| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Ricardo Gaspar | + 5386 word(s) | 5386 | 2020-11-11 09:59:07 | | | |
| 2 | Ricardo Gaspar | + 10 word(s) | 5396 | 2021-01-28 13:21:29 | | | | |
| 3 | Catherine Yang | -1962 word(s) | 3434 | 2021-01-29 07:08:52 | | |
Video Upload Options
An important advantage of Nucleic Acid (NA)-based therapies over synthetic drugs and protein treatments is the prospect of a more universal approach to designing therapies. Designing NAs with different sequences, for different targets, can be achieved by using the same technologies. This versatility and scalability of NA drug design and production on demand open the way for more efficient, affordable and personalized treatments in the future. However, the delivery of exogenous therapeutic NAs into the patients’ targeted cells is also challenging. Membrane-type lipids exhibiting permanent or transient cationic character have been shown to associate with NAs (anionic), forming nanosized lipid-NA complexes. These complexes form a wide variety of nanostructures, depending on the global formulation composition and properties of the lipids and NAs. Importantly, these different lipid-NA nanostructures interact with cells via different mechanisms and their therapeutic potential can be optimized to promising levels in vitro. The complexes are also highly customizable in terms of surface charge and functionalization to allow a wide range of targeting and smart-release properties. Most importantly, these synthetic particles offer possibilities for scaling-up and affordability for the population at large. Hence, the versatility and scalability of these particles seem ideal to accommodate the versatility that NA therapies offer.
1. Lipid-Nucleic Acid nanoparticle formation, Structure and Stability
Lipid-Nucleic Acid (NA) nanoparticles (or lipoplexes) readily form for a variety of lipids and NAs [1][2]. This constitutes one of their main advantages: ease of formation and versatility [3]. Here, we attempt to provide a simplified fundamental description about their formation, structure and stability.
2. Lipid-NA Complexation: Counterion Release as the Main Driving Force for Lipid-NA Association
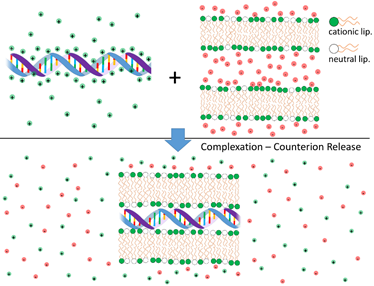
Cationic liposomes (CLs) and DNA interact strongly leading to the formation of complexes with well-organized structures. The electrostatic free energy driving complexation between the two species has two main contributions: (i) Coulomb attractions between cationic liposomes and NAs (anionic); and (ii) the large gain in entropy that results from the release of the inorganic counterions from both cationic lipid and DNA when these two species partially neutralize each other [4][5][6] (Figure 1). Indeed, this second component is the major driving force for complexation. These findings have been convincingly demonstrated through calorimetric measurements, where the enthalpy change involved in the complexation between CLs and DNA was found to be endothermic (thus requiring an increase in entropy to drive complex formation) [7], and by measurements of conductivity to count the number of released ions [8]. Even when used qualitatively, the concept of counterion release provides a strong insight into the overall lipoplex phase behavior. For instance, this framework helps explaining why anionic liposomes and DNA form complexes in the presence of divalent inorganic salt (eg. Ca2+) [9]. Since a divalent cation can simultaneously neutralize one negative charge in both the DNA and anionic liposome, two of the original monovalent counterions can be released, leading to a net gain in entropy and complexation. A similar mechanism can also explain the formation of complexes between zwitterionic liposomes and DNA in the presence of Ca2+ [10][11]. The formation of complexes driven by counterion release is a widespread phenomenon in colloid science [12] and in biology [6][13][14].
Figure 1. Schematic illustration of the release of inorganic counterions associated with cationic liposome-DNA complexation and consequent increase of the system entropy. On the top are shown segments of non-complexed DNA (anionic) and cationic bilayers. Shown are also their inorganic counterions, with limited mobility and in close proximity to the charged surfaces. On the bottom a segment of an isoelectric complex is shown. As the oppositely charged DNA and cationic bilayers neutralize each other, the inorganic counterions are released to the surrounding medium, leading to a high increase in the system entropy. This entropic gain is the main contribution to the complexation free energy.
3. Cationic-to-anionic charge ratio, overcharging and colloidal stability of Lipid-NA complexes
The cationic-to-anionic charge ratio, here represented as rchg, is a critical parameter influencing the structure, lipoplex colloidal stability and their interactions with biologic systems [3][15][16][17][18][19]. rchg is defined by the total number of cationic charges coming from the lipid system divided by the total number of anionic charges coming from the NA.
The counterion release in lipoplex formation is maximum, and almost complete when rchg ~1, around the particle isoelectric point. Such particles have a near net-zero charge, typically showing poor colloidal stability and formation of large aggregates [4]. For rchg greater or smaller than one, the lipoplexes become overcharged with a net positive or negative charge, respectively [20]. The extent of lipoplex overcharging depends on the lipoplex structure and lipid membrane charge density (σM). Typically, lipoplexes can become more positively overcharged if lipid membranes have higher σM. Conversely, lipid membranes with lower σM can become more negatively overcharged [20]. It is also plausible to expect differences in overcharging degree between DNA, siRNA and mRNA, as observed for different polyelectrolytes [21][22][23].
