| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Jason Huntley | + 5146 word(s) | 5146 | 2020-11-10 10:36:15 | | | |
| 2 | Rita Xu | -2218 word(s) | 2928 | 2021-01-15 08:47:33 | | |
Video Upload Options
Ticks can transmit a variety of infectious agents, including viruses, bacteria, and parasites, to humans and animals. Tularemia, caused by the Gram-negative bacterium Francisella tularensis, is a highly-infectious zoonotic agent that elicits flu-like symptoms in humans. In the United States, approximately half of tularemia cases are tick-associated. In a recent review by Tully and Huntley, the following points are highlighted: (1) Background information on tularemia and F. tularensis; (2) What is known about the four different tick vectors associated with tularemia, including their geographic ranges and other arthropod vectors historically associated with tularemia; (3) Physiological cues in ticks (compared to the mammalian host) that may prompt F. tularensis to modify its metabolism and protein expression to survive and persist in ticks.
1. Introduction
The most recent data from the United States (U.S.) Centers for Disease Control and Prevention (CDC) indicate that the reported number of tick-borne disease cases have more than doubled in the past 13 years and account for 77% of all vector-borne disease cases [1]. Although insecticides historically have been useful for controlling tick-borne diseases, resistance is becoming widespread in the U.S. [1][2]. Tick-borne diseases, including Lyme disease, spotted fever rickettsioses, babesiosis, anaplasmosis/ehrlichiosis, and tularemia, have been difficult to control because vaccines are not available. Additionally, reports of tick geographic range expansion and increases in wildlife populations that support ticks further complicate tick-borne disease control efforts [3][4][5]. Although Lyme disease accounted for 82% of all U.S. tick-borne disease cases between 2004 and 2016, tularemia cases have seen a resurgence in recent decades. Over 225 tularemia cases per year have been reported since 2015, with 314 reported cases in 2015—the most recorded since 1964
Francisella tularensis, the causative agent of tularemia, has been classified as a Tier 1 Select Agent by the CDC because of its low infectious dose, ease of aerosolization, rapid onset of severe disease, and high morbidity and mortality rates. Two F. tularensis subspecies cause human disease and, although they are closely related genetically, vary in their infectious dose and disease severity [6]. F. tularensis subsp. tularensis (Type A) has an extremely low infectious dose (<10 CFU) and is associated with severe, often lethal, disease. F. tularensis subsp. holarctica (Type B), has a slightly higher infectious dose (>100 CFU) and is associated with progressive disease with lower mortality rates [7]. Type A F. tularensis is further divided into three distinct clades, A1a, A1b and A2. Previous studies have shown that mice infected with A1b die earlier than those infected with A1a, A2 or Type B F. tularensis strains, demonstrating marked differences in F. tularensis virulence [6]. Through repeated subculturing of F. tularensis subsp. holarctica, a live attenuated strain was created by the former Soviet Union in the 1930s and has been designated as the live vaccine strain (LVS) [8]. Despite its name, LVS is not a licensed vaccine in the U.S. due to the unresolved questions about the mechanism(s) of attenuation, adverse effects in some immunized humans, and incomplete protection against Type A aerosol infection [7][8]. However, LVS has been proven to be a useful tool to study F. tularensis virulence, as it causes lethal disease in mice yet can be worked with using normal biosafety precautions (i.e., biosafety level 2; BSL2) [9]. A separate Francisella species, F. novicida, rarely associated with disease in immunocompromised humans, is used as a surrogate for F. tularensis in some studies because of reduced biosafety requirements and ease of genetic manipulation [10].
Beyond concerns over the potential use of F. tularensis as a bioweapon (i.e., Select Agent designation), approximately half of tularemia cases in the U.S. are associated with tick bites [11][12]. In contrast, European tularemia cases are generally associated with ingestion of contaminated water from wells, streams, rivers, ponds, and lakes [13][14]. However, in Sweden, most tularemia cases are associated with mosquito bites [15][16][17]. A recently-published model indicated that climate change may triple the number of European tularemia cases per year, due to increases in mosquitos, higher temperatures, and increased precipitation [18]. Although ticks are generally not considered to be major drivers of European tularemia infections, F. tularensis-infected ticks have been reported in Spain [19], Germany [20], Denmark [21] and Poland [22][23]. Conversely, other groups have not detected F. tularensis in ticks collected from Poland, France, or the Netherlands [24][25][26], suggesting that more studies are needed to understand the current and future risks of tick-transmitted tularemia in Europe.
In the U.S., where tularemia was first documented, field-collected Dermacentor andersoni ticks (Rocky Mountain wood tick) were shown to transmit lethal Bacterium tularense (now known as F. tularensis) to guinea pigs [27]. However, the true role of D. andersoni ticks in U.S. tularemia cases remains unknown. Data from the CDC indicate that U.S. tularemia infections more commonly stem from D. variabilis (American dog tick) and Amblyomma americanum (Lone star tick) ticks, which are known to vary in their geographic distribution and mammalian hosts, as well as less understood factors, including tick physiology, endosymbionts, and antimicrobial defenses [4][28]. Indeed, previous studies have reported that differences in the numbers of tick phagocytic cells and prevalence/type of endosymbionts in A. americanum, D. andersoni, and D. variabilis ticks affected the molting success (i.e., survival between tick life stages) of these three tick species [29][30][31].
