| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Hatim Boughanem | + 2462 word(s) | 2462 | 2021-01-27 05:07:52 | | | |
| 2 | Bruce Ren | -20 word(s) | 2442 | 2021-01-28 02:46:33 | | |
Video Upload Options
Evidence from observational and in vitro studies suggests that insulin growth-factor-binding pro-tein type 2 (IGFBP2) is a promising protein in non-communicable diseases, such as obesity, insulin resistance, metabolic syndrome, or type 2 diabetes. Accordingly, great efforts have been carried out to explore the role of IGFBP2 in obesity state and insulin-related diseases, which it is typically found decreased.
1. Introduction
Insulin-like growth factors (IGFs) are growth peptides, that are implicated in mammalian development, growth and cell proliferation and differentiation [1]. IGFs are usually bound to IGF-binding proteins (IGFBPs), and mediate their actions by regulating their bioavailability [2]. There are six well characterized high-affinity IGFBP members, designated IGFBP1 through 6. The IGFBP family members share similar structure and molecular organization, which is suggestive of similar mechanism of action, whereas they have different modes of regulation and distinct expression patterns [3]. In this wise, IGFBP2 is a key member that participates in different physiological and metabolic processes.
The physiological role of IGFBP2 on metabolic disorders are controversial and less defined, although there is growing evidence for a solid association [4]. Therefore, understanding metabolic regulation of IGFBP2 and its influence on metabolic diseases could provide new insights that can be applied as therapeutic targets.
2. Building the Molecular Structure of IGFBP2
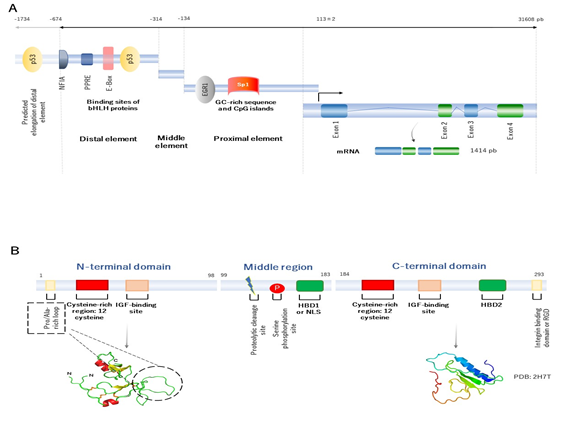
The human IGFBP2 gene is located in chromosome 2q35 and has four exons and three introns [5], [6]. The combination of exons and introns provides up to five splice variants. However, only the first transcript variant encodes for the functional isoform, that it is usually found to be involved in its main physiological role [7]. The IGFBP2 gene expression is modulated by the interaction of a number of transcription factors at the regulatory elements in the promoter region. The IGFBP2 promoter is located between the position -674 upstream up to the transcription start site (TSS), localized into 113±2 [5], [8]. In 2006, Sato and collaborators, by studying dedifferentiated lung cancer cells, suggested that the structure of the human IGFBP2 promoter can be organized into three elements, represented as the distal (localized between -674 and -314, which apparently acts as an enhancer element), middle (extends from -314 to -134, and seems to be an inhibitor/silencer), and proximal regulatory element (extends from -134 to the transcription start site, and apparently acts as a regulator of transcriptional activity). It is worth mentioning that the proximal regulatory element lacks TATA or CAAT boxes and contains high-GC sequences, meaning that this element may participate in epigenetic regulation (Figure 1A) [8].
Figure 1. Human representative of IGFBP2 genetic and molecular structure. (A) Human representative of genetic structure of IGFBP2. There are three regulatory elements, ranging from distal, middle to proximal elements. The structure gene contains four exons and three introns. The product is a mature mRNA of 1414 pb. (B) Human representative structure of IGFBP2 protein domain for the mature protein of 289 amino acids residues. There are three principal domains. The N-terminal domain, the middle domain or the middle region and the C-terminal domain. The N- (extracted from Galea C. et al. [9]) and C-terminal (PDB: 2H7T for the C-domain of IGFBP2) domain are presented. Both N- and C-terminal domains contain IGF binding sites, which suggests that these domains are highly conserved. However, the middle region has a proteolytic cleavage and phosphorylation sites, which indicates that this domain may be implicated in the regulation of bioavailability of IGF1 and stability of the protein structure.