4. Lipid-NA Structure
DNA is a rigid and long molecule. mRNA is typically somewhat shorter than DNA plasmids and significantly more flexible due to being a single strand. Further, because of the single-stranded nature, the exposed nucleobases make mRNA significantly hydrophobic. siRNA is a rigid but short molecule. All these NAs are highly charged. Lipids can have positive, neutral or negative spontaneous curvature (H0), with typical bending rigidities κ significantly larger than the thermal energy kBT. This means that lipids tend to organize in structures that favor their H0, such as micelles, bilayers and inverted micelles.
When cationic liposomes and NAs come into contact, they will maximize the gain of entropy from counterion release, while accommodating elastic requirements from both the lipid film and NAs. Given the diverse nature of lipidic systems and NAs, several structures are found for lipid-NA systems. Importantly, these structures result in different interactions with cells and different transfection efficiencies. Knowledge of the main factors leading to these structures and how to tune them, leads to knowledge of how to improve their therapeutic potential.
Overall, the electrostatic free energy (which accounts for both Coulomb attraction and counterion release) is maximized if the cationic lipid is able to fully wrap the NA double helix. However, the elastic energy required to bend the lipid membranes and NA chains also has to be taken into account. The resulting lipid-NA structures will represent a compromise between these three most-dominant energetic terms (electrostatic interactions and elasticity from both NA and lipid film). Due to the relatively high rigidity of the DNA double helix, structures that favor a suitable contact between the cationic lipids and DNA without bending DNA significantly will be favored. This is the case for the most common multilamellar phase (LaC) and normal and inverted hexagonal phase (HIC and HIIC) structures [24]. Regarding the lipid membranes, if their bending rigidity κ is significantly greater than the thermal energy (kBT), the elastic free energy of the lipid will also be significant and the structure of the complex will be governed by the symmetry of the lipid phase [25] (Figure 2). In other words, the curvature of the lipid membrane will define if the lipoplex will have a LaC, HIC or HIIC structure. If κ is significantly lower, close to kBT, deforming the lipid membrane is less costly and a HIIC structure is favored, since it optimizes the contact between the cationic lipids and DNA. If siRNA is used instead of DNA, since it is significantly shorter, it allows the formation of bicontinuous cubic phases if that is the preferred arrangement of the lipid phase [26]. Below we will overview most of the structures resolved so far for lipid-NA systems.
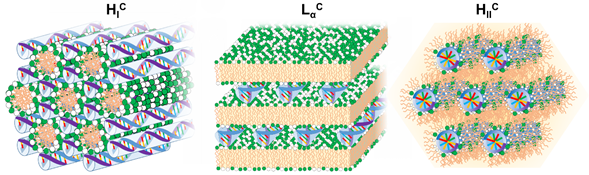
Figure 2. Schematic illustration of the most common structures found for cationic lipid-DNA assemblies. On the left the normal hexagonal phase (HIC) is shown, with lipid elongated micelles arranged on a hexagonal lattice and the DNA rods arranged on a honeycomb lattice in the interstices between the lipid micelles [27]. In the middle the lamellar phase (LαC), with alternating lipid bilayers and DNA monolayers sandwiched between them [4]. On the right the inverted hexagonal phase (HIIC) with lipid inverted micelles coating the DNA arranged on a hexagonal lattice [28].
5. Lamellar complexes – LaC
The LaC is favored by lipid membranes with spontaneous membrane curvature H0 = 0. This zero curvature can be accomplished with many different lipid combinations, including with lipids that want to impose negative curvature such as 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (DOPE) and MO. The LaC structure consists of stacked lipid membranes with DNA chains intercalated between them (Figure 2). This structure is able to provide favorable electrostatic interactions between the DNA and cationic membranes while satisfying the elastic requirements of both. That is, the DNA molecule is allowed to remain relatively rigid without excessive bending, and the lipid membranes are allowed to keep their zero mean curvature. The lipid membranes are typically composed of a mixture of cationic and neutral lipids. Even within this flat membrane arrangement, because the lipid chains are typically in the liquid state, cationic lipids tend to undergo local demixing and accumulate preferentially in the vicinity of the DNA rods. This leads to enhanced Coulomb interactions and maximizes entropy from counterion release at the cost of a smaller entropic penalty from the local demixing of the lipid [5][6]. The LaC structure was convincingly demonstrated by small-angle X-ray scattering (SAXS) measurements [4] and cryogenic transmission electron microscopy (cryo-TEM) [29][30][31]. The X-ray data further reveals that the DNA macromolecules between the lipid bilayers are organized in a 2D smectic phase [4][20][32] in which the average distance between neighboring DNA chains is tunable by the lipid membrane charge density σM and rchg. This is confirmed by fluorescence cross-correlation spectroscopy (FCCS) measurements, which show that for constant rchg, higher σM leads to a larger number of DNA plasmids per particle, while for fixed σM, a higher rchg leads to a smaller number of DNA plasmids per particle since there are more cationic particles on where to distribute the DNA [33]. The existence of divalent cations in solution can also lead to further condensation of DNA [34].
