| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Julian Markovich Rozenberg | + 8950 word(s) | 8950 | 2020-12-30 09:14:12 | | | |
| 2 | Rita Xu | -4705 word(s) | 4245 | 2021-01-07 06:58:22 | | |
Video Upload Options
Epithelial organs are the first barrier against microorganisms and genotoxic stress, in which the p53 family members p63 and p73 have both overlapping and distinct functions. Intriguingly, p73 displays a very specific localization to basal epithelial cells in human tissues, while p63 is expressed in both basal and differentiated cells. Investigations of p63 and p73 protein–protein interactions reveal distinct functions underlying the aforementioned distribution. The p73 and p63 cooperate in the genome stability surveillance in proliferating cells; p73 specific interactors contribute to the transcriptional repression, anaphase promoting complex and spindle assembly checkpoint, whereas p63 specific interactors play roles in the regulation of mRNA processing and splicing in both proliferating and differentiated cells suggesting diversification of the RNA and DNA specific functions within the p53 family.
1. Introduction
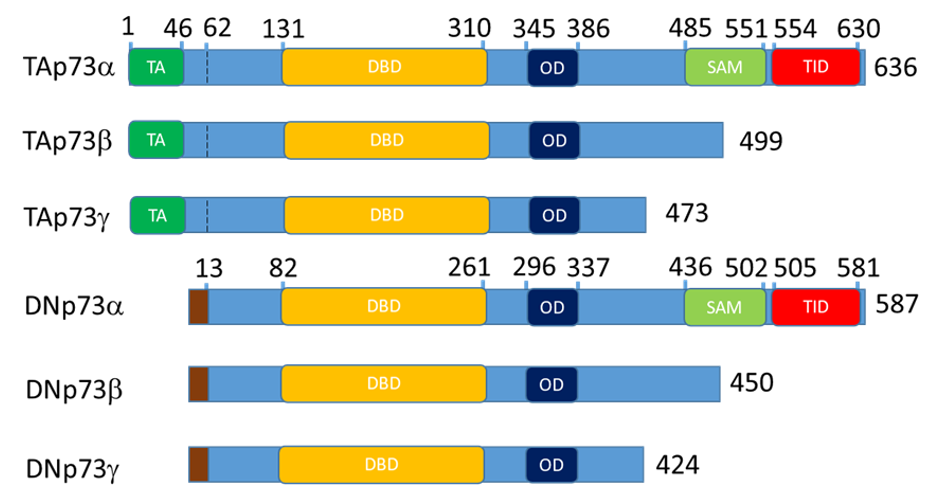
The p53-family is a group of proteins, which consists of p53, p63, and p73 [1][2]. p63 and p73 are paralogs of p53 and have similar structure: transactivation (TA) at the N-terminus following by DNA-binding, nuclear localization, tetramerization domains and sterile alpha motif (SAM) domain, a specific feature of the p63 and p73 at the C-terminus (Figure 1) [1][2][3].
p53, p63 and p73 proteins are represented by multiple isoforms that regulate each other’s transcription and functions [4][5][6][7][8][9]. Full length p73 and p63 contain TA domain and alternative intragenic transcriptional start sites between p63/p73 exons 3 and 4 produce N-truncated “DN” isoforms lacking the TA domain. Full length “-alfa” isoforms contain SAM domain and alternative C-terminal splicing produces beta, gamma, etc., isoforms lacking the SAM domain. Thus, an isoform nomenclature “TAp73α” means that a p73 isoform contains N-terminal TA domain with full length SAM domain at the C-terminal. Accordingly, DNp73β and above (gamma, zeta etc.) means that the activation domain is missing and the SAM domain is truncated. Thereby, combination of N- and C-terminal variants creates a variety of isoforms (Figure 1) [3][10].
Figure 1. Schematic representation of the p73 domain structure for the main isoform expressed in the epithelial tissue. The p63 follows the same nomenclature. Altogether, 13 isoform of p63 and 13 isoforms of p73 exist. DN- isoforms begin with the stretch of 13 amino acids that replace TA domain at 62 bp. The TAp73a coordinates are for the O15350 protein and DNp73a coordinates are for the BAB87244.1 protein. TA – transactivation domain, DBD – DNA binding domain, OD - oligomerization domain, SAM – sterile alpha motif domain and TID - transcription inhibitory domain.
In the full length TAp73α and TAp63a, N-terminal TA domain acts as the transcriptional activator whereas C- terminal containing SAM and inhibitory domains repress TA activity [11][12]. Generally, isoforms lacking TA domain DNp63 and DNp73 repress activity of isoforms containing TA domain [4][7]. Also, p73 isoforms can act as conditional activators or repressors [13][14]. For example, both TAp73β and DNp73β but neither p53, TAp73α, TAp63α, nor TAp63β induce the expression of caspase-2S [13]. Similarly, TAp73α co-activates c-Jun whereas TAp73β can act as a repressor of c-Jun mediated transactivation [14][15].
In particular, DNA-binding domain is the most conserved structural element across the p53-family (Figure 4), leading to identical DNA recognition [16][17][18][19], underlining the functional relatedness of these proteins as transcription factors. p53 functions as a transcription factor in charge of the activation of genes responsible for the cell-cycle arrest in G1 and G2/M checkpoints and apoptosis induction. At the same time, both p73 and p63 are known for regulation of the p53-responsive genes: MDM2, IGFBP3, p21 [20]. However, apart from the large amount of evidence that p53, p63 and p73 proteins have overlapping and interdependent functions in the apoptosis control and genome stability [21], p63 and p73 also have distinct specific roles in developmental processes [22][23][24][25][26], DNA repair [27][28][29] and ciliogenesis [30][31][32].
Specifically, p63 and p73 play distinct roles in epithelial morphogenesis. The p63 is a major player in skin biology and its role in epidermis was thoroughly investigated during the last two decades [33][34][35][36][37][38][39].
The p63 knockout in mice leads to E1 lethality and pups born with non-differentiated skin [33][34]. Total p63 knockout has been rescued by specific p63 isoforms expression under the control of KRT5 promoter [38]. Double TAp63a/DNp63a rescue showed regions of differentiated skin, while p63-/TAp63 did not rescue, whereas p63-/DNp63 showed only basal keratinocytes formation. Consistently, DNp63 regulated basal undifferentiated keratinocytes marker KRT14, whereas TAp63 regulated markers of differentiated keratinocytes (Ets-1, KRT1, transglutaminases, involucrin) [38]. In addition, p63 function is required for the establishment of the cell division perpendicular to the basal membrane at E18.5 day of the embryo [35][40]. Moreover, both β1-integrin and laminin are distributed around the cells and not localised to the basal membrane in the p63 knockout mice [40].
In contrast, the p73 knockout mice are characterised by runting phenotype, hippocampal dysgenesis and hydrocephalus, sterility, chronic inflammation and infection in the lungs, sinus, and ears [41]. Total p73 knockout mice do not have profound skin phenotype except the somewhat slower wound-healing response [42]. TAp73 knockout mice demonstrate sterility, hippocampal dysgenesis, hydrocephalus, premature aging, genomic instability, and increased frequency of tumors [24]. Finally DNp73 knockout causes neurodegeneration, specifically hippocampal dysgenesis and hydrocephalus [43].
In the human skin, some keratinocytes fail to divide and become polyploid upon differentiation that can be induced by the DNA damage and is associated with overexpression of Cyclin E [44][45]. Using fluorescence in situ hybridization for DNA, it was shown that human skin keratinocytes become multinucleated with their genomes increasing up to 4 fold [46][47].
Intriguingly, p73 displays a very specific localization to basal epithelial cells in all human tissue where it’s localization was determined by immunohistochemistry. In contrast, the p63 is expressed in both basal and differentiated cells [42][48].
The role of p63 in polyploidy has not been evaluated, however, the presence of p63 in differentiating keratinocytes suggests that p63 is not interfering with the polyploidy of these cells. At the same time, p73 suppresses polyploidy and aneuploidy in the absence of functional p53 [23][49]. Also, cells from TAp73−/− mice exhibit genomic instability associated with the enhanced aneuploidy [24]. Therefore, it is possible that the differential presence of the p63 and p73 in the differentiated and basal cells is related to the strict polyploidy control by the p73. Specifically, the absence of p73 allows cells to loosen the mitotic spindle assembly checkpoint and hence become polyploid and multinucleated.
2. p63 and p73 Distribution in the Normal Epithelial Tissues
According to the human protein and RNA expression atlas http://www.proteinatlas.org [50], the p53 protein could be detected at extremely low levels in the oral mucosa, oesophagus, urinary bladder, skin, and tonsils, and this expression is variable across samples and antibodies. In contrast, p63 and p73 proteins are consistently expressed. Moreover, proteins exhibit a characteristic expression pattern. In the skin, p63 levels are the highest in the basal cells, but also detected in the squamous, more differentiated cells whereas p73 is found only in the basal cells (Figure 2A,B). Similarly, in the nasopharynx (Figure 2D,E), vagina (not shown) and oral mucosa (Figure 2G,H) the p73 is restricted to the basal cells and p63 is expressed throughout the basal epithelium and differentiated cells. The p73 expression can be detected in a few cells of the basal layer of the salivary glands lumens, whereas p63 is determined in the different cells (not shown).
In the bronchial epithelium of the lungs, p63 and p73 staining patterns seem to be similar (Figure 2J–K). Whereas p63 antibody strongly stains these tissues, with the different staining patterns than described above, it also stains myoepithelial cells of the breast, urinary bladder, epididymis, seminal vesicles, prostate, and placenta. Interestingly, in these organs, p63 stained cells immediately next to the basal membranes, resembling the p73 staining pattern in the organs expressing both p63 and p73.