The complex life cycle of the tick (3 year progression from larvae to nymph to adult), including taking a blood meal from various hosts and molting to the next life stage after each blood meal, combined with varying lengths and severities of North American winters, indicates that upon infecting a tick, F. tularensis must undergo major changes over the course of >5 months to persist and replicate, before being transmitted to naive mammals. Laboratory experiments have confirmed that F. tularensis persists in ticks for >4 months, supporting the role of the tick as a potential environmental reservoir [32][33]. Additionally, F. tularensis has been shown to persist in ticks between molts (transstadial transmission) and replicate to high bacterial numbers in ticks, demonstrating that ticks serve as both a reservoir and an amplification vessel for F. tularensis in the environment [33][34]. Rabbits also have been implicated as a major environmental reservoir for tularemia, as studies have demonstrated that they can survive for 3–13 days following intradermal infection (mimicking a tick infection) with Type A1a, A1b or A2 F. tularensis and over 14 days for Type B F. tularensis infection [35]. This rabbit infection data, together with the 3–7 days that both D. variabilis and A. americanum ticks (varies depending on the tick life stage) take a blood meal, suggest that infected rabbits can potentially infect large numbers of ticks [4][33][34]. Despite those studies, we still do not understand whether different tick species acquire different levels of F. tularensis infections from different hosts, whether different tick species promote or restrict F. tularensis infections, what F. tularensis—tick interactions occur, how F. tularensis is maintained through the tick molt, or how infected ticks transmit F. tularensis to naïve hosts.
2. Epidemiology of F. tularensis Transmission by Ticks in the U.S.
Tularemia has been shown to be transmitted by at least four different ticks in the U.S., including D. variabilis, D. andersoni, D. occidentalis (Pacific coast tick), and A. americanum [27][33][34][36]. Although D. variabilis has been implicated as the primary vector, and A. americanum also appears to be an important vector for tularemia, we still do not know which tick(s) poses the greatest threat to human health in the U.S. [33][37]. A 1924 study noted that D. andersoni ticks could transmit virulent F. tularensis to guinea pigs [27] and a more recent study reported that D. andersoni ticks could be infected by and transmit F. novicida (rare infection of immunocompromised humans) [38]. However, very little is known about current infection rates of D. variabilis, A. americanum, or D. andersoni ticks with virulent strains of F. tularensis or about which ticks are commonly associated with current human tularemia cases. These, and other gaps in knowledge, have resulted in a call for new research studies on tick-borne F. tularensis [28]. In addition, although very few studies have explored F. tularensis–tick interactions, such studies could provide important information that could be used to develop new strategies to reduce F. tularensis in the environment [12].
One area of the U.S. that has been extensively studied to understand F. tularensis environmental persistence and tick transmission is Martha’s Vineyard. In both 1978 and 2000, two separate tularemia outbreaks occurred on Martha’s Vineyard, each involving 15 patients, with 1 fatality [39][40]. Although pneumonic disease was the most common symptom in both outbreaks, D. variabilis tick bites remain the only proven mode of transmission in most cases [41]. Sampling studies at Martha’s Vineyard have assessed F. tularensis infection rates in D. variabilis ticks, finding a median annual prevalence of 3.4% over four years, suggesting that F. tularensis infections are stable on the island [42]. Additionally, D. variabilis ticks infected with Type A F. tularensis have been reported to harbor over 108 genome equivalents/tick. Although genome equivalents may not accurately quantitate viable bacteria, those data indicate that D. variabilis ticks may sustain high F. tularensis numbers that cause significant transmission and disease in humans [37][43].
The majority of tularemia cases occur in the south-central U.S. [33][42][44]. In fact, four states accounted for 58% of tularemia cases in 2018 (most recent data from the CDC): Arkansas (24%), Oklahoma (19%), Kansas (8%) and Missouri (7%) [45]. In this region of the U.S., two major tick species exist: D. variabilis and A. americanum [46][47]. In one study from Missouri, >8500 A. americanum ticks were harvested over three years from various hosts, including white-tailed deer, fox, opossum and rabbits, with tick prevalence rates on these potential hosts ranging from 0.7% to 100%. Although the presence and absence of pathogenic bacteria were not assessed in these A. americanum ticks, >1100 A. americanum nymphs were collected from a single rabbit, a natural reservoir for tularemia, highlighting the ability of one infected host to spread tularemia to thousands of other hosts, including humans [48]. In Arkansas, approx. 92% of field-collected ticks were A. americanum and approx. 7% were D. variabilis. Interestingly, none of the D. variabilis ticks tested positive for F. tularensis (>2000 ticks tested), whereas approx. 4% of A. americanum ticks (>5000 ticks tested) were positive for F. tularensis [49]. A study of over 3500 field-collected D. variabilis ticks from Minnesota identified Type A F. tularensis in 3.6% of those ticks [50]. That infection rate was similar to what has been reported at Martha’s Vineyard, where the annual F. tularensis infection rates in D. variabilis ticks ranges from 2.7% to 4.3%, demonstrating that tick-borne tularemia infections are not restricted to the south-central U.S. [42]. In contrast, sampling studies in Washington state did not identify F. tularensis in either D. andersoni or D. variabilis ticks. However, that study examined less than 200 Dermacentor sp. ticks and only 25 tularemia cases were reported in Washington state between 2011 and 2016 [51]. Clearly, more studies are needed to assess F. tularensis infection rates in multiple tick vectors across the U.S. Further, given the geographic range expansion of various ticks throughout the U.S. [4], continuing studies will be needed to understand how this expansion will affect tick-borne disease transmission.