The LaC also forms when CLs are mixed with siRNA [35][36][37]. In contrast with lipid-DNA lipoplexes, the SAXS patterns of siRNA lipoplexes using DOTAP or multivalent cationic lipid MVL5 barely show siRNA-siRNA correlation peaks, which suggests that there is no end-to-end stacking. This is consistent with a 2D liquid-like correlation behavior of the short siRNA rods in contrast with the 2D smectic and 3D columnar arrangements from the long and short DNA systems [36]. Another interesting observation is that perhaps due to the 2D liquid-like nature of the siRNA organization embedded in the LaC, the overall structure seems to equilibrate more rapidly. This is suggested by the observation of narrower Bragg peaks with SAXS, which also indicates more ordered structures [36].
Regarding mRNA, despite their present enthusiasm for therapeutic applications (either for expression of therapeutic proteins [38] or cancer immunotherapy approaches [17][39]), studies on the structures of lipid-mRNA complexes are far scarcer [40][41][42][43]. In general, the behavior of lipid-mRNA assembly tends to follow the behavior observed for DNA, and lipid compositions favoring bilayers with zero curvature tend to favor LaC structures also in the cases of single stranded RNA [40][41]. However, as pointed out above, mRNA is significantly more flexible than DNA (Fig. 1); the charge density is ca. half of that of DNA, which is still highly charged; and the fact that the nucleobases are exposed may lead to additional hydrophobic interactions with the lipids, as it is observed with single-stranded DNA [44][45][46][47]. Cationic liposomes containing DOTAP and DOPC were shown to form well-ordered lamellar complexes with mRNA, with SAXS patterns clearly showing the first and second order Bragg peaks [41].
6. Inverted hexagonal complexes – HIIC
The inverted hexagonal structure (HIIC) is favored by lipid membranes with negative spontaneous membrane curvature H0 < 0, such as cationic membranes containing DOPE above a critical molar fraction [28]. These structures are highly efficient at transfecting cells in vitro, presumably due to their ease to fuse with the endosomal membrane, facilitating the lipoplex escape into the cytoplasm [28][48][49]. However, in most cases, the HIIC forms in a narrow lipid composition range, which makes it also somewhat more challenging to formulate reproducibly under broader experimental conditions – a challenge that needs to be taken into account for scaling-up. Whereas in LaC complexes, the structure can regulate itself by adjusting the separation between the DNA rods, ensuring an optimal matching of cationic charge on the membranes to the amount of DNA intercalated between them, in the HIIC complexes this degree of freedom does not exist [50]. The system therefore separates rather easily into two phases with a HIIC typically coexisting with an LaC phase, as is found in the DOTAP-DOPE system [28]. The HIIC also forms when mixing DNA with cationic membranes that naturally tend to favor bicontinuous cubic phases instead of inverted hexagonals such as monoolein-DOTAP systems [51]. This is because bicontinuous cubic bilayers would force DNA to bend, imposing an energetic penalty that disfavors the cubic and favors inverted hexagonal phases instead. As will be seen below, the bicontinuous cubic phase is recovered again at the expense of the hexagonal if siRNA [52] or non-sticky short DNA fragments [51] are used instead of DNA.
The HIIC also forms when CLs with H0 < 0 are mixed with siRNA [36]. In contrast with DNA complexes, HIIC siRNA complexes exhibited high toxicity and much lower target-specific gene silencing than their lamellar analogues.
7. Inverse bicontinuous cubic complexes – QIIC
Inverse bicontinuous cubic phases occur in some natural lipid systems, such as monoolein- and phytantriol-water systems [53][54][55][56]. These structures are favored by lipids with positive saddle splay modulus κG, which drives the formation of membranes with negative Gaussian curvature C1C2 (with a saddle-like appearance). Typically these phases are based on the Ia3d, Pn3m, or Im3m space groups, with the lipids arranged on a continuous periodic minimal surface separating two independent continuous water networks. The term “bicontinuous” reflects the fact that both lipid and water domains are continuous in the three spatial dimensions. These systems can be doped with cationic lipids [57], making them amenable for complexation with NAs. However, when complexed with DNA, the cubic phase is destabilized and the system forms an inverted hexagonal phase instead [52]. This is likely because in order to provide a good overlap between the cationic lipid membrane and the DNA, the DNA would need to be substantially bent to follow the topology of the 3D water channels, creating an elastic energy penalty. This is reinforced by the study of complexation between short DNA fragments and cationic bicontinuous cubic phases of the gyroid type (space group Ia3d) [51]. Short DNA strands with 5 base pairs with blunt ends engage in end-to-end stacking destabilizing the cubic phase and forming HIIC complexes instead. However, if the DNA ends are composed by non-sticky overhangs, end-to-end stacking is diminished and a transition to a QIIC phase is observed. Overall these findings demonstrate that the large rigidity of DNA destabilizes the bicontinuous cubic structure, favoring inverted hexagonal phases instead, but if the NAs are short enough, the cubic phase can become stable again. Hence, the QIIC complex structure represents an example where besides the lipid elastic properties, the NA rigidity and length are crucial at determining the final structure of the complex. This type of insight about the physical properties of building blocks (lipids and NAs) and the crucial interactions between them has led to the development of siRNA-lipid bicontinuous cubic complexes [52], since the siRNA small length and no tendency for end-to-end stacking are amenable to fit the bicontinuous cubic unit cell dimensions [26].