In vivo, mitosis mostly occurs in the basal layer, where the p73 is, but a few cells go through mitosis just above the basal layer (Figure 2C,F,I,L) marking the last asymmetric division, leading to the terminal differentiation for one cell and an ability to proliferate for another [35][55][57].
How do these data correlate with RNA expression levels of p63 and p73?
According to GTEx portal RNA-seq data, expression of p73 is quite tissue specific, with relatively high levels of DNp73 in epithelial tissue: skin, vagina, esophagus (similar to the protein Atlas data) and TAp73 is expressed in the brain https://www.gtexportal.org/home/gene/TP73.
Similarly to the immunohistochemistry images, the level of tp73 expression in skin is 7.8 transcripts per million (TPM), whereas the level of tp63 is 127 TPM and tp53 is 36 TPM (Supplementary Excel File). In comparison, MDM2 is 11 TPM and MDM4 is 24 TPM and CDKN1A (p21) is 225 TPM. According to the single cell RNA-seq of the human skin, the p73 RNA is under detection threshold [55]. However, to release single cells, the skin is digested at 37 °C for 2 h, which can lead to RNA degradation, especially for the unstable transcripts [58][59].
Given the low p73 RNA levels in skin and other epithelial organs, the question arises: how reliable is p73 antibody staining? We have thoroughly analyzed the available data using different antibodies and conclude that p73 expression has the same characteristic patterns in skin and other epithelial organs, as defined by experiments using antibodies raised against distinct p73 epitopes in independent studies.
Analysis of the literature and web resources revealed that there are 4 different antibodies against different epitopes that generate exactly the same p73 staining pattern in skin. Chemicon rabbit polyclonal AB7824 [4], Leica Biosystems mouse monoclonal (Leica Biosystems Cat# NCL-p73, RRID:AB_563940), and two Abcam’s: EP436Y (aa 50–150) [42][48], EPR19884 (C-terminus, skin IHC image is at the Abcam website). Reactivity of the Abcam’s antibodies to p73 is validated by the knockout/knockdown and overexpression experiments either by manufacturer or by independent investigators using immunohistochemistry and immunoblotting [42][60]. The main concern is that p73 antibodies can cross react with p63. There are several lines of evidence against it. Firstly, p63 immunostaining appeared earlier during skin development than EP436Y p73 staining [42]. Secondly, during oocyte development, EP436Y p73 antibody stains granulosa cells and these staining disappear in the p73 −/− mice. In contrast, p63 stains primordial follicles in both p73+/+ and p73 −/− mice [60]. These data were supported by immunoblotting of the wild type and p73 −/− ovaries. Third, the p73 EP436Y antibody does not react with p63 by immunohistochemistry [48] as it stains a subset of the p63 positive cells in human and mouse epidermis [42][48]. Moreover, closer examination revealed that a few cells have brighter p73 staining than p63 [42][48]. The absence of reactivity of the EP436Y p73 antibody to p63 by immunoblotting was also demonstrated [42]. Therefore, it is possible to conclude that at least EP436Y p73 antibody is reliable.
In addition, several tissues display strong p63 staining and no AB7824 or RRID:AB_563940 p73 antibody staining (breast, prostate, placenta, seminal vesicles) was observed, whereas other epithelial organs display the same staining of the basal cells as in skin (Figure 2) http://www.proteinatlas.org) [50].
Thus, p73 protein is expressed in the basal layer of epithelial organs, whereas p63 appears in the basal as well as in the differentiated cells.
3. Common and Distinct Multiprotein Complexes of p53 Family Members and Their Distribution in the Epithelial Organs.
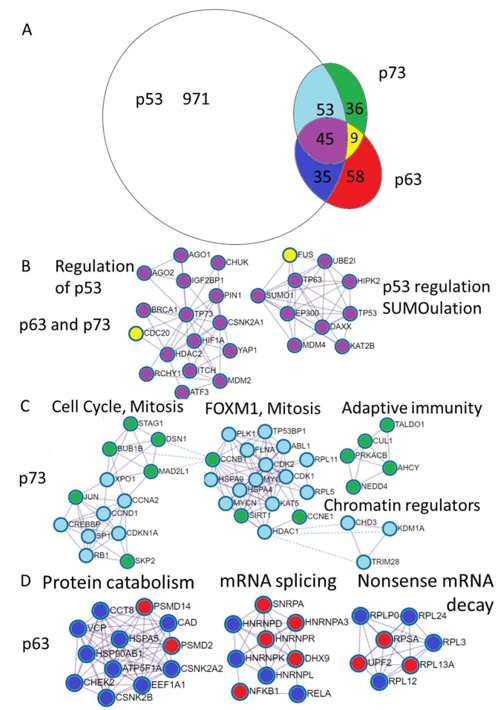
Differential and common functions of p63 and p73 are reflected by the proteins they interact with. In order to have the available information of their binding partners, we downloaded protein-protein interactions for the p53, p63 and p73 from the BioGrid database. We excluded interactions from the genetic screens, keeping only physical associations. Shared and unique interactions are presented as an Euler diagram (Figure 3A).
Furthermore, we annotated shared and unique p53, p63 and p73 interactions using Metascape software [53]. The MCODE algorithm within Metascape detects densely interconnected regions in protein-protein interaction networks that may represent distinct molecular complexes (Figure 3B–D). In addition, Metascape applies enrichment analysis of pathways and processes to identified networks of protein interactions and uses the top three enriched terms for the annotation of biological roles of the putative molecular complexes. In addition, we compared protein interaction data with mRNA expression using publicly available TCGA RNA-seq datasets and single cell sequencing of human skin (Supplementary File) [55]. Below we review what is known about the functional impact of the p53 family protein interactions focusing on the biology of the epithelial tissue.
Figure 3. Annotation of the unique p73 and p63 protein interactions reveal distinct physiological functions. (A). Euler diagram of the p53, p63, and p73 protein interactions from the BioGrid database. Coloring of the Euler diagram corresponds to the coloring of dots representing proteins in the subsequent sections of the figure (B–D) describing protein interactions. (B). Annotation of proteins uniquely interacting with p63 revealed putative complexes involved in the protein catabolism and synthesis including protein folding and transport (CCT8, HSPA5, HSP90AB, VCP, CCT8, ATP5F1A), translation—EEF1A1 and degradation (PSMD2, PSMD14). The second complex contains ribonucleoproteins involved in the mRNA splicing and Nf-kb transcription factors and the third represents ribosomal proteins involved in translation and nonsense mRNA decay (UPF2). (C). Annotation of proteins interacting with p73 revealed putative complexes involved in the regulation of the cell cycle (Rb, CDKN1A (p21), Cyclin D1, Cyclin A2, SPK2) as well as mitosis (BUB1B, MAD2L1, DSN1, STAG1). The next complex is FOXM1 mediated cell-cycle regulation. Interestingly, the adaptive immunity cluster is composed exclusively of the p73 interacting proteins and three chromatin regulators can bind both p53 and p73. (D). Annotation of proteins interacting with both p63 and p73 revealed putative complexes involved in the regulation (repression) of the p53 stability (MDM2, MDM4,RCHY1, ITCH, SUMO1, UBE2I) and transcriptional activity (HDAC2) transcriptional coactivators (EP300, KAT2B, HIPK2), DNA repair (BRCA1), DNA binding transcription factors (HIF1a, YAP, ATF3, p63, p73). Two proteins that are bound by p63 and p73 and not p53 are CDC20—a pivotal protein in the mitosis and FUS—the splicing regulating protein.
3.1. Interaction between p63 and p73 in Gene Regulation
It is widely accepted that transcription is regulated by the interplay of the multiprotein complexes [61][62][63][64][65]. Moreover, due to the presence of dominant activators or repressors co-occupying DNA regulatory sites, functions of a particular transcription factor might be limited to a subset of its binding sites [61][64][66][67][68]. For the p53 transcription factors family, interactions with distinct proteins may skew the binding of p53, p63 and p73 isoforms towards different sites in the genome and determine their specific activities [14][21][43][63][69][70][71].
As discussed above, p63 and p73 colocalize in the basal layers of the epithelial organs, while p73 is not present in the differentiating cells. Indeed, the p63 and p73 can hetero-oligomerize through their tetramerization domains, and the resulting hetero-tetramer consisting of two p63 and two p73 molecules is a thermodynamically preferred structure [48].
Calcium differentiation of the keratinocytes induces p73 protein and promotes its interaction with p63 at days 2–4 of calcium-induced differentiation [48]. The interaction between p63 and p73 is initiated just before cell culture stratification and might play a role in the induction of differentiation markers. However, it is important to mention that at 48 h in calcium-enriched media, cultures of human keratinocytes typically contain a mixture of proliferating and differentiated cells, thereby, it is possible that interaction between p63 and p73 occurs in the proliferating cells, consistent with in vivo p73/p63 colocalization in the basal cells.
In support of this data, it was shown that in HaCat and NHEK cells p73 delta and gamma isoforms as well as p63 were able to activate involucrin and loricrin promoters suggesting that both of these genes may play a role in the induction of differentiation [72].