In human, a wide number of transcription factors can bind to the IGFBP2 promoter and activate its transcription activity in vitro. In 2006, Sato et al. and Grimberg et al. found by analyzing lung cancer cells, that the IGFBP2 promoter has putative binding sites for early growth response protein 1 (EGR1) and transformation-related protein 53 (p53), that are crucial for its transactivation [8], [10]. In addition, in small-cell lung cancer cells, Yazawa et al. (2009) demonstrated that neurogenic differentiation (NEUROD)—a helix-loop-helix (HLH) transcription factor—binds to the enhancer box (E-box) sequence, located at the distal element and regulates its activation [11]. Further, Mireuta and colleagues (2010) argued that the proximal regulatory element is a potential target for specificity protein 1 (Sp1) [12]. In 2017, Lee and collaborators showed in MCF-7 epithelial breast tumor cells, that nuclear factor IA (NFIA) binds to the E-box sequence, to potentially enhance IGFBP2 transcription as well [13]. Recently, our group, by studying nuclear extracts from visceral adipose tissue, demonstrated that peroxisome-proliferator activated receptor γ2 (PPARγ2) (through PPARγ2-retinoid X receptor α (RXRα) heterodimer) physically interacts with the IGFBP2 promoter through PPAR responsive element (PPRE) domain [14].
Accordingly, comparative studies in animal models reveal a large number of metabolic transcription factors that can bind to the igfbp2 promoter, such as multiple endocrine neoplasia type 1 (MEN1), Sp1, PPARα, CCAAT-enhancer-binding protein α (C/EBPα), or hypoxia-inducible factor 1 (HIF1) and nuclear factor kappa B (NFκB) [15]. Therefore, these transcription factors could predict the physiological role of IGFBP2, at least at transcriptional and transductional levels. The fact that the IGFBP2 has PPAR, NEUROD, and NFκB binding sites places it as an important mediator of metabolic and inflammatory processes. Conversely, the notion of the interaction with transcription factors, such as p53, sp1, EGR1, and MEN1, provides an idea of its role in cell cycle, cell proliferation, and growth.
Overall, the IGFBP2 gene encodes for a final mature IGFBP2 protein, that contains 289 amino acid residues of 31.4 kDa of molecular weight. Its structure has three principal domains: the N-terminal, middle region, and the C-terminal domain. The N- and C-terminal domains are conserved regions between the IGFBP family members, whereas the middle region shows high variability. This region, therefore, contributes for a singular identity and specific functions (Figure 1B).
The N-terminal domain of IGFBP2 protein, encoded by the exon 1, contains 98 amino acid residues and shares approximately 58% of sequence between the IGFBP family members. This domain is characterized by containing a highly conserved cysteine-rich region, which allows to form up to six disulphide bonds. The N-domain contains an additional 16-residue Pro/Ala-rich, situated between Pro21 and Arg36 residues, that is not found in the other members. This motif forms a solvent-exposed loop on the three-dimensional structure of the protein, and may comprise a potential -SH3 domain interaction site for binding to other proteins. Within this domain, another local motif (GCGCCxxC) is found, which may be crucial in the interaction of IGFBP2 with IGFs. The N-terminal domain has an IGFs-binding domain, that is involved in high-affinity binding to IGFs.
