
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Nurbubu Moldogazieva | + 5212 word(s) | 5212 | 2021-05-14 07:52:32 | | | |
| 2 | Bruce Ren | -21 word(s) | 5191 | 2021-05-28 03:52:04 | | |
Video Upload Options
It has been long recognized that cancer cells reprogram their metabolism under hypoxia conditions due to a shift from oxidative phosphorylation (OXPHOS) to glycolysis in order to meet elevated requirements in energy and nutrients for proliferation, migration, and survival. However, data accumulated over recent years has increasingly provided evidence that cancer cells can revert from glycolysis to OXPHOS and maintain both reprogrammed and oxidative metabolism, even in the same tumor. This phenomenon, denoted as cancer cell metabolic plasticity or hybrid metabolism, depends on a tumor micro-environment that is highly heterogeneous and influenced by an intensity of vasculature and blood flow, oxygen concentration, and nutrient and energy supply, and requires regulatory interplay between multiple oncogenes, transcription factors, growth factors, and reactive oxygen species (ROS), among others. Hypoxia-inducible factor-1 (HIF-1) and AMP-activated protein kinase (AMPK) are key modulators of the switch between reprogrammed and oxidative metabolism. Our review focuses on cross-talks between HIF-1, glucose transporters (GLUTs), and AMPK, and other regulatory proteins including oncogenes such as c-Myc, p53, and KRAS along with growth factor-initiated protein kinase B (PKB)/Akt, phosphatidyl-3-kinase (PI3K), and mTOR signaling pathways in controlling cancer cell metabolism.
1. Introduction
2. Hypoxia-Inducible Factors
3. An interplay between HIF-1 and Facilitative Glucose Transporters
4. Role of HIF-1 in Metabolic Reprogramming of Cancer Cells
4.1. Enhancement of Glycolysis
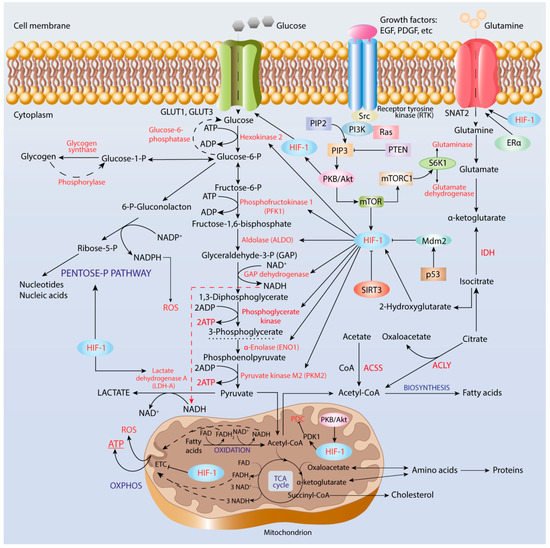
4.2. Pentose Phosphate Pathway
4.3. Cancer Acidification and Its Role in the Reverse to OXPHOS
4.4. Lipid Biosynthesis
References
- Smith, K.A.; Waypa, G.B.; Schumacker, P.T. Redox signaling during hypoxia in mammalian cells. Redox Biol. 2017, 13, 228–234.
- Ralph, S.J.; Rodriguez-Enriguez, S.; Neuzil, J.; Saavedra, E.; Moreno-Sanchez, R. The causes of cancer revisited: “Mitochondrial malignancy” and ROS-induced oncogenic transformation—Why mitochondria are targets for cancer therapy. Mol. Asp. Med. 2010, 31, 145–170.
- DeBerardinis, R.J.; Chandel, N.S. Fundamentals of cancer metabolism. Sci. Adv. 2016, 2, e1600200.
- Vaupel, P.; Multhoff, G. Hypoxia-/HIF-1α-driven factors of the tumor microenvironment impeding antitumor immune response and promoting malignant progression. Adv. Exp. Med. Biol. 2018, 1072, 171–175.
- Caino, M.C.; Altieri, D.C. Molecular pathways: Mitochondrial reprogramming in tumor progression and therapy. Clin. Cancer Res. 2016, 22, 540–545.
- Francis, A.; Venkatesh, G.H.; Zaarour, R.F.; Zeinelabdin, N.A.; Nawafleh, H.H.; Prasad, P.; Buart, S.; Terry, S.; Chouaib, S. Tumor hypoxia: A key determinant of microenvironment hostility and major checkpoint during the antitumor response. Crit. Rev. Immunol. 2018, 38, 505–524.
- Gu, Q.; He, Y.; Ji, J.; Yao, Y.; Shen, W.; Luo, J.; Zhu, W.; Cao, H.; Geng, Y.; Zhang, S.; et al. Hypoxia-inducible factor-1α (HIF-1α) and reactive oxygen species (ROS) mediates radiation-induced invasiveness through the SDF-1α/CXCR4 pathway in non-small cell lung carcinoma cells. Oncotarget 2015, 6, 10893–10907.
- Choudhry, H.; Harria, A. Advances in hypoxia-inducible factor biology. Cell Metabol. 2018, 27, 281–298.
- Warburg, O.; Wind, F.; Negelein, E. The metabolism of tumors in the body. J. Gen. Physiol. 1927, 8, 519–530.
- Warburg, O. On respiratory impairment in cancer cells. Science 1956, 124, 269–270.
- Racker, E. Bioenergetics and the problem of tumor growth: An understanding of the mechanism of the generation and control of biological energy may shed light on the problem of tumor growth. Am. Sci. 1972, 60, 56–63.
- Guzy, R.D.; Hoyos, B.; Robin, E.; Chen, H.; Liu, L.; Mansfield, K.D.; Simon, M.C.; Hammerling, U.; Schumacker, P.T. Mitochondrial complex III is required for hypoxia-induced ROS production and cellular oxygen sensing. Cell Metab. 2005, 1, 401–408.