Although this review is focused on F. tularensis infections of ticks, historical data suggest that biting flies also can transmit tularemia [52][53]. However, no recent data from the CDC have linked tularemia infections with biting flies. One study used Drosophila as an arthropod model for F. tularensis infections, finding that doses as low as 200 CFU killed >90% of fruit flies injected with F. tularensis LVS [54], bringing into question whether flies play a significant role in tularemia transmission.
3. Factors that Affect F. tularensis Infections of Ticks
Tick-borne pathogens must be able to efficiently transition from mammalian to arthropod hosts following tick feeding [55][56]. F. tularensis has been reported to infect and cause disease in over 300 animal species, including humans, highlighting the zoonotic potential and plasticity of F. tularensis [57]. Although not well studied, F. tularensis likely undergoes substantial changes, including major changes in protein expression profiles, between the mammalian host and the tick vector. Factors such as temperature and pH have been shown to be important cues when Borrelia burgdorferi, the causative agent of Lyme disease, transitions between mammalian hosts and ticks. These stimuli result in modifications of bacterial surface proteins to enhance B. burgdorferi acquisition by ticks [58][59]. Two surface-exposed lipoproteins, OspA and OspC, are among the most well-characterized proteins that are differentially expressed in B. burgdorferi. OspA is highly expressed under conditions that resemble the tick environment (pH 7.5 and 23 °C) [60]. Conversely, during a tick blood meal, the pH and temperature of the tick midgut change to 6.8 and 35 °C, respectively, triggering the upregulation of OspC, and promoting the migration of B. burgdorferi through the tick salivary gland to the mammalian host [61][62].
In mammals, F. tularensis is an intracellular pathogen, infecting cell types ranging from macrophages, to neutrophils, to epithelial cells, to erythrocytes [63]. Many previous studies have identified F. tularensis virulence factors and have examined F. tularensis pathogenesis mechanisms using macrophage infection models [64]. When comparing macrophages and ticks, F. tularensis encounters low pH in both the macrophage phagosome and tick midgut [62][65]. In fact, both Type A F. tularensis and LVS are resistant to acid stress and viable at pH 3 [66]. While it has been reported that F. tularensis responds to low pH by upregulating genes in the Francisella pathogenicity island (FPI) to escape from the phagosome [67][68], we are not aware of any study that directly examines whether F. tularensis uses low pH as an indicator of the transition between mammalian and arthropod hosts.
Conversely, because iron is extremely limited in the macrophage phagosome but is readily available in replete ticks (through hemolysis), it is possible that F. tularensis may successfully transition from mammalian to arthropod hosts by sensing changes in iron and/or altering expression of iron-regulated genes in the tick (Figure 1) [69]. Iron-regulated genes have been shown to be important for the regulation of virulence in B. burgdorferi [70]. One study found that F. tularensis LVS differentially regulated over 70 genes in iron-limiting conditions, many of which were shown to be associated with virulence or intracellular replication [71]. Although that study did not explore F. tularensis gene regulation under iron-replete or high-iron conditions, similar to the tick midgut after a blood meal, such data could provide important information about how F. tularensis initially responds and adapts to life inside a tick.
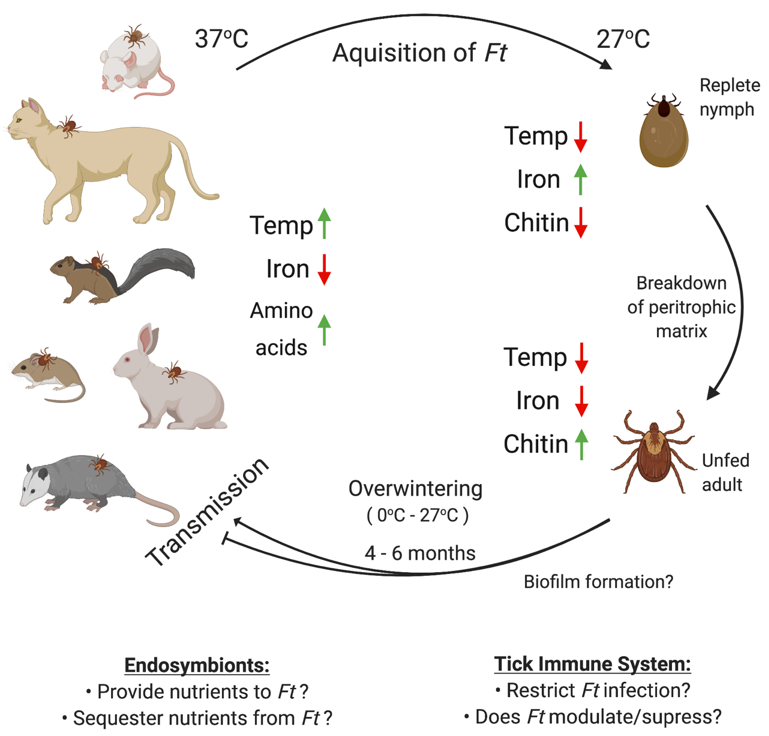
Figure 1. Possible factors affecting F. tularensis infection, persistence, and transmission in ticks. Ticks acquire F. tularensis (Ft) by taking a blood meal from an infected host (e.g., mouse, cat, squirrel, vole, rabbit, or opossum). Upon infecting ticks, F. tularensis likely regulates gene expression based on changes in temperature (37 to 27 °C) and iron (low to high). Following processing of the blood meal, nutrients are likely limited in the tick. However, during the tick molting process, chitin fragments may be made available via chitin remodeling and/or breakdown of the peritrophic matrix. Subsequently, F. tularensis must overwinter in ticks, a process which may require biofilm formation, evasion/modulation of the tick immune system, and/or interactions with tick endosymbionts. However, previous studies have not examined any of these factors. Upon transmission from infected ticks to a new mammalian host, F. tularensis likely senses mammalian cues (e.g., increased temperature, low iron, higher concentrations of amino acids) and may alter its gene expression, including virulence genes, to promote infection. Figure created using Biorender.com.