The middle region (called L-domain) is encoded by the exon 2 and links the N- to the C-terminal domains. This domain extends approximately for 85 amino acids residues, and its sequence appears to be unique and singular among the IGFBP members. It is susceptible to proteolytic cleavage and inactivation of IGFBP2 protein, since specific proteases separate the N- and C-domains, and decrease the affinity of IGFBP2 by IGFs. The middle region has various predicted phosphorylated sites [16], and it is often phosphorylated in serine at the position 106 [17]. The purpose of phosphorylation is still unknown, but may contribute to the resistance to proteolysis. Within the L-domain, there is another subdomain called heparin-binding domain 1 (HBD1), located between 179 and 184. The principal function of the HBD1 domain is to bind to extracellular matrix components (as glycosaminoglycans, integrins, etc.,). The HBD1 domain can act as a ligand, by binding to the tyrosine phosphatase β (RPTPβ) receptor [18], and mimic the nuclear localization signal (NLS) to translocate into the nucleus as well [19]. Therefore, this motif is required for specific recognition and interaction binding.
Finally, the C-terminal region is encoded by the exon 3 and 4 and contains 107 amino acid residues. Interestingly, the sequence of this domain shares high similarity with the thyroglobulin-type-I domain [20]. This domain is conserved between IGFBP members and characterized by highly conserved six cysteine residues [21]. The main function of C-domain is involved in high-affinity binding to IGFs, which contains a specific region to bind to IGFs. Within this domain, an RGD (Arg-Gly-Asp) sequence, located on 265 and 267 residues, can bind to integrins [22], and activate many physiological and cell cycle processes, such as cell invasion and proliferation, by inducing β-catenin and further affecting Wnt signaling [23]. The C-terminal domain contains an HBD motif as well, called HBD2. The interaction of HBD2 to proteoglycans leads to regulate many of cell cycle processes as cell adhesion, proliferation, and migration [24].
Classically, the molecular structure of IGFBP2 is designed to bind to IGFs, but also can act as a ligand, through different domains located within the protein. This fact places IGFBP2 as a versatile molecule that could participate in a wide range of metabolic and cell cycle processes. Nonetheless, despite the aforementioned findings, many of the molecular pathways of IGFBP2 remain unknown. Investigating the molecular pathways of IGFBP2 could help to better understand the role that IGFBP2 plays in metabolic diseases.
3. Mechanism of Action of IGFBP2 and its Physiological Role
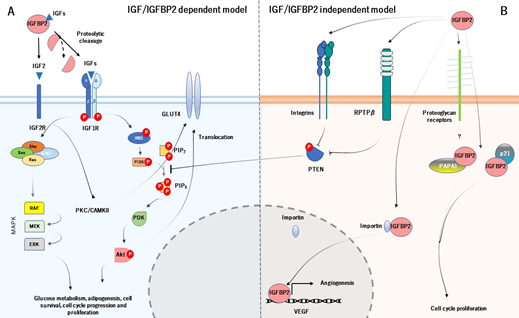
IGFBP2 acts systematically following two main action models, known as IGF-dependent model (IGFBP2 binds and regulates the bioavailability of IGFs) and IGF-independent model (IGFBP2 acts as a ligand by binding to receptors at the cell surface and to extracellular matrix) (Figure 2) [25]. Likewise, the principal function of IGFBP2 starts with its binding to IGFs (they form a binary complex) and modulates their actions at systemic levels. The colocalization of IGF type 1 or 2 with IGFBP2 has frequently been questioned, suggesting a possible leaning for each one. Interestingly, IGFBP2 has more pronounced affinity for IGF2, up to 10-20-fold greater than IGF1, which may be its primary ligand [26]. The binding of IGFBP2 to IGF1 seems to locally modulate IGF1 signaling, under various conditions [27]. However, as discussed below, the biological meaning of the IGF2/IGFBP2 complex is to increase its affinity by the extracellular matrix [28].
Figure 2. The cellular action of IGFBP2 is mediated by two principal models. The IGF-dependent model (A) modulates the activity and bioavailability of IGFs, which prevents the binding of IGFs to their receptor. The IGF-independent model (B) constitutes the physical interaction between IGFBP2 and specific receptors at the cell surface and extracellular matrix, which acts as a ligand. The main function of both models is regulating cell cycle, insulin, and glucose metabolism and angiogenesis.