- Yuan, G.; Nanduri, J.; Khan, S.; Semenza, G.L.; Prabhakar, N.R. Induction of HIF-1alpha expression by intermittent hypoxia: Involvement of NADPH oxidase, Ca2þ signaling, prolyl hydroxylases, and mTOR. J. Cell Physiol. 2008, 217, 674–685.
- Mihaylova, M.M.; Shaw, R.J. Metabolic reprogramming by class I and II histone deacetylases. Trends Endocrinol. Metab. 2013, 24, 48–57.
- Marusyk, A.; Polyak, K. Tumor heterogeneity: Causes and consequences. Biochim. Biophys. Acta 2010, 1805, 105–117.
- van der Heijden, M.; Miedema, D.M.; Waclaw, B.; Veenstra, V.L.; Lecca, M.C.; Nijman, L.E.; van Dijk, E.; van Neerven, S.M.; Lodestijin, S.C.; Lenos, K.J.; et al. Spatiotemporal regulation of clonogenicity in colorectal cancer xenografts. Proc. Natl. Acad. Sci. USA 2019, 116, 6140–6145.
- Campbell, P.J.; Pleasance, E.D.; Stephens, P.J.; Dicks, E.; Rance, R.; Goodhead, I.; Follows, G.A.; Green, A.R.; Futreal, P.A.; Stratton, M.R. Subclonal phylogenetic structures in cancer revealed by ultra-deep sequencing. Proc. Natl. Acad. Sci. USA 2008, 105, 13081–13086.
- Russo, M.; Siravegna, G.; Blaszkowsky, L.S.; Corti, G.; Crisafulli, G.; Ahronian, L.G.; Mussolin, B.; Kwak, E.L.; Buscarino, M.; Lazzari, L.; et al. Tumor heterogeneity and lesion-specific response to targeted therapy in colorectal cancer. Cancer Discov. 2016, 6, 147–153.
- Scott, D.A.; Richardson, A.D.; Filipp, F.V.; Knutzen, C.A.; Chiang, G.G.; Ronai, Z.A.; Osterman, A.L.; Smith, J.W. Comparative metabolic flux profiling of melanoma cell lines: Beyond the Warburg effect. J. Biol. Chem. 2011, 286, 42626–42634.
- Gentric, G.; Mieulet, V.; Mechta-Grigorou, F. Heterogeneity in cancer metabolism: New concepts in an old field. Antioxid Redox Signal 2017, 26, 462–485.
- Martinez-Outschoorn, U.; Sotgia, F.; Lisanti, M.P. Tumor microenvironment and metabolic synergy in breast cancers: Critical importance of mitochondrial fuels and function. Semin. Oncol. 2014, 41, 195–216.
- Mihaylova, M.M.; Shaw, R.J. The AMPK signaling pathway coordinates cell growth, autophagy and metabolism. Nat. Cell Biol. 2011, 13, 1016–1023.
- Jia, D.; Lu, M.; Jung, K.H.; Park, J.H.; Yu, L.; Onuchic, J.N.; Kaipparettu, B.A.; Levine, H. Elucidating cancer metabolic plasticity by coupling gene regulation with metabolic pathways. Proc. Natl. Acad. Sci. USA 2019, 116, 3909–3918.
- Moldogazieva, N.T.; Lutsenko, S.V.; Terentiev, A.A. Reactive oxygen and nitrogen species-induced protein modifications: Implication in carcinogenesis and anti-cancer therapy. Cancer Res. 2018, 78, 6040–6047.
- Hielscher, A.; Gerecht, S. Hypoxia and free radicals: Role in tumor progression and the use of engineering-based platforms to address these relationships. Free Radic. Biol. Med. 2015, 79, 281–291.
- Semenza, G.L. Oxygen sensing, hypoxia-inducible factors, and disease pathophysiology. Annu. Rev. Pathol. 2014, 9, 47–71.
- Samanta, D.; Semenza, G.L. Maintenance of redox homeostasis by hypoxia-inducible factors. Redox Biol. 2017, 13, 331–335.
- Wang, G.L.; Jiang, B.H.; Rue, E.A.; Semenza, G.L. Hypoxia-inducible factor 1 is a basic-helix-loop-helix heterodimer that is regulated by cellular O2 tension. Proc. Natl. Acad. Sci. USA 1995, 92, 5510–5514.
- Ke, Q.; Costa, M. Hypoxia-inducible factor-1 (HIF-1). Mol. Pharmacol. 2006, 70, 1469–1480.
- Saito, S.; Lin, Y.-C.; Tsai, M.-H.; Lin, C.-S.; Murayama, Y.; Sato, R.; Yokoyama, K.K. Emerging roles of hypoxia-inducible factors and reactive oxygen species in cancer and pluripotent stem cells. Kaohsiung J. Med. Sci. 2015, 31, 279–286.
- Solaini, G.; Baracca, A.; Lenaz, G.; Sgarbi, G. Hypoxia and mitochondrial oxidative metabolism. Biochim. Biophys. Acta 2010, 1797, 1171–1177.
- Solaini, G.; Sgarbi, G.; Baracca, A. Oxidative phosphorylation in cancer cells. Biochim. Biophys. Acta 2010, 1807, 534–542.
- Berra, E.; Benizri, E.; Ginouves, A.; Volmat, V.; Roux, D.; Pouyssegur, J. HIF prolyl-hydroxylase 2 is the key oxygen sensor setting low steady-state levels of HIF-1 alpha in normoxia. EMBO J. 2003, 22, 4082–4090.
- Ginouves, A.; Ilc, K.; Macías, N.; Pouysségur, J.; Berra, E. PHDs overactivation during chronic hypoxia “desensitizes” HIFα and protects cells from necrosis. Proc. Natl. Acad. Sci. USA 2008, 105, 4745–4750.
- Maxwell, P.H.; Wiesener, M.S.; Chang, G.W.; Clifford, S.C.; Vaux, E.C.; Cockman, M.E.; Wykoff, C.C.; Pugh, C.W.; Maher, E.R.; Ratcliffe, P.J. The tumour suppressor protein VHL targets hypoxia-inducible factors for oxygen-dependent proteolysis. Nature 1999, 399, 271–275.
