Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Yves Briers | + 884 word(s) | 884 | 2021-07-01 08:05:37 | | | |
| 2 | Yves Briers | + 674 word(s) | 1558 | 2021-07-23 12:32:43 | | | | |
| 3 | Rita Xu | Meta information modification | 1558 | 2021-07-26 03:34:40 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Briers, Y. PhaLP. Encyclopedia. Available online: https://encyclopedia.pub/entry/12361 (accessed on 15 April 2026).
Briers Y. PhaLP. Encyclopedia. Available at: https://encyclopedia.pub/entry/12361. Accessed April 15, 2026.
Briers, Yves . "PhaLP" Encyclopedia, https://encyclopedia.pub/entry/12361 (accessed April 15, 2026).
Briers, Y. (2021, July 23). PhaLP. In Encyclopedia. https://encyclopedia.pub/entry/12361
Briers, Yves . "PhaLP." Encyclopedia. Web. 23 July, 2021.
Copy Citation
Phage lytic proteins are a clinically advanced class of novel enzyme-based antibiotics, so-called enzybiotics. PhaLP is a database of phage lytic proteins, which serves as an open portal to facilitate the development of phage lytic proteins. PhaLP is a comprehensive, easily accessible and automatically updated database.
phage lytic proteins
endolysins
machine learning
biological database
conserved protein domains
1. Introduction
Bacteriophages are infective viral particles targeting bacterial cells. During their lytic replication cycle, phages face twice the challenge to cross the bacterial cell wall of their host [1]. Therefore, phages make use of two types of phage lytic proteins: virion-associated lysins (VALs) and endolysins [2]. A VAL is part of the virion and forms a local and small pore in the peptidoglycan layers at the site of infection, while the cell remains intact. The phage genome is ejected through this pore. VALs are mostly a structural part of the virion but can also occur as internal capsid proteins [3][4]. Endolysins are responsible for the massive degradation of the peptidoglycan layer at the end of the lytic cycle. In the canonical phage lysis system, they accumulate in large numbers in the cytoplasm and their release into the periplasmic space is timed by pore-forming holins in the cytoplasmic membrane. The sudden degradation of the peptidoglycan, assisted by the high osmotic pressure inside the cell, causes cell lysis and the concomitant release of newly matured phage particles [5][6].
Phage lytic proteins comprise one or more functional domains categorized into two classes: enzymatically active domains (EADs) and cell wall binding domains (CBDs) [7]. Additionally, VALs contain domains with a function other than peptidoglycan degradation such as structural anchoring to the viral particle. At the biochemical level all phage lytic proteins have the same purpose, i.e., peptidoglycan degradation, yet variation in environment and host has led to highly diverse domains and architectures [1][3][4][8]. This variety largely springs from the diversity of peptidoglycan chemotypes among bacterial species, along with the high diversity of secondary cell wall-associated carbohydrate polymers [9][10]. Endolysins have either a globular (one EAD) or modular architecture (multiple domains; at least one EAD and optionally one or more CBDs) [11]. CBDs target the glycan or peptide moieties of peptidoglycan, or specific components of (lipo)teichoic acids, increasing the proximity of EAD to its substrate [12][13]. Five classes of EADs are distinguished based on the bond they cleave in peptidoglycan (Figure 1). (i) N-acetylmuramoyl-L-alanine amidases (EC 3.5.1.28) hydrolyze the amide bond between N-acetylmuramic acid (MurNAc) and L-alanine residues, effectively cleaving the peptide moiety from the glycan strand. (ii) N-acetyl-β-D-glucosaminidases catalyze the hydrolysis of glycosidic β-1,4 linkages between N-acetylglucosamine (GlcNac) and MurNAc. (iii) N-acetyl-β-D-muramidases (EC 3.2.1.17) and (iv) lytic transglycosylases (EC 4.2.2.n2) cleave these linkages between MurNAc and GlcNAc. N-acetyl-β-D-glucosaminidases and N-acetyl-β-D-muramidases are glycosidases (EC 3.2.1.-) that use a hydrolytic mechanism resulting in a terminal reducing GlcNAc or MurNAc residue, respectively. Lytic transglycosylases, on the other hand, use an intramolecular mechanism that creates a 1,6-anhydro bond at the MurNAc residue. Peptidases (EC 3.4.-.-) cleave the bond between two amino acids within the peptidoglycan stem peptide, cross-link or cross-bridge [14]. The specific epitope and chemical bond targeted by the CBD and the EAD, respectively, brings about a well-defined spectrum of activity for phage lytic proteins [15][16]. The modularity of phage lytic proteins, along with the diverse range of domains, has been exploited by protein engineers to modulate the specificity, activity and solubility by domain swapping [11][17].