The IGF-dependent model is responsible for the regulation of the phosphatidylinositol 3-kinase (PI3K)/ alpha serine/threonine-protein kinase (Akt) signalling pathway. The activation of PI3K/Akt pathway starts with the specific proteolysis and release of IGFBP2 from IGF1 or IGF2 and the binding of IGFs to IGF1 receptor (IGF1R) [29]. The complex IGFs/IGF1R leads to conformational changes in the IGF1R, which auto-phosphorylates their subunits β. The auto-phosphorylation of the IGF1R causes the activation of PI3K, which further induces the phosphorylation and activation of Akt. The active Akt has a wide number of target genes related with cell cycle and glucose metabolism, such as mechanistic target of rapamycin complex 1 (mTORC1), BCL2 associated agonist of cell death (BAD) and translocation of glucose transporter 4 (GLUT4), and also inhibits a large number of genes, as tuberous sclerosis 1 (TSC1) or glycogen synthase kinase 3 β (GSK3β) [30]. On the other hand, the phosphorylated IGF1R can also induce the recruitment of SHC adaptor protein (Shc), growth factor receptor-bound protein 2 (Grb2), Son of Sevenless (SOS), and rat sarcoma (Ras), which leads to stimulate the mitogen activated protein kinase (MAPK) signalling pathway. The MAPK pathway is responsible for growth processes, such as development, cell proliferation, differentiation, and migration, as well as inhibition of apoptosis [31] (Figure 2A). As for the IGF2R, it has been thought that it does not have apparent intracellular signalling activity and it is believed to act as a scavenger receptor for IGF2 [32]. However, a study carried out by Chu et al. (2008), showed that IGF2R in H9c2 cardiomyoblast cell cultures, activates the protein kinase C (PKC)/calcium/calmodulin-dependent protein kinase II (CaMKII) signalling pathway, which is dependent on the IGF2R [33]. This may result in GLUT4 translocation to the plasma membrane, characterizing an insulin-independent pathway [34].
As for the IGF-independent model, IGFBP2 acts as a potential ligand that binds to different targets. In particular, IGFBP2 binds to integrins through RGD domain, and stimulates cell migration and proliferation, adhesion, as well as cell differentiation [35]. The activation of IGFBP2/integrin complex increases the phosphorylation and inactivation of phosphatase and tensin homolog (PTEN), which leads to the stimulation of Akt signaling [36] (it is worth noting that despite the IGFBP2/integrin inactivates PTEN, there are also other mechanisms that regulate PTEN in this context). Moreover, IGFBP2 binds to extracellular matrix, which this interaction is mediated by HBD. Its biological function is to modulate the bioavailability of IGFs in the extracellular space and also acts as an active and local reservoir of IGFs to regulate cell adhesion, migration, proliferation, and differentiation [37]. The HBD domain is used to bind to RPTPβ receptor, being a receptor with tyrosine phosphatase activity. The IGFBP2/RPTPβ leads to its dimerization and further inactivates its phosphatase function. The inactivation of RPTPβ, cooperatively with IGF1 triggers to increase the phosphorylation of PTEN and further stimulate Akt [38]. In addition, the binding of IGF2 to IGFBP2 greatly increases the affinity for O-sulfated glycosaminoglycans, heparin, and heparan sulfate. Accordingly, Lund et al. (2014) demonstrated in vitro that the HBD1 and HBD2 contribute differentially to glycosaminoglycans binding and increase the affinity of both free IGFBP2 and the IGF2/IGFBP2 protein complex for heparin.
