
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Angela Scala | + 1298 word(s) | 1298 | 2020-06-10 10:05:17 | | | |
| 2 | Catherine Yang | Meta information modification | 1298 | 2020-06-12 05:51:25 | | | | |
| 3 | Catherine Yang | -16 word(s) | 1282 | 2020-10-27 10:54:17 | | |
Video Upload Options
The occurrence of new pandemic viruses, such as SARS-CoV-1 (2003), MERS-CoV (2012) and SARS-CoV-2 (2019) has indicated an urgent need for diagnostic tools able to reliably identify infected individuals and to determine if the infection is in the acute phase. Although nanotechnology based on graphene has been poorly applied for the rapid diagnosis of viral diseases, the extraordinary properties of graphene have been recently exploited for the diagnosis of COVID-19. Novel graphene-based field-effect transistor (GFET) biosensors were developed for the quantitative detection of viral RNA and viral spike protein. The fabrication of COVID-19 FET sensor for spike protein recognition is based on the integration of the SARS-CoV-2 spike antibody with graphene, whereas the GFET sensor for viral RNA recognition exploited the CRISPR/Cas biotechnology.
1. Introduction
Direct methods, exploiting graphene nanotechnology, for the rapid virus detection, have been only marginally investigated in the past, and no critical discussion has been reported in successive literature reviews [1][2].
This attitude was unchanging even during SARS–CoV-1 emergency that was responsible for the 2003 severe acute respiratory syndrome (SARS) infection in Asia, causing about 8000 cases and 774 deaths, also during the Middle East respiratory distress syndrome (MERS) of 2013, which affected Saudi Arabia causing close to 858 deaths [3][4][5][6]. Advances in nanotechnology have begun to play an important role in viral detection, to improve the detection limit, operational simplicity of viral diagnostics [7].
A coplanar-gate graphene field-effect transistors (GFETs) [8] have been proposed for the detection of HIV-1 (human immunodeficiency virus 1) and MLV (murine leukemia virus) viruses using antibodies of vesicular stomatitis Indiana virus (VSV) as biorecognition element. VSV antibodies are immobilized on the G layer using 1-pyrenebutanoic acid succinimidyl ester (PASE). PASE binds G by π-π interactions, anchoring the antibody’s primary amine groups by the opposite succinimidyl group. The formation of the virus-antibody complex leads to a downward shift of the Dirac point voltage, regardless of the types of detected viruses. The proposed platform has worked in a wide range of concentrations (from 47.8 aM to 10.55 nM), but the lack of virus specificity appears the main limitation of this strategy.
An surface plasmon resonance (SPR) sensor based on an polyamidoamine-functionalized rGO(composite, with monoclonal antibodies immobilized on self-assembled dithiobis (succinimidyl undecanoate, DSU) for the detection/quantification of Dengue virus (DENV), has been recently described [2].
The specificity and the sensibility of the sensor have been achieved by anchoring a stable biorecognition element (antibodies (IgM) against Dengue type 2 envelope proteins) on the gold surface of the sensor. The specific binding of antibody-DENV 2 E-protein allows a significant change in the angle of the reflectivity minimum that is correlated to Dengue virus detection. The proposed sensor has shown a sensitive and selective response towards DENV 2 E-proteins compared to DENV 1 E-proteins and ZIKV (Zika virus) E-proteins. Although no G materials have been integrated into the above-described sensor [2], the criteria used for its fabrication were included in this review since the strategy could be extended to other viruses, and the performance of SPR noble metal could be improved in the presence of G [9].
2. Graphene-Based Strategies in the Diagnosis of Viral Diseases
Differently from the past, the current sanitary pandemic emergency caused by the new type of coronavirus (SARS–CoV-2) is characterized by global effort to identify biomarkers that predict the severity of COVID-19 patients and to develop diagnostic tools for the rapid detection of SARS–CoV-2 infection [10].