A third environmental cue that F. tularensis might use to successfully transition to ticks may be temperature changes (Figure 1). One study reported that 11% of the F. tularensis LVS genome was differentially regulated when the bacterium was switched from ambient (26 °C) to mammalian (37 °C) temperature. Up to 40% of those identified genes were known to be important for virulence or intracellular replication, suggesting that temperature changes prime F. tularensis for pathogenicity in mammals [72]. Although that study may have provided information relevant to F. tularensis transmission from ticks to mammals, no studies have been performed to identify F. tularensis genes differentially regulated when transitioning from mammals to ticks. Another study found that F. tularensis modifies its lipopolysaccharide (LPS) structure in response to temperature changes, including altering expression of acetyltransferases, which add shorter or longer acyl chains to lipid A under ambient (18 °C) or mammalian (37 °C) temperatures, respectively [73]. Those LPS modifications were shown to promote bacterial survival and growth in cold conditions, which could be speculated to help F. tularensis survive and persist in ticks during the winter. However, experiments to confirm whether these LPS modifications promote F. tularensis persistence in ticks have not been conducted.
Likely because of the difficulties and biosafety risks of working with infected ticks, tick cell lines offer a simplified tool to understand how variables, such as temperature and tick species, can affect bacterial infection and persistence in arthropods. In one study, cell lines derived from D. andersoni and Ixodes scapularis ticks were infected with F. novicida, finding that at 34 °C, F. novicida infected and replicated 2-logs higher in D. andersoni-derived cells, compared to I. scapularis-derived cells. However, F. novicida infection killed up to 25% of D. andersoni cells, compared to I. scapularis cells that appeared to be unaffected by F. novicida up to 6 days post-infection. At 24 °C, F. novicida infected and replicated to similar levels in D. andersoni-derived cells, compared to I. scapularis-derived cells. However, approximately 15% less D. andersoni cell death was detected at the lower temperature, despite elevated bacterial numbers, indicating that low temperatures may decrease bacterial virulence, while still supporting bacterial replication in tick cells [5]. Although the above highlighted studies provided important insights into how F. tularensis may sense environmental cues and promote tick infections, there are still major gaps in our understanding of how virulent F. tularensis strains infect ticks, how F. tularensis adapts to life in the tick, and which ticks pose the greatest risk for tularemia transmission.
References
- Rosenberg, ; Lindsey, N.P.; Fischer, M.; Gregory, C.J.; Hinckley, A.F.; Mead, P.S.; Paz-Bailey, G.; Waterman, S.H.; Drexler, N.A.; Kersh, G.J.; et al. Vital Signs: Trends in Reported Vectorborne Disease Cases United States and Territories, 2004–2016. Mmwr Morb. Mortal. Wkly. Rep. 2018, 67, 496–501, doi:10.15585/mmwr.mm6717e1.
- Ostfeld, S.; Canham, C.D.; Oggenfuss, K.; Winchcombe, R.J.; Keesing, F. Climate, deer, rodents, and acorns as determinants of variation in lyme-disease risk. PLoS Biol. 2006, 4, e145, doi:10.1371/journal.pbio.0040145.
- Molaei, ; Little, E.A.H.; Williams, S.C.; Stafford, K.C. Bracing for the Worst‐Range Expansion of the Lone Star Tick in the Northeastern United States. N. Engl. J. Med. 2019, 381, 2189–2192, doi:10.1056/NEJMp1911661.
- Sonenshine, E. Range Expansion of Tick Disease Vectors in North America: Implications for Spread of Tick-Borne Disease. Int. J. Environ. Res. Public Health 2018, 15, 478, doi:10.3390/ijerph15030478.
- Reif, E.; Ujczo, J.K.; Alperin, D.C.; Noh, S.M. Francisella tularensis novicida infection competence differs in cell lines derived from United States populations of Dermacentor andersoni and Ixodes scapularis. Sci. Rep. 2018, 8, 12685, doi:10.1038/s41598-018-30419-4.