Interestingly, IGFBP2 can be located within the cells as well. A study pointed out that IGFBP2 intracellularly interacts with cyclin-dependent kinase inhibitor 1 (p21), to modulate cell proliferation in vivo in mouse lung epithelial cell line [39]. IGFBP2 also interacts with Pim-1-associated protein-1 (PAPA1) in the human prostate cancer cell line (LNCaP). The binding of IGFBP2 to PAPA1 seems to have a role in growth-promoting effect. The inhibition of PAPA1 enhances cell growth, suggesting that the proliferative effect of IGFBP2 may be regulated by its intracellular interaction with PAPA1 [40]. But IGFBP2 may act at intranuclear levels. A study demonstrated that IGFBP2 can be located into the nucleus in several common cancer cells. This is possible because of the HBD1 motif (HBD1 can mimic the NLS domain). Within the nucleus, IGFBP2 can act as a transcription modulator, which contributes to the activation of the expression of vascular endothelial growth factor (VEGF), an essential gene for angiogenesis (Figure 2B).
As already noted, IGFBP2 modulates the IGF signaling, binds to the extracellular matrix, and interacts with the cell surface receptor to modulate the PI3K/Akt signaling pathway. Given that IGFBP2 regulates the bioavailability of IGFs, it is worth noting that IGFBP2 is a key factor in cell proliferation and growth. The IGFs/IGFBP2 is closely related with glucose/insulin metabolism (through PTEN/Akt signaling) and cell cycle (trough MAPK pathway) as well, which could explain the dual role of IGFBP2 to act in such processes. Thus, the metabolic role of IGFBP2 could place it as a potential regulator in metabolic disorders, either by IGFs/IGFBP2-dependent and independent model.
References
- Rosenfeld, R.G. Insulin-like Growth Factors and the Basis of Growth. New Engl. J. Med. 2003, 349, 2184–2186, doi:10.1056/nejmp038156.
- Riedemann, J.; Macaulay, V.M. IGF1R signalling and its inhibition. Endocr. Relat. Cancer 2006, 13, S33–S43, doi:10.1677/erc.1.01280.
- Allard, J.B.; Duan, C. IGF-Binding Proteins: Why Do They Exist and Why Are There So Many? Front. Endocrinol. 2018, 9, 117, doi:10.3389/fendo.2018.00117.
- Heald, A.H.; Kaushal, K.; Siddals, K.W.; Rudenski, A.S.; Anderson, S.G.; Gibson, J.M. Insulin-like Growth Factor Binding Protein-2 (IGFBP-2) is a Marker for the Metabolic Syndrome. Exp. Clin. Endocrinol. Diabetes 2006, 114, 371–376, doi:10.1055/s-2006-924320.
- Binkert, C.; Margot, J.B.; Landwehr, J.; Heinrich, G.; Schwander, J. Structure of the human insulin-like growth factor binding protein-2 gene. Mol. Endocrinol. 1992, 6, 826–836, doi:10.1210/mend.6.5.1376411.
- Zapf, J.; Schmid, C.; Guler, H.P.; Waldvogel, M.; Hauri, C.; Futo, E.; Hossenlopp, P.; Binoux, M.; Froesch, E.R. Regulation of binding proteins for insulin-like growth factors (IGF) in humans. Increased expression of IGF binding protein 2 during IGF I treatment of healthy adults and in patients with extrapancreatic tumor hypoglycemia. J. Clin. Investig. 1990, 86, 952–961, doi:10.1172/jci114797.
- Cunningham, F.; Achuthan, P.; Akanni, W.; Allen, J.; Amode, M.R.; Armean, I.M.; Bennett, R.; Bhai, J.; Billis, K.; Boddu, S.; et al. Ensembl 2019. Nucleic Acids Res. 2019, 47, D745–D751, doi:10.1093/nar/gky1113.
- Sato, H.; Yazawa, T.; Suzuki, T.; Shimoyamada, H.; Okudela, K.; Ikeda, M.; Hamada, K.; Yamada-Okabe, H.; Yao, M.; Ku-bota, Y.; et al. Growth Regulation via Insulin-Like Growth Factor Binding Protein-4 and −2 in Association with Mutant K-ras in Lung Epithelia. Am. J. Pathol. 2006, 169, 1550–1566, doi:10.2353/ajpath.2006.051068.