Interactions of DNp63a and TAp73β led to reduced TAp73β transactivation of the luciferase reporter in neuroblastoma cells [48]. However, in skin and other epithelia p73 and p63 are mostly represented by the DN- isoforms, whose transcriptional activation is determined by interaction with other transcription factors [14][21][71]. The distinction between DN- isoforms of p63 and p73 is demonstrated by co-expression of the KLF4 with DNp73β (but not DNp63α or either KLF4 or DNp73β alone) that dramatically induced fibroblast–keratinocytes reprogramming and expression of keratinocyte specific marks (KRT5, KRT14, FLG, SPN) in both fibroblasts and mesenchymal triple-negative breast cancer cell line MDA-MB-231 [42].
Several signaling transcription factors/pathways cooperate with the p63/p73 to regulate gene expression. Of these, the most studied are CEBP/AP1/NfKb transcription factors that play a pivotal role in the epithelial differentiation [61][73].
It is likely that in skin cells DNp63a and DNp73α interact with either c-Jun or c-Rel and localise to the AP-1 binding sites similarly to what was observed in neck squamous cell carcinoma and osteosarcoma Saos cells [14][21]. In the squamous cancer cells, proinflammatory cytokine TNF-α modulates interaction of DNp63α/TAp73 with c-REL and induces redistribution of TAp73 from the p53 to AP-1 DNA binding sites repressing TAp73 pro-apoptotic functions and inducing an oncogenic gene expression in the absence of functional p53 [21][71]. Thus, TAp73 in complex with AP-1 versus DNp63a switch from anti- to pro-oncogenic activities [15][21][71].
Accordingly, a consensus p53 binding motif is found next to the composite CEBP/c-Jun binding site enriched in the promoters bound by CEBPb and c-Jun in mice basal keratinocytes [61]. Notably, promoters with differentiation induced RNA polymerase 2 binding are preferentially occupied by CEBPb and c-Jun. However, the combinatorial p53/CEBP/AP1 motif was overrepresented in the CEBPb bound promoters also active in the basal keratinocytes.
In addition, it was shown that p63 interacts with CEBPb and Nf-kb in lung cancer cells upon cigarette smoke extract exposure and regulates COX-2 expression, supporting possible functional consequence of the p53/CEBP/AP1 binding sites proximity [74].
The localization of the p63 binding sites as suggested by the ChIP-seq data does not change upon the keratinocytes differentiation and modulation of the activity of the p63-bound enhancers is likely determined by other protein complexes that cooperate with p63 [66]. Upon progression through the cell cycle during epidermal differentiation, high expression of the chromatin remodelers occurs in the mitotic phase of the cell cycle, concomitant with induction of the early differentiation markers KRT1 and KRT10 [55][57], possibly marking a subset of epidermal asymmetrical division, in which p63 plays a pivotal role [35]. Accordingly, p73 interacts with chromatin remodelers and transcription factors (Jun, RB1, SP1, HDAC1) involved in the cell cycle regulation, FOXM1 pathway and mitosis (Figure 3), suggesting that the p73 specific interactions might modulate p63 activity either by promoting or repressing transcription [15][63][75][76][77].
Indeed, additional evidence from the chromatin immunoprecipitation sequencing (ChIP-seq) data suggests that p63 and p73 occupy a similar set of promoters in cancer cell lines [42][78]. Even though genome-wide colocalization of the p63 and p73 in the primary keratinocytes or in skin has not been yet analyzed, it is assumed that p63 and p73 will be localized similarly [42]. Thus, both hetero-tetramerization mediated and distinct DNp63 and DNp73 DNA binding are likely to occur in the epithelial organs. While a plausible hypothesis would be that these complexes differentially regulate transcription in the basal epithelial and in the differentiating cells, details of the regulation are just beginning to emerge [42][48][66][74].
3.2. Common p63 and p73 Interactors
Complexes of 52 proteins that interact with both p63 and p73 correspond to the shared pathways regulating the p53 family. These include regulators of the p53 stability and transcriptional activation by ubiquitination and sumoylation: MDM2, MDM4, RCHY1, ITCH, SUMO1, UBE2I [22][79][80][81][82][83][84][85], transcriptional co-regulators: HDAC2, EP300, KAT2B, HIPK2 [77][86][87][88][89][90], DNA repair gene BRCA1 that interacts with p63 [91] while repressing TAp73 expression and chemoresistance [92], DNA binding transcription factors: HIF1a, YAP, ATF3, p63, p73 [93][94][95]. Two proteins that are bound by p63 and p73 and not p53 are CDC20—a pivotal protein in the mitosis [96][97] and splicing regulating protein FUS [98][99].
It was shown that HDAC1 and HDAC2 depletion phenocopy p63 knockout in mice derepressing negatively regulated DNp63 targets CDKN1A(p21), SFN(14-3-3σ), and CDKN2A [37][100]. Interestingly, classical p53 target and cell cycle inhibitor CDKN1A [101] is expressed strictly in the suprabasal cells in human epithelial organs, especially in the esophagus and vagina. A p53/p73/CDKN1A inhibitor MDM2 is also in this group with unrestricted expression in all skin cells [22][84][102].
3.3. p73 Interactions
Analysis of 89 p73 or p53 interacting proteins revealed two putative protein complexes involved in the cell-cycle progression, chromatin modification complex and adaptive immunity complex (Figure 3C).
Analysis of 36 p73-only interacting proteins revealed clusters of interactors pointing to their potential role in the mitotic prometaphase, and resolution of the sister chromatin cohesion and segregation. These include protein of the spindle assembly checkpoint complex (Figure 3C, complex 1) (CCNB1, MAD2L1, Bub1B, STAG1, DSN1) [96][103][104][105][106][107][108].
In turn, 53 p73 interacting proteins that are also known to bind p53 but not p63, represent histone modifiers including: histone deacetylases: HDAC1, CHD3 [85][109], histone acetylases: KAT5, CREBBP [110][111], H3K4me2 or H3K9me2 lysine demethylase KDM1A [112]. The p73 and p53 interact with proteins involved in G1/S transition of the cell cycle (SP1, MYC, CCND1, RB1) [76][106][113][114][115] as well as proteins involved in the G2 to M transition Cyclin A and CDK1 [106][116]. Also, p73 and p53 interact with Aurora A kinase that represses p73 mediated transcriptional activation and SAC functions [96][117][118].
This group includes a classical p53/p73 target p21 (CDKN1A)—a major regulator of the G1/S cell cycle progression that acts by dephosphorylation and repression of multiple proteins involved in the cell cycle progression [101]. Accordingly its expression is restricted to the suprabasal layer of skin, vagina and other epithelial organs with no proliferation [101][119]. However, p21 protein does not interact with p73 directly, only p73 binding to the p21 promoter has been described, although p21 interacts with many proteins that do interact with p73, including SP1, RB1, CyclinD1 and CyclinA2.
Analysis of 58 p53/p73 interacting proteins that are highly expressed in the skin [55] revealed putative protein complexes involved in the cell cycle regulation and FOXM1 pathway: CCND1, CCNA2, CCNB1, CDK1, CDK2, Rb, transcriptional regulation: MYC, HDAC1, KAT5, CREBBP, cellular response to stress: HSPA4, HSPA9, NFYB, microtubule organizing center and G2/M transition:PLK1, AURKA.
3.4. p63 Interactors
Gene annotation of 187 p63-only interacting proteins reveals significant fraction of them being involved in RNA splicing and TGFb response. Chrondate and nervous system developmental pathways are highly represented in this group as well.
The TAp73 interacting with APC/C(CDC20) repress progression to anaphase and aberrant chromosome segregation, while APC/C(CDC20) interaction with DNp63a promotes DNp63a degradation and is required for keratinocytes differentiation [97]. Notably, p63 interacting APC/C components CDC20 and ANAPC2 are located strictly in a few cells of the basal and suprabasal layer, likely marking the last division of the cells committed to differentiation.
However, in the suprabasal layer, after keratinocytes differentiate, p63 protein is abundant suggesting that DNp63a isoform that is sensitive to the APC/C (CDC20, CDH1) mediated degradation is more restricted to the basal keratinocytes, whereas DNp63 present in the differentiated layers is likely other isoforms such as DNp63b and DNp63g [97].
Interestingly, one of the p63 interacting proteins is chaperonin containing TCP1, subunit 8 (theta) (CCT8) [98], that according to high throughput screening in turn interacts with gamma tubulin1 [120][121][122] and component of SAC complex BUB1 [123], suggesting yet unexplored implication of the p63 in the spindle checkpoint that might interact with or compete with p73.
Among proteins interacting with p63 are WWP1, C/EBPb, SMAD4 and KLF5. The WWP1 is an E3 ubiquitin ligase that is required for p63 transcriptional activity and WWP1 depletion causes cell cycle arrest [124]. Consistently, WWP1 displays an interesting staining pattern in skin, being nuclear in the basal cells and strictly peri-nuclear in the differentiated keratinocytes. Moreover, WWP1 ubiquitinates and leads to degradation and repression of KLF5 [125][126] that is highly expressed in the nuclei and cytoplasm of the basal and differentiated keratinocytes and of SMAD4 [127] that is localised to the cytoplasm in the epithelial tissue.
C/EBPb and C/EBPa are pivotal factors in the induction of genes during keratinocytes differentiation [61][73]. While, interaction between C/EBP and DNp63 leads to the induction of COX-2 expression in the lung epithelial cells upon smoke extract exposure [74], COX-2 induction in the skin keratinocytes leads to PGE2 induction and inflammation in response to ultraviolet irradiation [128]. Therefore, the functions of WWP1 interacting proteins is this subgroup are likely to be a switch of transcriptional regulation from basal to differentiated program.
