| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Sorina Dinescu | + 4364 word(s) | 4364 | 2020-11-23 04:13:23 |
Video Upload Options
Considered to be highly lethal if not diagnosed in early stages, cutaneous malignant melanoma is among the most aggressive and treatment-resistant human cancers, and its incidence continues to rise, largely due to ultraviolet radiation exposure, which is the main carcinogenic factor. Over the years, researchers have started to unveil the molecular mechanisms by which malignant melanoma can be triggered and sustained, in order to establish specific, reliable biomarkers that could aid the prognosis and diagnosis of this fatal disease, and serve as targets for development of novel efficient therapies. The high mutational burden and heterogeneous nature of melanoma shifted the main focus from the genetic landscape to epigenetic and epitranscriptomic modifications, aiming at elucidating the role of non-coding RNA molecules (ncRNAs) in the fine tuning of melanoma progression. Studies have shown that ncRNAs, among them microRNAs and lncRNAs, play a role in melanoma invasion, metastasis and acquired resistance to treatment. In addition, they could serve as prognostic/diagnostic biomarkers and potential targets for promising therapeutic strategies.
1. Introduction
Non-coding RNAs (ncRNAs) have been observed to tamper with the efficiency of molecular therapies, in addition to facilitating melanoma invasion and metastasis [1][2]. They represent important epigenetic and epitranscriptomic regulators, classified and differentiated into long (lncRNAs) and small (sncRNAs) molecules, according to their length which can be longer or smaller than 200 bps. While multiple types of RNAs can be found in the small category, particular interest has been devoted to studying the deregulation of microRNAs (miRNAs/miR) in melanoma [3]. miRNAs can modulate the gene expression at post-transcriptional level mostly by interfering with the 3′UTR region of mRNAs, and substantial evidence points to their involvement in each stage of melanoma progression [1][4]. In contrast, the epigenetic control exerted by lncRNAs depends on the recruitment of regulatory proteins at specific DNA target regions, to silence or activate gene promoters, and their role in malignant melanoma is more elusive [5][6]. Some lncRNAs have been described to stand as decoys or sequesters of miRNAs, while others play a role in stabilizing the translational ribosomal machinery [7].
2. MicroRNAs Modulate Melanoma Invasion and Metastasis
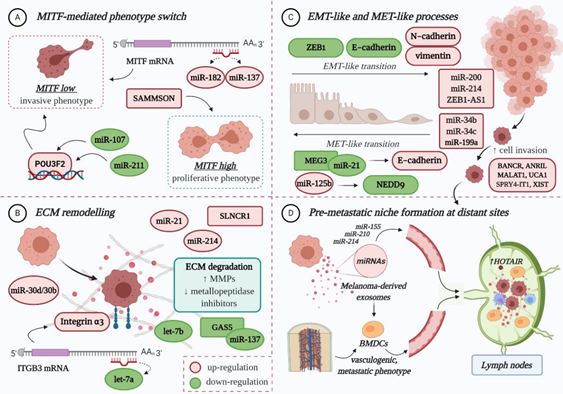
The role of miRNAs in melanoma progression is very tangled, therefore excellent reviews can be consulted here: [1][8][9][10]. In principle, miRNAs influence the evolution of melanoma to secondary sites mainly by: (A) regulating the expression of MITF-M, an essential melanocyte-and melanoma-specific transcription factor that operates as a “switch” in the establishment of an invasive or proliferative phenotype; (B) remodelling the extracellular matrix (ECM); (C) promoting epithelial-to-mesenchymal transition (EMT) and its reverse, mesenchymal-to-epithelial transition (MET); and (D) preparing the formation of the pre-metastatic niche [1][9][11][12][13] (Figure 1).
Figure 1. The role of non-coding RNAs in melanoma invasion and metastasis: (A) Both miRNAs and lncRNAs modulate the expression of MITF-M, directly or indirectly, therefore influencing the switch towards an invasive or proliferative phenotype; (B) The deregulated expression of ncRNAs facilitates ECM remodelling, by targeting integrins, increasing matrix metalloproteinases (MMPs) and diminishing the metallopeptidase inhibitors (TIMPs), therefore promoting ECM degradation and decreasing tumour cell-matrix adhesion; (C) Multiple miRNAs and lncRNAs favour the EMT-like process of melanoma cells, which enhances their migratory capacity, mainly by affecting the expression of specific adhesion molecules. Some ncRNAs also promote the MET-like process of malignant melanocytes, which leads to enhanced proliferation and colony formation, thus favouring metastasis; (D) The formation of a pre-metastatic niche at distant sites is mainly influenced by melanoma-derived exosomes, enriched in miRNA cargo (e.g., miR-155, miR-210, miR-214). In sentinel lymph nodes, they instigate microenvironmental changes that facilitate the recruitment, trapping and growth of circulating tumour cells. They also prime bone-marrow-derived cells (BMDCs), which achieve a vasculogenic, metastatic phenotype, and facilitate their recruitment to distant sites, where BMDCs will contribute to the formation of a pre-metastatic niche. LncRNA HOTAIR is enriched in lymph node metastases compared to primary tumours, which may suggest a role in the pre-metastatic niche formation. POU3F2: POU-domain class 3 transcription factor 2; NEDD9: Neural precursor cell expressed developmentally down-regulated protein 9; ZEB1: zinc finger E-box-binding homeobox 1. Colour code: red for upregulation and green for downregulation. Figure created with BioRender.com.
Reduced expression of MITF-M in melanoma cells determines the acquisition of an invasive phenotype [12]. Several miRNAs have been found to regulate the activity of this lineage-restricted gene at a post-transcriptional level, among them miR-182, miR-137, miR-211 and miR-107 [13][14][15][16][17]. Overexpression of miR-182 stimulates melanoma cell migration and invasion through the direct downregulation of MITF and FOXO3 expression; studies on melanoma cell lines and tissue samples have found that its expression increases gradually from primary to metastatic stage [15]. Similarly, high levels of miR-137 expression have been correlated with poor survival in advanced melanoma patients [18]. In contrast, decreased expression of miR-211 can be observed in highly invasive melanoma cell lines [19]. Normally, miR-211 prevents the loss of cell adhesion through the negative modulation of NUAK1 [20], thereby inhibiting the migratory and invasive capacity of melanoma cells [21]. Additionally, it has been proposed that miR-211 can transcriptionally repress POU3F2 (POU-domain class 3 transcription factor 2), also known as BRN2 (brain-specific homeobox 2), which is a known suppressor of MITF. As such, loss of miR-211 can increase the expression of BRN2, and therefore inhibit MITF, ensuring that malignant melanocytes are kept in a pro-invasive, dedifferentiated state [16]. Recently, Zhao et al. reported the downregulation of miR-107, another tumour suppressor that inhibits melanoma cell invasion by targeting POU3F2 [17]. In BRAF-mutant cells, overexpression of BRN2 contributes to cytoskeletal rearrangement and increased cell invasion [22].
Several miRNAs play a role in ECM remodelling [23]. In melanoma, loss of let-7a, which is a negative regulator of ITGB3, leads to elevated levels of integrin α3 [24], while suppression of let-7b indirectly enhances the production of matrix metalloproteinase (MMP)-9 [25], facilitating collagen degradation and cell invasion. Meanwhile, miR-21 is thought to induce an invasive phenotype in melanoma cells by targeting the mRNA of tissue inhibitor of metalloproteinases (TIMP)-3 [26]. Lastly, the increased expression of miR-30d/miR-30b cluster stimulates the invasive and metastatic potential of melanoma cells, both in vitro and in vivo, potentially by silencing GALNT7 (polypeptide N-acetylgalactosaminyltransferase 7), which strongly affects the O-glycosylation of membrane proteins, and subsequent interaction with the ECM [27].
True EMT in melanoma is not possible as melanocytes are not epithelial cells, however some miRNAs are implicated in an EMT-like process that promotes the invasion and metastasis of tumour cells. One of them is the cluster miR-224/miR-452, which silences a metastatic suppressor that normally inhibits E2F1, and which has been shown to facilitate the cytoskeletal rearrangement of less aggressive cells, resulting in increased migratory and invasive propensity [28]. Also considered of particular relevance, the miR-200 family (miR-200a, miR-200b, miR-200c, and miR-141) drives the EMT-like process by upregulating the expression of Bmi-1 oncogene, which in turn promotes the activation of PI3K/AKT and MAPK cascades. This negatively influences the expression of ZEB1 (zinc finger E-box-binding homeobox 1) and E-Cadherin, at the same time stimulating N-Cadherin and vimentin expression [29]. High levels of miR-214 have also been associated with enhanced cell motility and metastatic potential, as this specific miRNA indirectly downregulates the expression of some adhesion molecules (MCAM, E-cadherin) and metallopeptidase inhibitors (TIMP1, TIMP2) [30].