- Galea, C.A.; Mobli, M.; McNeil, K.A.; Mulhern, T.D.; Wallace, J.C.; King, G.F.; Forbes, B.E.; Norton, R.S. Insulin-like growth factor binding protein-2: NMR analysis and structural characterization of the N-terminal domain. Biochimie 2012, 94, 608–616, doi:10.1016/j.biochi.2011.09.012.
- Grimberg, A.; Coleman, C.M.; Shi, Z.; Burns, T.F.; MacLachlan, T.K.; Wang, W.; El-Deiry, W.S. Insulin-like growth factor binding protein-2 is a novel mediator of p53 inhibition of insulin-like growth factor signaling. Cancer Biol. Ther. 2006, 5, 1408–1414, doi:10.4161/cbt.5.10.3455.
- Yazawa, T.; Sato, H.; Shimoyamada, H.; Okudela, K.; Woo, T.; Tajiri, M.; Ogura, T.; Ogawa, N.; Suzuki, T.; Mitsui, H.; et al. Neuroendocrine Cancer-Specific Up-Regulating Mechanism of Insulin-Like Growth Factor Binding Protein-2 in Small Cell Lung Cancer. Am. J. Pathol. 2009, 175, 976–987, doi:10.2353/ajpath.2009.081004.
- Mireuta, M.; Darnel, A.; Pollak, M. IGFBP-2 expression in MCF-7 cells is regulated by the PI3K/AKT/mTOR pathway through Sp1-induced increase in transcription. Growth Factors 2010, 28, 243–255, doi:10.3109/08977191003745472.
- Lee, C.-C.; Chen, P.-H.; Ho, K.-H.; Shih, C.-M.; Cheng, C.-H.; Lin, C.-W.; Cheng, K.-T.; Liu, A.-J.; Chen, K.-C. The mi-croRNA-302b-inhibited insulin-like growth factor-binding protein 2 signaling pathway induces glioma cell apoptosis by tar-geting nuclear factor IA. PLoS ONE 2017, 12, e0173890, doi:10.1371/journal.pone.0173890.
- Boughanem, H.; Cabrera-Mulero, A.; Millán-Gómez, M.; Garrido-Sánchez, L.; Cardona, F.; Tinahones, F.J.; Moreno-Santos, I.; Macías-González, M.; Mulero, C.-; Millán-Gómez, M.; et al. Transcriptional Analysis of FOXO1, C/EBP- and PPAR-2 Genes and Their association with Obesity-Related Insulin Resistance. Genes 2019, 10, 706, doi:10.3390/genes10090706.
- Badinga, L.; Song, S.; Simmen, R.C.M.; A Simmen, F. A Distal Regulatory Region of the Insulin-Like Growth Factor Binding Protein-2 (IGFBP-2) Gene Interacts with the Basic Helix-Loop-Helix Transcription Factor, AP-4. Endocrine 1998, 8, 281–290, doi:10.1385/endo:8:3:281.
- Firth, S.M.; Baxter, R.C. Cellular Actions of the Insulin-Like Growth Factor Binding Proteins. Endocr. Rev. 2002, 23, 824–854, doi:10.1210/er.2001-0033.
- Graham, M.E.; Kilby, D.M.; Firth, S.M.; Robinson, P.J.; Baxter, R.C. Thein VivoPhosphorylation and Glycosylation of Human Insulin-like Growth Factor-binding Protein-5. Mol. Cell. Proteom. 2007, 6, 1392–1405, doi:10.1074/mcp.m700027-mcp200.
- Shen, X.; Xi, G.; Maile, L.A.; Wai, C.; Rosen, C.J.; Clemmons, D.R. Insulin-Like Growth Factor (IGF) Binding Protein 2 Func-tions Coordinately with Receptor Protein Tyrosine Phosphatase and the IGF-I Receptor To Regulate IGF-I-Stimulated Signal-ing. Mol. Cell. Biol. 2012, 32, 4116–4130, doi:10.1128/MCB.01011-12.