Currently, nucleic acid testing on respiratory specimens is the reference gold standard method for the diagnosis of COVID-19 infected patients [11]. The test requires a series of laboratory procedures: (a) viral RNA extraction; (b) addition to a master mix containing nuclease-free water, reverse primers, a fluorophore-quencher probe, and a reaction mix (i.e., polymerase, reverse transcriptase, magnesium, nucleotides, and additives); (c) loading of extracted RNA/master mix into a PCR thermocycler; (d) several cycles at settled temperature. During the RT-PCR cycles, the cleavage of the fluorophore-quencher probe generates a fluorescent signal detected and recorded in real-time [10].
RT-PCR uses respiratory samples to genetically detect SARS–CoV-2; some data have suggested that 20–34% of COVID-19 patients resulted negative in the test despite being infected. This variance in the sensitivity could be mainly attributed to low viral load (i.e., patients tested in the early stage of the viral disease) [11]. Other RT-PCR issues include the time consuming and expensive analysis and the technical expertise in carrying out the text.
Other technologies, such as point-of-care technologies and serologic immunoassays, are rapidly emerging to address these deficiencies [7].
Analytic methods to assess prior infection and immunity to SARS–CoV-2 by antibody identification are essential for epidemiologic studies, although sensibility and specificity of the tests currently available in the market remain undefined. Cross-reactivity of antibody to non–SARS–CoV-2 coronavirus proteins is the main issue of these serologic tests [10][11]. The development of an antigen detection test [11] could take advantage of progress in the production of monoclonal antibodies against the nucleocapsid protein of SARS–CoV-2. The global effort to increase SARS–CoV-2 testing capacity takes advantage of the most recent advances in chemistry, molecular biology, genome technology, and nanotechnology. Several projects are ongoing in this direction, and some results are already reported in the literature [12][13].
The detection of SARS–CoV-2 in respiratory samples has been achieved by LSPR biosensor, combining the photothermal effect and plasmonic sensing transduction for SARS–CoV-2 viral nucleic acid [13].
A field-effect transistor (FET)-based biosensing device for detecting SARS–CoV-2 spike protein (S) in clinical samples was reported by Seo et al. [14]. Antibodies against S protein were anchored to the graphene sheet (external coating of FET) by 1-pyrenebutanoic acid succinimidyl ester (PBASE, Figure 1).
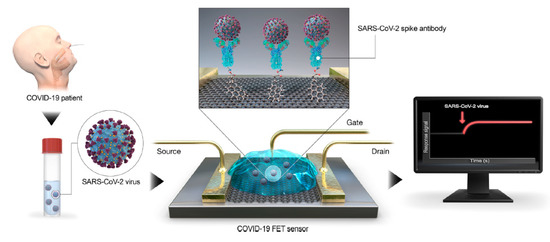
Figure 1. Coronavirus disease 2019 (COVID-19) field-effect transistor (FET)-sensor. Graphene is selected as sensing material and is decorated with the SARS–CoV-2 spike antibody using 1-pyrenebutanoic acid succinimidyl ester (PBASE) as interfacing molecule and probe linker. Reprinted with permission from reference [14], Copyright © 2020, American Chemical Society. https://pubs.acs.org/doi/abs/10.1021/acsnano.0c02823 Further permissions related to the material excerpted should be directed to the ACS.
The performance of the sensor is determined using antigen protein, cultured virus, and nasopharyngeal swab specimens from COVID-19 patients. The device could detect S protein at concentrations of 1 fg/mL in PBS and 100 fg/mL in the clinical transport medium, and it could distinguish the SARS–CoV-2 antigen protein from those of MERS-CoV. The successful fabrication of a COVID-19 FET sensor based on the integration of the SARS–CoV-2 spike antibody with graphene suggests the key role of G for diagnostic scope [15]. Specifically, the functionalization of G with diverse functional molecules [16][17][18][19]could be the key element to tailor its properties and to obtain advanced diagnostic tools for the SARS–CoV-2 diagnosis. Meanwhile, for the revision of this manuscript, some works dealing with sensors for COVID-19 diagnosis based on graphene are reported in the literature [20], and, although further researches are undoubtedly necessary, the leading role of G in the world’s fight against COVID-19 is clearly coming out [21].