siRNA has been shown to form complexes with an inverted bicontinuous cubic structure of the gyroid type (QII,GC) when mixed with DOTAP-monoolein systems. The bicontinuous cubic structure and inclusion of siRNA has been convincingly demonstrated by a combination of SAXS and dual-color fluorescence colocalization microscopy [52]. In addition, the cubic unit cell dimension is seen to grow from ~11 nm (before) to ~15 nm (after) mixing with siRNA, while keeping the same symmetry, showing a small adjustment of the native lipid curvature to accommodate siRNA within the two water channel networks. Importantly, this structure was found to have promising gene silencing activity while keeping non-specific gene knockout and toxicity relatively low.
8. Normal bicontinuous cubic complexes – QIC
Another type of bicontinuous cubic phase – the normal bicontinuous cubic (QI) – can be formed by surfactants or lipids with a spontaneous curvature that is slightly positive, typically between values that would favor the normal hexagonal or the lamellar phase [53][58][59]. In surfactant/lipid systems, the normal bicontinuous cubic usually belongs to the space group Ia3d. In this case the periodic minimal surface (gyroid) is the water layer, which separates two independent branched micelle networks composed by the surfactant/lipid.
Despite the difficulty of forming inverted bicontinuous cubic phases complexed with DNA (as discussed above), a normal cubic phase with Ia3d symmetry composed of DNA, dodecyltrimethylammonium (DTA+) and lecithin, has been convincingly described [60][61] by means of SAXS and phase diagram determinations.
9. Normal columnar complexes – HIC and SIC
Besides the normal bicontinuous cubic phase, DNA has been reported to form also normal hexagonal (HIC) and square (SIC) phases along with related phases of lower symmetry.
HIC phases were convincingly demonstrated by SAXS, cryo-TEM and phase behavior considerations for mixtures of DOPC with double-tailed lipids (dioleoyl chains) bearing highly charged dendritic headgroups (nominal charge of +8 and +16) [27][62]. The high charge and size of the dendritic headgroups increases the spontaneous curvature of the lipid films into positive values favoring the formation of highly-charged rod-like micelles. These micelles are arranged in a hexagonal lattice with DNA rods located in their interstices in a honeycomb-like lattice [27]. As the dendritic lipid fraction over DOPC increases, the rod-like micelles are thought to become shorter and more asymmetric due to the higher preponderance of the dendritic lipid high positive curvature. This is thought to be the cause of the deformation of the hexagonal lattice into a structure of lower symmetry as the dendritic lipid fraction increases [62]. Ongoing with the distorted columnar phases, in the presence of brine and cell-culture media a highly packed neat DNA columnar phase is also observed, forming probably due to depletion forces arising from the dendritic lipid micelles. Importantly, these normal hexagonal phases assembled from dendritic lipids were shown to have an in vitro transfection efficiency as high as the optimized lamellar and inverted hexagonal phases, with the advantage that the high efficiency is less dependent on the lipid charge density and is particularly high in cell cultures that are traditionally difficult to transfect, such as mouse embryonic fibroblasts [27]. The insensitivity of the efficiency on the membrane charge density hints at a different mechanism of transfection, possibility related with the continuous DNA substructure, which likely facilitates the release of DNA once a part of it becomes exposed to the cytosol [27][62].
Normal hexagonal phases with some degree of distortion are also though to form in single-tailed cationic surfactant-DNA systems [45][63][64][65][66][67][68][69]. Small-angle X-ray and neutron scattering show structures consistent with hexagonal symmetry. While frequently the scattering patterns do not allow a clear distinction between normal and inverted hexagonal phases (see e.g. Ref. [69]), the relative position of these phases on the phase diagrams of surfactant-DNA complex salts (see e.g. Ref. [61]) make them more consistent with normal than inverse structures.
10. Other normal phases
Single-tailed cationic surfactants (e.g. CTAB and DTAB) tend to form spherical micelles in the absence of DNA. The formation of columnar phases with DNA (as discussed above) in which the surfactant is likely arranged in a rod-like micelle results from the high-rigidity of DNA. If long single-stranded DNA is used instead, the hexagonal phase can be replaced by a normal discrete cubic phase (IIC), presumably because now the much higher flexibility of ssDNA is able to accommodate the natural curvature of the cationic surfactant [69]. This and related structures are also frequently observed in other flexible polyelectrolyte-surfactant systems [70][71][72]. Alternatively, as already pointed out above, if the ssDNA is very short, it can induce the formation of lamellar phases instead, in a base-sequence specific manner, likely due to hydrophobic interactions between the bases and the surfactant [45].