The MET-like process of malignant melanocytes is favoured by miR-125b overexpression, who directly targets a transcript of NEDD9 (neural precursor cell expressed developmentally down-regulated protein 9) [31][32], and whose in vitro inhibition was found to decrease the invasive potential of aggressive melanoma cells [31]. Other miRNAs (miR-34b, miR-34c, and miR-199a-3p) also contribute to this mesenchymal movement by targeting the mRNA of tyrosine-protein kinase Met (c-MET), whose increased expression facilitates melanoma cell migration and metastasis [33][34].
miRNAs also participate in the formation of a pre-metastatic niche at distant organs that will enable the implantation and survival of tumour cells. Melanoma-derived exosomes, with their enriched miRNA cargo (e.g., miR-155, miR-210, miR-214), are primarily involved in this intercellular communication (for review: [35][36]). For instance, exosomes recruited into sentinel lymph nodes promote the upregulation of proteases that degrade the ECM and enhance the expression of pro-angiogenic factors (TNFα, VEGF, etc.) to facilitate the recruitment, trapping and growth of malignant melanocytes within the metastatic niche [37][38]. They are also capable of priming bone-marrow-derived cells (BMDCs) to achieve a vasculogenic, metastatic phenotype, and facilitate their recruitment to metastatic sites where they will contribute to the establishment of a suitable microenvironment for trapping circulating melanoma cells [39].
3. LncRNAs Modulate Melanoma Invasion and Metastasis
Despite the emerging interest and growing evidence of the contribution of lncRNAs in cancer [40][41], little is known about the impact of deregulated expression of lncRNAs in the invasion and metastasis of MM [2][42] (Figure 1).
The first lncRNA characterized was SPRY4-IT1, a transcript derived from an intron of the SPRY4 gene, whose expression is increased in melanoma [43]. It was identified as a regulator of several processes, as suppression of SPRY4-IT1 resulted in abnormal cell growth, differentiation and apoptosis, as well as decreased invasion capacity of melanoma cell lines [43][44]. Although the molecular mechanisms that interfere with the invasion of MM are not clear, a study concerning non-small cell lung carcinoma revealed a possible role in the activation of EMT by modulating both E-cadherin and vimentin expression, leading to cell proliferation and metastasis [45]. Interestingly, another lncRNA with a potential role in regulating EMT was discovered by Siena et al., while exploring the lncRNA gene expression patterns across melanocytes, primary and metastatic melanoma cells. They found a significant upregulation of ZEB1 antisense RNA 1 (ZEB1-AS1) in melanoma cells. Data analysis from TCGA confirmed the increased expression of ZEB1-AS1 in metastatic melanoma and its association with hotspot mutations in BRAF and RAS family genes. Additionally, enrichment analysis correlated ZEB1-AS1 with the gene expression of zinc finger E-box binding homeobox 1 (ZEB1), an essential EMT marker, suggesting a possible role in melanoma invasiveness and phenotype switching [46].
TCGA data analysis also confirmed the clinical relevance of SRA-like non-coding RNA1 (SLNCR1), a lncRNA whose highly conserved sequence is strikingly similar to that of lncRNA steroid receptor RNA activator 1 (SRA1), and whose increased expression is associated with shorter overall survival in melanoma patients. Functional and mechanistic studies revealed SLNCR1 promotes melanoma invasion through upregulation of MMP9 (involved in ECM degradation) in cooperation with the brain-specific homeobox protein 3a (Brn3a) and the androgen receptor (AR) [47].
A screen of differentially expressed lncRNAs in BRAF V600E mutated melanoma cells has led to the identification of BRAF-activated non-coding RNA (BANCR) as a putative regulator of cell proliferation and motility. Overexpression of BANCR in MM promotes the activation of the extracellular signal-regulated kinases 1/2 (ERK1/2) and c-Jun N-terminal kinase (JNK) components of the MAPK pathway, while its knockdown affects the migratory capacity of tumour cells [48][49]. A positive feedback mechanism with the V600E mutation is thought to induce BANCR overexpression, subsequent upregulation of chemokines and increase in melanoma cell motility [49][50].
The oncogenic activity of antisense non-coding RNA in the INK4A locus (ANRIL) in melanoma revolves around the regulation of its encoding locus that also harbours the tumour suppressor genes INK4A and INK4B. In cutaneous melanoma tissue samples and cell lines, strong levels of ANRIL negatively modulated the expression of CDKN2A/2B proteins [51][52]. In addition, knockdown experiments managed to restore INK4A and INK4B expression, at the same time suppressing in vitro colony formation and migration [53]. Meanwhile, the encoding gene of survival associated mitochondrial melanoma-specific oncogenic lncRNA (SAMMSON) also harbours the melanoma-specific oncogene MITF and it was demonstrated that SAMMSON is frequently co-amplified with it [54]. As shown by functional assays, knockdown of this lineage-specific lncRNA drastically affects cell proliferation and viability, even sensitizing melanoma to MAPK-targeted drugs. Mechanistically, SAMMSON interacts with mitochondrial protein p32, a critical regulator of tumour metabolism [54].
Although the molecular mechanisms are currently unknown, HOX transcript antisense intergenic RNA (HOTAIR) is perceived as a regulator of melanoma invasion and metastatic progression, considering it was found particularly enriched in lymph-node metastases compared to primary lesions and knockdown of HOTAIR in vitro suppressed melanoma cell motility and invasion [55]. In a similar manner, a significantly higher expression of metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) and urothelial carcinoma associated 1 (UCA1) was observed in advanced stages of MM, than in early stages, suggesting the potential role of MALAT1 and UCA1 in melanoma invasion and metastasis [56]. Moreover, a subsequent mechanistic study demonstrated that UCA1 can interact with miR-507 and promote the inhibition of FOXM1 expression, leading to increased invasiveness and clonogenic potential of melanoma cells [57].
Recently, the upregulated expression of X-inactive-specific transcript (XIST) in MM tissues and resistant cell lines was reported, with XIST being proposed as a crucial regulator in melanoma progression [58]. Through the use of bioinformatics, XIST was revealed to act as a molecular sponge for miR-21, which targets PI3KR1, a regulatory subunit of PI3K. Functional studies concerning XIST repressed PI3KR1 and AKT expression, leading to inhibition of melanoma cell proliferation and migration, at the same time increasing sensitivity to oxaliplatin [58].
Lastly, lncRNAs can also act as tumour suppressors in melanoma, however only growth arrest-specific transcript 5 (GAS5) and maternally expressed gene 3 (MEG3) have been reported so far [59][60]. Lentiviral-mediated overexpression of GAS5 diminished the expression of MMP2, a specific protein involved in ECM degradation, and reduced the migratory and invasive capacity of human MM cells [61]. Interestingly, GAS5 seems to repress melanoma tumorigenesis via miR-137, while MEG3 inhibits tumour growth and metastasis by modulating miR-21/E-cadherin axis [60][62].
4. Non-Coding RNAs in Drug Resistance
Continuous treatment with target-based therapies remains unsuccessful and leads to melanoma relapse due to acquisition of drug resistance. Even though BRAF monotherapy (BRAFi), or its combination with MEK inhibitors (MAPKi) instigate a profound regression in patients with BRAF-mutated metastatic melanoma, their effect is only temporary [63][64]. The same issue arises with the use of immunotherapy [65]. The role of ncRNAs in the development of melanoma drug resistance has been questioned and while the contribution of miRNAs is well studied [10], the role of lncRNAs in such resistance is still largely unknown.