- Azar, W.J.; Zivkovic, S.; A Werther, G.; Russo, V.C. IGFBP-2 nuclear translocation is mediated by a functional NLS sequence and is essential for its pro-tumorigenic actions in cancer cells. Oncogene 2014, 33, 578–588, doi:10.1038/onc.2012.630.
- Sala, A.; Capaldi, S.; Campagnoli, M.; Faggion, B.; Labò, S.; Perduca, M.; Romano, A.; Carrizo, M.E.; Valli, M.; Visai, L.; et al. Structure and Properties of the C-terminal Domain of Insulin-like Growth Factor-binding Protein-1 Isolated from Human Amniotic Fluid. J. Biol. Chem. 2005, 280, 29812–29819, doi:10.1074/jbc.m504304200.
- Mark, S.; Kübler, B.; Höning, S.; Oesterreicher, S.; John, H.; Braulke, T.; Forssmann, W.-G.; Ständker, L.; Kuebler, B. Diversity of Human Insulin-like Growth Factor (IGF) Binding Protein-2 Fragments in Plasma: Primary Structure, IGF-Binding Proper-ties, and Disulfide Bonding Pattern†. Biochemie 2005, 44, 3644–3652, doi:10.1021/bi0478401.
- Kuang, Z.; Yao, S.; Keizer, D.W.; Wang, C.C.; Leon, B.; Forbes, B.E.; Wallace, J.C.; Norton, R.S. Structure, Dynamics and Heparin Binding of the C-terminal Domain of Insulin-like Growth Factor-binding Protein-2 (IGFBP-2). J. Mol. Biol. 2006, 364, 690–704, doi:10.1016/j.jmb.2006.09.006.
- Patil, S.S.; Gokulnath, P.; Bashir, M.; Shwetha, S.D.; Jaiswal, J.; Shastry, A.H.; Arimappamagan, A.; Santosh, V.; Kondaiah, P. Insulin-like growth factor binding protein-2 regulates β-catenin signaling pathway in glioma cells and together contributes to poor patient prognosis. Neuro-Oncology 2016, 18, now053–1497, doi:10.1093/neuonc/now053.
- Russo, V.C.; SchüttB.S.; Andaloro, E.; Ymer, S.I.; Hoeflich, A.; Ranke, M.B.; Bach, L.A.; Werther, G.A. Insulin-Like Growth Factor Binding Protein-2 Binding to Extracellular Matrix Plays a Critical Role in Neuroblastoma Cell Proliferation, Migration, and Invasion. Endocrinology 2005, 146, 4445–4455, doi:10.1210/en.2005-0467.
- Wheatcroft, S.B.; Kearney, M.T. IGF-dependent and IGF-independent actions of IGF-binding protein-1 and -2: Implications for metabolic homeostasis. Trends Endocrinol. Metab. 2009, 20, 153–162, doi:10.1016/j.tem.2009.01.002.
- Rosenfeld, R.G.; Pham, H.; Conover, C.A.; Hintz, R.L.; Baxter, R.C. Structural and Immunological Comparison of Insu-lin-Like Growth Factor Binding Proteins of Cerebrospinal and Amniotic Fluids. J. Clin. Endocrinol. Metab. 1989, 68, 638–646, doi:10.1210/jcem-68-3-638.
- Alkharobi, H.; Alhodhodi, A.; Hawsawi, Y.; Alkafaji, H.; Devine, D.; El-Gendy, R.; Beattie, J. IGFBP-2 and -3 co-ordinately regulate IGF1 induced matrix mineralisation of differentiating human dental pulp cells. Stem Cell Res. 2016, 17, 517–522, doi:10.1016/j.scr.2016.09.026.
- Lund, J.; Søndergaard, M.T.; Conover, C.A.; Overgaard, M.T. Heparin-binding mechanism of the IGF2/IGF-binding protein 2 complex. J. Mol. Endocrinol. 2014, 52, 345–355, doi:10.1530/jme-13-0184.