Another protein interacting with p63 and WWP1 is KLF5. Both KLF5 and KlF4—a well-known pluripotency Yamanaka factor [129]—are expressed in skin with distinct staining patterns. The KLF5 decorates nuclear borders just like WWP1, whereas KLF4 is highly expressed throughout the differentiated cells. KLF4 is not known to interact directly with p63, however, it was shown that p63 regulates KLF4 expression and combined p63/KLF4 expression differentiate fibroblasts into keratinocytes [130][131]. In contrast, the role of p73 in reprogramming is controversial, because while one study used p73 depletion technique to demonstrate that p73 has no role in reprogramming [131], the other concluded that overexpression of DNp73β and KLF4 leads to reprogramming [42].
Common p63 and p53 interacting proteins relate to autophagy and protein folding (CHEK2, RELA, HSP90AB1) Figure 3D. Among p63 interacting proteins are several 60S ribosomal subunits including RPLP0, RPL3, RPL12, RPL13A, RPL24 as well as 40S ribosomal protein SA and UPF2 that controls nonsense-mediated mRNA decay [98]. In addition, p63 binds several human ribonucleoproteins HNRNPK, HNRNPL HNRNPA3, HNRNPR [98], HNRNPD [132] and HNRNPAB [133] suggesting that p63 isoforms function in the splicing regulation (Figure 3D).
3.5. p53 Interactions
It was not possible to annotate 971 unique p53 interactions by the Metascape. When we limited our analysis to 604 proteins that are detected in skin by the single cell RNA-seq [55], the Metascape recognized 11 putative protein complexes with functions ranging from the DNA repair to mRNA splicing and metabolism (Supplementary Excel File) consistently with known p53 functions in the genotoxic stress responses, transcription and RNA regulation pathways [121][134][135][136].
References
- Yang, A.; Kaghad, M.; Wang, Y.; Gillett, E.; Fleming, M.D.; Dötsch, V.; Andrews, N.C.; Caput, D.; McKeon, F. P63, a p53 homolog at 3q27-29, encodes multiple products with transactivating, death-inducing, and dominant-negative activities. Cell 1998, 2, 305–316, doi:10.1016/s1097-2765(00)80275-0.
- Kaghad, M.; Bonnet, H.; Yang, A.; Creancier, L.; Biscan, J.C.; Valent, A.; Minty, A.; Chalon, P.; Lelias, J.M.; Dumont, X.; et al. Monoallelically expressed gene related to p53 at 1p36, a region frequently deleted in neuroblastoma and other human cancers. Cell 1997, 90, 809–819, doi:10.1016/s0092-8674(00)80540-1.
- Scoumanne, A.; Harms, K.L.; Chen, X. Structural basis for gene activation by p53 family members. Cancer Ther. 2005, 4, 1178–1185, doi:10.4161/cbt.4.11.2254.
- Grob, T.J.; Novak, U.; Maisse, C.; Barcaroli, D.; Lüthi, A.U.; Pirnia, F.; Hügli, B.; Graber, H.U.; De Laurenzi, V.; Fey, M.F.; et al. Human delta Np73 regulates a dominant negative feedback loop for TAp73 and p53. Cell Death Differ. 2001, 8, 1213–1223, doi:10.1038/sj.cdd.4400962.
- Waltermann, A.; Kartasheva, N.N.; Dobbelstein, M. Differential regulation of p63 and p73 expression. Oncogene 2003, 22, 5686–5693, doi:10.1038/sj.onc.1206859.
- Nakagawa, T.; Takahashi, M.; Ozaki, T.; Watanabe, K.; Hayashi, S.; Hosoda, M.; Todo, S.; Nakagawara, A. Negative autoregulation of p73 and p53 by DeltaNp73 in regulating differentiation and survival of human neuroblastoma cells. Cancer Lett. 2003, 197, 105–109, doi:10.1016/s0304-3835(03)00090-9.
- Nakagawa, T.; Takahashi, M.; Ozaki, T.; Watanabe Ki, K.; Todo, S.; Mizuguchi, H.; Hayakawa, T.; Nakagawara, A. Autoinhibitory regulation of p73 by Delta Np73 to modulate cell survival and death through a p73-specific target element within the Delta Np73 promoter. Cell. Biol. 2002, 22, 2575–2585, doi:10.1128/mcb.22.8.2575-2585.2002.
- Lanza, M.; Marinari, B.; Papoutsaki, M.; Giustizieri, M.L.; D’Alessandra, Y.; Chimenti, S.; Guerrini, L.; Costanzo, A. Cross-talks in the p53 family: deltaNp63 is an anti-apoptotic target for deltaNp73alpha and p53 gain-of-function mutants. Cell Cycle 2006, 5, 1996–2004, doi:10.4161/cc.5.17.3188.
- Tophkhane, C.; Yang, S.-H.; Jiang, Y.; Ma, Z.; Subramaniam, D.; Anant, S.; Yogosawa, S.; Sakai, T.; Liu, W.-G.; Edgerton, S.; et al. P53 inactivation upregulates p73 expression through E2F-1 mediated transcription. PLoS ONE 2012, 7, e43564, doi:10.1371/journal.pone.0043564.
- Vikhreva, P.; Melino, G.; Amelio, I. P73 Alternative Splicing: Exploring a Biological Role for the C-Terminal Isoforms. Mol. Biol. 2018, 430, 1829–1838, doi:10.1016/j.jmb.2018.04.034.
- Nozell, S.; Wu, Y.; McNaughton, K.; Liu, G.; Willis, A.; Paik, J.C.; Chen, X. Characterization of p73 functional domains necessary for transactivation and growth suppression. Oncogene 2003, 22, 4333–4347, doi:10.1038/sj.onc.1206470.
- Liu, G.; Chen, X. The C-terminal sterile alpha motif and the extreme C terminus regulate the transcriptional activity of the alpha isoform of p73. Biol. Chem. 2005, 280, 20111–20119, doi:10.1074/jbc.M413889200.
- Toh, W.H.; Logette, E.; Corcos, L.; Sabapathy, K. TAp73beta and DNp73beta activate the expression of the pro-survival caspase-2S. Nucleic Acids Res. 2008, 36, 4498–4509, doi:10.1093/nar/gkn414.
- Koeppel, M.; van Heeringen, S.J.; Kramer, D.; Smeenk, L.; Janssen-Megens, E.; Hartmann, M.; Stunnenberg, H.G.; Lohrum, M. Crosstalk between c-Jun and TAp73alpha/beta contributes to the apoptosis-survival balance. Nucleic Acids Res. 2011, 39, 6069–6085, doi:10.1093/nar/gkr028.
- Vikhanskaya, F.; Toh, W.H.; Dulloo, I.; Wu, Q.; Boominathan, L.; Ng, H.H.; Vousden, K.H.; Sabapathy, K. P73 supports cellular growth through c-Jun-dependent AP-1 transactivation. Cell Biol. 2007, 9, 698–705, doi:10.1038/ncb1598.
- Tichý, V.; Navrátilová, L.; Adámik, M.; Fojta, M.; Brázdová, M. Redox state of p63 and p73 core domains regulates sequence-specific DNA binding. Biophys. Res. Commun. 2013, 433, 445–449, doi:10.1016/j.bbrc.2013.02.097.
- Patel, S.; Bui, T.T.T.; Drake, A.F.; Fraternali, F.; Nikolova, P.V. The p73 DNA binding domain displays enhanced stability relative to its homologue, the tumor suppressor p53, and exhibits cooperative DNA binding. Biochemistry 2008, 47, 3235–3244, doi:10.1021/bi7023207.
- Cai, B.-H.; Chao, C.-F.; Lu, M.-H.; Lin, H.-C.; Chen, J.-Y. A half-site of the p53-binding site on the keratin 14 promoter is specifically activated by p63. Biochem. 2012, 152, 99–110, doi:10.1093/jb/mvs053.
- Klein, C.; Georges, G.; Künkele, K.P.; Huber, R.; Engh, R.A.; Hansen, S. High thermostability and lack of cooperative DNA binding distinguish the p63 core domain from the homologous tumor suppressor p53. Biol. Chem. 2001, 276, 37390–37401, doi:10.1074/jbc.M103801200.
- Di Como, C.J.; Gaiddon, C.; Prives, C. P73 function is inhibited by tumor-derived p53 mutants in mammalian cells. Cell. Biol. 1999, 19, 1438–1449, doi:10.1128/mcb.19.2.1438.
- Si, H.; Lu, H.; Yang, X.; Mattox, A.; Jang, M.; Bian, Y.; Sano, E.; Viadiu, H.; Yan, B.; Yau, C.; et al. TNF-α modulates genome-wide redistribution of ΔNp63α/TAp73 and NF-κB cREL interactive binding on TP53 and AP-1 motifs to promote an oncogenic gene program in squamous cancer. Oncogene 2016, 35, 5781–5794, doi:10.1038/onc.2016.112.
- Zdzalik, M.; Pustelny, K.; Kedracka-Krok, S.; Huben, K.; Pecak, A.; Wladyka, B.; Jankowski, S.; Dubin, A.; Potempa, J.; Dubin, G. Interaction of regulators Mdm2 and Mdmx with transcription factors p53, p63 and p73. Cell Cycle 2010, 9, 4584–4591, doi:10.4161/cc.9.22.13871.
- Talos, F.; Nemajerova, A.; Flores, E.R.; Petrenko, O.; Moll, U.M. P73 suppresses polyploidy and aneuploidy in the absence of functional p53. Cell 2007, 27, 647–659, doi:10.1016/j.molcel.2007.06.036.