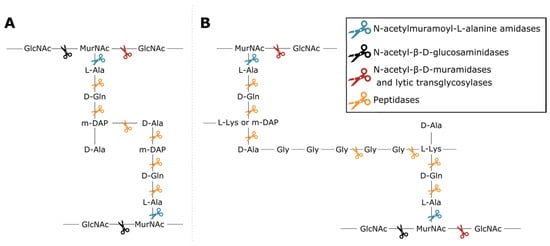
Figure 1. Cleavage sites of the different enzymatic classes of EADs on the primary structures of two common peptidoglycan types. (A) peptidoglycan chemotype A1γ is the most common in Gram-negative bacteria; (B) peptidoglycan chemotype A3, either α or γ depending on the presence of L-Lys or mDAP in the third position of the peptide subunit, is a type example for Gram-positive bacteria such as Staphylococcus aureus [10]. The colored scissors indicate cleavage sites of different classes of EADs. GlcNAc and MurNAc in the glycan strands of each structure refer to N-acetylglucosamine and N-acetylmuramic acid, respectively.
As early as 1957, it was observed that phage lytic proteins can cause “lysis from without” upon exogenous addition to bacteria [18][19]. It was not until 2001 that their use as enzyme-based antibacterial agents, coined “enzybiotics”, was demonstrated in a murine model against Gram-positive bacteria [20]. The presence of an outer membrane was initially prohibitive for the use of phage lytic proteins as enzybiotics against Gram-negative bacteria, but meanwhile various protein engineering approaches have been developed to overcome this barrier, including the use of outer membrane permeabilizing peptides (Artilysins®) [11], bacteriocin domains (lysocins) [21] and phage receptor binding proteins (Innolysins) [22]. Today, enzybiotics are considered the most advanced alternative class of antibacterials under clinical investigation [23]. They offer a necessary response to the alarming threat of antibiotic resistance across global health care systems. Their mode of action is fundamentally different from any existing class of antibiotics. Enzybiotics actively degrade the peptidoglycan component without the need for an active bacterial metabolism, unlike classic antibiotics. This is reflected in a faster cell death and effectiveness against metabolically inactive cells like persisters. In addition, their spectrum is typically narrower (genus, species or strain level) compared to classic antibiotics, causing less harm to beneficial microflora. Finally, the conserved molecular target makes enzybiotics less prone to the inevitable fate of many traditional antibiotics: the emergence and spread of resistance mechanisms [11][24]. A growing community of researchers and companies is therefore investigating their applications, including clinical trials, and engineering their properties to kill a broad diversity of bacteria [8]. Phage lytic proteins have the inherent potential to be developed against any bacterial species, thus representing an unprecedented extensive class of antibiotics with narrow spectrum.
2. PhaLP Database
The MySQL-based PhaLP database (https://www.phalp.org) integrates nine data types (proteins, phages, hosts, conserved domains, coding sequences (CDSs), GO annotations, enzymatic activities (ECs), tertiary structures, experimental evidence) originating from multiple sources databases (UniProt, UniParc [25], NCBI taxonomy, Virus-Host DB [26], InterPro [27], GenBank, QuickGO, ExPASy ENZYME database, PDB and PubMed). Figure 2 provides an overview of these data types and their mutual relationships.
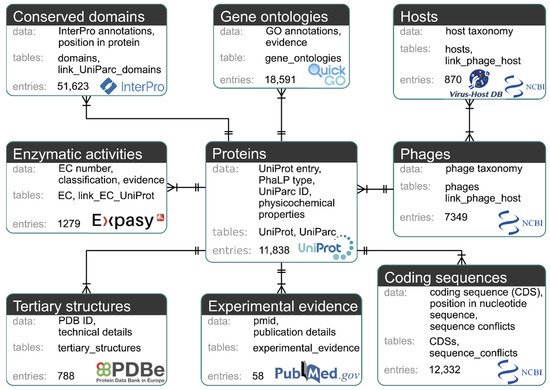
Figure 2. Diagram of the nine data types of PhaLP. Each data type is represented by a box containing a description of the data, the corresponding MySQL table(s), the number of entries in PhaLP v2019_10 and the source database. Relationships between data types are indicated with a crow’s foot notation. A relationship is indicated by a line between two data types with a double perpendicular line at a “one” side and a crow’s foot at a “many” side. The “one-to-many” relationship between for example “phages” and “proteins” can be interpreted as: one phage entry can be linked to multiple protein entries, but each protein entry can only be associated with one phage entry. The “many-to-many” relationship between “hosts” and “phages” can be interpreted as: a phage can have multiple hosts, but a host can also be associated with multiple phages.