4.1. Resistance to BRAF or MAPK Inhibitors
The deregulated expression of several miRNAs, namely upregulation of onco-miRNAs and downregulation of oncosuppressors, has been shown to contribute to the acquisition of drug resistance to BRAFi or MAPKi-based therapy in melanoma (Table 1). For example, overexpression of miR-31a, miR-100 and miR-125b stimulates tumour cell proliferation and apoptosis escape, decreasing drug sensitivity in patients treated with vemurafenib. Their expression is associated with the chemokine monocyte chemoattractant protein-1 (CCL2), which promotes melanoma progression in BRAFi-resistant cells [66]. Inhibition of miR-125a leads to the partial drug resensitization of melanoma in a subset of BRAFi-resistant cell lines [67]. Also, it has been concluded that miR-204 and miR-211 facilitate the emergence of resistance to vemurafenib, as higher expression levels were found in resistant melanoma cells compared to their drug-naïve counterparts [68][69]. On top of that, low levels of miR-579-3p can affect not only the BRAF/MAPK signalling pathway, but also the MDM2/p53 pathway, causing uncontrolled cell proliferation and migration, coupled with inhibition of apoptosis, thus contributing to the development of MAPKi resistance [70]. Of note, overexpression of miR-579-3p was able to prevent colony formation in cells exposed to vemurafenib and impair the growth of BRAFi-resistant melanoma cells in combination with the MEK inhibitor (MEKi) trametinib [70].
Recently, Sanlorenzo et al. identified MIRAT (MAPK Inhibitor Resistance Associated Transcript), a novel cytoplasmic lncRNA, which is significantly overexpressed in melanoma cells with acquired resistance to MAPKi, and modulates MAPK signalling by binding to the MEK scaffold protein IQGAP1 [71]. Knockdown of SAMMSON was shown to sensitize melanoma to MAPK-targeting therapeutics as well, but its underlying mechanism is unknown [54]. Meanwhile, Joung et al. performed a genome-scale activation screen and found EMICERI (EQTN MOB3B IFNK C9orf72 enhancer RNA I), a novel lncRNA that confers resistance to vemurafenib through upregulation of MOB3B and subsequent activation of the Hippo signalling pathway [72].
Table 1. Non-coding RNAs that contribute to resistance or sensitivity to targeted therapy with BRAF or MAPK inhibitors.
|
Induced Effect |
Overexpression of ncRNAs |
Target(s) |
Drugs |
References |
|
Drug resistance |
miR-34a, miR-100 and miR-125b |
CCL-2 |
Vemurafenib |
[66] |
|
miR-125a |
BAK1, MLK3 |
[67] |
||
|
miR-204 and miR-211 |
NUAK1/ARK5, IGFBP5, TGF-bRII, Slug, CHD5 |
|||
|
miR-514a |
NF1 |
[73] |
||
|
MIRAT |
IQGAP1/MAPK signalling |
Trametinib (MEKi) and PLX4720 (BRAFi) |
[71] |
|
|
SAMMSON |
p32 mitochondrial protein |
Vemurafenib and pimasertib (MEKi) |
[54] |
|
|
EMICERI |
MOB3B/LATS1/Hippo signalling axis |
Vemurafenib |
[72] |
|
|
|
|
|
|
|
|
Drug sensitivity |
miR-7 |
EGFR/IGF-1R/CRAF |
Vemurafenib |
[74] |
|
miR-32 |
MCL-1 |
[75] |
||
|
miR-200c |
Bmi-1 |
Vemurafenib or analog PLX4720 |
[76] |
|
|
miR-579-3p |
BRAF, MDM2 |
Vemurafenib and trametinib |
[70] |
4.2. Resistance to Immunotherapy
To date, results regarding the effect of ncRNAs on melanoma response to immune checkpoint blockers are sparse [77][78]. In order for studies to gain momentum in this area, some suggested the use of single-cell RNAseq approach to distinguish between the expression of cancer cells and that of the immune components [79]. Nevertheless, some miRNAs have been recognised as being involved in the conversion of monocytes into immunosuppressive MDSCs (let-7e, miR-99b, miR-100, miR-125a, miR-125b, miR-146a, miR-146b, miR-155) [80], while some interfere with PD-1 (miR-28) or PD-L1 (miR-17-5p) at a post-transcriptional level [81][82], facilitating resistance to immunotherapy (Table 2). LncRNAs have also been identified as potential modulators of myeloid cell differentiation towards an immunosuppressive phenotype (olfr29-ps1, lnc-chop) [83][84], however it seems that one particular polycistronic lncRNA, namely melanoma-overexpressed antigen (MELOE), can unexpectedly improve antigen presentation after being translated into short polypeptides, potentially increasing melanoma immunotherapy efficiency [85] (Table 2).
Table 2. Non-coding RNAs that interfere with immunotherapy in a negative or positive manner.
|
Name of ncRNAs |
Function |
Immune Response |
References |
|
miR-28 |
Blocks immune checkpoint Silences PD-1 by binding to its 3′UTR region |
Negative |
[81] |
|
miR-17-5p |
Blocks immune checkpoint ligand Binds to PD-L1 and contributes to melanoma resistance |
Negative |
[82] |
|
Let-7e, miR-99b, miR-100, miR-125a, miR-125b, miR-146a, miR-146b, miR-155 |
Control of MDSC Favours myeloid cell differentiation and polarization towards an immunosuppressive phenotype |
Negative |
[80] |
|
Olfr29-ps1 |
Control of MDSC Promotes MDSCs’ differentiation and function via de m6A-modified Olfr29-ps1/miR-214-3p/MyD88 regulatory network |
Negative |
[83] |
|
Lnc-chop |
Control of MDSC Promotes the differentiation and function of MDSCs |
Negative |
[84] |
|
MELOE |
Antigen presentation Produces immunogenic antigens (MELOE-1 and -2) that are recognized by cytotoxic T cells |
Positive |
[85] |
5. Clinical Applications of Non-Coding RNA Molecules in Melanoma Management
The clinical utilities and implications of ncRNAs in melanoma management are not fully established and future investigations are needed to clarify this aspect [2][79], however some promising results that recommend miRNAs and lncRNAs as tools for diagnosing, monitoring and treating cutaneous melanoma are presented in the following sections.
5.1. Non-Coding RNAs as Circulating Biomarkers for Cutaneous Melanoma
Even though lots of therapeutic progress has been made throughout the past years in finding alternative options for melanoma prognosis, it still remains challenging to find minimal invasive methods to follow up cancer progression and metastasis. Thus, liquid biopsies rise as a useful tool for the detection of circulating cancer biomarkers, among them ncRNAs [86][87][88].
Up to date lots of effort and work has been put in identifying a landscape of miRNAs as circulating biomarkers for melanoma diagnosis and prognosis (Table 3). Among the first to address this matter, Leidinger et al. have performed high throughput methods to validate a spectrum of 16-miRNAs which could be identified only in melanoma positive patients [89]. Several studies have indicated an increased level of multiple miRNAs, such as miR-19a, miR-149 and miR-126, in the plasma of metastatic melanoma patients as compared to healthy controls, suggesting these molecules to be actively involved in melanoma progression [90][91][92]. Interestingly, the co-detection of miR-185 and miR-1246 in liquid samples could offer an accurate identification of patients with metastatic melanoma, which could allow an early cancer diagnosis [93]. Additionally, Van Laar et al. have proposed a 38-miRNA signature (MEL38) in order to designate melanoma from normal plasma samples and an 18-miRNA signature (MEL18) which has the means to differentiate non-metastatic (stage I/II) and metastatic (stage III/IV) melanoma subjects [94]. A study done by Li et al. has detected miR-221 in serums samples from cutaneous MM patients, making this molecule a potential biomarker for melanoma evolution [95]. Another 4 miRNAs molecules (miR-30d, miR-15b, miR-150 and miR-425) have been proposed as potential biomarkers in order to discriminate between low and high-risk cases of recurrence [96]. Furthermore, Stark et al. indicated a 7-miRNA panel (MELmiR-7) including miR-16, miR-509-5p, miR-4706 and miR-211-5p which could be successfully implemented in the prediction of melanoma growth and recurrence, thus having a significant impact on melanoma prognosis and diagnosis [97].
In a similar manner, lncRNA molecules have also been identified as potential circulating biomarkers for melanoma (Table 3). In this context, it has been demonstrated that plasma lncRNA SPRY4-IT is significantly higher in tumour samples as compared to healthy controls [98]. Furthermore, Tang et al. observed the overexpression of HOTAIR in melanoma serum samples in comparison to non-cancer probes [55], while Cantile et al. found a significantly higher expression of HOTAIR in serum samples taken from patients with advanced melanoma [99]. These promising reports support the fact that HOTAIR has strong implications in the carcinogenesis of melanoma, making it a potential prognostic and diagnostic marker for MM.