- Tomasini, R.; Tsuchihara, K.; Wilhelm, M.; Fujitani, M.; Rufini, A.; Cheung, C.C.; Khan, F.; Itie-Youten, A.; Wakeham, A.; Tsao, M.-S.; et al. TAp73 knockout shows genomic instability with infertility and tumor suppressor functions. Genes Dev. 2008, 22, 2677–2691, doi:10.1101/gad.1695308.
- Nicolai, S.; Rossi, A.; Di Daniele, N.; Melino, G.; Annicchiarico-Petruzzelli, M.; Raschellà, G. DNA repair and aging: The impact of the p53 family. Aging (Albany NY) 2015, 7, 1050–1065, doi:10.18632/aging.100858.
- Fujitani, M.; Sato, R.; Yamashita, T. Loss of p73 in ependymal cells during the perinatal period leads to aqueductal stenosis. Rep. 2017, 7, 12007, doi:10.1038/s41598-017-12105-z.
- Lin, Y.-L.; Sengupta, S.; Gurdziel, K.; Bell, G.W.; Jacks, T.; Flores, E.R. P63 and p73 transcriptionally regulate genes involved in DNA repair. PLoS Genet. 2009, 5, e1000680, doi:10.1371/journal.pgen.1000680.
- Liu, J.; Lin, M.; Zhang, C.; Wang, D.; Feng, Z.; Hu, W. TAp63γ enhances nucleotide excision repair through transcriptional regulation of DNA repair genes. DNA Repair (Amst) 2012, 11, 167–176, doi:10.1016/j.dnarep.2011.10.016.
- Zaika, E.; Wei, J.; Yin, D.; Andl, C.; Moll, U.; El-Rifai, W.; Zaika, A.I. P73 protein regulates DNA damage repair. FASEB J. 2011, 25, 4406–4414, doi:10.1096/fj.11-192815.
- Jackson, P.K.; Attardi, L.D. P73 and foxj1: Programming multiciliated epithelia. Trends Cell Biol. 2016, 26, 239–240, doi:10.1016/j.tcb.2016.03.001.
- Marshall, C.B.; Mays, D.J.; Beeler, J.S.; Rosenbluth, J.M.; Boyd, K.L.; Santos Guasch, G.L.; Shaver, T.M.; Tang, L.J.; Liu, Q.; Shyr, Y.; et al. P73 is required for multiciliogenesis and regulates the Foxj1-associated gene network. Cell Rep. 2016, 14, 2289–2300, doi:10.1016/j.celrep.2016.02.035.
- Fuertes-Alvarez, S.; Maeso-Alonso, L.; Villoch-Fernandez, J.; Wildung, M.; Martin-Lopez, M.; Marshall, C.; Villena-Cortes, A.J.; Diez-Prieto, I.; Pietenpol, J.A.; Tissir, F.; et al. P73 regulates ependymal planar cell polarity by modulating actin and microtubule cytoskeleton. Cell Death Dis. 2018, 9, 1183, doi:10.1038/s41419-018-1205-6.
- Yang, A.; Schweitzer, R.; Sun, D.; Kaghad, M.; Walker, N.; Bronson, R.T.; Tabin, C.; Sharpe, A.; Caput, D.; Crum, C.; et al. P63 is essential for regenerative proliferation in limb, craniofacial and epithelial development. Nature 1999, 398, 714–718, doi:10.1038/19539.
- Mills, A.A.; Zheng, B.; Wang, X.J.; Vogel, H.; Roop, D.R.; Bradley, A. P63 is a p53 homologue required for limb and epidermal morphogenesis. Nature 1999, 398, 708–713, doi:10.1038/19531.
- Lechler, T.; Fuchs, E. Asymmetric cell divisions promote stratification and differentiation of mammalian skin. Nature 2005, 437, 275–280, doi:10.1038/nature03922.
- Yi, R.; Poy, M.N.; Stoffel, M.; Fuchs, E. A skin microRNA promotes differentiation by repressing “stemness”. Nature 2008, 452, 225–229, doi:10.1038/nature06642.
- LeBoeuf, M.; Terrell, A.; Trivedi, S.; Sinha, S.; Epstein, J.A.; Olson, E.N.; Morrisey, E.E.; Millar, S.E. Hdac1 and Hdac2 act redundantly to control p63 and p53 functions in epidermal progenitor cells. Cell 2010, 19, 807–818, doi:10.1016/j.devcel.2010.10.015.
- Candi, E.; Rufini, A.; Terrinoni, A.; Dinsdale, D.; Ranalli, M.; Paradisi, A.; De Laurenzi, V.; Spagnoli, L.G.; Catani, M.V.; Ramadan, S.; et al. Differential roles of p63 isoforms in epidermal development: Selective genetic complementation in p63 null mice. Cell Death Differ. 2006, 13, 1037–1047, doi:10.1038/sj.cdd.4401926.
- Shalom-Feuerstein, R.; Lena, A.M.; Zhou, H.; De La Forest Divonne, S.; Van Bokhoven, H.; Candi, E.; Melino, G.; Aberdam, D. ΔNp63 is an ectodermal gatekeeper of epidermal morphogenesis. Cell Death Differ. 2011, 18, 887–896, doi:10.1038/cdd.2010.159.
- Poulson, N.D.; Lechler, T. Robust control of mitotic spindle orientation in the developing epidermis. Cell Biol. 2010, 191, 915–922, doi:10.1083/jcb.201008001.
- Yang, A.; Walker, N.; Bronson, R.; Kaghad, M.; Oosterwegel, M.; Bonnin, J.; Vagner, C.; Bonnet, H.; Dikkes, P.; Sharpe, A.; et al. P73-deficient mice have neurological, pheromonal and inflammatory defects but lack spontaneous tumours. Nature 2000, 404, 99–103, doi:10.1038/35003607.
- Beeler, J.S.; Marshall, C.B.; Gonzalez-Ericsson, P.I.; Shaver, T.M.; Santos Guasch, G.L.; Lea, S.T.; Johnson, K.N.; Jin, H.; Venters, B.J.; Sanders, M.E.; et al. P73 regulates epidermal wound healing and induced keratinocyte programming. PLoS ONE 2019, 14, e0218458, doi:10.1371/journal.pone.0218458.
- Wilhelm, M.T.; Rufini, A.; Wetzel, M.K.; Tsuchihara, K.; Inoue, S.; Tomasini, R.; Itie-Youten, A.; Wakeham, A.; Arsenian-Henriksson, M.; Melino, G.; et al. Isoform-specific p73 knockout mice reveal a novel role for delta Np73 in the DNA damage response pathway. Genes Dev. 2010, 24, 549–560, doi:10.1101/gad.1873910.
- Alonso-Lecue, P.; de Pedro, I.; Coulon, V.; Molinuevo, R.; Lorz, C.; Segrelles, C.; Ceballos, L.; López-Aventín, D.; García-Valtuille, A.; Bernal, J.M.; et al. Inefficient differentiation response to cell cycle stress leads to genomic instability and malignant progression of squamous carcinoma cells. Cell Death Dis. 2017, 8, e2901, doi:10.1038/cddis.2017.259.
- Freije, A.; Ceballos, L.; Coisy, M.; Barnes, L.; Rosa, M.; De Diego, E.; Blanchard, J.M.; Gandarillas, A. Cyclin E drives human keratinocyte growth into differentiation. Oncogene 2012, 31, 5180–5192, doi:10.1038/onc.2012.22.
- Gandarillas, A.; Sanz-Gómez, N.; Freije, A. Polyploidy and the mitosis path to epidermal cell fate. Cell Cycle 2019, 18, 359–362, doi:10.1080/15384101.2019.1568766.
- Zanet, J.; Freije, A.; Ruiz, M.; Coulon, V.; Sanz, J.R.; Chiesa, J.; Gandarillas, A. A mitosis block links active cell cycle with human epidermal differentiation and results in endoreplication. PLoS ONE 2010, 5, e15701, doi:10.1371/journal.pone.0015701.
- Gebel, J.; Luh, L.M.; Coutandin, D.; Osterburg, C.; Löhr, F.; Schäfer, B.; Frombach, A.-S.; Sumyk, M.; Buchner, L.; Krojer, T.; et al. Mechanism of TAp73 inhibition by ΔNp63 and structural basis of p63/p73 hetero-tetramerization. Cell Death Differ. 2016, 23, 1930–1940, doi:10.1038/cdd.2016.83.
- Aylon, Y.; Oren, M. P53: Guardian of ploidy. Oncol. 2011, 5, 315–323, doi:10.1016/j.molonc.2011.07.007.
- Thul, P.J.; Åkesson, L.; Wiking, M.; Mahdessian, D.; Geladaki, A.; Ait Blal, H.; Alm, T.; Asplund, A.; Björk, L.; Breckels, L.M.; et al. A subcellular map of the human proteome. Science 2017, 356, doi:10.1126/science.aal3321.
- Oughtred, R.; Stark, C.; Breitkreutz, B.-J.; Rust, J.; Boucher, L.; Chang, C.; Kolas, N.; O’Donnell, L.; Leung, G.; McAdam, R.; et al. The BioGRID interaction database: 2019 update. Nucleic Acids Res. 2019, 47, D529–D541, doi:10.1093/nar/gky1079.
- Micallef, L.; Rodgers, P. EulerAPE: Drawing area-proportional 3-Venn diagrams using ellipses. PLoS ONE 2014, 9, e101717, doi:10.1371/journal.pone.0101717.