- Maxwell, P.H.; Pugh, C.W.; Ratcliffe, P.J. Activation of the HIF pathway in cancer. Curr. Opin. Genet. Dev. 2001, 11, 293–299.
- Blancher, C.; Moore, J.W.; Robertson, N.; Harris, A.L. Effects of ras and von Hippel-Lindau (VHL) gene mutations on hypoxia-inducible factor (HIF)-1alpha, HIF-2alpha, and vascular endothelial growth factor expression and their regulation by the phosphatidylinositol 3’-kinase/Akt signaling pathway. Cancer Res. 2001, 61, 7349–7355.
- Lando, D.; Peet, D.J.; Gorman, J.J.; Whelan, D.A.; Whitelaw, M.L.; Bruick, R.K. FIH-1 is an asparaginyl hydroxylase enzyme that regulates the transcriptional activity of hypoxia-inducible factor. Genes Dev. 2002, 16, 1466–1471.
- Kelly, C.; Smallbone, K.; Brady, M. Tumour glycolysis: The many faces of HIF. J. Theor. Biol. 2008, 254, 508–513.
- Shaw, R.J.; Cantley, L.C. Ras, PI(3)K and mTOR signalling controls tumour cell growth. Nature 2006, 441, 424–430.
- Ma, W.; Sung, H.J.; Park, J.Y.; Matoba, S.; Hwang, P.M. A pivotal role for p53: Balancing aerobic respiration and glycolysis. J. Bioenerg. Biomembr. 2007, 39, 243–246.
- Karnoub, A.E.; Weinberg, R.A. Ras oncogenes: Split personalities. Nat. Rev. Mol. Cell Biol. 2008, 9, 517–531.
- Vaseva, A.V.; Moll, U.M. The mitochondrial p53 pathway. Biochim. Biophys. Acta 2009, 1787, 414–420.
- Sauer, H.; Engel, S.; Milosevic, N.; Sharifpanah, F.; Wartenberg, M. Activation of AMP-kinase by AICAR induces apoptosis of DU-145 prostate cancer cells through generation of reactive oxygen species and activation of c-Jun N-terminal kinase. Int. J. Oncol. 2012, 40, 501–508.
- Stine, Z.P.; Walton, Z.E.; Altman, B.J.; Hsieh, A.L.; Dang, C.V. MYC, metabolism, and cancer. Cancer Discov. 2015, 5, 1024–1039.
- Dang, C.V. A time for MYC: Metabolism and therapy. Cold Spring Harb. Symp. Quant. Biol. 2016, 81, 79–83.
- Madan, E.; Parker, T.M.; Pelham, C.J.; Palma, A.M.; Peixoto, M.L.; Nagane, M.; Chandaria, A.; Tomás, A.R.; Canas-Marques, R.; Henriques, V.; et al. HIF-transcribed p53 chaperones HIF-1α. Nucleic Acids Res. 2019, 47, 10212–10234.
- Robertson, E.D.; Semenchenko, K.; Wasylyk, B. Crosstalk between Mdm2, p53 and HIF-1α: Distinct responses to oxidative stress and implications for tumour hypoxia. Subcell. Biochem. 2014, 85, 199–214.
- Bartoletti-Stella, A.; Mariani, E.; Kurelac, I.; Maresca, A.; Caratozzolo, M.F.; Iommarini, L.; Carelli, V.; Eusebi, L.H.; Guido, A.; Cenacchi, G.; et al. Gamma rays induce a p53-dependent mitochondrial biogenesis that is counter-regulated by HIF-1α. Cell Death Dis. 2013, 4, e663.
- Chun, S.Y.; Johnson, C.; Washburn, J.G.; Cruz-Correa, M.R.; Dang, D.T.; Dang, L.H. Oncogenic KRAS modulates mitochondrial metabolism in human colon cancer cells by inducing HIF-1α and HIF-2α target genes. Mol. Cancer. 2010, 9, 293.
- Ogrunc, R.; Di Micco, R.; Liontos, M.; Bombardelli, L.; Mione, M.; Fumagalli, M.; Gorgoulis, V.G.; d’Adda di Fagagna, F. Oncogene-induced reactive oxygen species fuel hyperproliferation and DNA damage response activation. Cell Death Differ. 2014, 21, 998–1012.
- Saitoh, M.; Pullen, N.; Brennan, P.; Cantrell, D.; Dennis, P.B.; Thomas, G. Regulation of an activated S6 kinase 1 variant reveals a novel mammalian target of rapamycin phosphorylation site. J. Biol. Chem. 2002, 277, 20104–20112.
- Miller, D.M.; Thomas, S.D.; Islam, A.; Muench, D.; Sedoris, K. c-Myc and cancer metabolism. Clin. Cancer Res. 2012, 18, 5546–5553.
- Gong, T.; Cui, L.; Wang, H.; Wang, H.; Han, N. Knockdown of KLF5 suppresses hypoxia-induced resistance to cisplatin in NSCLC cells by regulating HIF-1α-dependent glycolysis through inactivation of the PI3K/Akt/mTOR pathway. J. Transl. Med. 2018, 16, 164.
- Chae, Y.C.; Vaira, V.; Caino, M.C.; Tang, H.Y.; Seo, J.H.; Kossenkov, A.V.; Ottobrini, L.; Martelli, C.; Lucignani, G.; Bertolini, I.; et al. Mitochondrial Akt regulation of hypoxic tumor reprogramming. Cancer Cell 2016, 30, 257–272.
- Kiwai, M.T.; Kitadai, Y.; Tanaka, S.; Onogawa, S.; Matsutani, N.; Kaio, E.; Ito, M.; Chayama, K. Expression of hypoxia-inducible factor-1alpha is associated with tumor vascularization in human colorectal carcinoma. Int. J. Cancer 2003, 105, 176–181.