The protein data form the central hub of PhaLP and describe single phage lytic proteins, corresponding to UniProt entries. UniProt was chosen as primary data source because it provides high-quality, curated and functionally annotated sequence data [25]. To collect a set of phage lytic proteins, UniProt is programmatically queried. The query is carefully constructed to include as much phage lytic proteins as possible without including other proteins. The resulting dataset was manually curated.
Due to increasing sequencing efforts, the amount of available sequence data in biological databases is increasing at an exponential rate [25]. A common problem with many so-called secondary databases is that, while their data sources (primary databases) keep expanding, the authors stop updating the database, resulting in outdated secondary databases. Therefore, the algorithm to gather data from primary databases for PhaLP is automated in a Python script. With every eight-weekly UniProt release, the algorithm is rebooted, adding new entries and updating data that have changed in the source databases. Curation of the new entries remains essential for continuous fine-tuning of the initial query. The latter task can be facilitated by users through the online contact form by reporting new, non-curated entries that are suspected of not being actual phage lytic proteins.
3. Use of PhaLP
PhaLP can be consulted through two user-friendly web interfaces. The first is a basic searchable and sortable table that displays basic info on each protein entry and the phage encoding the protein (https://www.phalp.org/database; accessed on 25 June 2021). Upon clicking on the UniProt accession number, the user is sent to an overview page with all data linked to that entry. Additionally, the page contains links to the original data sources, as well as two interactive graphical viewers: a representation of the conserved domains on the sequence and a representation of the genomic neighborhood for every CDS linked to the protein.
The second interface is a BioMart that allows the user to customize the selection of attributes that is shown from all tables in the database as well as filter on all these attributes (https://www.phalp.org/biomart; accessed on 25 June 2021). The resulting customized dataset is provided in a tabular format that can be viewed in the interface or downloaded as a “tab-separated values” file. The latter can be loaded into any software of choice to perform further analyses. To allow even more advanced querying, the database is available for download as a MySQL dump file, making it possible to integrate the PhaLP database in customized workflows. Both user interfaces will display the latest version of PhaLP. Older versions will remain available for download as a MySQL dump file.
References
- Fernandes, S.; São-José, C. Enzymes and Mechanisms Employed by Tailed Bacteriophages to Breach the Bacterial Cell Barriers. Viruses 2018, 10, 396.
- Gutiérrez, D.; Fernández, L.; Rodríguez, A.; García, P. Are phage lytic proteins the secret weapon to kill Staphylococcus aureus? MBio 2018, 9, e01923-17.
- Oliveira, H.; São-José, C.; Azeredo, J. Phage-derived peptidoglycan degrading enzymes: Challenges and future prospects for in vivo therapy. Viruses 2018, 10, 292.
- Latka, A.; Maciejewska, B.; Majkowska-Skrobek, G.; Briers, Y.; Drulis-Kawa, Z. Bacteriophage-encoded virion-associated enzymes to overcome the carbohydrate barriers during the infection process. Appl. Microbiol. Biotechnol. 2017, 101, 3103–3119.
- Cahill, J.; Young, R. Phage Lysis: Multiple Genes for Multiple Barriers. Adv. Virus Res. 2019, 103, 33–70.
- Catalão, M.J.; Gil, F.; Moniz-Pereira, J.; São-José, C.; Pimentel, M. Diversity in bacterial lysis systems: Bacteriophages show the way. FEMS Microbiol. Rev. 2013, 37, 554–571.
- Loessner, M.J.; Kramer, K.; Ebel, F.; Scherer, S. C-terminal domains of Listeria monocytogenes bacteriophage murein hydrolases determine specific recognition and high-affinity binding to bacterial cell wall carbohydrates. Mol. Microbiol. 2002, 44, 335–349.
- São-José, C. Engineering of phage-derived lytic enzymes: Improving their potential as antimicrobials. Antibiotics 2018, 7, 29.
- Briers, Y.; Volckaert, G.; Cornelissen, A.; Lagaert, S.; Michiels, C.W.; Hertveldt, K.; Lavigne, R. Muralytic activity and modular structure of the endolysins of Pseudomonas aeruginosa bacteriophages φKZ and EL. Mol. Microbiol. 2007, 65, 1334–1344.
- Schleifer, K.H.; Kandler, O. Peptidoglycan types of bacterial cell walls and their taxonomic implications. Bacteriol. Rev. 1972, 36, 407–477.