- Zhou, Y.; Zhou, B.; Pache, L.; Chang, M.; Khodabakhshi, A.H.; Tanaseichuk, O.; Benner, C.; Chanda, S.K. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Commun. 2019, 10, 1523, doi:10.1038/s41467-019-09234-6.
- Bader, G.D.; Hogue, C.W.V. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinformatics 2003, 4, 2, doi:10.1186/1471-2105-4-2.
- Cheng, J.B.; Sedgewick, A.J.; Finnegan, A.I.; Harirchian, P.; Lee, J.; Kwon, S.; Fassett, M.S.; Golovato, J.; Gray, M.; Ghadially, R.; et al. Transcriptional programming of normal and inflamed human epidermis at single-cell resolution. Cell Rep. 2018, 25, 871–883, doi:10.1016/j.celrep.2018.09.006.
- Huang, K.-Y.; Lee, T.-Y.; Kao, H.-J.; Ma, C.-T.; Lee, C.-C.; Lin, T.-H.; Chang, W.-C.; Huang, H.-D. DbPTM in 2019: Exploring disease association and cross-talk of post-translational modifications. Nucleic Acids Res. 2019, 47, D298–D308, doi:10.1093/nar/gky1074.
- Finnegan, A.; Cho, R.J.; Luu, A.; Harirchian, P.; Lee, J.; Cheng, J.B.; Song, J.S. Single-cell transcriptomics reveals spatial and temporal turnover of keratinocyte differentiation regulators. Genet. 2019, 10, 775, doi:10.3389/fgene.2019.00775.
- Trost, A.; Bauer, J.W.; Lanschützer, C.; Laimer, M.; Emberger, M.; Hintner, H.; Onder, K. Rapid, high-quality and epidermal-specific isolation of RNA from human skin. Dermatol. 2007, 16, 185–190, doi:10.1111/j.1600-0625.2006.00534.x.
- Reimann, E.; Abram, K.; Kõks, S.; Kingo, K.; Fazeli, A. Identification of an optimal method for extracting RNA from human skin biopsy, using domestic pig as a model system. Rep. 2019, 9, 20111, doi:10.1038/s41598-019-56579-5.
- Santos Guasch, G.L.; Beeler, J.S.; Marshall, C.B.; Shaver, T.M.; Sheng, Q.; Johnson, K.N.; Boyd, K.L.; Venters, B.J.; Cook, R.S.; Pietenpol, J.A. P73 is required for ovarian follicle development and regulates a gene network involved in cell-to-cell adhesion. iScience 2018, 8, 236–249, doi:10.1016/j.isci.2018.09.018.
- Rozenberg, J.M.; Bhattacharya, P.; Chatterjee, R.; Glass, K.; Vinson, C. Combinatorial recruitment of CREB, C/EBPβ and c-Jun determines activation of promoters upon keratinocyte differentiation. PLoS ONE 2013, 8, e78179, doi:10.1371/journal.pone.0078179.
- Suda, N.; Itoh, T.; Nakato, R.; Shirakawa, D.; Bando, M.; Katou, Y.; Kataoka, K.; Shirahige, K.; Tickle, C.; Tanaka, M. Dimeric combinations of MafB, cFos and cJun control the apoptosis-survival balance in limb morphogenesis. Development 2014, 141, 2885–2894, doi:10.1242/dev.099150.
- Subramanian, D.; Bunjobpol, W.; Sabapathy, K. Interplay between TAp73 protein and selected activator protein-1 (AP-1) Family Members Promotes AP-1 target gene activation and cellular growth. Biol. Chem. 2015, 290, 18636–18649, doi:10.1074/jbc.M115.636548.
- Rozenberg, J.M.; Taylor, J.M.; Mack, C.P. RBPJ binds to consensus and methylated cis elements within phased nucleosomes and controls gene expression in human aortic smooth muscle cells in cooperation with SRF. Nucleic Acids Res. 2018, 46, 8232–8244, doi:10.1093/nar/gky562.
- Zhao, J.; Li, X.; Guo, M.; Yu, J.; Yan, C. The common stress responsive transcription factor ATF3 binds genomic sites enriched with p300 and H3K27ac for transcriptional regulation. BMC Genomics 2016, 17, 335, doi:10.1186/s12864-016-2664-8.
- Kouwenhoven, E.N.; Oti, M.; Niehues, H.; van Heeringen, S.J.; Schalkwijk, J.; Stunnenberg, H.G.; van Bokhoven, H.; Zhou, H. Transcription factor p63 bookmarks and regulates dynamic enhancers during epidermal differentiation. EMBO Rep. 2015, 16, 863–878, doi:10.15252/embr.201439941.
- Qu, J.; Yi, G.; Zhou, H. P63 cooperates with CTCF to modulate chromatin architecture in skin keratinocytes. Epigenetics Chromatin 2019, 12, 31, doi:10.1186/s13072-019-0280-y.
- Qu, J.; Tanis, S.E.J.; Smits, J.P.H.; Kouwenhoven, E.N.; Oti, M.; van den Bogaard, E.H.; Logie, C.; Stunnenberg, H.G.; van Bokhoven, H.; Mulder, K.W.; et al. Mutant p63 affects epidermal cell identity through rewiring the enhancer landscape. Cell Rep. 2018, 25, 3490-3503, doi:10.1016/j.celrep.2018.11.039.
- Deb, D.; Lanyi, A.; Scian, M.; Keiger, J.; Brown, D.R.; Le Roith, D.; Deb, S.P.; Deb, S. Differential modulation of cellular and viral promoters by p73 and p53. J. Oncol. 2001, 18, 401–409.
- Martin, A.G.; Trama, J.; Crighton, D.; Ryan, K.M.; Fearnhead, H.O. Activation of p73 and induction of Noxa by DNA damage requires NF-kappa B. Aging (Albany NY) 2009, 1, 335–349.
- Lu, H.; Yang, X.; Duggal, P.; Allen, C.T.; Yan, B.; Cohen, J.; Nottingham, L.; Romano, R.-A.; Sinha, S.; King, K.E.; et al. TNF-α promotes c-REL/ΔNp63α interaction and TAp73 dissociation from key genes that mediate growth arrest and apoptosis in head and neck cancer. Cancer Res. 2011, 71, 6867–6877, doi:10.1158/0008-5472.CAN-11-2460.
- De Laurenzi, V.; Rossi, A.; Terrinoni, A.; Barcaroli, D.; Levrero, M.; Costanzo, A.; Knight, R.A.; Guerrieri, P.; Melino, G. P63 and p73 transactivate differentiation gene promoters in human keratinocytes. Biophys. Res. Commun. 2000, 273, 342–346, doi:10.1006/bbrc.2000.2932.
- Rishi, V.; Bhattacharya, P.; Chatterjee, R.; Rozenberg, J.; Zhao, J.; Glass, K.; Fitzgerald, P.; Vinson, C. CpG methylation of half-CRE sequences creates C/EBPalpha binding sites that activate some tissue-specific genes. Natl. Acad. Sci. USA 2010, 107, 20311–20316, doi:10.1073/pnas.1008688107.
- Ratovitski, E.A. LKB1/PEA3/ΔNp63 pathway regulates PTGS-2 (COX-2) transcription in lung cancer cells upon cigarette smoke exposure. Med. Cell. Longev. 2010, 3, 317–324, doi:10.4161/oxim.3.5.13108.
- Innocente, S.A.; Lee, J.M. p73 is a p53-independent, Sp1-dependent repressor of cyclin B1 transcription. Biophys. Res. Commun. 2005, 329, 713–718, doi:10.1016/j.bbrc.2005.02.028.
- Ozaki, T.; Watanabe, K.; Nakagawa, T.; Miyazaki, K.; Takahashi, M.; Nakagawara, A. Function of p73, not of p53, is inhibited by the physical interaction with RACK1 and its inhibitory effect is counteracted by pRB. Oncogene 2003, 22, 3231–3242, doi:10.1038/sj.onc.1206382.
- Zhang, J.; Chen, X. DeltaNp73 modulates nerve growth factor-mediated neuronal differentiation through repression of TrkA. Cell. Biol. 2007, 27, 3868–3880, doi:10.1128/MCB.02112-06.
- Yang, A.; Zhu, Z.; Kettenbach, A.; Kapranov, P.; McKeon, F.; Gingeras, T.R.; Struhl, K. Genome-wide mapping indicates that p73 and p63 co-occupy target sites and have similar dna-binding profiles in vivo. PLoS ONE 2010, 5, e11572, doi:10.1371/journal.pone.0011572.
- Wu, H.; Zeinab, R.A.; Flores, E.R.; Leng, R.P. Pirh2, a ubiquitin E3 ligase, inhibits p73 transcriptional activity by promoting its ubiquitination. Cancer Res. 2011, 9, 1780–1790, doi:10.1158/1541-7786.MCR-11-0157.
- Oberst, A.; Malatesta, M.; Aqeilan, R.I.; Rossi, M.; Salomoni, P.; Murillas, R.; Sharma, P.; Kuehn, M.R.; Oren, M.; Croce, C.M.; et al. The Nedd4-binding partner 1 (N4BP1) protein is an inhibitor of the E3 ligase Itch. Natl. Acad. Sci. USA 2007, 104, 11280–11285, doi:10.1073/pnas.0701773104.
- Rossi, M.; Aqeilan, R.I.; Neale, M.; Candi, E.; Salomoni, P.; Knight, R.A.; Croce, C.M.; Melino, G. The E3 ubiquitin ligase Itch controls the protein stability of p63. Natl. Acad. Sci. USA 2006, 103, 12753–12758, doi:10.1073/pnas.0603449103.
