulator in early stages of liver fibrosis and cirrhosis. Cell Death Dis. 2018, 9, 958.
- Hall, C.; Ehrlich, L.; Meng, F.; Invernizzi, P.; Bernuzzi, F.; Lairmore, T.C.; Alpini, G.; Glaser, S. Inhibition of microRNA-24 increases liver fibrosis by enhanced menin expression in Mdr2(-/-) mice. J. Surg. Res. 2017, 217, 160–169.
- Chen, L.; Chen, R.; Kemper, S.; Cong, M.; You, H.; Brigstock, D.R. Therapeutic effects of serum extracellular vesicles in liver fibrosis. J. Extracell. Vesicles 2018, 7, 1461505.
- Guo, C.J.; Pan, Q.; Xiong, H.; Qiao, Y.Q.; Bian, Z.L.; Zhong, W.; Sheng, L.; Li, H.; Shen, L.; Hua, J.; et al. Therapeutic potential of microRNA: A new target to treat intrahepatic portal hypertension? Biomed. Res. Int. 2014, 2014, 797898.
- Breitkopf, K.; Godoy, P.; Ciuclan, L.; Singer, M.V.; Dooley, S. TGF-beta/Smad signaling in the injured liver. Z. Gastroenterol. 2006, 44, 57–66.
- Liang, J.; Deng, X.; Lin, Z.X.; Zhao, L.C.; Zhang, X.L. Attenuation of portal hypertension by natural taurine in rats with liver cirrhosis. World J. Gastroenterol. 2009, 15, 4529–4537.
- Thabut, D.; Shah, V. Intrahepatic angiogenesis and sinusoidal remodeling in chronic liver disease: New targets for the treatment of portal hypertension? J. Hepatol. 2010, 53, 976–980.
- Chawla, Y.K.; Bodh, V. Portal vein thrombosis. J. Clin. Exp. Hepatol. 2015, 5, 22–40.
- Vemuganti, R.; Silva, V.R.; Mehta, S.L.; Hazell, A.S. Acute liver failure-induced hepatic encephalopathy s associated with changes in microRNA expression rofiles in cerebral cortex of the mouse [corrected]. Metab. Brain Dis. 2014, 29, 891–899.
- Baker, L.; Lanz, B.; Andreola, F.; Ampuero, J.; Wijeyesekera, A.; Holmes, E.; Deutz, N. New technologies—New insights into the pathogenesis of hepatic encephalopathy. Metab. Brain Dis. 2016, 31, 1259–1267.
- Oenarto, J.; Karababa, A.; Castoldi, M.; Bidmon, H.J.; Gorg, B.; Haussinger, D. Ammonia-induced miRNA expression changes in cultured rat astrocytes. Sci. Rep. 2016, 6, 18493.
- de Boer, Y.S.; van Gerven, N.M.; Zwiers, A.; Verwer, B.J.; van Hoek, B.; van Erpecum, K.J.; Beuers, U.; van Buuren, H.R.; Drenth, J.P.; den Ouden, J.W.; et al. Genome-wide association study identifies variants associated with autoimmune hepatitis type 1. Gastroenterology 2014, 147, 443–452.e5.
- Ueno, K.; Aiba, Y.; Hitomi, Y.; Shimoda, S.; Nakamura, H.; Gervais, O.; Kawai, Y.; Kawashima, M.; Nishida, N.; Kohn, S.S.; et al. Integrated GWAS and mRNA microarray analysis identified IFNG and CD40L as the central upstream regulators in primary biliary cholangitis. Hepatol. Commun. 2020, 4, 724–738.