Table 3. Promising results that recommend non-coding RNAs as circulating biomarkers for potential clinical applications.
|
Potential Role |
Non-Coding RNAs |
Sample Type |
References |
|
Circulating biomarkers for prognosis and diagnosis |
16 miRNA panel (among them miR-30d and miR-17) |
Blood |
[89] |
|
miR-19a miR-126 miR-149 |
Plasma |
||
|
miR-185 miR-1246 |
[93] |
||
|
18 miRNA panel (among them miR-199b-5p and let-7e) |
[94] |
||
|
miR-221 |
Serum |
[95] |
|
|
miR-15b miR-30d miR-150 miR-425 |
[96] |
||
|
7 miRNA panel (among them miR-16, miR-211-5p, miR-4706 and miR-509) |
[97] |
||
|
SPRY4-IT |
Plasma |
[98] |
|
|
HOTAIR |
Serum |
5.2. Non-Coding RNAs as Targets for Promising Therapeutic Strategies
The potential of targeting ncRNAs to develop novel anticancer therapeutic strategies or to increase the efficacy of already existing ones has been pointed out in several studies [79][100][101].
A number of ways in which miRNAs could be used have already been analysed, among which (1) synthetic miRNA mimetic agents, that could replace lost miRNA, (2) small-molecule inhibitors of miRNA, used for suppressing miRNA biogenesis or interaction with its target, (3) anti-miRNA oligomers, which are competitive inhibitors of miRNAs, leading to an upregulation of the target mRNA, or even (4) directly targeting miRNAs in the course of their transport within the tumour milieu or to other sites [102]. For example, miR-200c has been described as a potential therapeutic target for overcoming resistance to BRAFi therapy. In BRAFi-resistant cell lines and more importantly in clinical samples, low levels of miR-200c are correlated with acquired resistance. Restoration of miR-200c expression or knockdown of its molecular target favours the effect of inhibitory drugs and impairs the establishment of resistance [76] (Table 4). However, the use of miRNAs in therapy is hampered by their poor intracellular uptake, as well as rapid degradation in biological fluids. Strategies to deliver tumour-suppressive miRNAs or interfere with tumour-promoting miRNAs are still under development [103]. For instance, Fattore et al. developed lipid nanoparticles to encapsulate miR-204-5p and miR-199b-5p, either individually or in combination, and tested them on in vitro drug resistant models. Their results showed that these lipid nanoparticles loaded with oncosuppressor miRNAs are highly efficient in impairing melanoma cell proliferation and viability, affecting key signalling cascades involved in cell survival, in addition to positively influencing the efficacy of BRAF and MEK inhibitory drugs [104] (Table 4).
The involvement of lncRNAs in melanoma has promising therapeutic implications and selective knockdown of specific lncRNAs could lead to the development of reliable therapeutic strategies (Table 4). Out of the current methods available for studying lncRNAs, small interference RNA (siRNA)-dependent knockdown is used the most [105]. For instance, siRNA-mediated knockdown of SPRY4-IT1 in melanoma cell lines prevented tumour cell growth and limited invasion [43][44]. Similar results were obtained for knockdown of HOTAIR and UCA1, including inhibition of cell motility and invasive capacity [55][57]. MALAT1 knockdown was followed by a decrease in melanoma cell migration [56], whereas colony formation and metastatic ability of cancer cells were diminished in the absence of ANRIL [53]. Knockdown of SLNCR1 decreased the invasiveness of melanoma cells, although cell proliferation and motility were not affected [47]. Administration of GAS5, a tumour suppressor lncRNA, to nude mice inhibited melanoma growth, however further studies focused on the therapeutic value of GAS5 are needed [59].
The combination of a SAMMSON-specific antisense oligonucleotide with the BRAFi dabrafenib exerted a synergistic anti-tumour effect and induced apoptosis in a patient derived tumour xenograft preclinical model, whereas dabrafenib could only restrain melanoma growth (Table 4). Moreover, no significant difference was found between the toxicity levels of administrating SAMMSON-specific antisense oligonucleotide with BRAF inhibitors in comparison to administration of BRAF or MEK inhibitors alone [54].
Table 4. Promising results that recommend non-coding RNAs as therapeutic targets for potential clinical applications.
|
Potential Role |
Functional Studies |
Research Model |
Therapeutic Effect(s) |
References |
|
Targets for promising therapeutic strategies |
Lentiviral overexpression of miR-200c |
BRAFi-resistant cell lines |
Restores sensitivity to BRAFi therapies |
[76] |
|
Lipid nanoparticles loaded with miR-204-5p and/or miR-199b-5p |
In vitro drug resistant models |
Impair melanoma cell proliferation and viability Positively influence the effect of MAPKi |
[104] |
|
|
siRNA-mediated knockdown of SPRY4-IT1 |
Malignant melanoma cell lines |
Prevents tumour cell growth and limits invasion |
||
|
siRNA-mediated knockdown of HOTAIR |
Inhibits cell motility and decreases invasion |
[55] |
||
|
siRNA-mediated knockdown of UCA1 |
Inhibits cell proliferation and invasion Induces cell cycle arrest |
[57] |
||
|
siRNA-mediated knockdown of MALAT1 |
Impairs melanoma cell migration |
[56] |
||
|
siRNA-mediated knockdown of ANRIL |
Diminishes colony formation and metastatic ability |
[43] |
||
|
siRNA-mediated knockdown of SLNCR1 |
Decreases invasiveness of melanoma cells |
[47] |
||
|
Lentiviral overexpression of GAS5 |
In vitro and in vivo models |
Inhibits melanoma growth and cell migration |
[59] |
|
|
SAMMSON-specific antisense oligonucleotide |
Patient- derived xenograft |
Induces apoptosis Exerts a synergistic anti-tumour effect with dabrafenib |
[54] |
6. Conclusions
Research concerning the molecular landscape of malignant melanoma has brought impressive results in terms of patients’ overall survival in metastatic disease, due to its contribution to the development of targeted-based drugs and immunotherapy. Nevertheless, acquired resistance to therapy still remains a challenge, reflecting upon the poor prognosis of a significant number of patients. In this regard, therapeutic strategies aimed to modulate ncRNAs in combination with targeted agents and/or immunotherapy may represent a more efficient solution, considering that miRNAs and lncRNAs are not only involved in melanoma invasion and metastasis, but also facilitate resistance against currently available molecular therapeutic approaches. Strategies to deliver tumour-suppressive ncRNAs or interfere with tumour-promoting ncRNAs are searched for and future directions for the development of innovative treatment modalities include the use of intelligent nanocarriers loaded with ncRNAs for selective gene therapy. Although miRNAs and lncRNAs seem very promising biomarkers, their translation into the clinical area requires further studies and future investigative clinical trials.
References
- Gajos-Michniewicz, A.; Czyz, M.; Role of miRNAs in Melanoma Metastasis. Cancers 2019, 11, 326, 10.3390/cancers11030326..
- Yu, X.; Zheng, H.; Tse, G.; Chan, M.T.V.; Wu, W.K.K.; Long non-coding RNAs in melanoma. Cell Prolif 2018, 51, e12457, 10.1111/cpr.12457..
- Jarroux, J.; Morillon, A.; Pinskaya, M.; History, Discovery, and Classification of lncRNAs. Adv. Exp. Med. Biol. 2017, 1008, 1-46, 10.1007/978-981-10-5203-3_1.
- Luo, C.; Weber, C.E.M.; Osen,W.; Bosserhoff, A.-K.; Eichmüller, S.B.; The role of microRNAs in melanoma. Eur J Cell Biol 2014, 93, 11-22, 10.1016/j.ejcb.2014.02.001.
- Hirota, K.; Miyoshi, T.; Kugou, K.; Hoffman, C.S.; Shibata, T.; Ohta, K.; Stepwise chromatin remodelling by a cascade of transcription initiation of non-coding RNAs. Nature 2008, 456, 130-134, 10.1038/nature07348.
- Brockdorff, N.; Noncoding RNA and Polycomb recruitment. RNA 2013, 19, 429-442, 10.1261/rna.037598.112.
- Hansen, T.B.; Jensen, T.I.; Clausen, B.H.; Bramsen, J.B.; Finsen, B.; Damgaard, C.K.; Kjems, J.; Natural RNA circles function as efficient microRNA sponges. Nature 2013, 495, 384-388, 10.1038/nature11993.