- Molins, R.; Delorey, M.J.; Yockey, B.M.; Young, J.W.; Sheldon, S.W.; Reese, S.M.; Schriefer, M.E.; Petersen, J.M. Virulence differences among Francisella tularensis subsp. tularensis clades in mice. PLoS ONE 2010, 5, e10205, doi:10.1371/journal.pone.0010205.
- Ellis, ; Oyston, P.C.; Green, M.; Titball, R.W. Tularemia. Clin. Microbiol. Rev. 2002, 15, 631–646, doi:10.1128/cmr.15.4.631–646.2002.
- Oyston, C.; Quarry, J.E. Tularemia vaccine: Past, present and future. Antonie Van Leeuwenhoek 2005, 87, 277–281.
- Eigelsbach, T.; Downs, C.M. Prophylactic effectiveness of live and killed tularemia vaccines. I. Production of vaccine and evaluation in the white mouse and guinea pig. J. Immunol. 1961, 87, 415–425.
- Kingry, C.; Petersen, J.M. Comparative review of Francisella tularensis and Francisella novicida. Front. Cell. Infect. Microbiol. 2014, 4, 35, doi:10.3389/fcimb.2014.00035.
- Dennis, T.; Inglesby, T.V.; Henderson, D.A.; Bartlett, J.G.; Ascher, M.S.; Eitzen, E.; Fine, A.D.; Friedlander, A.M.; Hauer, J.; Layton, M.; et al. Tularemia as a biological weapon: Medical and public health management. JAMA 2001, 285, 2763–2773, doi:10.1001/jama.285.21.2763.
- Zellner, ; Huntley, J.F. Ticks and Tularemia: Do We Know What We Don’t Know? Front. Cell. Infect. Microbiol. 2019, 9, 146, doi:10.3389/fcimb.2019.00146.
- Larssen, W.; Bergh, K.; Heier, B.T.; Vold, L.; Afset, J.E. All-time high tularaemia incidence in Norway in 2011: Report from the national surveillance. Eur. J. Clin. Microbiol. Infect. Dis. 2014, 33, 1919–1926, doi:10.1007/s10096-014-2163-2.
- Hennebique, ; Boisset, S.; Maurin, M. Tularemia as a waterborne disease: A review. Emerg. Microbes Infect. 2019, 8, 1027–1042, doi:10.1080/22221751.2019.1638734.
- Dryselius, ; Hjertqvist, M.; Makitalo, S.; Lindblom, A.; Lilja, T.; Eklof, D.; Lindstrom, A. Large outbreak of tularaemia, central Sweden, July to September 2019. Eurosurveillance 2019, 24, doi:10.2807/1560¨C7917.ES.2019.24.42.1900603.
- Ryden, ; Bjork, R.; Schafer, M.L.; Lundstrom, J.O.; Petersen, B.; Lindblom, A.; Forsman, M.; Sjostedt, A.; Johansson, A. Outbreaks of tularemia in a boreal forest region depends on mosquito prevalence. J. Infect. Dis. 2012, 205, 297–304, doi:10.1093/infdis/jir732.
- Eliasson, ; Broman, T.; Forsman, M.; Back, E. Tularemia: Current epidemiology and disease management. Infect. Dis. Clin N. Am. 2006, 20, 289–311, ix, doi:10.1016/j.idc.2006.03.002.
- Ma, ; Bring, A.; Kalantari, Z.; Destouni, G. Potential for Hydroclimatically Driven Shifts in Infectious Disease Outbreaks: The Case of Tularemia in High-Latitude Regions. Int. J. Environ. Res. Public Health 2019, 16, 3713, doi:10.3390/ijerph16193717.
- Lopes de Carvalho, ; Toledo, A.; Carvalho, C.L.; Barandika, J.F.; Respicio-Kingry, L.B.; Garcia-Amil, C.; Garcia-Perez, A.L.; Olmeda, A.S.; Ze-Ze, L.; Petersen, J.M.; et al. Francisella species in ticks and animals, Iberian Peninsula. Ticks Tick Borne Dis. 2016, 7, 159–165, doi:10.1016/j.ttbdis.2015.10.009.
- Appelt, ; Koppen, K.; Radonic, A.; Drechsel, O.; Jacob, D.; Grunow, R.; Heuner, K. Genetic Diversity and Spatial Segregation of Francisella tularensis Subspecies holarctica in Germany. Front. Cell. Infect. Microbiol. 2019, 9, 376, doi:10.3389/fcimb.2019.00376.
- Haulrig, B.; Mathiasen, G.; Nielsen, R.M.; Kromann, C.B.; Krogfelt, K.A.; Wiese, L. Two cases of tick-borne transmitted tularemia on Southern Zealand, Denmark. APMIS 2020, 128, 61–64, doi:10.1111/apm.13008.
- Wojcik-Fatla, ; Zajac, V.; Sawczyn, A.; Cisak, E.; Sroka, J.; Dutkiewicz, J. Occurrence of Francisella spp. in Dermacentor reticulatus and Ixodes ricinus ticks collected in eastern Poland. Ticks Tick Borne Dis. 2015, 6, 253–257, doi:10.1016/j.ttbdis.2015.01.005.
- Bielawska-Drozd, ; Cieslik, P.; Zakowska, D.; Glowacka, P.; Wlizlo-Skowronek, B.; Zieba, P.; Zdun, A. Detection of Coxiella burnetii and Francisella tularensis in Tissues of Wild-living Animals and in Ticks of North-west Poland. Pol. J. Microbiol. 2018, 67, 529–534, doi:10.21307/pjm-2018-059.