Some cationic gemini surfactants forming normal micelles were found to complex with siRNA and seem to form a type of condensed micellar phase with siRNA sandwiched between gemini surfactant micelles [73]. In this study, the assembly time was restricted to 15min after mixing between siRNA and the surfactant since it is the time needed for complex formation and administration into biological systems. It is possible that at longer times (hours or days) these complexes could arrange into ordered phases like the ones discussed above (e.g. normal cubic or normal hexagonal). In a related gemini surfactant family, composed of bis-imidazolium gemini surfactants, although a condensed micellar phase in the presence of siRNA was also the most prevalent structure, in some cases, depending on the spacer length and charge ratio rchg, additional Bragg peaks in the scattering patterns were consistent with more ordered structures, possibly hexagonal and cubic [74].
References
- Felgner, P.L.; Gadek, T.R.; Holm, M.; Roman, R.; Chan, H.W.; Wenz, M.; Northrop, J.P.; Ringold, G.M.; Danielsen, M. Lipofection: A Highly efficient, lipid-mediated DNA-transfection procedure. Proc. Natl. Acad.Sci. USA 1987, 84, 7413–7417.
- Felgner, P.L.; Rhodes, G. Gene therapeutics. Nature 1991, 349, 351–352.
- Ewert, K.K.; Ahmad, A.; Evans, H.M.; Safinya, C.R. Cationic lipid-DNA complexes for non-viral gene therapy: Relating supramolecular structures to cellular pathways. Expert Opin. Biol. Ther. 2005, 5, 33–53.
- Rädler, J.O.; Koltover, I.; Salditt, T.; Safinya, C.R. Structure of DNA-cationic liposome complexes: DNA intercalation in multilamellar membranes in distinct interhelical packing regimes. Science 1997, 275, 810–814.
- Harries, D.; May, S.; Gelbart, W.M.; Ben-Shaul, A. Structure, Stability, and Thermodynamics of Lamellar DNA-Lipid Complexes. Biophys. J. 1998, 75, 159–173.
- Harries, D.; May, S.; Ben-shaul, A. Counterion release in membrane–biopolymer interactions. Soft Matter 2013, 9, 9268–9284.
- Barreleiro, P.C.A.; Olofsson, G.; Alexandridis, P. Interaction of DNA with cationic vesicles: A calorimetric study. J. Phys. Chem. B 2000, 104, 7795–7802.
- Wagner, K.; Harries, D.; May, S.; Kahl, V.; Rädler, J.O.; Ben-Shaul, A. Direct evidence for counterion release upon cationic lipid-DNA condensation. Langmuir 2000, 16, 303–306.
- Liang, H.; Harries, D.; Wong, G.C.L. Polymorphism of DNA-anionic liposome complexes reveals hierarchy of ion-mediated interactions. Proc. Natl. Acad. Sci. USA 2005, 102, 11173–11178.
- Mcmanus, J.J.; Rädler, J.O.; Dawson, K.A. Does Calcium Turn a Zwitterionic Lipid Cationic? J. Phys. Chem. B 2003, 107, 9869–9875.
- McManus, J.J.; Rädler, J.O.; Dawson, K.A. Observation of a rectangular columnar phase in aDNA-calcium-zwitterionic lipid complex. J. Am. Chem. Soc. 2004, 126, 15966–15967.
- Khan, A.; Marques, E.F. Synergism and polymorphism in mixed surfactant systems. Curr. Opin. Colloid Interface Sci. 2000, 4, 4002–4410.
- Wong, G.C.L.; Pollack, L. Electrostatics of Strongly Charged Biological Polymers: Ion-Mediated Interactions and Self-Organization in Nucleic Acids and Proteins. Annu. Rev. Phys. Chem. 2010, 61, 171–189.
- DeHaseth, P.L.; Lohman, T.M.; Record, M.T. Nonspecific Interaction of lac Repressor with DNA:An Association Reaction Driven by Counterion Release. Biochemistry 1977, 16, 4783–4790.
- Ewert, K.K.; Zidovska, A.; Ahmad, A.; Bouxsein, N.F.; Evans, H.M.; McAllister, C.S.; Samuel, C.E.; Safinya, C.R. Cationic Lipid–Nucleic Acid Complexes for Gene Delivery and Silencing: Pathways and Mechanisms for Plasmid DNA and siRNA. Top. Curr. Chem. 2010, 296, 191–226.
- Majzoub, R.N.; Ewert, K.K.; Safinya, C.R. Cationic liposome—Nucleic acid nanoparticle assemblies with applications in gene delivery and gene silencing. Philos. Trans. R. Soc. A 2016, 374, 20150129.
- Kranz, L.M.; Diken, M.; Haas, H.; Kreiter, S.; Loquai, C.; Reuter, K.C.; Meng, M.; Fritz, D.; Vascotto, F.; Hefesha, H.; et al. Systemic RNA delivery to dendritic cells exploits antiviral defence for cancer immunotherapy. Nature 2016, 534, 396–401.