In summary, the biomolecules till now used to target SARS–CoV-2 includes the viral RNA, the viral spike proteins, and the specific immunoglobulins produced by the host immune system. The biosensing community is actively working to improve portability, time, and cost of PCR-based SARS–CoV-2 detection, as well as to create manufacturable PCR-based microfluidic devices. Recently, also the gene-editing technology (CRISPR/Cas) has been developed to overcome the issues of PCR-based systems. Two different detection modes have been proposed in CRISPR technology, i.e., binding- or cleavage-based [22]. The sensor is developed by immobilization on a graphene-based field-effect transistor (GFET) of Cas9 with a sgRNA, specific to the target sequence of SARS COV-2; the electrical signal originated by the binding of the target nucleic acid by the Cas9–sgRNA complex is recorded via a simple handheld device without amplification.
References
- Jin Woo Kim; Seunghyun Kim; Yoon-Ha Jang; Kwang-Il Lim; Wi Hyoung Lee; Attomolar detection of virus by liquid coplanar-gate graphene transistor on plastic.. Nanotechnology 2019, 30, 345502, 10.1088/1361-6528/ab0f52.
- Nur Alia Sheh Omar; Yap Wing Fen; Jaafar Abdullah; Amir Sadrolhosseini; Yasmin Mustapha Kamil; Nurul Fauzi; Hazwani Hashim; M.A. Mahdi; Quantitative and Selective Surface Plasmon Resonance Response Based on a Reduced Graphene Oxide–Polyamidoamine Nanocomposite for Detection of Dengue Virus E-Proteins. Nanomaterials 2020, 10, 569, 10.3390/nano10030569.
- Jin-Ho Lee; Soo-Jeong Park; Jeong-Woo Choi; Electrical Property of Graphene and Its Application to Electrochemical Biosensing. Nanomaterials 2019, 9, 297, 10.3390/nano9020297.
- Elisabeth Mahase; Coronavirus covid-19 has killed more people than SARS and MERS combined, despite lower case fatality rate.. BMJ 2020, 368, m641.
- Davide Gentile; Vincenzo Patamia; Angela Scala; Maria Teresa Sciortino; Anna Piperno; Antonio Rescifina; Putative Inhibitors of SARS-CoV-2 Main Protease from A Library of Marine Natural Products: A Virtual Screening and Molecular Modeling Study. Marine Drugs 2020, 18, 225, 10.3390/md18040225.
- Catherine A. Hogan; Chelsea Caya; J. Papenburg; Rapid and simple molecular tests for the detection of respiratory syncytial virus: a review. Expert Review of Molecular Diagnostics 2018, 18, 617-629, 10.1080/14737159.2018.1487293.
- Lena Gorgannezhad; Muhammad Umer; Nazmul Islam; Anh V. Nguyen; Muhammad J. A. Shiddiky; Circulating tumor DNA and liquid biopsy: opportunities, challenges, and recent advances in detection technologies. Lab on a Chip 2018, 18, 1174-1196, 10.1039/c8lc00100f.
- Xiaoni Fang; Yaokai Duan; Gary Adkins; Songqin Pan; Hua Wang; Yang Liu; Wenwan Zhong; Highly Efficient Exosome Isolation and Protein Analysis by an Integrated Nanomaterial-Based Platform. Analytical Chemistry 2018, 90, 2787-2795, 10.1021/acs.analchem.7b04861.
- Elisa Lenzi; Dorleta Jimenez De Aberasturi; Luis M. Liz-Marzán; Surface-Enhanced Raman Scattering Tags for Three-Dimensional Bioimaging and Biomarker Detection. ACS Sensors 2019, 4, 1126-1137, 10.1021/acssensors.9b00321.
- Buddhisha Udugama; Pranav Kadhiresan; Hannah N. Kozlowski; Ayden Malekjahani; Matthew Osborne; Vanessa Y. C. Li; Hongmin Chen; Samira Mubareka; Jonathan B. Gubbay; Warren C. W. Chan; et al. Diagnosing COVID-19: The Disease and Tools for Detection. ACS Nano 2020, 14, 3822-3835, 10.1021/acsnano.0c02624.
- Matthew P. Cheng; Jesse Papenburg; Michaël Desjardins; Sanjat Kanjilal; Caroline Quach; Michael Libman; Sabine Dittrich; Cedric P. Yansouni; Diagnostic Testing for Severe Acute Respiratory Syndrome–Related Coronavirus-2. Annals of Internal Medicine 2020, 172, 726-734, 10.7326/m20-1301.