- Forminska, ; Wolkowicz, T.; Brodzik, K.; Stefanoff, P.; Golab, E.; Masny, A.; Osiak, B.; Cieslik, P.; Bielawska-Drozd, A.; Zasada, A.A. Genetic diversity of Francisella tularensis in Poland with comments on MLVA genotyping and a proposition of a novel rapid v4-genotyping. Ticks Tick Borne Dis. 2020, 11, 101322, doi:10.1016/j.ttbdis.2019.101322.
- Michelet, ; Joncour, G.; Devillers, E.; Torina, A.; Vayssier-Taussat, M.; Bonnet, S.I.; Moutailler, S. Tick species, tick-borne pathogens and symbionts in an insular environment off the coast of Western France. Ticks Tick Borne Dis. 2016, 7, 1109–1115, doi:10.1016/j.ttbdis.2016.08.014.
- Janse, ; Maas, M.; Rijks, J.M.; Koene, M.; van der Plaats, R.Q.; Engelsma, M.; van der Tas, P.; Braks, M.; Stroo, A.; Notermans, D.W.; et al. Environmental surveillance during an outbreak of tularaemia in hares, the Netherlands, 2015. Eurosurveillance 2017, 22, 30607, doi:10.2807/1560¨C7917.ES.2017.22.35.30607.
- Parker, R.; Spencer, R.R.; Francis, E. Tularæmia: XI. Tularæmia Infection in Ticks of the Species Dermacentor Andersoni Stiles in the Bitterroot Valley, Mont. Public Health Rep. (1896–1970) 1924, 39, 1057–1073, doi:10.2307/4577151.
- Eisen, A call for renewed research on tick-borne Francisella tularensis in the Arkansas-Missouri primary national focus of tularemia in humans. J. Med. Entomol. 2007, 44, 389–397, doi:10.1603/0022-2585(2007)44[389:acfrro]2.0.co;2.
- Nagamori, ; Payton, M.; Coburn, L.; Thomas, J.E.; Reichard, M. Nymphal engorgement weight predicts sex of adult Amblyomma americanum, Amblyomma maculatum, Dermacentor andersoni, Dermacentor variabilis, and Rhipicephalus sanguineus ticks. Exp. Appl. Acarol. 2019, 77, 401–410, doi:10.1007/s10493-019-00346-8.
- Oliver, D.; Dusty Loy, J.; Parikh, G.; Bartholomay, L. Comparative analysis of hemocyte phagocytosis between six species of arthropods as measured by flow cytometry. J. Invertebr. Pathol. 2011, 108, 126–130, doi:10.1016/j.jip.2011.07.004.
- Duron, ; Binetruy, F.; Noel, V.; Cremaschi, J.; McCoy, K.D.; Arnathau, C.; Plantard, O.; Goolsby, J.; Perez de Leon, A.A.; Heylen, D.J.A.; et al. Evolutionary changes in symbiont community structure in ticks. Mol. Ecol. 2017, 26, 2905–2921, doi:10.1111/mec.14094.
- Hopla, E. Experimental studies on tick transmission of tularemia organisms. Am. J. Hyg. 1953, 58, 101–118, doi:10.1093/oxfordjournals.aje.a119585.
- Mani, J.; Metcalf, J.A.; Clinkenbeard, K.D. Amblyomma americanum as a Bridging Vector for Human Infection with Francisella tularensis. PLoS ONE 2015, 10, e0130513, doi:10.1371/journal.pone.0130513.
- Coburn, ; Maier, T.; Casey, M.; Padmore, L.; Sato, H.; Frank, D.W. Reproducible and quantitative model of infection of Dermacentor variabilis with the live vaccine strain of Francisella tularensis. Appl. Environ. Microbiol. 2015, 81, 386–395, doi:10.1128/AEM.02917-14.
- Brown, R.; Adney, D.R.; Bielefeldt-Ohmann, H.; Gordy, P.W.; Felix, T.A.; Olea-Popelka, F.J.; Bowen, R.A. Pathogenesis and Immune Responses of Francisella Tularensis Strains in Wild-Caught Cottontail Rabbits (Sylvilagus Spp.). J. Wildl. Dis. 2015, 51, 564–575, doi:10.7589/2015-02-030.
- Parker, R.; Brooks, C.S.; Marsh, H. The Occurrence of Bacterium tularense in the Wood Tick, Dermacentor occidentalis, in California. Public Health Rep. (1896–1970) 1929, 44, 328–335, doi:10.2307/4579265.
- Reese, M.; Petersen, J.M.; Sheldon, S.W.; Dolan, M.C.; Dietrich, G.; Piesman, J.; Eisen, R.J. Transmission efficiency of Francisella tularensis by adult american dog ticks (Acari: Ixodidae). J. Med. Entomol. 2011, 48, 884–890, doi:10.1603/me11005.
- Reif, E.; Palmer, G.H.; Ueti, M.W.; Scoles, G.A.; Margolis, J.J.; Monack, D.M.; Noh, S.M. Dermacentor andersoni transmission of Francisella tularensis subsp. novicida reflects bacterial colonization, dissemination, and replication coordinated with tick feeding. Infect. Immun. 2011, 79, 4941–4946, doi:10.1128/IAI.05676-11.