- Huang, G.W.; Yang, L.Y.; Lu, W.Q. Expression of hypoxia-inducible factor 1alpha and vascular endothelial growth factor in hepatocellular carcinoma: Impact on neovascularization and survival. World J. Gastroenterol. 2005, 11, 1705–1708.
- Naruse, T.; Kawasaki, G.; Yanamoto, S.; Mizuno, A.; Umeda, M. Immunochemical study of VEGF expression in oral squamous cell carcinomas: Correlation with the mTOR-HIF-1α pathway. Anticancer Res. 2011, 31, 4429–4437.
- Shao, J.B.; Li, Z.; Zhang, N.; Yang, F.; Gao, W.; Sun, Z.G. Hypoxia-inducible factor-1α in combination with vascular endothelial growth factor could predict the prognosis of postoperative patients with oesophageal squamous cell carcinoma. Pol. J. Pathol. 2019, 70, 84–90.
- Fu, Z.; Chen, D.; Cheng, H.; Wang, F. Hypoxia-inducible factor-1α protects cervical carcinoma cells from apoptosis induced by radiation via modulation of vascular endothelial growth factor and p53 under hypoxia. Med. Sci. Monit. 2015, 21, 318–325.
- Zhang, Y.; Chen, C.; Zhang, J. Effects and significance of formononetin on expression levels of HIF-1α and VEGF in mouse cervical cancer tissue. Oncol. Lett. 2019, 18, 2248–2253.
- Chen, W.; Hill, H.; Christie, A.; Kim, M.S.; Holloman, E.; Pavia-Jimenez, A.; Homayoun, F.; Ma, Y.; Patel, N.; Yell, P.; et al. Targeting renal cell carcinoma with a HIF-2 antagonist. Nature 2016, 539, 112–117.
- Zhu, Y.; Yan, Y.; Gius, D.R.; Vassilpoulos, A. Metabolic regulation of sirtuins upon fasting and the implication for cancer. Curr. Opn. Oncol. 2013, 25, 630–636.
- Liu, H.; Li, S.; Liu, X.; Chen, Y.; Denog, H. SIRT3 overexpression inhibits growth of kidney tumor cells and enhances mitochondrial biogenesis. J. Proteome. Res. 2018, 17, 3143–3152.
- Zhang, C.Z.; Liu, L.; Cai, M.; Pan, Y.; Fu, J.; Cao, Y.; Yun, J. Low SIRT3 expression correlates with poor differentiation and unfavorable prognosis in primary hepatocellular carcinoma. PLoS ONE 2012, 7, e51703.
- Kweon, K.H.; Lee, C.R.; Jung, S.J.; Ban, E.J.; Kang, S.-W.; Jeong, J.J.; Nam, K.-H.; Jo, Y.S.; Lee, J.; Chung, W.Y. Sirt1 induction confers resistance to etoposide-induced genotoxic apoptosis in thyroid cancers. Int. J. Oncol. 2014, 45, 2065–2075.
- Finley, L.W.; Haigis, M.C. Metabolic regulation by SIRT3: Implications for tumorigenesis. Trends Mol. Med. 2012, 18, 516–523.
- Finley, L.W.; Carracedo, A.; Lee, J.; Souza, A.; Egia, A.; Zhang, J.; Teruya-Feldstein, J.; Moreira, P.I.; Cardoso, S.M.; Clish, C.B.; et al. SIRT3 opposes reprogramming of cancer cell metabolism through HIF1α destabilization. Cancer Cell 2011, 19, 416–428.
- Xiong, Y.; Wang, L.; Wang, S.; Wang, M.; Zhao, J.; Zhang, Z.; Li, X.; Jia, L.; Han, Y. SIRT3 deacetylates and promotes degradation of PTEN-defective non-small cell lung cancer. J. Cancer Res. Clin. Oncol. 2018, 144, 189–198.
- Scheepers, A.; Joost, H.-G.; Schurmann, A. The glucose transporter families SGLT and GLUT: Molecular basis of normal and aberrant function. JPEN J. Parenter. Enteral Nutr. 2004, 28, 364–371.
- An, D.; Toyoda, T.; Taylor, E.B.; Yu, H.; Fujii, N.; Hirshman, M.F.; Goodyear, L.J. TBC1D1 regulates insulin- and contraction-induced glucose transport in mouse skeletal muscle. Diabetes 2010, 59, 1358–1365.
- Mafakheri, S.; Flörke, R.R.; Kanngieber, S.; Hartwig, S.; Espelage, L.; De Wendt, C.; Schönberger, T.; Hamker, N.; Lehr, S.; Chadt, A.; et al. AKT and AMP-activated protein kinase regulate TBC1D1 through phosphorylation and its interaction with the cytosolic tail of insulin-regulated aminopeptidase IRAP. J. Biol. Chem. 2018, 16293, 17853–17862.
- Ancey, P.-B.; Contat, C.; Meylan, E. Glucose transporters in cancer—From tumor cells to the tumor microenvironment. FEBS J. 2018, 285, 2926–2943.
- Jóźwiak, P.; Krzeŝlak, A.; Pomorski, L.; Lipińka, A. Expression of hypoxia-related glucose transporters GLUT1 and GLUT3 in benign, malignant and non-neoplastic thyroid lesions. Mol. Med. Rep. 2012, 6, 601–606.
- Krzeŝlak, A.; Wojcik-Krowiranda, K.; Forma, E.; Jóźwiak, P.; Romanowicz, H.; Bienkiewicz, A.; Brys, M. Expression of GLUT1 and GLUT3 glucose transporters in endometrial and breast cancers. Pathol. Oncol. Res. 2012, 18, 721–728.
- Kuo, C.C.; Ling, H.H.; Chiang, M.C.; Chung, C.H.; Lee, W.Y.; Chu, C.Y.; Wu, Y.C.; Chen, C.H.; Lai, Y.W.; Tsai, I.L.; et al. Metastatic colorectal cancer rewrites metabolic program through a Glut3-YAP-dependent signaling. Theranostics 2019, 9, 2526–2540.