- Kim, H.-S. Role of insulin-like growth factor binding protein-3 in glucose and lipid metabolism. Ann. Pediatr. Endocrinol. Metab. 2013, 18, 9–12, doi:10.6065/apem.2013.18.1.9.
- Pollak, M. The insulin and insulin-like growth factor receptor family in neoplasia: An update. Nat. Rev. Cancer 2012, 12, 159–169, doi:10.1038/nrc3215.
- Choi, Y.-S.; Cho, H.-Y.; Hoyt, K.R.; Naegele, J.R.; Obrietan, K. IGF-1 receptor-mediated ERK/MAPK signaling couples status epilepticus to progenitor cell proliferation in the subgranular layer of the dentate gyrus. Glia 2008, 56, 791–800, doi:10.1002/glia.20653.
- Ghosh, P.; Dahms, N.M.; Kornfeld, S. Mannose 6-phosphate receptors: New twists in the tale. Nat. Rev. Mol. Cell Biol. 2003, 4, 202–213, doi:10.1038/nrm1050.
- Chu, C.-H.; Tzang, B.-S.; Chen, L.-M.; Kuo, C.-H.; Cheng, Y.-C.; Chen, L.-Y.; Tsai, F.-J.; Tsai, C.-H.; Kuo, W.; Huang, C.-Y. IGF-II/mannose-6-phosphate receptor signaling induced cell hypertrophy and atrial natriuretic peptide/BNP expression via Gαq interaction and protein kinase C-α/CaMKII activation in H9c2 cardiomyoblast cells. J. Endocrinol. 2008, 197, 381–390, doi:10.1677/joe-07-0619.
- Kappel, V.D.; Zanatta, L.; Postal, B.G.; Silva, F.R.M.B. Rutin potentiates calcium uptake via voltage-dependent calcium channel associated with stimulation of glucose uptake in skeletal muscle. Arch. Biochem. Biophys. 2013, 532, 55–60, doi:10.1016/j.abb.2013.01.008.
- Ruoslahti, E. RGD and other recognition sequences for integrins. Annu. Rev. Cell Dev. Biol. 1996, 12, 697–715, doi:10.1146/annurev.cellbio.12.1.697.
- Perks, C.M.; Vernon, E.G.; Rosendahl, A.H.; Tonge, D.; Holly, J.M.P. IGF-II and IGFBP-2 differentially regulate PTEN in hu-man breast cancer cells. Oncogene 2007, 26, 5966–5972, doi:10.1038/sj.onc.1210397.
- Russo, V.C. Insulin-Like Growth Factor Binding Protein-2 Binds to Cell Surface Proteoglycans in the Rat Brain Olfactory Bulb. Endocrinology 1997, 138, 4858–4867, doi:10.1210/en.138.11.4858.
- Xi, G.; Wai, C.; DeMambro, V.; Rosen, C.J.; Clemmons, D.R. IGFBP-2 Directly Stimulates Osteoblast Differentiation. J. Bone Miner. Res. 2014, 29, 2427–2438, doi:10.1002/jbmr.2282.
- Terrien, X.; Bonvin, E.; Corroyer, S.; Tabary, O.; Clément, A.; Henrion-Caude, A. Intracellular colocalization and interaction of IGF-binding protein-2 with the cyclin-dependent kinase inhibitor p21CIP1/WAF1 during growth inhibition. Biochem. J. 2005, 392, 457–465, doi:10.1042/bj20050517.
- Miyako, K.; Cobb, L.J.; Francis, M.; Huang, A.; Peng, B.; Pintar, J.E.; Ariga, H.; Cohen, P. PAPA-1 Is a Nuclear Binding Part-ner of IGFBP-2 and Modulates Its Growth-Promoting Actions. Mol. Endocrinol. 2009, 23, 169–175, doi:10.1210/me.2008-0168.