- Gerstmans, H.; Criel, B.; Briers, Y. Synthetic biology of modular endolysins. Biotechnol. Adv. 2018, 36, 624–640.
- Loessner, M.J. Bacteriophage endolysins—Current state of research and applications. Curr. Opin. Microbiol. 2005, 8, 480–487.
- Bustamante, N.; Campillo, N.E.; García, E.; Gallego, C.; Pera, B.; Diakun, G.P.; Sáiz, J.L.; García, P.; Díaz, J.F.; Menéndez, M. Cpl-7, a lysozyme encoded by a pneumococcal bacteriophage with a novel cell wall-binding motif. J. Biol. Chem. 2010, 285, 33184–33196.
- Vermassen, A.; Leroy, S.; Talon, R.; Provot, C.; Popowska, M.; Desvaux, M. Cell wall hydrolases in bacteria: Insight on the diversity of cell wall amidases, glycosidases and peptidases toward peptidoglycan. Front. Microbiol. 2019, 10, 1–27.
- Eugster, M.R.; Haug, M.C.; Huwiler, S.G.; Loessner, M.J. The cell wall binding domain of Listeria bacteriophage endolysin PlyP35 recognizes terminal GlcNAc residues in cell wall teichoic acid. Mol. Microbiol. 2011, 81, 1419–1432.
- Oliveira, H.; Melo, L.D.R.; Santos, S.B.; Nobrega, F.L.; Ferreira, E.C.; Cerca, N.; Azeredo, J.; Kluskens, L.D. Molecular aspects and comparative genomics of bacteriophage endolysins. J. Virol. 2013, 87, 4558–4570.
- Yang, H.; Yu, J.; Wei, H. Engineered bacteriophage lysins as novel anti-infectives. Front. Microbiol. 2014, 5.
- Maxted, W.R. The active agent in nascent phage lysis of streptococci. J. Gen. Microbiol. 1957, 16, 584–595.
- Krause, R.M. Studies on bacteriophages of hemolytic streptococci. I. Factors influencing the interaction of phage and susceptible host cell. J. Exp. Med. 1957, 106, 365–384.
- Nelson, D.; Loomis, L.; Fischetti, V.A. Prevention and elimination of upper respiratory colonization of mice by group A streptococci by using a bacteriophage lytic enzyme. Proc. Natl. Acad. Sci. USA 2001, 98, 4107–4112.
- Heselpoth, R.D.; Euler, C.W.; Schuch, R.; Fischetti, V.A. Lysocins: Bioengineered antimicrobials that deliver lysins across the outer membrane of Gram-negative bacteria. Antimicrob. Agents Chemother. 2019, 63, 453–457.
- Zampara, A.; Sørensen, M.C.H.; Grimon, D.; Antenucci, F.; Vitt, A.R.; Bortolaia, V.; Briers, Y.; Brøndsted, L. Exploiting phage receptor binding proteins to enable endolysins to kill Gram-negative bacteria. Sci. Rep. 2020, 10.
- Czaplewski, L.; Bax, R.; Clokie, M.; Dawson, M.; Fairhead, H.; Fischetti, V.A.; Foster, S.; Gilmore, B.F.; Hancock, R.E.W.; Harper, D.; et al. Alternatives to antibiotics—A pipeline portfolio review. Lancet Infect. Dis. 2016, 16, 239–251.
- Grishin, A.V.; Karyagina, A.S.; Vasina, D.V.; Vasina, I.V.; Gushchin, V.A.; Lunin, V.G. Resistance to peptidoglycan-degrading enzymes. Crit. Rev. Microbiol. 2020, 46, 703–726.
- The UniProt Consortium UniProt: A worldwide hub of protein knowledge. Nucleic Acids Res. 2019, 47, D506–D515.
- Mihara, T.; Nishimura, Y.; Shimizu, Y.; Nishiyama, H.; Yoshikawa, G.; Uehara, H.; Hingamp, P.; Goto, S.; Ogata, H. Linking virus genomes with host taxonomy. Viruses 2016, 8, 66.
- Blum, M.; Chang, H.Y.; Chuguransky, S.; Grego, T.; Kandasaamy, S.; Mitchell, A.; Nuka, G.; Paysan-Lafosse, T.; Qureshi, M.; Raj, S.; et al. The InterPro protein families and domains database: 20 years on. Nucleic Acids Res. 2021, 49, D344–D354.
More
Information
Subjects:
Virology
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.6K
Revisions:
3 times
(View History)
Update Date:
26 Jul 2021
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