- Wu, N.; Zheng, B.; Shaywitz, A.; Dagon, Y.; Tower, C.; Bellinger, G.; Shen, C.-H.; Wen, J.; Asara, J.; McGraw, T.E. AMPK-dependent degradation of TXNIP upon energy stress leads to enhanced glucose uptake via GLUT1. Mol. Cell 2013, 49, 1167–1175.
- Tanegashiwa, K.; Sato-Miyata, Y.; Funakoshi, M.; Nishito, Y.; Aigaki, T.; Hara, T. Epigenetic regulation of the glucose transporter gene Slc2a1 by the β-hydroxybutyrate underlies preferential glucose supply to the brain of fasted mice. Genes Cells 2017, 22, 71–83.
- Masin, M.; Vazquez, J.; Rossi, S.; Groeneveld, S.; Samson, N.; Schwalie, P.C.; Deplanck, B.; Frawley, L.E.; Gouttenoire, J.; Moradpour, D.; et al. GLUT3 is induced during epithelial-mesenchymaal transition and promotes tumor cell proliferation in non-small cell lung cancer. Cancer Metabol. 2014, 2, 11.
- Ali, A.; Levantini, E.; Fhu, C.W.; Teo, J.T.; Clohessy, J.G.; Goggi, J.L.; Wu, C.-S.; Chen, L.; Chin, T.M.; Tenen, D.G. CAV1-GLUT3 signaling is important for cellular energy and can be targeted by atorvastatin in non-small cell lung cancer. Theranostics 2019, 9, 6157–6174.
- Aida, E.; Yasuda, M.; Miyazawa, M.; Fujita, M.; Osamura, R.Y.; Harisawa, T.; Muramatsu, T.; Murakami, M.; Saito, K.; Mikami, M. Hypoxic status in ovarian serous and mucinous tumors: Relationship between histological characteristics and HIF-1alpha/GLUT-1 expression. Arch. Gynecol. Obstet. 2008, 277, 539–546.
- Wincewicz, A.; Sulkowska, M.; Koda, M.; Sulkowski, S. Clinicopathological significance and linkage of the distribution of HIF-1 in human primary colorectal cancer. Pathol. Oncol. Res. 2007, 13, 15–20.
- Hao, L.S.; Liu, Q.; Tian, C.; Zhang, D.X.; Wang, B.; Zhou, D.X.; Li, Z.P.; Yuan, Z.X. Correlation and expression analysis of hypoxia-inducible factor 1α, glucose transporter 1 and lactate dehydrogenase 5 in human gastric cancer. Oncol. Lett. 2019, 18, 1431–1441.
- Yang, T.Y.; Hao, L.S.; Guo, D.Z. Expression of hypoxia-inducible factor 1α, glucose transporter 1 and lactate dehydrogenase 5 in colorectal cancer and clinicopathological significance. Zhonghua Bing Li Xue Za Zhi 2017, 46, 93–97.
- Bobarykina, A.Y.; Minchenko, D.O.; Opentanova, I.L.; Moenner, M.; Caro, J.; Esumi, H.; Minchenko, O.H. Hypoxic regulation of PFKFB and PFKFB-4 gene expression in gastric and pancreatic cell lines and expression of PFKFB genes in gastric cancers. Acta Biochim. Pol. 2006, 53, 789–799.
- Cori, C.F.; Cori, G.T. The carbohydrate metabolism of tumors: II. Changes in the sugar, lactic acid, and CO2-combining power of blood passing through a tumor. J. Biol. Chem. 1925, 65, 397–405.
- Ristow, M. Oxidative metabolism in cancer growth. Curr. Opin. Clin. Nutr. Metab. Care 2006, 9, 339–345.
- Shoshan, M. On mitochondrial metabolism in tumor biology. Curr. Opin. Oncol. 2017, 29, 48–54.
- Cancemi, P.; Buttacavoli, N.; Roz, E.; Feo, S. Expression of alpha-enolase (ENO1), Myc promoter-binding protein-1 (MBP-1) and matrix metalloproteinases (MMP-2 and MMP-9) reflect the nature and aggressiveness of breast tumors. Int. J. Mol. Sci. 2016, 20, 3952.
- Jiang, Z.; Wang, X.; Li, J.; Yang, H.; Lin, X. Aldolase A as a prognostic factor and mediator of progression via inducing epithelial-to-mesenchymal transition in gastric cancer. J. Cell Mol. Med. 2018, 22, 4377–4386.
- Yasuda, A.S.; Mori, A.; Isobe, N.; Yang, W.; Oe, H.; Fujimoto, A.; Yonenaga, Y.; Sakashita, H.; Imamura, M. Hexokinase II and VEGF expression in liver tumors: Correlation with hypoxia-inducible factor 1 alpha and its significance. J. Hepatol. 2004, 40, 117–123.
- Jin, Z.; Gu, J.; Xin, X.; Li, Y.; Wang, H. Expression of hexokinase 2 in epithelial ovarian tumors and its clinical significance in serous ovarian cancer. Eur. J. Gynaecol. Oncol. 2014, 35, 519–524.
- Guzman, G.; Chennuri, R.; Chan, A.; Rea, B.; Quintana, A.; Patel, R.; Xu, P.Z.; Xie, H.; Hay, N. Evidence for heightened hexokinase II immunoexpression in hepatocyte dysplasia and hepatocellular carcinoma. Dig. Dis. Sci. 2015, 60, 420–426.
- Bartrons, R.; Simon-Molas, H.; Rodriguez-Garcia, A.; Castano, E.; Navarro-Sabaté, A.; Manzano, A.; Martinez-Outschoom, U.E. Fructose 2.6-bisphosphate in cancer cell metabolism. Front. Oncol. 2018, 8, 331.
- Yang, H.; Shu, Z.; Jiang, Y.; Mao, W.; Pang, L.; Redwood, A.; Jeter-Jones, S.L.; Jennings, N.B.; Ometas, A.; Zhou, J.; et al. 6-Phosphofructo-2-kinase/Fructose-2,6-bisphosphatase-2 regulates TP53-dependent paclitaxel sensitivity in ovarian and breast cancers. Clin. Cancer Res. 2019, 25, 5702–5716.