- Segura, M.F.; Greenwald, H.S.; Hanniford, D.; Osman, I.; Hernando, E.; MicroRNA and cutaneous melanoma: from discovery to prognosis and therapy. Carcinogenesis 2012, 33, 1823-1832, 10.1093/carcin/bgs205.
- Mannavola, F.; Tucci, M.; Felici, C.; Stucci, S.; Silvestris, F.; miRNAs in melanoma: a defined role in tumor progression and metastasis. Exp Rev Clin Immunol 2015, 12, 79-89, 10.1586/1744666x.2016.1100965.
- Varrone, F.; Caputo, E.; The miRNAs Role in Melanoma and in Its Resistance to Therapy. Int J Molec Sci 2020, 21, 878, 10.3390/ijms21030878.
- Garraway, L.A.; Widlund, H.R.; Rubin, M.A.; Getz, G.; Berger, A.J.; Ramaswamy, S.; Beroukhim, R.; Milner, D.A.; Granter, S.R.; Du, J.; et al.et al. Integrative genomic analyses identify MITF as a lineage survival oncogene amplified in malignant melanoma. Nature 2005, 436, 117-122, 10.1038/nature03664.
- Carreira, S.; Goodall, J.; Denat, L.; Rodriguez, M.; Nuciforo, P.; Hoek, K.S.; Testori, A.; Larue, L.; Goding, C.R.; Mitf regulation of Dia1 controls melanoma proliferation and invasiveness. Genes Dev. 2006, 20, 3426-3439, 10.1101/gad.406406.
- Bell, R.E.; Levy, C.; The three M’s: melanoma, microphthalmia-associated transcription factor and microRNA. Pigment Cell Melanoma Res 2011, 24, 1088-1106, 10.1111/j.1755-148x.2011.00931.x.
- Bemis, L.T.; Chen, R.; Amato, C.M.; Classen, E.H.; Robinson, S.E.; Coffey, D.G.; Erickson, P.F.; Shellman, G.; Robinson, W.A.F.; MicroRNA-137 Targets Microphthalmia-Associated Transcription Factor in Melanoma Cell Lines. Cancer Res 2008, 68, 1362-1368, 10.1158/0008-5472.can-07-2912.
- Segura, M.F.; Hanniford, D.; Menendez, S.; Reavie, L.; Zou, X.; Alvarez-Diaz, S.; Zakrzewski, J.; Blochin, E.; Rose, A.; Bogunovic, D.; et al.et al. Aberrant miR-182 expression promotes melanoma metastasis by repressing FOXO3 and microphthalmia-associated transcription factor. Proc. Natl. Acad. Sci. USA 2009, 106, 1814-1819, 10.1073/pnas.0808263106.
- Boyle, G.M.; Woods, S.L.; Bonazzi, V.F.; Stark, M.S.; Hacker, E.; Aoude, L.G.; Dutton-Regester, K.; Cook, A.L.; Sturm, R.A.; Hayward, N.K.; et al. Melanoma cell invasiveness is regulated by miR-211 suppression of the BRN2 transcription factor. Pigment Cell Melanoma Res 2011, 24, 525-537, 10.1111/j.1755-148x.2011.00849.x.
- Zhao, G.; Wei, Z.; Guo, Y.; MicroRNA-107 is a novel tumor suppressor targeting POU3F2 in melanoma.. Biol Res 2020, 53, 11-10, 10.1186/s40659-020-00278-3.
- Luo, C.; Tetteh, P.W.; Merz, P.R.; Dickes, E.; Abukiwan, A.; Hotz-Wagenblatt, A.; Holland-Cunz, S.; Sinnberg, T.; Schittek, B.; Schadendorf, D.; et al.et al. miR-137 Inhibits the Invasion of Melanoma Cells through Downregulation of Multiple Oncogenic Target Genes. J Investig Derm 2013, 133, 768-775, 10.1038/jid.2012.357.
- Mueller, D.W.; Rehli, M.; Bosserhoff, A.K.; miRNA Expression Profiling in Melanocytes and Melanoma Cell Lines Reveals miRNAs Associated with Formation and Progression of Malignant Melanoma. J Investig Derm 2009, 129, 1740-1751, 10.1038/jid.2008.452.
- Bell, R.E.; Khaled, M.; Netanely, D.; Schubert, S.; Golan, T.; Buxbaum, A.; Janas, M.M.; Postolsky, B.; Goldberg, M.S.; Shamir, R.; et al.et al. Transcription Factor/microRNA Axis Blocks Melanoma Invasion Program by miR-211 Targeting NUAK1. J Investig Derm 2014, 134, 441-451, 10.1038/jid.2013.340.
- Mazar, J.; DeYoung, K.; Khaitan, D.; Meister, E.; Almodovar, A.; Goydos, J.; Ray, A.; Perera, R.J.; The Regulation of miRNA-211 Expression and Its Role in Melanoma Cell Invasiveness. PLoS ONE 2010, 5, e13779, 10.1371/journal.pone.0013779.
- Arozarena, I.; Sanchez-Laorden, B.; Packer, L.; Hidalgo-Carcedo, C.; Hayward, R.; Viros, A.; Sahai, E.; Marais, R.; Oncogenic BRAF Induces Melanoma Cell Invasion by Downregulating the cGMP-Specific Phosphodiesterase PDE5A. Cancer Cell 2011, 19, 45-57, 10.1016/j.ccr.2010.10.029.
- Pan, Z.; Tian, Y.; Niu, G.; Cao, C.; Role of microRNAs in remodeling the tumor microenvironment (Review). Int J Oncol 2020, 56, 407-416, 10.3892/ijo.2019.4952.
- Müller, D.W.; Bosserhoff, A.K.; Integrin β3 expression is regulated by let-7a miRNA in malignant melanoma. Oncogene 2008, 27, 6698-6706, 10.1038/onc.2008.282.
- Fu, T.Y.; Chang, C.C.; Lin, C.T.; Lai, C.H.; Peng, S.Y.; Ko, Y.J.; Tang, P.C.; Let-7b-mediated suppression of basigin expression and metastasis in mouse melanoma cells. Exp Cell Res 2011, 317, 445-451, 10.1016/j.yexcr.2010.11.004.
- Martin del Campo, S.E.; Latchana, N.; Levine, K.M.; Grignol, V.P.; Fairchild, E.T.; Jaime-Ramirez, A.C.; Dao, T.V.; Karpa, V.I.; Carson, M.; Ganju, A.; et al.et al. MiR-21 Enhances Melanoma Invasiveness via Inhibition of Tissue Inhibitor of Metalloproteinases 3 Expression: In Vivo Effects of MiR-21 Inhibitor. PLOS ONE 2015, 10, e0115919, 10.1371/journal.pone.0115919.
- Gaziel-Sovran, A.; Segura, M.F.; Di Micco, R.; Collins, M.K.; Hanniford, D.; Vega-Saenz de Miera, E.; Rakus, J.F.; Dankert, J.F.; Shang, S.; Kerbel, R.S.; et al.et al. miR-30b/30d Regulation of GalNAc Transferases Enhances Invasion and Immunosuppression during Metastasis. Cancer Cell 2011, 20, 104-118, 10.1016/j.ccr.2011.05.027.
- Knoll, S.; Fürst, K.; Kowtharapu, B.; Schmitz, U.; Marquardt, S.; Wolkenhauer, O.; Martin, H.; Pützer, B.M.; E2F1 induces miR‐224/452 expression to drive EMT through TXNIP downregulation. EMBO Rep. 2014, 15, 1315-1329, 10.15252/embr.201439392.
- Zhong, X.; Zheng, L.; Shen, J.; Zhang, D.; Xiong, M.; Zhang, Y.; He, X.; Tanyi, J.L.; Yang, F.; Montone, K.T.; et al.et al. Suppression of MicroRNA 200 Family Expression by Oncogenic KRAS Activation Promotes Cell Survival and Epithelial-Mesenchymal Transition in KRAS-Driven Cancer. Mol Cell Biol 2016, 36, 2742-2754, 10.1128/mcb.00079-16.
- Penna, E.; Orso, F.; Cimino, D.; Tenaglia, E.; Lembo, A.; Quaglino, E.; Poliseno, L.; Haimovic, A.; Osella-Abate, S.; De Pittà, C.; et al.et al. microRNA-214 contributes to melanoma tumour progression through suppression of TFAP2C. EMBO J. 2011, 30, 1990-2007, 10.1038/emboj.2011.102.