- Kugeler, J.; Mead, P.S.; Janusz, A.M.; Staples, J.E.; Kubota, K.A.; Chalcraft, L.G.; Petersen, J.M. Molecular Epidemiology of Francisella tularensis in the United States. Clin. Infect. Dis. 2009, 48, 863–870, doi:10.1086/597261.
- Goethert, K.; Shani, I.; Telford, S.R., 3rd. Genotypic diversity of Francisella tularensis infecting Dermacentor variabilis ticks on Martha’s Vineyard, Massachusetts. J. Clin. Microbiol. 2004, 42, 4968–4973, doi:10.1128/JCM.42.11.4968–4973.2004.
- Goethert, K.; Telford, S.R., 3rd. A new Francisella (Beggiatiales: Francisellaceae) inquiline within Dermacentor variabilis say (Acari: Ixodidae). J. Med. Entomol. 2005, 42, 502–505, doi:10.1093/jmedent/42.3.502.
- Goethert, K.; Telford, S.R., 3rd. Nonrandom distribution of vector ticks (Dermacentor variabilis) infected by Francisella tularensis. Plos Pathog. 2009, 5, e1000319, doi:10.1371/journal.ppat.1000319.
- Goethert, K.; Telford, S.R., 3rd. Quantum of infection of Francisella tularensis tularensis in host-seeking Dermacentor variabilis. Ticks Tick Borne Dis. 2010, 1, 66–68, doi:10.1016/j.ttbdis.2010.01.001.
- Montales, T.; Beebe, A.; Chaudhury, A.; Haselow, D.; Patil, S.; Weinstein, S.; Taffner, R.; Patil, N. A Clinical Review of Tick-Borne Diseases in Arkansas. J. Ark. Med. Soc. 2016, 112, 254–258.
- Centers for Disease Control and Prevention: Tularemia: https://www.cdc.gov/tularemia/statistics/index.html (accessed on 23 September 2020).
- James, M.; Burdett, C.; McCool, M.J.; Fox, A.; Riggs, P. The geographic distribution and ecological preferences of the American dog tick, Dermacentor variabilis (Say), in the U.S.A. Med. Vet. Entomol. 2015, 29, 178–188, doi:10.1111/mve.12099.
- Monzon, D.; Atkinson, E.G.; Henn, B.M.; Benach, J.L. Population and Evolutionary Genomics of Amblyomma americanum, an Expanding Arthropod Disease Vector. Genome Biol. Evol. 2016, 8, 1351–1360, doi:10.1645/0022-3395(2000)086[1156:HAASAO]2.0.CO;2.
- Kollars, M., Jr.; Oliver, J.H., Jr.; Durden, L.A.; Kollars, P.G. Host association and seasonal activity of Amblyomma americanum (Acari: Ixodidae) in Missouri. J. Parasitol. 2000, 86, 1156–1159, doi:10.1645/0022-3395.
- Calhoun, L. Natural occurrence of tularemia in the lone star tick, Amblyomma americanus (Linn.), and in dogs in Arkansas. Am. J. Trop. Med. Hyg. 1954, 3, 360–366, doi:10.4269/ajtmh.1954.3.360.
- Whitten, ; Demontigny, C.; Bjork, J.; Foss, M.; Peterson, M.; Scheftel, J.; Neitzel, D.; Sullivan, M.; Smith, K. Prevalence of Francisella tularensis in Dermacentor variabilis Ticks, Minnesota, 2017. Vector Borne Zoonotic Dis. 2019, 19, 596–603, doi:10.1089/vbz.2018.2388.
- Dykstra, A.; Oltean, H.N.; Kangiser, D.; Marsden-Haug, N.; Rich, S.M.; Xu, G.; Lee, M.K.; Morshed, M.G.; Graham, C.B.; Eisen, R.J. Ecology and Epidemiology of Tickborne Pathogens, Washington, USA, 2011–2016. Emerg. Infect. Dis. 2020, 26, 648–657, doi:10.3201/eid2604.191382.
- McCoy, A plague-like disease of rodents. Public Health Bull. 1911, 19, 61-72.
- Jellison, L. Tularemia; geographical distribution of deerfly fever and the biting fly, Chrysops discalis Williston. Public Health Rep. 1950, 65, 1321–1329.
- Vonkavaara, ; Telepnev, M.V.; Ryden, P.; Sjostedt, A.; Stoven, S. Drosophila melanogaster as a model for elucidating the pathogenicity of Francisella tularensis. Cell. Microbiol. 2008, 10, 1327–1338, doi:10.1111/j.1462-5822.2008.01129.x.
- Murfin, E.; Kleinbard, R.; Aydin, M.; Salazar, S.A.; Fikrig, E. Borrelia burgdorferi chemotaxis toward tick protein Salp12 contributes to acquisition. Ticks Tick Borne Dis. 2019, 10, 1124–1134, doi:10.1016/j.ttbdis.2019.06.002.
- Galletti, F.; Fujita, A.; Rosa, R.D.; Martins, L.A.; Soares, H.S.; Labruna, M.B.; Daffre, S.; Fogaca, A.C. Virulence genes of Rickettsia rickettsii are differentially modulated by either temperature upshift or blood-feeding in tick midgut and salivary glands. Parasites Vectors 2016, 9, 331, doi:10.1186/s13071-016-1581-7.