- Fiorillo, A.; Petrosino, M.; Ilari, A.; Pasquo, A.; Cipollone, A.; Maggi, M.; Chiaraluce, R.; Consalvi, V. The phosphoglycerate kinase 1 variants found in carcinoma cells display different catalytic activity and conformational stability compared to native enzyme. PLoS ONE 2018, 13, e0199191.
- Israelsen, W.J.; Heiden, M.G.V. Pyruvate kinase: Function, regulation and role in cancer. Semin. Cell Dev. Biol. 2015, 43, 43–51.
- Li, H.; Xu, H.; Xing, R.; Pan, Y.; Li, W.; Cui, J.; Lu, Y. Pyruvate kinase M2 contributes to cell growth in gastric cancer via aerobic glycolysis. Pathol. Res. Pract. 2019, 215, 152409.
- Li, S.; Ji, X.; Wang, R.; Miao, Y. Follicle-stimulating hormone promoted pyruvate kinase isozyme type M2-induced glycolysis and proliferation of ovarian cancer cells. Arch. Gynecol. Obstet. 2019, 299, 1443–1451.
- Sun, Q.; Chen, X.; Ma, J.; Peng, H.; Wang, F.; Zha, X.; Wang, Y.; Jing, Y.; Chen, R.; Chang, L.; et al. Mammalian target of rapamycin up-regulation of pyruvate kinase isoenzyme type M2 is critical for aerobic glycolysis and tumor growth. Proc. Natl. Acad. Sci. USA 2011, 108, 4129–4134.
- Said, H.M.; Polat, B.; Hagemann, C.; Anacker, J.; Fientje, M.; Vordermark, D. Absence of GAPDH regulation in tumor cells of different origin under hypoxic conditions in vitro. BMC Res. Notes 2009, 2, 8.
- Sanders, E.; Diehl, S. Analysis and interpretation of transcriptomic data obtained from extended Warburg effect genes in patients with clear cell renal carcinoma. Oncoscience 2015, 2, 151–186.
- Ortega, A.D.; Sanchez-Arago, M.; Giner-Sanchez, D.; Sanchez-Cenizo, L.; Willers, I.; Cuezva, J.M. Glucose avidity of carcinomas. Cancer Lett. 2009, 276, 125–135.
- Opperman, H.; Birkemeyer, C.; Meixensberger, J.; Gaunitz, F. Non-enzymatic reaction of carnosine and glycerol-3-phosphate accompanies metabolic changes in the pentose phosphate pathway. Cell Prolif. 2019, e12702.
- Preuss, J.; Richardson, A.D.; Pinkerton, A.; Hedrick, M.; Sergienko, E.; Rahlfs, S.; Becker, K.; Bode, L. Identification and characterization of novel human glucose-6-phosphate dehydrogenase Inhibitors. J. Biomol. Screen. 2013, 18, 286–297.
- Hong, W.; Cai, P.; Xu, C.; Cao, D.; Yu, W.; Zhao, Z.; Huang, M.; Jin, J. Inhibition of glucose-6-phosphate dehydrogenase reverses cisplatin resistance in lung cancer cells via the redox system. Front. Pharmacol. 2018, 9, 43.
- Lucarelli, G.; Galleggiante, V.; Rutigliano, M.; Sanguedolce, F.; Cagiano, S.; Bufo, P.; Lastilla, G.; Maiorano, E.; Ribatti, D.; Giglio, A.; et al. Metabolomic profile of glycolysis and the pentose phosphate pathway identifies the central role of glucose-6-phosphate dehydrogenase in clear cell-renal cell carcinoma. Oncotarget 2015, 6, 13371–13386.
- Zheng, W.; Feng, Q.; Liu, J.; Guo, Y.; Gao, L.; Li, R.; Xu, M.; Yan, G.; Yin, Z.; Zhang, S.; et al. Inhibition of 6-phosphogluconate dehydrogenase reverses cisplatin resistance in ovarian and lung cancer. Front. Pharmacol. 2017, 8, 421.
- Zhang, H.; Zhang, H.; Wang, S.; Ni, Z.; Wang, T. 1-Hydroxy-8-methoxy-anthraquinon reverses cisplatin resistance by inhibiting 6PGD in cancer cells. Open Life Sci. 2019, 14, 454–461.
- Diaz-Moralli, S.; Tarrado-Castellarnau, M.; Alenda, C.; Castells, A.; Cascante, M. Transketolase-like 1 expression is modulated during colorectal cancer progression and metastasis formation. PLoS ONE 2011, 6, e25323.
- Benito, A.; Polat, I.H.; Noé, V.; Ciudad, C.J.; Marin, S.; Cascante, M. Glucose-6-phosphate dehydrogenase and transketolase modulate breast cancer cell metabolic reprogramming and correlate with poor patient outcome. Oncotarget 2017, 8, 106693–106706.
- Chao, Y.K.; Peng, T.L.; Chuang, W.Y.; Yeh, C.J.; Li, Y.L.; Lu, Y.C.; Cheng, A.J. Transketolase serves a poor prognosticator in esophageal cancer by promoting cell invasion via epithelial-mesenchymal transition. J. Cancer 2016, 7, 1804–1811.
- Sun, W.; Liu, Y.; Glazer, C.A.; Shao, C.; Bhan, S.; Demokan, S.; Zhao, M.; Rudek, M.A.; Ha, P.K.; Califano, J.A. TKTL1 is activated by promoter hypomethylation and contributes to head and neck squamous cell carcinoma carcinogenesis through increased aerobic glycolysis and HIF1alpha stabilization. Clin. Cancer Res. 2010, 16, 857–866.
- Fritz, P.; Coy, J.F.; Mürdter, T.E.; Ott, G.; Alscher, M.D.; Friedel, G. TKTL-1 expression in lung cancer. Pathol. Res. Pract. 2012, 208, 203–209.