- Rambow, F.; Bechadergue, A.; Luciani, F.; Gros, G.; Domingues, M.; Bonaventure, J.; Meurice, G.; Marine, J.C.; Larue, L.; Regulation of Melanoma Progression through the TCF4/miR-125b/NEDD9 Cascade. J Investig Derm 2016, 136, 1229-1237, 10.1016/j.jid.2016.02.803.
- Domingues, M.J.; Rambow, F.; Job, B.; Papon, L.; Liu, W.; Larue, L.; Bonaventure, J.; Beta-catenin Inhibitor ICAT Modulates the Invasive Motility of Melanoma Cells. Cancer Res 2014, 74, 1983-1995, 10.1158/0008-5472.can-13-0920.
- Pencheva, N.; Tran, H.; Buss, C.; Huh, D.; Drobnjak, M.; Busam, K.; Tavazoie, S.F.; Convergent Multi-miRNA Targeting of ApoE Drives LRP1/LRP8-Dependent Melanoma Metastasis and Angiogenesis. Cell 2012, 151, 1068-1082, 10.1016/j.cell.2012.10.028.
- Migliore, C.; Petrelli, A.; Ghiso, E.; Corso, S.; Capparuccia, L.; Eramo, A.; Comoglio, P.M.; Giordano, S.; MicroRNAs Impair MET-Mediated Invasive Growth. Cancer Res 2008, 68, 10128-10136, 10.1158/0008-5472.can-08-2148.
- Tucci, M.; Mannavola, F.; Passarelli, A.; Stucci, L.S.; Cives, M.; Silvestris, F.; Exosomes in melanoma: a role in tumor progression, metastasis and impaired immune system activity. Oncotarget 2018, 9, 20826-20837, 10.18632/oncotarget.24846.
- Gowda, R.; Robertson, B.M.; Iyer, S.; Barry, J.; Dinavahi, S.S.; Robertson, G.P.; The role of exosomes in metastasis and progression of melanoma. Cancer Treat Rev 2020, 85, 101975, 10.1016/j.ctrv.2020.101975.
- Hood, J.L.; San, R.S.; Wickline, S.A.; Exosomes Released by Melanoma Cells Prepare Sentinel Lymph Nodes for Tumor Metastasis. Cancer Res 2011, 71, 3792-3801, 10.1158/0008-5472.can-10-4455.
- Hood, J.L.; Melanoma exosomes enable tumor tolerance in lymph nodes.. Med Hypotheses 2016, 90, 11-3, 10.1016/j.mehy.2016.02.018.
- Peinado, H.; Zhang, H.; Matei, I.R.; Costa-Silva, B.; Hoshino, A.; Rodrigues, G.; Psaila, B.; Kaplan, R.N.; Bromberg, J.F.; Kang, Y.; et al.et al. Pre-metastatic niches: organ-specific homes for metastases. Nat Rev Cancer 2017, 17, 302-317, 10.1038/nrc.2017.6.
- Bhan, A.; Soleimani, M.; Mandal, S.S.; Long Noncoding RNA and Cancer: A New Paradigm. Cancer Res 2017, 77, 3965-3981, 10.1158/0008-5472.can-16-2634.
- Dinescu, S.; Ignat, S.; Lazar, A.D.; Constantin, C.; Neagu, M.; Costache, M.; Epitranscriptomic Signatures in lncRNAs and Their Possible Roles in Cancer. Genes 2019, 10, 52, 10.3390/genes10010052.
- Dalay, N.; Role of the lncRNAs in malignant melanoma and their involvement in metastasis. Transl Cancer Res 2016, 5, S758-S764, 10.21037/tcr.2016.10.93.
- Khaitan, D.; Dinger, M.E.; Mazar, J.; Crawford, J.; Smith, M.A.; Mattick, J.S.; Perera, R.J.; The Melanoma-Upregulated Long Noncoding RNA SPRY4-IT1 Modulates Apoptosis and Invasion. Cancer Res 2011, 71, 3852-3862, 10.1158/0008-5472.can-10-4460.
- Mazar, J.; Zhao, W.; Khalil, A.M.; Lee, B.; Shelley, J.; Govindarajan, S.S.; Yamamoto, F.; Ratnam, M.; Aftab, M.N.; Collins, S.; et al.et al. The Functional Characterization of Long Noncoding RNA SPRY4-IT1 in Human Melanoma Cells. Oncotarget 2014, 5, 8959-8969, 10.18632/oncotarget.1863.
- Sun, M.; Liu, X.H.; Lu, K.H.; Nie, F.Q.; Xia, R.; Kong, R.; Yang, J.S.; Xi, T.P.; Liu, Y.W.; Zou, Y.F.; et al.et al. EZH2-mediated epigenetic suppression of long noncoding RNA SPRY4-IT1 promote s NSCLC cell proliferation and metastasis by affecting the epithelial–mesenchymal transition. Cell Death Dis 2014, 5, e1298-e1298, 10.1038/cddis.2014.256.
- Siena, Á.D.D.; Plaça, J.R.; Araújo, L.F.; Ichihara de Barros, I.; Peronni, K.; Molfetta, G.; Oliveira de Biagi, C.A., Jr.; Espreafico, E.M.; Sousa, J.F.; Silva, W.A., Jr.; et al. Whole transcriptome analysis reveals correlation of long noncoding RNA ZEB1-AS1 with invasive profile in melanoma. Sci Rep 2019, 9, 1-11, 10.1038/s41598-019-47363-6.
- Schmidt, K.; Joyce, C.E.; Buquicchio, F.; Brown, A.; Ritz, J.; Distel, R.J.; Yoon, C.H.; Novina, C.D.; The lncRNA SLNCR1 Mediates Melanoma Invasion through a Conserved SRA1-like Region. Cell Rep 2016, 15, 2025-2037, 10.1016/j.celrep.2016.04.018.
- Hombach, S.; Kretz, M.; The non-coding skin: Exploring the roles of long non-coding RNAs in epidermal homeostasis and disease. BioEssays 2013, 35, 1093-1100, 10.1002/bies.201300068.
- Flockhart, R.J.; Webster, D.E.; Qu, K.; Mascarenhas, N.; Kovalski, J.; Kretz, M.; Khavari, P.A.; BRAFV600Eremodels the melanocyte transcriptome and inducesBANCRto regulate melanoma cell migration. Genome Res 2012, 22, 1006-1014, 10.1101/gr.140061.112.
- Akhbari, P.; Whitehouse, A.; Boyne, J.R.; Long non-coding RNAs drive metastatic progression in melanoma (Review). Int J Oncol 2014, 45, 2181-2186, 10.3892/ijo.2014.2691.
- Pasmant, E.; Laurendeau, I.; Heron, D.; Vidaud, M.; Vidaud, D.; Bieche, I.; Characterization of a Germ-Line Deletion, Including the Entire INK4/ARF Locus, in a Melanoma-Neural System Tumor Family: Identification of ANRIL, an Antisense Noncoding RNA Whose Expression Coclusters with ARF. Cancer Res 2007, 67, 3963-3969, 10.1158/0008-5472.can-06-2004.
- Xie, H.; Rachakonda, P.S.; Heidenreich, B.; Nagore, E.; Sucker, A.; Hemminki, K.; Schadendorf, D.; Kumar, R.; Mapping of deletion breakpoints at the CDKN2A locus in melanoma: detection of MTAP-ANRIL fusion transcripts. Oncotarget 2016, 7, 16490-16504, 10.18632/oncotarget.7503.
- Xu, S.; Wang, H.; Pan, H.; Shi, Y.; Li, T.; Ge, S.; Jia, R.; Zhang, H.; Fan, X.; ANRIL lncRNA triggers efficient therapeutic efficacy by reprogramming the aberrant INK4-hub in melanoma. Cancer Lett 2016, 381, 41-48, 10.1016/j.canlet.2016.07.024.
- Leucci, E.; Vendramin, R.; Spinazzi, M.; Laurette, P.; Fiers, M.; Wouters, J.; Radaelli, E.; Eyckerman, S.; Leonelli, C.; Vanderheyden, K.; et al.et al. Melanoma addiction to the long non-coding RNA SAMMSON. Nature 2016, 531, 518-522, 10.1038/nature17161.
- Tang, L.; Zhang, W.; Su, B.; Yu, B.; Long Noncoding RNA HOTAIR Is Associated with Motility, Invasion, and Metastatic Potential of Metastatic Melanoma. BioMed Res Int 2013, 2013, 1-7, 10.1155/2013/251098.