- Buck, J.; Grossen, P.; Cullis, P.R.; Huwyler, J.; Witzigmann, D. Lipid-Based DNA Therapeutics: Hallmarks of Non-Viral Gene Delivery. ACS Nano 2019, 13, 3754–3782.
- Majzoub, R.N.; Chan, C.-L.; Ewert, K.K.; Silva, B.F.B.; Liang, K.S.; Jacovetty, E.L.; Carragher, B.; Potter, C.S.;Safinya, C.R. Uptake and transfection efficiency of PEGylated cationic liposome-DNA complexes with and without RGD-tagging. Biomaterials 2014, 35, 4996–5005.
- Koltover, I.; Salditt, T.; Safinya, C.R. Phase diagram, stability, and overcharging of lamellar cationic lipid-DNA self-assembled complexes. Biophys. J. 1999, 77, 915–924.
- Jonsson, M.; Linse, P. Polyelectrolyte-macroion complexation. I. Effect of linear charge density, chain length, and macroion charge. J. Chem. Phys. 2001, 115, 3406–3418.
- Jonsson, M.; Linse, P. Polyelectrolyte-macroion complexation. II. Effect of chain flexibility. J. Chem. Phys. 2001, 115, 10975–10985.
- Stornes, M.; Linse, P.; Dias, R.S. Monte Carlo Simulations of Complexation between Weak Polyelectrolytes and a Charged Nanoparticle. Influence of Polyelectrolyte Chain Length and Concentration. Macromolecules 2017, 50, 5978–5988.
- Safinya, C.R.; Ewert, K.K.; Majzoub, R.N.; Leal, C. Cationic liposome–nucleic acid complexes for gene delivery and gene silencing. New J. Chem. 2014, 38, 5164–5172.
- Raviv, U.; Needleman, D.J.; Li, Y.; Miller, H.P.; Wilson, L.; Safinya, C.R. Cationic-liposome microtubule complexes: Pathways to the formation of two-state lipid-protein nanotubes with open or closed ends. Proc. Natl. Acad. Sci. USA 2005, 102, 11167–11172.
- Kang, M.; Kim, H.; Leal, C. Self-organization of nucleic acids in lipid constructs. Curr. Opin. ColloidInterface Sci. 2016, 26, 58–65.
- Ewert, K.K.; Evans, H.M.; Zidovska, A.; Bouxsein, N.F.; Ahmad, A.; Safinya, C.R. A columnar phase of dendritic lipid-based cationic liposome-DNA complexes for gene delivery: Hexagonally ordered cylindrical micelles embedded in a DNA honeycomb lattice. J. Am. Chem. Soc. 2006, 128, 3998–4006.
- Koltover, I.; Salditt, T.; Rädler, J.O.; Safinya, C.R. An Inverted Hexagonal Phase of Cationic Liposome-DNA Complexes Related to DNA Release and Delivery. Science 1998, 281, 78–81.
- Lasic, D.D.; Strey, H.; Stuart, M.C.A.; Podgornik, R.; Frederik, P.M. The Structure of DNA-LiposomeComplexes. J. Am. Chem. Soc. 1997, 119, 832–833.
- Battersby, B.J.; Grimm, R.; Huebner, S.; Cevc, G. Evidence for three-dimensional interlayer correlations in cationic lipid-DNA complexes as observed by cryo-electron microscopy. Biochim. Biophys. Acta 1998, 1372, 379–383.
- Schmutz, M.; Durand, D.; Debin, A.; Palvadeau, Y.; Etienne, A.; Thierry, A.R. DNA packing in stable lipid complexes designed for gene transfer imitates DNA compaction in bacteriophage. Proc. Natl. Acad. Sci. USA 1999, 96, 12293–12298.
- Salditt, T.; Koltover, I.; Rädler, J.O.; Safinya, C.R. Two-dimensional smectic ordering of linear DNA chains inself-assembled dna-cationic liposome mixtures. Phys. Rev. Lett. 1997, 79, 2582–2585.
- Gómez-Varela, A.I.; Gaspar, R.; Miranda, A.; Assis, J.L.; Valverde, R.R.H.F.; Einicker-Lamas, M.; Silva, B.F.B.; De Beule, P.A.A. Fluorescence Cross-Correlation Spectroscopy as a valuable tool to characterize cationic liposome-DNA nanoparticle assembly. J. Biophotonics 2020, e202000200.
- Koltover, I.;Wagner, K.; Safinya, C.R. DNA Condensation in Two-Dimensions. Proc. Natl. Acad. Sci. USA 2000, 97, 14046–14052.
- Desigaux, L.; Sainlos, M.; Lambert, O.; Chevre, R.; Letrou-Bonneval, E.; Vigneron, J.P.; Lehn, P.; Lehn, J.-M.; Pitard, B. Self-assembled lamellar complexes of siRNA with lipidic aminoglycoside derivatives promote efficient siRNA delivery and interference. Proc. Natl. Acad. Sci. USA 2007, 104, 16534–16539.
- Bouxsein, N.F.; McAllister, C.S.; Ewert, K.K.; Samuel, C.E.; Safinya, C.R. Structure and gene silencing activities of monovalent and pentavalent cationic lipid vectors complexed with siRNA. Biochemistry 2007, 46, 4785–4792.