- Jiang, P.; Du, W.; Mancuso, A.; Wellen, K.E.; Yang, X. Reciprocal regulation of p53 and malic enzymes modulates metabolism and senescence. Nature 2013, 493, 689–693.
- Bedard, K.; Krause, K.H. The NOX family of ROS-generating NADPH oxidases: Physiology and pathophysiology. Physiol. Rev. 2007, 87, 245–313.
- Keeley, T.P.; Mann, G.E. Defining physiological normoxia for improved translation of cell physiology to animal models and humans. Physiol. Rev. 2019, 99, 161–234.
- McKeown, S.R. Defining normoxia, physoxia and hypoxia in tumors—Implications for treatment response. Br. J. Radiol. 2014, 87, 20130676.
- Carreau, A.; El Hafny-Rahbi, B.; Matejuk, A.; Grillon, C.; Kieda, C. Why is the partial oxygen pressure of human tissues a crucial parameter? Small molecules and hypoxia. J. Cell. Mol. Med. 2011, 15, 1239–1253.
- San-Millan, I.; Brooks, G. Reexamining cancer metabolism: Lactate production for carcinogenesis could be the purpose and explanation of the Warburg effect. Carcinogenesis 2017, 38, 119–133.
- Hamaguchi, T.; Izuka, N.; Tsunedomi, R.; Haamamoto, Y.; Miyamoto, T.; Iida, M.; Tokuhisa, Y.; Sakamoto, K.; Takashima, M.; Tamesa, M.; et al. Glycolysis module activated by hypoxia-inducible factor 1alpha is related to the aggressive phenotype of hepatocellular carcinoma. Int. J. Oncol. 2008, 33, 725–731.
- Sanchez-Sanchez, A.M.; Antolin, I.; Puente-Moncada, N.; Suarez, S.; Gomez-Lobo, M.; Rodrigez, C.; Martin, V. Melatonin cytotoxicity is associated to Warburg effect inhibition in Ewing sarcoma cells. PLoS ONE 2015, 10, e0135420.
- Kato, Y.; Ozawa, S.; Miyamoto, C.; Maehata, Y.; Suzuki, A.; Maeda, T.; Baba, Y. Acidic extracellular microenvironment and cancer. Cancer Cell Int. 2013, 13, 89.
- Pouysségur, J.; Sardet, C.; Franchi, A.; L’Allemain, G.; Paris, S. A specific mutation abolishing Na+/H+ antiport activity in hamster fibroblasts precludes growth at neutral and acidic pH. Proc. Natl. Acad. Sci. USA 1984, 81, 4833–4837.
- Park, H.J.; Lyons, J.C.; Ohtsubo, T.; Song, C.W. Acidic environment causes apoptosis by increasing caspase activity. Br. J. Cancer 1999, 80, 1892–1897.
- San-Millán, I.; Julian, C.G.; Matarazzo, C.; Martinez, J.; Brooks, G.A. Is lactate an oncometabolite? Evidence supporting a role for lactate in the regulation of transcriptional activity of cancer-related genes in MCF7 breast cancer cells. Front. Oncol. 2020, 9, 1536.
- Cassim, S.; Pouyssegur, J. Tumor microenvironment: A metabolic player that shapes the immune response. Int. J. Mol. Sci. 2020, 21, 157.
- Ding, J.; Karp, J.E.; Emadi, A. Elevated lactate dehydrogenase (LDH) can be a marker of immune suppression in cancer: Interplay between hematologic and solid neoplastic clones and their microenvironments. Cancer Biomark. 2017, 19, 353–363.
- Husain, Z.; Huang, Y.; Seth, P.; Sukhatme, V.P. Tumor-derived lactate modifies antitumor immune response: Effect on myeloid-derived suppressor cells and NK cells. J. Immunol. 2013, 191, 1486–1495.
- Chiche, J.; Ilc, K.; Laferrière, J.; Trottier, E.; Dayan, F.; Mazure, N.M.; Brahimi-Horn, M.C.; Pouysségur, J. Hypoxia-inducible carbonic anhydrase IX and XII promote tumor cell growth by counteracting acidosis through the regulation of the intracellular pH. Cancer Res. 2009, 69, 358–368.
- Lobo, R.C.; Hubbard, N.E.; Damonte, P.; Mori, H.; Penzvalto, Z.; Pham, C.; Koehne, A.L.; Go, A.C.; Anderson, S.E.; Cala, P.M.; et al. Glucose uptake and intracellular pH in a mouse model of ductal carcinoma in situ (DCIS) suggests metabolic heterogeneity. Front. Cell Dev. Biol. 2016, 4, 93.
- Parks, S.K.; Chiche, J.; Pouyssegur, J. pH control mechanisms of tumor survival and growth. J. Cell. Physiol. 2011, 226, 299–308.
- Spugnini, E.P.; Sonveaux, P.; Stock, C.; Perez-Sayans, M.; De Milito, A.; Avnet, S.; Garcia, A.G.; Harguindey, S.; Fais, S. Proton channels and exchangers in cancer. Biochim. Biophys. Acta 2015, 1848, 2715–2726.
- Sonveaux, P.; Végran, F.; Schroeder, T.; Wergin, M.C.; Verrax, J.; Rabbani, Z.N.; De Saedeleer, C.J.; Kennedy, K.M.; Diepart, C.; Jordan, B.F.; et al. Targeting lactate-fueled respiration selectively kills hypoxic tumor cells in mice. J. Clin. Investig. 2008, 118, 3930–3942.
- Brooks, G.A. Cell-cell and intracellular lactate shuttles. J. Physiol. 2009, 587, 5591–5600.
- Faubert, B.; Li, K.Y.; Cai, L.; Hensley, C.T.; Kim, J.; Zacharias, L.G.; Yang, C.; Do, Q.N.; Doucette, S.; Burguete, D.; et al. Lactate metabolism in human lung tumors. Cell 2017, 171, 358–371.