- Tian, Y.; Zhang, X.; Hao, Y.; Fang, Z.; He, Y.; Potential roles of abnormally expressed long noncoding RNA UCA1 and Malat-1 in metastasis of melanoma. Melanoma Res 2014, 24, 335-341, 10.1097/cmr.0000000000000080.
- Wei, Y.; Sun, Q.; Zhao, L.; Wu, J.; Chen, X.; Wang, Y.; Zang, W.; Zhao, G.; LncRNA UCA1-miR-507-FOXM1 axis is involved in cell proliferation, invasion and G0/G1 cell cycle arrest in melanoma. Med Oncol 2016, 33, 1-9, 10.1007/s12032-016-0804-2.
- Pan, B.M.; Lin, X.; Zhang, L.; Hong, W.; Zhang, Y.; Long noncoding RNA X-inactive specific transcript promotes malignant melanoma progression and oxaliplatin resistance. Melanoma Res 2019, 29, 254-262, 10.1097/cmr.0000000000000560.
- Chen, L.; Yang, H.; Xiao, Y.; Tang, X.; Li, Y.; Han, Q.; Fu, J.; Yang, Y.; Zhu, Y.; LncRNA GAS5 is a critical regulator of metastasis phenotype of melanoma cells and inhibits tumor growth in vivo. OncoTargets 2016, 9, 4075-4087, 10.2147/OTT.S98203.
- Wu, L.; Zhu, L.; Li, Y.; Zheng, Z.; Lin, X.; Yang, C.; LncRNA MEG3 promotes melanoma growth, metastasis and formation through modulating miR-21/E-cadherin axis. Cancer Cell Int 2020, 20, 1-14, 10.1186/s12935-019-1087-4.
- Chen, L.; Yang, H.; Xiao, Y.; Tang, X.; Li, Y.; Han, Q.; Fu, J.; Yang, Y.; Zhu, Y.; Lentiviral-mediated overexpression of long non-coding RNA GAS5 reduces invasion by mediating MMP2 expression and activity in human melanoma cells. Int J Oncol 2016, 48, 1509-1518, 10.3892/ijo.2016.3377.
- Bian, D.; Shi, W.; Shao, Y.; Li, P.; Song, G.; Long non-coding RNA GAS5 inhibits tumorigenesis via miR-137 in melanoma. Am J Transl Res 2017, 9, 1509-1520.
- Rizos, H.; Menzies, A.M.; Pupo, G.M.; Carlino, M.S.; Fung, C.; Hyman, J.; Haydu, L.E.; Mijatov, B.; Becker, T.M.; Boyd, S.C.; et al.et al. BRAF Inhibitor Resistance Mechanisms in Metastatic Melanoma: Spectrum and Clinical Impact. Clin Cancer Res 2014, 20, 1965-1977, 10.1158/1078-0432.ccr-13-3122.
- Gatzka, M.V.; Targeted Tumor Therapy Remixed—An Update on the Use of Small-Molecule Drugs in Combination Therapies. Cancers 2018, 10, 155, 10.3390/cancers10060155.
- Ribas, A.; Wolchok, J.D.; Cancer immunotherapy using checkpoint blockade. Science 2018, 359, 1350-1355, 10.1126/science.aar4060.
- Vergani, E.; Di Guardo, L.; Dugo, M.; Rigoletto, S.; Tragni, G.; Ruggeri, R.; Perrone, F.; Tamborini, E.; Gloghini, A.; Arienti, F.; et al.et al. Overcoming melanoma resistance to vemurafenib by targeting CCL2-induced miR-34a, miR-100 and miR-125b. Oncotarget 2015, 7, 4428-4441, 10.18632/oncotarget.6599.
- Koetz-Ploch, L.; Hanniford, D.; Dolgalev, I.; Sokolova, E.; Zhong, J.; Diaz-Martinez, M.; Bernstein, E.; Darvishian, F.; Flaherty, K.T.; Chapman, P.B.; et al.et al. MicroRNA-125a promotes resistance to BRAF inhibitors through suppression of the intrinsic apoptotic pathway.. Pigment Cell Melanoma Res 2017, 30, 328-338, 10.1111/pcmr.12578.
- Díaz-Martínez, M.; Benito-Jardón, L.; Alonso, L.; Koetz-Ploch, L.; Hernando, E.; Teixidó, J.; miR-204-5p and miR-211-5p Contribute to BRAF Inhibitor Resistance in Melanoma. Cancer Res 2017, 78, 1017-1030, 10.1158/0008-5472.can-17-1318.
- Vitiello, M.; D’Aurizio, R.; Poliseno, L.; Biological role of miR-204 and miR-211 in melanoma. Oncoscience 2018, 5, 248-251, 10.18632/oncoscience.443.
- Fattore, L.; Mancini, R.; Acunzo, M.; Romano, G.; Lagana, A.; Pisanu, M.E.; Malpicci, D.; Madonna, G.; Mallardo, D.; Capone, M.; et al.et al. miR-579-3p controls melanoma progression and resistance to target therapy. Proceedings of the National Academy of Sciences 2016, 113, E5005-E5013, 10.1073/pnas.1607753113.
- Sanlorenzo, M.; Vujic, I.; Esteve-Puig, R.; Lai, K.; Vujic, M.; Lin, K.; Posch, C.; Dimon, M.; Moy, A.; Zekhtser, M.; et al.et al. The lincRNA MIRAT binds to IQGAP1 and modulates the MAPK pathway in NRAS mutant melanoma.. Sci Rep 2018, 8, 10902, 10.1038/s41598-018-27643-3.
- Joung, J.; Engreitz, J.M.; Konermann, S.; Abudayyeh, O.O.; Verdine, V.K.; Aguet, F.; Genome-scale activation screen identifies a lncRNA locus regulating a gene neighbourhood. Nature 2017, 548, 343-346, 10.1038/nature23451.
- Stark, M.S.; Bonazzi, V.F.; Boyle, G.M.; Palmer, J.M.; Symmons, J.; Lanagan, C.M.; Schmidt, C.W.; Herington, A.C.; Ballotti, R.; Pollock, P.M.; et al.et al. miR-514a regulates the tumour suppressor NF1 and modulates BRAFi sensitivity in melanoma. Oncotarget 2015, 6, 17753-17763, 10.18632/oncotarget.3924.
- Sun, X.; Li, J.; Sun, Y.; Zhang, Y.; Dong, L.; Shen, C.; Yang, L.; Yang, M.; Li, Y.; Shen, G.; et al.et al miR-7 reverses the resistance to BRAFi in melanoma by targeting EGFR/IGF-1R/CRAF and inhibiting the MAPK and PI3K/AKT signaling pathways. Oncotarget 2016, 7, 53558-53570, 10.18632/oncotarget.10669.
- Mishra, P.J.; Merlino, G.; Integrated Genomics Identifies miR-32/MCL-1 Pathway as a Critical Driver of Melanomagenesis: Implications for miR-Replacement and Combination Therapy. PLOS ONE 2016, 11, e0165102, 10.1371/journal.pone.0165102.
- Liu, S.; Tetzlaff, M.T.; Wang, T.; Yang, R.; Xie, L.; Zhang, G.; Krepler, C.; Xiao, M.; Beqiri, M.; Xu, W.; et al.et al. miR-200c/Bmi1 axis and epithelial-mesenchymal transition contribute to acquired resistance to BRAF inhibitor treatment. Pigment Cell Melanoma Res 2015, 28, 431-441, 10.1111/pcmr.12379.
- Vishnubalaji, R.; Hibah, S.; Elango, R.; Alajez, N.M; Noncoding RNAs as potential mediators of resistance to cancer immunotherapy. Semin Cancer Biol 2020, 65, 65-79, 10.1016/j.semcancer.2019.11.006.
- Zhou, Y.; Zhu, Y.; Xie, Y.; Ma, X.; The Role of Long Non-coding RNAs in Immunotherapy Resistance. Front Oncol 2019, 9, 1292, 10.3389/fonc.2019.01292.
- Lorusso, C.; De Summa, S.; Pinto, R.; Danza, K.; Tommasi, S.; miRNAs as Key Players in the Management of Cutaneous Melanoma. Cells 2020, 9, 415, 10.3390/cells9020415.