- Keim, ; Johansson, A.; Wagner, D.M. Molecular epidemiology, evolution, and ecology of Francisella. Ann. N. Y. Acad. Sci. 2007, 1105, 30–66, doi:10.1196/annals.1409.011.
- Brooks, S.; Hefty, P.S.; Jolliff, S.E.; Akins, D.R. Global analysis of Borrelia burgdorferi genes regulated by mammalian host-specific signals. Infect. Immun. 2003, 71, 3371–3383, doi:10.1128/iai.71.6.3371–3383.2003.
- Carroll, A.; Cordova, R.M.; Garon, C.F. Identification of 11 pH-regulated genes in Borrelia burgdorferi localizing to linear plasmids. Infect. Immun. 2000, 68, 6677–6684, doi:10.1128/iai.68.12.6677–6684.2000.
- Yang, F.; Pal, U.; Alani, S.M.; Fikrig, E.; Norgard, M.V. Essential role for OspA/B in the life cycle of the Lyme disease spirochete. J. Exp. Med. 2004, 199, 641–648, doi:10.1084/jem.20031960.
- Pal, ; Li, X.; Wang, T.; Montgomery, R.R.; Ramamoorthi, N.; Desilva, A.M.; Bao, F.; Yang, X.; Pypaert, M.; Pradhan, D.; et al. TROSPA, an Ixodes scapularis receptor for Borrelia burgdorferi. Cell 2004, 119, 457–468, doi:10.1016/j.cell.2004.10.027.
- Yang, ; Goldberg, M.S.; Popova, T.G.; Schoeler, G.B.; Wikel, S.K.; Hagman, K.E.; Norgard, M.V. Interdependence of environmental factors influencing reciprocal patterns of gene expression in virulent Borrelia burgdorferi. Mol. Microbiol. 2000, 37, 1470–1479, doi:10.1046/j.1365-2958.2000.02104.x.
- Celli, ; Zahrt, T.C. Mechanisms of Francisella tularensis intracellular pathogenesis. Cold Spring Harb. Perspect. Med. 2013, 3, a010314, doi:10.1101/cshperspect.a010314.
- Steiner, J.; Furuya, Y.; Jordan, M.B.; Metzger, D.W. Protective Role for Macrophages in Respiratory Francisella tularensis Infection. Infect. Immun. 2017, 85, doi:10.1128/IAI.00064-17.
- Canton, ; Khezri, R.; Glogauer, M.; Grinstein, S. Contrasting phagosome pH regulation and maturation in human M1 and M2 macrophages. Mol. Biol. Cell. 2014, 25, 3330–3341, doi:10.1091/mbc.E14-05-0967.
- KuoLee, ; Zhao, X.; Austin, J.; Harris, G.; Conlan, J.W.; Chen, W. Mouse model of oral infection with virulent type A Francisella tularensis. Infect. Immun. 2007, 75, 1651–1660, doi:10.1128/IAI.01834-06.
- Fuller, R.; Kijek, T.M.; Taft-Benz, S.; Kawula, T.H. Environmental and intracellular regulation of Francisella tularensis ripA. Bmc Microbiol. 2009, 9, 216, doi:10.1186/1471-2180-9-216.
- Chong, ; Wehrly, T.D.; Nair, V.; Fischer, E.R.; Barker, J.R.; Klose, K.E.; Celli, J. The early phagosomal stage of Francisella tularensis determines optimal phagosomal escape and Francisella pathogenicity island protein expression. Infect. Immun. 2008, 76, 5488–5499, doi:10.1128/IAI.00682-08.
- Pan, ; Tamilselvam, B.; Hansen, E.J.; Daefler, S. Modulation of iron homeostasis in macrophages by bacterial intracellular pathogens. Bmc Microbiol. 2010, 10, 64, doi:10.1186/1471-2180-10-64.
- Ouyang, ; Deka, R.K.; Norgard, M.V. BosR (BB0647) controls the RpoN-RpoS regulatory pathway and virulence expression in Borrelia burgdorferi by a novel DNA-binding mechanism. Plos Pathog. 2011, 7, e1001272, doi:10.1371/journal.ppat.1001272.
- Deng, ; Blick, R.J.; Liu, W.; Hansen, E.J. Identification of Francisella tularensis genes affected by iron limitation. Infect. Immun. 2006, 74, 4224–4236, doi:10.1128/IAI.01975-05.
- Horzempa, ; Carlson, P.E., Jr.; O’Dee, D.M.; Shanks, R.M.; Nau, G.J. Global transcriptional response to mammalian temperature provides new insight into Francisella tularensis pathogenesis. Bmc Microbiol. 2008, 8, 172, doi:10.1186/1471-2180-8-172.
- Li, ; Powell, D.A.; Shaffer, S.A.; Rasko, D.A.; Pelletier, M.R.; Leszyk, J.D.; Scott, A.J.; Masoudi, A.; Goodlett, D.R.; Wang, X.; et al. LPS remodeling is an evolved survival strategy for bacteria. Proc. Natl. Acad. Sci. USA. 2012, 109, 8716–8721, doi:10.1073/pnas.1202908109.