- Hui, S.; Ghergurovich, J.M.; Morscher, R.J.; Jang, C.; Teng, X.; Lu, W.; Esparza, L.A.; Reya, T.; Zhang, L.; Guo, J.Y.; et al. Glucose feeds the TCA cycle via circulating lactate. Nature 2017, 551, 115–118.
- Wu, H.; Ying, M.; Hu, X. Lactic acidosis switches cancer cells from aerobic glycolysis back to dominant oxidative phosphorylation. Oncotarget 2016, 7, 40621–40629.
- Menendez, J.A.; Lupu, R. Fatty acid synthase and the lipogenic phenotype in cancer pathogenesis. Nat. Rev. Cancer 2007, 7, 763–777.
- Guillaumond, F.; Bidaut, G.; Ouaissi, M.; Servais, S.; Gouirand, V.; Olivares, O.; Lac, S.; Borge, L.; Roques, J.; Gayet, O.; et al. Cholesterol uptake disruption, in association with chemotherapy, is a promising combined metabolic therapy for pancreatic adenocarcinoma. Proc. Natl. Acad. Sci. USA 2015, 112, 2473–2478.
- Qin, C.; Yang, G.; Yang, J.; Ren, B.; Wang, H.; Chen, G.; Zhao, F.; You, L.; Wang, W.; Zhao, Y. Metabolism of pancreatic cancer: Paving the way to better anticancer strategies. Mol. Cancer 2020, 19, 50.
- Zhao, S.; Torres, A.; Henry, R.A.; Trefely, S.; Wallace, M.; Lee, J.V.; Carrer, A.; Sengupta, A.; Campbell, S.L.; Kuo, Y.M.; et al. ATP-citrate lyase controls a glucose-to-acetate metabolic switch. Cell Rep. 2016, 17, 1037–1052.
- Yoshii, Y.; Furukawa, T.; Saga, T.; Fujibayashi, Y. Acetate/acetyl-CoA metabolism associated with cancer fatty acid synthesis: Overview and application. Cancer Lett. 2015, 356, 211–216.
- Zu, X.Y.; Zhang, Q.H.; Liu, J.H.; Cao, R.X.; Zhong, J.; Yi, G.H.; Quan, Z.H.; Pizzorno, G. ATP citrate lyase inhibitors as novel cancer therapeutic agents. Recent Patents Anticancer. Drug Discov. 2012, 7, 154–167.
- Wellen, K.E.; Hatzivassiliou, G.; Sachdeva, U.M.; Bui, T.V.; Cross, J.R.; Thompson, C.B. ATP-citrate lyase links cellular metabolism to histone acetylation. Science 2009, 22, 1076–1080.
- Li, Y.; Gruber, J.J.; Litzenburger, U.M.; Zhou, Y.; Miao, Y.R.; LaGory, E.L.; Li, A.M.; Hu, Z.; Yip, M.; Hart, L.S.; et al. Acetate supplementation restores chromatin accessibility and promotes tumor cell differentiation under hypoxia. Cell Death Dis. 2020, 11, 102.
- Schug, Z.T.; Peck, B.; Jones, D.T.; Zhang, Q.; Grosskurth, S.; Alam, I.S.; Goodwin, L.M.; Smethurst, E.; Mason, S.; Blyth, K.; et al. Acetyl-CoA synthetase 2 promotes acetate utilization and maintains cancer cell growth under metabolic stress. Cancer Cell 2015, 27, 57–71.
- Mashimo, T.; Pichumani, K.; Vemireddy, V.; Hatanpaa, K.J.; Singh, D.K.; Sirasanagandla, S.; Nannepaga, S.; Piccirillo, S.G.; Kovacs, Z.; Foong, C.; et al. Acetate is a bioenergetic substrate for human glioblastoma and brain metastases. Cell 2014, 159, 1603–1614.
- Tretter, L.; Adam-Vizi, V. Alpha-ketoglutarate dehydrogenase: A target and generator of oxidative stress. Philos. Trans. R. Soc. Lond. B Biol Sci. 2005, 360, 2335–2345.
- Sun, R.C.; Denko, N.C. Hypoxic regulation of glutamine metabolism through HIF1 and SIAH2 supports lipid synthesis that is necessary for tumor growth. Cell Metab. 2014, 19, 285–292.
- Lien, E.C.; Vander Heiden, M.G. A framework for examining how diet impacts tumour metabolism. Nat. Rev. Cancer 2019, 19, 651–661.
- Peck, B.; Schulze, A. Lipid metabolism at the nexus of diet and tumor microenvironment. Trends Cancer 2019, 5, 693–703.
- Sunami, Y.; Rebelo, A.; Kleeff, J. Lipid metabolism and lipid droplets in pancreatic cancer and stellate cells. Cancers 2017, 10, 3.
- Atoum, M.F.; Alzoughool, F.; Al-Hourani, H. Linkage between obesity. Leptin and breast cancer. Breast Cancer 2020, 14.
- Bhaskaran, K.; Douglas, I.; Forbes, H.; dos-Santos-Silva, I.; Leon, D.A.; Smeeth, L. Body-mass index and risk of 22 specific cancers: A population-based cohort study of 5·24 million UK adults. Lancet 2014, 384, 755–765.
- Philip, B.; Roland, C.L.; Daniluk, J.; Liu, Y.; Chatterjee, D.; Gomez, S.B.; Ji, B.; Huang, H.; Wang, H.; Fleming, J.B.; et al. A high-fat diet activates oncogenic Kras and COX2 to induce development of pancreatic ductal adenocarcinoma in mice. Gastroenterology 2013, 145, 1449–1458.
- Luo, Y.; Yang, Y.; Liu, M.; Wang, D.; Wang, F.; Bi, Y.; Ji, J.; Li, S.; Liu, Y.; Chen, R.; et al. Oncogenic KRAS reduces expression of FGF21 in acinar cells to promote pancreatic tumorigenesis in mice on a high-fat diet. Gastroenterology 2019, 157, 1413–1428.