- Huber, V.; Vallacchi, V.; Fleming, V.; Hu, X.; Cova, A.; Dugo, M.; Shahaj, E.; Sulsenti, R.; Vergani, E.; Filipazzi, P.; et al.et al. Tumor-derived microRNAs induce myeloid suppressor cells and predict immunotherapy resistance in melanoma. J Clin Investig 2018, 128, 5505-5516, 10.1172/jci98060.
- Li, Q.; Johnston, N.; Zheng, X.; Wang, H.; Zhang, X.; Gao, D.; Min, W.; miR-28 modulates exhaustive differentiation of T cells through silencing programmed cell death-1 and regulating cytokine secretion. Oncotarget 2016, 7, 53735-53750, 10.18632/oncotarget.10731.
- Audrito, V.; Serra, S.; Stingi, A.; Orso, F.; Gaudino, F.; Bologna, C.; Neri, F.; Garaffo, G.; Nassini, R.; Baroni, G.; et al.et al. PD-L1 up-regulation in melanoma increases disease aggressiveness and is mediated through miR-17-5p. Oncotarget 2017, 8, 15894-15911, 10.18632/oncotarget.15213.
- Shang, W.; Gao, Y.; Tang, Z.; Zhang, Y.; Yang, R.; The Pseudogene Olfr29-ps1 Promotes the Suppressive Function and Differentiation of Monocytic MDSCs. Cancer Immunol Res 2019, 7, 813-827, 10.1158/2326-6066.cir-18-0443.
- Gao, Y.; Wang, T.; Li, Y.; Zhang, Y.; Yang, R; Lnc-chop Promotes Immunosuppressive Function of Myeloid-Derived Suppressor Cells in Tumor and Inflammatory Environments. J Immunol 2018, 200, 2603-2614, 10.4049/jimmunol.1701721.
- Charpentier, M.; Croyal, M.; Carbonnelle, D.; Fortun, A.; Florenceau, L.; Rabu, C.; Krempf, M.; Labarriere, N.; Lang, F.; IRES-dependent translation of the long non coding RNA meloe in melanoma cells produces the most immunogenic MELOE antigens. Oncotarget 2016, 7, 59704-59713, 10.18632/oncotarget.10923.
- Palmirotta, R.; Lovero, D.; Cafforio, P.; Felici, C.; Mannavola, F.; Pellè, E.; Quaresmini, D.; Tucci, M.; Silvestris, F.; Liquid biopsy of cancer: a multimodal diagnostic tool in clinical oncology. Adv Med Oncol 2018, 10, 758835918794630, 10.1177/1758835918794630.
- Schwarzenbach, H.; Hoon, D.S.B.; Pantel, K.; Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer 2011, 11, 426-437, 10.1038/nrc3066.
- Sole, C.; Arnaiz, E.; Manterola, L.; Otaegui, D.; Lawrie, C.H.; The circulating transcriptome as a source of cancer liquid biopsy biomarkers. Semin Cancer Biol 2019, 58, 100-108, 10.1016/j.semcancer.2019.01.003.
- Leidinger, P.; Keller, A.; Borries, A.; Reichrath, J.; Rass, K.; Jager, S.V.; Lenhof, H.-P.; Meeseet, E.; High-throughput miRNA profiling of human melanoma blood samples. BMC Cancer 2010, 10, 262-262, 10.1186/1471-2407-10-262.
- Mumford, S.L.; Towler, B.P.; Pashler, A.L.; Gilleard, O.; Martin, Y.; Newbury, S.F.; Circulating MicroRNA Biomarkers in Melanoma: Tools and Challenges in Personalised Medicine. Biomolecules 2018, 8, 21, 10.3390/biom8020021.
- Pfeffer, S.R.; Grossmann, K.F.; Cassidy, P.B.; Yang, C.H.; Fan, M.; Kopelovich, L.; Leachman, S.A.; Pfeffer, L.M.; Detection of Exosomal miRNAs in the Plasma of Melanoma Patients. J Clin Med 2015, 4, 2012-2027, 10.3390/jcm4121957.
- Mannavola, F.; D’Oronzo, S.; Cives, M.; Stucci, L.S.; Ranieri, G.; Silvestris, F.; Tucci, M.; Extracellular Vesicles and Epigenetic Modifications Are Hallmarks of Melanoma Progression. Int J Mol Sci 2020, 21, 52, 10.3390/ijms21010052.
- Armand-Labit, V.; Meyer, N.; Casanova, A.; Bonnabau, H.; Platzer, V.; Tournier, E.; Sansas, B.; Verdun, S.; Thouvenot, B.; Hilselberger, B.; et al.et al. Identification of a Circulating MicroRNA Profile as a Biomarker of Metastatic Cutaneous Melanoma. Acta Derm Venereol 2016, 96, 29-34, 10.2340/00015555-2156.
- Van Laar, R.; Lincoln, M.; Van Laar, B.; Development and validation of a plasma-based melanoma biomarker suitable for clinical use.. Br J Cancer 2018, 118, 857-866, 10.1038/bjc.2017.477.
- Li, P.; He, Q.Y.; Luo, C.Q.; Qian, L.Y.; Circulating miR-221 Expression Level and Prognosis of Cutaneous Malignant Melanoma. Med Sci Monit 2014, 20, 2472-2477, 10.12659/MSM.891327.
- Fleming, N.H.; Zhong, J.; Da Silva, I.P.; De Miera, E.V.S.; Brady, B.; Han, S.W.; Hanniford, D.; Wang, J.; Shapiro, R.L.; Hernando, E.; et al.et al Serum-based miRNAs in the prediction and detection of recurrence in melanoma patients. Cancer 2014, 121, 51-59, 10.1002/cncr.28981.
- Stark, M.S.; Klein, K.; Weide, B.; Haydu, L.E.; Pflugfelder, A.; Tang, Y.H.; Palmer, J.M.; Whiteman, D.C.; Scolyer, R.A.; Mann, G.J.; et al.et al. The Prognostic and Predictive Value of Melanoma-related MicroRNAs Using Tissue and Serum: A MicroRNA Expression Analysis. EBioMedicine 2015, 2, 671-680, 10.1016/j.ebiom.2015.05.011.
- Liu, T.; Shen, S.K.; Xiong, J.G.; Xu, Y.; Zhang, H.Q.; Liu, H.J.; Lu, Z.G.; Clinical significance of long noncoding RNA SPRY 4‐ IT 1 in melanoma patients. FEBS Open Bio 2016, 6, 147-154, 10.1002/2211-5463.12030.
- Cantile, M.; Scognamiglio, G.; Marra, L.; Aquino, G.; Botti, C.; Falcone, M.; Malzone, M.G.; Liguori, G.; Di Bonito, M.; Franco, R.; et al.et al. HOTAIR role in melanoma progression and its identification in the blood of patients with advanced disease. J Cell Physiol 2017, 232, 3422-3432, 10.1002/jcp.25789.
- Esteller, M.; Non-coding RNAs in human disease. Nat Rev Genet 2011, 12, 861-874, 10.1038/nrg3074.
- Hulstaert, E.; Brochez, L.; Volders, P.J.; Vandesompele, J.; Mestdagh, P.; Long non-coding RNAs in cutaneous melanoma: clinical perspectives. Oncotarget 2017, 8, 43470-43480, 10.18632/oncotarget.16478.
- Stenvang, J.; Petri, A.; Lindow, M.; Obad, S.; Kauppinen, S.; Inhibition of microRNA function by antimiR oligonucleotides. Silence 2012, 3, 1, 10.1186/1758-907x-3-1.
- Hosseinahli, N.; Aghapour, M.; Duijf, P.H.G.; Baradaran, B.; Treating cancer with microRNA replacement therapy: A literature review. J Cell Physiol 2018, 233, 5574-5588, 10.1002/jcp.26514.
- Fattore, L.; Campani, V.; Ruggiero, C.F.; Salvati, V.; Liguoro, D.; Scotti, L.; Botti, G.; Ascierto, P.A.; Mancini, R.; De Rosa, G.; et al.et al. In Vitro Biophysical and Biological Characterization of Lipid Nanoparticles Co-Encapsulating Oncosuppressors miR-199b-5p and miR-204-5p as Potentiators of Target Therapy in Metastatic Melanoma. Int J Molec Sci 2020, 21, 1930, 10.3390/ijms21061930.
- Aftab, M.N.; Dinger, M.E.; Perera, R.J.; The role of microRNAs and long non-coding RNAs in the pathology, diagnosis, and management of melanoma. Arch Biochem Biophys 2014, 563, 60-70, 10.1016/j.abb.2014.07.022.