- Huayu Huang; Wanqiao Bai; Changxun Dong; Rui Guo; Zhihua Liu; An ultrasensitive electrochemical DNA biosensor based on graphene/Au nanorod/polythionine for human papillomavirus DNA detection. Biosensors and Bioelectronics 2015, 68, 442-446, 10.1016/j.bios.2015.01.039.
- Guangyu Qiu; Zhibo Gai; Yile Tao; Jean Schmitt; Gerd A. Kullak-Ublick; Jing Wang; Dual-Functional Plasmonic Photothermal Biosensors for Highly Accurate Severe Acute Respiratory Syndrome Coronavirus 2 Detection. ACS Nano 2020, 14, 5268-5277, 10.1021/acsnano.0c02439.
- Giwan Seo; GeonHee Lee; Mi Jeong Kim; Seung-Hwa Baek; Minsuk Choi; Keun Bon Ku; Chang-Seop Lee; Sangmi Jun; Daeui Park; Hong Gi Kim; et al.Seong-Jun KimJeong-O. LeeBum Tae KimEdmond Changkyun ParkSeung Il Kim Rapid Detection of COVID-19 Causative Virus (SARS-CoV-2) in Human Nasopharyngeal Swab Specimens Using Field-Effect Transistor-Based Biosensor. ACS Nano 2020, 14, 5135-5142, 10.1021/acsnano.0c02823.
- Noemi Bellassai; Giuseppe Spoto; Biosensors for liquid biopsy: circulating nucleic acids to diagnose and treat cancer. Analytical and Bioanalytical Chemistry 2016, 408, 7255-7264, 10.1007/s00216-016-9806-3.
- Anna Piperno; Angela Scala; Antonino Mazzaglia; Giulia Neri; Rosamaria Pennisi; Maria Teresa Sciortino; Giovanni Grassi; Cellular Signaling Pathways Activated by Functional Graphene Nanomaterials. International Journal of Molecular Sciences 2018, 19, 3365, 10.3390/ijms19113365.
- Giulia Neri; Angela Scala; Enza Fazio; Placido Giuseppe Mineo; Antonio Rescifina; Anna Piperno; Giovanni Grassi; Repurposing of oxazolone chemistry: gaining access to functionalized graphene nanosheets in a top-down approach from graphite. Chemical Science 2015, 6, 6961-6970, 10.1039/c5sc02576a.
- Davide Barreca; Giulia Neri; Angela Scala; Enza Fazio; Davide Gentile; Antonio Rescifina; Anna Piperno; Giullia Neri; Covalently immobilized catalase on functionalized graphene: effect on the activity, immobilization efficiency, and tetramer stability. Biomaterials Science 2018, 6, 3231-3240, 10.1039/c8bm00850g.
- Anna Piperno; Antonino Mazzaglia; Angela Scala; Rosamaria Pennisi; Roberto Zagami; Giulia Neri; Serena Maria Torcasio; Consolato Rosmini; Placido Giuseppe Mineo; Monica Potara; et al.Monica FocsanSimion AstileanGrace Guoying ZhouMaria Teresa Sciortino Casting Light on Intracellular Tracking of a New Functional Graphene-Based MicroRNA Delivery System by FLIM and Raman Imaging. ACS Applied Materials & Interfaces 2019, 11, 46101-46111, 10.1021/acsami.9b15826.
- Eden Morales-Narváez Can Dincer; The impact of biosensing in a pandemic outbreak: COVID-19. Biosensors and Bioelectronics 2020, 163, 112274, 10.1016/j.bios.2020.112274.
- Palmieri, V; Papi, M.; Can graphene take part in the fight against covid-19?. NanoToday 2020, 33, 100883, 10.1016/j.bios.2015.01.039.
- Richard Bruch; Gerald A. Urban; Can Dincer; Unamplified gene sensing via Cas9 on graphene.. Nature Biomedical Engineering 2019, 3, 419-420, 10.1038/s41551-019-0413-4.