- Kulkarni, J.A.; Witzigmann, D.; Leung, J.; Tam, Y.Y.C.; Cullis, P.R. On the role of helper lipids in lipid nanoparticle formulations of siRNA. Nanoscale 2019, 11, 21733–21739.
- Kormann, M.S.D.; Hasenpusch, G.; Aneja, M.K.; Nica, G.; Flemmer, A.W.; Herber-jonat, S.; Huppmann, M.; Mays, L.E.; Illenyi, M.; Schams, A.; et al. Expression of therapeutic proteins after delivery of chemically modified mRNA in mice. Nat. Biotechnol. 2011, 29, 154–157.
- Sahin, U.; Türeci, Ö. Personalized vaccines for cancer immunotherapy. Science 2018, 359, 1355–1360.
- Thierry, A.R.; Norris, V.; Molina, F.; Schmutz, M. Lipoplex nanostructures reveal a general self-organization of nucleic acids. Biochim. Biophys. Acta Gen. Subj. 2009, 1790, 385–394.
- Ziller, A.; Nogueira, S.S.; Hühn, E.; Funari, S.S.; Brezesinski, G.; Hartmann, H.; Sahin, U.; Haas, H.;Langguth, P. Incorporation of mRNA in Lamellar Lipid Matrices for Parenteral Administration. Mol. Pharm. 2018, 15, 642–651.
- Arteta, M.Y.; Kjellman, T.; Bartesaghi, S.; Wallin, S.; Wu, X.; Kvist, A.J.; Dabkowska, A.; Székely, N.; Radulescu, A.; Bergenholtz, J.; et al. Successful reprogramming of cellular protein production through mRNA delivered by functionalized lipid nanoparticles. Proc. Natl. Acad. Sci. USA 2018, 115, E3351–E3360.
- Patel, S.; Ashwanikumar, N.; Robinson, E.; Xia, Y.; Mihai, C.; Grith, J.P.; Hou, S.; Esposito, A.A.; Ketova, T.; Welsher, K.; et al. Naturally-occurring cholesterol analogues in lipid nanoparticles induce polymorphic shape and enhance intracellular delivery of mRNA. Nat. Commun. 2020, 11, 983.
- Rosa, M.; Dias, R.; da Graça Miguel, M.; Lindman, B. DNA—Cationic surfactant interactions are different for double- and single-stranded DNA. Biomacromolecules 2005, 6, 2164–2171.
- Liu, X.; Abbott, N.L. Characterization of the nanostructure of complexes formed by single- or double-stranded oligonucleotides with a cationic surfactant. J. Phys. Chem. B 2010, 114, 15554–15564.
- Neumann, T.; Gajria, S.; Bouxsein, N.F.; Jaeger, L.; Tirrell, M. Structural responses of DNA-DDAB films to varying hydration and temperature. J. Am. Chem. Soc. 2010, 132, 7025–7037.
- Cuomo, F.; Mosca, M.; Murgia, S.; Avino, P.; Ceglie, A.; Lopez, F. Evidence for the role of hydrophobic forces on the interactions of nucleotide-monophosphates with cationic liposomes. J. Colloid Interface Sci. 2013, 410, 146–151.
- Lin, A.; Slack, N.; Ahmad, A.; George, C.; Samuel, C.; Safinya, C.R. Three-dimensional Imaging of Lipid Gene-Carriers: Membrane Charge Density Controls Universal Transfection Behavior in Lamellar Cationic Liposome-DNA Complexes. Biophys. J. 2003, 84, 3307–3316.
- Semple, S.C.; Akinc, A.; Chen, J.; Sandhu, A.P.; Mui, B.L.; Cho, C.K.; Sah, D.W.Y.; Stebbing, D.; Crosley, E.J.; Yaworski, E.; et al. Rational design of cationic lipids for siRNA delivery. Nat. Biotechnol. 2010, 28, 172–176.
- Gelbart,W.M.; Bruinsma, R.F.; Pincus, P.A.; Parsegian, V.A. DNA-Inspired Electrostatics. Phys. Today 2000, 53, 38–44.
- Leal, C.; Ewert, K.K.; Bouxsein, N.F.; Shirazi, R.S.; Li, Y.; Safinya, C.R. Stacking of short DNA induces the gyroid cubic-to-inverted hexagonal phase transition in lipid-DNA complexes. Soft Matter 2013, 9, 795–804.
- Leal, C.; Bouxsein, N.F.; Ewert, K.K.; Safinya, C.R. Highly Efficient Gene Silencing Activity of siRNA Embedded in a Nanostructured Gyroid Cubic Lipid Matrix. J. Am. Chem. Soc. 2010, 132, 16841–16847.
- Marques, E.F.; Silva, B.F.B. Surfactant Self-Assembly. Encycl. Colloid Interface Sci. 2013, 1202–1241.
- Seddon, J.M.; Templer, R.H. Cubic phases of self-assembled amphiphilic aggregates. Philos. Trans. R. Soc. A Math. Phys. Eng. Sci. 1993, 344, 377–401.