- Huang, Y.-P.; Wu, G.; Guo, Z.; Osada, M.; Fomenkov, T.; Park, H.L.; Trink, B.; Sidransky, D.; Fomenkov, A.; Ratovitski, E.A. Altered sumoylation of p63alpha contributes to the split-hand/foot malformation phenotype. Cell Cycle 2004, 3, 1587–1596, doi:10.4161/cc.3.12.1290.
- Wang, X.; Arooz, T.; Siu, W.Y.; Chiu, C.H.; Lau, A.; Yamashita, K.; Poon, R.Y. MDM2 and MDMX can interact differently with ARF and members of the p53 family. FEBS Lett. 2001, 490, 202–208, doi:10.1016/s0014-5793(01)02124-x.
- Bálint, E.; Bates, S.; Vousden, K.H. Mdm2 binds p73 alpha without targeting degradation. Oncogene 1999, 18, 3923–3929, doi:10.1038/sj.onc.1202781.
- Minty, A.; Dumont, X.; Kaghad, M.; Caput, D. Covalent modification of p73alpha by SUMO-1. Two-hybrid screening with p73 identifies novel SUMO-1-interacting proteins and a SUMO-1 interaction motif. Biol. Chem. 2000, 275, 36316–36323, doi:10.1074/jbc.M004293200.
- Asatsuma-Okumura, T.; Ando, H.; De Simone, M.; Yamamoto, J.; Sato, T.; Shimizu, N.; Asakawa, K.; Yamaguchi, Y.; Ito, T.; Guerrini, L.; et al. P63 is a cereblon substrate involved in thalidomide teratogenicity. Chem. Biol. 2019, 15, 1077–1084, doi:10.1038/s41589-019-0366-7.
- Liao, J.-M.; Zhang, Y.; Liao, W.; Zeng, S.X.; Su, X.; Flores, E.R.; Lu, H. IκB kinase β (IKKβ) inhibits p63 isoform γ (TAp63γ) transcriptional activity. Biol. Chem. 2013, 288, 18184–18193, doi:10.1074/jbc.M113.466540.
- MacPartlin, M.; Zeng, S.; Lee, H.; Stauffer, D.; Jin, Y.; Thayer, M.; Lu, H. P300 regulates p63 transcriptional activity. Biol. Chem. 2005, 280, 30604–30610, doi:10.1074/jbc.M503352200.
- Strano, S.; Monti, O.; Pediconi, N.; Baccarini, A.; Fontemaggi, G.; Lapi, E.; Mantovani, F.; Damalas, A.; Citro, G.; Sacchi, A.; et al. The transcriptional coactivator Yes-associated protein drives p73 gene-target specificity in response to DNA Damage. Cell 2005, 18, 447–459, doi:10.1016/j.molcel.2005.04.008.
- Zhao, L.Y.; Liu, Y.; Bertos, N.R.; Yang, X.-J.; Liao, D. PCAF is a coactivator for p73-mediated transactivation. Oncogene 2003, 22, 8316–8329, doi:10.1038/sj.onc.1206916.
- Buckley, N.E.; Conlon, S.J.; Jirstrom, K.; Kay, E.W.; Crawford, N.T.; O’Grady, A.; Sheehan, K.; Mc Dade, S.S.; Wang, C.-W.; McCance, D.J.; et al. The DeltaNp63 proteins are key allies of BRCA1 in the prevention of basal-like breast cancer. Cancer Res. 2011, 71, 1933–1944, doi:10.1158/0008-5472.CAN-10-2717.
- Ibrahim, N.; He, L.; Leong, C.-O.; Xing, D.; Karlan, B.Y.; Swisher, E.M.; Rueda, B.R.; Orsulic, S.; Ellisen, L.W. BRCA1-associated epigenetic regulation of p73 mediates an effector pathway for chemosensitivity in ovarian carcinoma. Cancer Res. 2010, 70, 7155–7165, doi:10.1158/0008-5472.CAN-10-0668.
- Strano, S.; Munarriz, E.; Rossi, M.; Castagnoli, L.; Shaul, Y.; Sacchi, A.; Oren, M.; Sudol, M.; Cesareni, G.; Blandino, G. Physical interaction with Yes-associated protein enhances p73 transcriptional activity. Biol. Chem. 2001, 276, 15164–15173, doi:10.1074/jbc.M010484200.
- Senoo, M.; Matsumura, Y.; Habu, S. TAp63gamma (p51A) and dNp63alpha (p73L), two major isoforms of the p63 gene, exert opposite effects on the vascular endothelial growth factor (VEGF) gene expression. Oncogene 2002, 21, 2455–2465, doi:10.1038/sj.onc.1205330.
- Amelio, I.; Inoue, S.; Markert, E.K.; Levine, A.J.; Knight, R.A.; Mak, T.W.; Melino, G. TAp73 opposes tumor angiogenesis by promoting hypoxia-inducible factor 1α degradation. Natl. Acad. Sci. USA 2015, 112, 226–231, doi:10.1073/pnas.1410609111.
- Katayama, H.; Wang, J.; Treekitkarnmongkol, W.; Kawai, H.; Sasai, K.; Zhang, H.; Wang, H.; Adams, H.P.; Jiang, S.; Chakraborty, S.N.; et al. Aurora kinase-A inactivates DNA damage-induced apoptosis and spindle assembly checkpoint response functions of p73. Cancer Cell 2012, 21, 196–211, doi:10.1016/j.ccr.2011.12.025.
- Rokudai, S.; Li, Y.; Otaka, Y.; Fujieda, M.; Owens, D.M.; Christiano, A.M.; Nishiyama, M.; Prives, C. STXBP4 regulates APC/C-mediated p63 turnover and drives squamous cell carcinogenesis. Natl. Acad. Sci. USA 2018, 115, E4806–E4814, doi:10.1073/pnas.1718546115.
- Amoresano, A.; Di Costanzo, A.; Leo, G.; Di Cunto, F.; La Mantia, G.; Guerrini, L.; Calabrò, V. Identification of DeltaNp63alpha protein interactions by mass spectrometry. Proteome Res. 2010, 9, 2042–2048, doi:10.1021/pr9011156.
- Wang, T.; Jiang, X.; Chen, G.; Xu, J. Interaction of amyotrophic lateral sclerosis/frontotemporal lobar degeneration-associated fused-in-sarcoma with proteins involved in metabolic and protein degradation pathways. Aging 2015, 36, 527–535, doi:10.1016/j.neurobiolaging.2014.07.044.
- Su, X.; Cho, M.S.; Gi, Y.-J.; Ayanga, B.A.; Sherr, C.J.; Flores, E.R. Rescue of key features of the p63-null epithelial phenotype by inactivation of Ink4a and Arf. EMBO J. 2009, 28, 1904–1915, doi:10.1038/emboj.2009.151.
- Engeland, K. Cell cycle arrest through indirect transcriptional repression by p53: I have a DREAM. Cell Death Differ. 2018, 25, 114–132, doi:10.1038/cdd.2017.172.
- Enge, M.; Bao, W.; Hedström, E.; Jackson, S.P.; Moumen, A.; Selivanova, G. MDM2-dependent downregulation of p21 and hnRNP K provides a switch between apoptosis and growth arrest induced by pharmacologically activated p53. Cancer Cell 2009, 15, 171–183, doi:10.1016/j.ccr.2009.01.019.
- Tomasini, R.; Tsuchihara, K.; Tsuda, C.; Lau, S.K.; Wilhelm, M.; Rufini, A.; Tsao, M.; Iovanna, J.L.; Jurisicova, A.; Melino, G.; et al. TAp73 regulates the spindle assembly checkpoint by modulating BubR1 activity. Natl. Acad. Sci. USA 2009, 106, 797–802, doi:10.1073/pnas.0812096106.
- Gebel, J.; Tuppi, M.; Krauskopf, K.; Coutandin, D.; Pitzius, S.; Kehrloesser, S.; Osterburg, C.; Dötsch, V. Control mechanisms in germ cells mediated by p53 family proteins. Cell Sci. 2017, doi:10.1242/jcs.204859.
- Mikulenkova, E.; Neradil, J.; Zitterbart, K.; Sterba, J.; Veselska, R. Overexpression of the ∆Np73 isoform is associated with centrosome amplification in brain tumor cell lines. Tumour Biol. 2015, 36, 7483–7491, doi:10.1007/s13277-015-3474-3.
- Gaiddon, C.; Lokshin, M.; Gross, I.; Levasseur, D.; Taya, Y.; Loeffler, J.-P.; Prives, C. Cyclin-dependent kinases phosphorylate p73 at threonine 86 in a cell cycle-dependent manner and negatively regulate p73. Biol. Chem. 2003, 278, 27421–27431, doi:10.1074/jbc.M300251200.
- Lunardi, A.; Di Minin, G.; Provero, P.; Dal Ferro, M.; Carotti, M.; Del Sal, G.; Collavin, L. A genome-scale protein interaction profile of Drosophila p53 uncovers additional nodes of the human p53 network. Natl. Acad. Sci. USA 2010, 107, 6322–6327, doi:10.1073/pnas.1002447107.
- Huttlin, E.L.; Ting, L.; Bruckner, R.J.; Gebreab, F.; Gygi, M.P.; Szpyt, J.; Tam, S.; Zarraga, G.; Colby, G.; Baltier, K.; et al. The bioplex network: A systematic exploration of the human interactome. Cell 2015, 162, 425–440, doi:10.1016/j.cell.2015.06.043.