- Larsson, K. Aqueous dispersions of cubic lipid-water phases. Curr. Opin. Colloid Interface Sci. 2000, 5, 64–69.
- Chang, D.P.; Barauskas, J.; Dabkowska, A.P.; Wadsäter, M.; Tiberg, F.; Nylander, T. Non-lamellar lipid liquid crystalline structures at interfaces. Adv. Colloid Interface Sci. 2015, 222, 135–147.
- Kim, H.; Song, Z.; Leal, C. Super-swelled lyotropic single crystals. Proc. Natl. Acad. Sci. USA 2017, 114, 10834–10839.
- Leal, C.; Ewert, K.K.; Shirazi, R.S.; Bouxsein, N.F.; Safinya, C.R. Nanogyroids Incorporating Multivalent Lipids: Enhanced Membrane Charge Density and Pore Forming Ability for Gene Silencing. Langmuir 2011, 27, 7691–7697.
- Hyde, S.T. Microstructure of Bicontinuous Surfactant Aggregates. J. Phys. Chem. 1989, 93, 1458–1464.
- Seddon, J.M.; Templer, R.H. Polymorphism of Lipid-Water Systems. In Handbook of Biological Physics; Lipowsky, R., Sackmann, E., Eds.; Elsevier Science B. V.: Amsterdam, The Netherlands, 1995; pp. 97–160. ISBN 978-0-444-81975-8.
- Bilalov, A.; Olsson, U.; Lindman, B. A cubic DNA-lipid complex. Soft Matter 2009, 5, 3827–3830.
- Bilalov, A.; Olsson, U.; Lindman, B. DNA–lipid self-assembly: Phase behavior and phase structures of a DNA–surfactant complex mixed with lecithin and water. Soft Matter 2011, 7, 730–742.
- Zidovska, A.; Evans, H.M.; Ewert, K.K.; Quispe, J.; Carragher, B.; Potter, C.S.; Safinya, C.R. Liquid Crystalline Phases of Dendritic Lipid–DNA Self-Assemblies: Lamellar, Hexagonal and DNA Bundles. J. Phys. Chem. B 2009, 113, 3694–3703.
- Ghirlando, R.; Wachtel, E.J.; Arad, T.; Minsky, A. DNA Packaging Induced by Micellar Aggregates: A Novel in Vitro DNA Condensation System. Biochemistry 1992, 31, 7110–7119.
- Mel’nikov, S.M.; Sergeyev, V.G.; Yoshikawa, K.; Takahashi, H.; Hatta, I. Cooperativity or phase transition? Unfolding transition of DNA cationic surfactant complex. J. Chem. Phys. 1997, 107, 6917-6924.
- Krishnaswamy, R.; Raghunathan, V.A.; Sood, A.K. Reentrant phase transitions of DNA-surfactant complexes. Phys. Rev. E 2004, 69, 031905.
- Krishnaswamy, R.; Pabst, G.; Rappolt, M.; Raghunathan, V.A.; Sood, A.K. Structure of DNA-CTAB-hexanol complexes. Phys. Rev. E 2006, 73, 031904.
- Leal, C.; Moniri, E.; Pegado, K.; Wennerström, H. Electrostatic Attraction between DNA and a Cationic Surfactant Aggregate. The Screening Effect of Salt. J. Phys. Chem. B 2007, 111, 5999–6005.
- Zhou, S.; Liang, D.; Burger, C.; Yeh, F.; Chu, B. Nanostructures of complexes formed by calf thymus DNA interacting with cationic surfactants. Biomacromolecules 2004, 5, 1256–1261.
- Hubˇcík, L.; Galliková, D.; Pullmannová1, P.; Lacinová, L.; Sulová, Z.; Hanulová, M.; Funari, S.S.; Devínsky, F.; Uhríková, D. DNA–DOPE–gemini surfactants complexes at low surface charge density: From structure to transfection efficiency. Gen. Physiol. Biophys. 2018, 37, 57–69.
- Subramanian, G.; Hjelm, R.P.; Deming, T.J.; Smith, G.S.; Li, Y.; Safinya, C.R. Structure of complexes of cationic lipids and poly(glutamic acid) polypeptides: A pinched lamellar phase. J. Am. Chem. Soc. 2000, 122, 26–34.
- Piculell, L.; Norrman, J.; Svensson, A.V.; Lynch, I.; Bernardes, J.S.; Loh,W. Ionic surfactants with polymeric counterions. Adv. Colloid Interface Sci. 2009, 147–148, 228–236.
- Chiappisi, L.; Hoffmann, I.; Gradzielski, M. Complexes of oppositely charged polyelectrolytes andsurfactants — Recent developments in the field of biologically derived polyelectrolytes. Soft Matter 2013, 9, 3896–3909.
- Falsini, S.; Ristori, S.; Ciani, L.; Di Cola, E.; Supuran, C.T.; Arcangeli, A.; In, M. Time resolved SAXS to study the complexation of siRNA with cationic micelles of divalent surfactants. Soft Matter 2014, 10, 2226–2233.