- Uramoto, H.; Wetterskog, D.; Hackzell, A.; Matsumoto, Y.; Funa, K. P73 competes with co-activators and recruits histone deacetylase to NF-Y in the repression of PDGF beta-receptor. Cell Sci. 2004, 117, 5323–5331, doi:10.1242/jcs.01384.
- Lemasson, I.; Nyborg, J.K. Human T-cell leukemia virus type I tax repression of p73beta is mediated through competition for the C/H1 domain of CBP. Biol. Chem. 2001, 276, 15720–15727, doi:10.1074/jbc.M100131200.
- Kim, J.-W.; Song, P.I.; Jeong, M.-H.; An, J.-H.; Lee, S.-Y.; Jang, S.-M.; Song, K.-H.; Armstrong, C.A.; Choi, K.-H. TIP60 represses transcriptional activity of p73beta via an MDM2-bridged ternary complex. Biol. Chem. 2008, 283, 20077–20086, doi:10.1074/jbc.M800161200.
- Tsai, W.-W.; Nguyen, T.T.; Shi, Y.; Barton, M.C. p53-targeted LSD1 functions in repression of chromatin structure and transcription in vivo. Cell. Biol. 2008, 28, 5139–5146, doi:10.1128/MCB.00287-08.
- Chau, B.N.; Chen, T.-T.; Wan, Y.Y.; DeGregori, J.; Wang, J.Y.J. Tumor necrosis factor alpha-induced apoptosis requires p73 and c-ABL activation downstream of RB degradation. Cell. Biol. 2004, 24, 4438–4447, doi:10.1128/mcb.24.10.4438-4447.2004.
- Koutsodontis, G.; Vasilaki, E.; Chou, W.-C.; Papakosta, P.; Kardassis, D. Physical and functional interactions between members of the tumour suppressor p53 and the Sp families of transcription factors: Importance for the regulation of genes involved in cell-cycle arrest and apoptosis. J. 2005, 389, 443–455, doi:10.1042/BJ20041980.
- Uramoto, H.; Izumi, H.; Ise, T.; Tada, M.; Uchiumi, T.; Kuwano, M.; Yasumoto, K.; Funa, K.; Kohno, K. P73 Interacts with c-Myc to regulate Y-box-binding protein-1 expression. Biol. Chem. 2002, 277, 31694–31702, doi:10.1074/jbc.M200266200.
- Fulco, M.; Costanzo, A.; Merlo, P.; Mangiacasale, R.; Strano, S.; Blandino, G.; Balsano, C.; Lavia, P.; Levrero, M. P73 is regulated by phosphorylation at the G2/M transition. Biol. Chem. 2003, 278, 49196–49202, doi:10.1074/jbc.M304921200.
- Dar, A.A.; Belkhiri, A.; Ecsedy, J.; Zaika, A.; El-Rifai, W. Aurora kinase A inhibition leads to p73-dependent apoptosis in p53-deficient cancer cells. Cancer Res. 2008, 68, 8998–9004, doi:10.1158/0008-5472.CAN-08-2658.
- Tentler, J.J.; Ionkina, A.A.; Tan, A.C.; Newton, T.P.; Pitts, T.M.; Glogowska, M.J.; Kabos, P.; Sartorius, C.A.; Sullivan, K.D.; Espinosa, J.M.; et al. P53 family members regulate phenotypic response to aurora kinase A inhibition in triple-negative breast cancer. Cancer Ther. 2015, 14, 1117–1129, doi:10.1158/1535-7163.MCT-14-0538-T.
- Kim, E.M.; Jung, C.-H.; Kim, J.; Hwang, S.-G.; Park, J.K.; Um, H.-D. The p53/p21 complex regulates cancer cell invasion and apoptosis by targeting Bcl-2 family proteins. Cancer Res. 2017, 77, 3092–3100, doi:10.1158/0008-5472.CAN-16-2098.
- Hein, M.Y.; Hubner, N.C.; Poser, I.; Cox, J.; Nagaraj, N.; Toyoda, Y.; Gak, I.A.; Weisswange, I.; Mansfeld, J.; Buchholz, F.; et al. A human interactome in three quantitative dimensions organized by stoichiometries and abundances. Cell 2015, 163, 712–723, doi:10.1016/j.cell.2015.09.053.
- Fogeron, M.-L.; Müller, H.; Schade, S.; Dreher, F.; Lehmann, V.; Kühnel, A.; Scholz, A.-K.; Kashofer, K.; Zerck, A.; Fauler, B.; et al. LGALS3BP regulates centriole biogenesis and centrosome hypertrophy in cancer cells. Commun. 2013, 4, 1531, doi:10.1038/ncomms2517.
- Liu, X.; Salokas, K.; Tamene, F.; Jiu, Y.; Weldatsadik, R.G.; Öhman, T.; Varjosalo, M. An AP-MS- and BioID-compatible MAC-tag enables comprehensive mapping of protein interactions and subcellular localizations. Commun. 2018, 9, 1188, doi:10.1038/s41467-018-03523-2.
- Huttlin, E.L.; Bruckner, R.J.; Paulo, J.A.; Cannon, J.R.; Ting, L.; Baltier, K.; Colby, G.; Gebreab, F.; Gygi, M.P.; Parzen, H.; et al. Architecture of the human interactome defines protein communities and disease networks. Nature 2017, 545, 505–509, doi:10.1038/nature22366.
- Peschiaroli, A.; Scialpi, F.; Bernassola, F.; El Sherbini, E.S.; Melino, G. The E3 ubiquitin ligase WWP1 regulates ΔNp63-dependent transcription through Lys63 linkages. Biophys. Res. Commun. 2010, 402, 425–430, doi:10.1016/j.bbrc.2010.10.050.
- Chen, C.; Sun, X.; Guo, P.; Dong, X.-Y.; Sethi, P.; Cheng, X.; Zhou, J.; Ling, J.; Simons, J.W.; Lingrel, J.B.; et al. Human Kruppel-like factor 5 is a target of the E3 ubiquitin ligase WWP1 for proteolysis in epithelial cells. Biol. Chem. 2005, 280, 41553–41561, doi:10.1074/jbc.M506183200.
- Ge, F.; Chen, W.; Qin, J.; Zhou, Z.; Liu, R.; Liu, L.; Tan, J.; Zou, T.; Li, H.; Ren, G.; et al. Ataxin-3 like (ATXN3L), a member of the Josephin family of deubiquitinating enzymes, promotes breast cancer proliferation by deubiquitinating Krüppel-like factor 5 (KLF5). Oncotarget 2015, 6, 21369–21378, doi:10.18632/oncotarget.4128.
- Morén, A.; Imamura, T.; Miyazono, K.; Heldin, C.-H.; Moustakas, A. Degradation of the tumor suppressor Smad4 by WW and HECT domain ubiquitin ligases. Biol. Chem. 2005, 280, 22115–22123, doi:10.1074/jbc.M414027200.
- Tripp, C.S.; Blomme, E.A.G.; Chinn, K.S.; Hardy, M.M.; LaCelle, P.; Pentland, A.P. Epidermal COX-2 induction following ultraviolet irradiation: Suggested mechanism for the role of COX-2 inhibition in photoprotection. Invest. Dermatol. 2003, 121, 853–861, doi:10.1046/j.1523-1747.2003.12495.x.
- Takahashi, K.; Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006, 126, 663–676, doi:10.1016/j.cell.2006.07.024.
- Chen, Y.; Mistry, D.S.; Sen, G.L. Highly rapid and efficient conversion of human fibroblasts to keratinocyte-like cells. Invest. Dermatol. 2014, 134, 335–344, doi:10.1038/jid.2013.327.
- Alexandrova, E.M.; Petrenko, O.; Nemajerova, A.; Romano, R.A.; Sinha, S.; Moll, U.M. ΔNp63 regulates select routes of reprogramming via multiple mechanisms. Cell Death Differ. 2013, 20, 1698–1708, doi:10.1038/cdd.2013.122.
- Kumar, M.; Matta, A.; Masui, O.; Srivastava, G.; Kaur, J.; Thakar, A.; Shukla, N.K.; RoyChoudhury, A.; Sharma, M.; Walfish, P.G.; et al. Nuclear heterogeneous nuclear ribonucleoprotein D is associated with poor prognosis and interactome analysis reveals its novel binding partners in oral cancer. Transl. Med. 2015, 13, 285, doi:10.1186/s12967-015-0637-3.
- Fomenkov, A.; Huang, Y.-P.; Topaloglu, O.; Brechman, A.; Osada, M.; Fomenkova, T.; Yuriditsky, E.; Trink, B.; Sidransky, D.; Ratovitski, E. P63 alpha mutations lead to aberrant splicing of keratinocyte growth factor receptor in the Hay-Wells syndrome. Biol. Chem. 2003, 278, 23906–23914, doi:10.1074/jbc.M300746200.
- Wu, C.; Cui, Y.; Liu, X.; Zhang, F.; Lu, L.-Y.; Yu, X. The RNF20/40 complex regulates p53-dependent gene transcription and mRNA splicing. Mol. Cell Biol. 2020, 12, 113–124, doi:10.1093/jmcb/mjz045.
- Shekhar, S.; Dey, S. Induction of p73, Δ133p53, Δ160p53, pAKT lead to neuroprotection via DNA repair by 5-LOX inhibition. Biol. Rep. 2020, 47, 269–274, doi:10.1007/s11033-019-05127-5.
- Hall, C.; Muller, P.A.J. The diverse functions of mutant 53, its family members and isoforms in cancer. J. Mol. Sci. 2019, 20, doi:10.3390/ijms20246188.