1000/1000
Hot
Most Recent

Quantitative parameters of FDG-PET, such as SUV and TLG have been used to evaluate therapeutic response. Recent advancement in anti-cancer therapeutics showed that tumor response to molecular-targeted drugs and immune-checkpoint inhibitors is different from conventional chemotherapy in terms of temporal metabolic alteration and morphological change after the course of effective therapy. Metabolic changes and temporal enlargement due to immune cell infiltration seen after immune-checkpoint inhibitors, such as anti-programmed cell death-1 (PD-1) and anti-programmed cell death ligand 1 (PD-L1) antibodies facilitated the modification of conventional Response Evaluation Criteria in Solid Tumor and FDG-PET response evaluation criteria. Tumor microenvironment including cancer stem cells (CSCs) that is thought to be a root cause of tumor heterogeneity; is considered a target of novel and effective therapy.
Accumulation of FDG reflects glucose metabolism of both cancer cells and immunologically competent cells in the tumor microenvironment. Immunological reaction to the therapy differs among patients according to the individual immune function. Considering the heterogeneity of tumor tissue and individual variation in tumor response to immunotherapy, radiomics approach combines quantitative image features with deep learning algorithm have the potentials to improve response assessment in more personalized treatment.
Stromal cell-derived factor 1 (SDF-1)/C-X-C chemokine receptor type 4 (CXCR4)-targeted α-particle therapy has been introduced, because SDF-1/CXCR4 axis is known to increase epithelial-mesenchymal transition to facilitate invasion and metastasis, and regulate immune response by accelerating T cell proliferation as well as PD-1 and PD-L1 expression in cancer cells and cytotoxic T lymphocytes, respectively. Prominent energy profile and biological effect of α-particles are promising as an alternative in targeted radionuclide therapy (TRT). Radiation dosimetry according to the theranostics approach will permit accurate TRT and artificial intelligence-based treatment decision making and precise response evaluation would be a precision nuclear medicine in the future.
Positron emission tomography (PET) has become an indispensable procedure for the initial assessment and post-therapeutic evaluation in clinical oncology, using dedicated radiopharmaceuticals targeting cellular metabolism and tumor-specific receptors [1]. PET as a means of molecular-based imaging is able to characterize biological processes associated with disease progression and therapeutic response quantitatively at the cellular and molecular levels. The outcome of a therapy cannot be interpreted properly without a surrogate biomarker to assess the efficacy of every therapeutic modality.
Therapeutic response is objectively evaluable by means of imaging. Conventional response evaluation criteria use morphological parameters; on the other hand, 2-[18F] fluoro-2-deoxy-d-glucose (FDG)-PET-based criteria use metabolic parameters. Histological response to anti-cancer therapy depends on the therapeutic modalities; cancer immunotherapy shows the distinctive phenomenon of immune-related tumor responses. Emerging observational data of immune-related response patterns have determined modification of the conventional response criteria. The current approaches to anti-cancer therapy target the tumor microenvironment as well as anti-tumor immunity.
It has been appreciated for nearly 100 years that cancer cells are metabolically distinct from other cells. All cells fundamentally require nutrients to meet metabolic demands for energy generation and biosynthesis. Metabolic demands of cell proliferation, differentiation, and biosynthesis of proteins, lipids, and nucleotides are different in tumor cells.
Elevated glucose uptake and cellular metabolism were thought to be the biochemical characteristics of cancer [2]. FDG-PET could disclose a high glycolytic rate and pyruvate oxidation in the mitochondria, depending on the cell proliferation. These altered metabolisms, including metabolic switch from aerobic to anaerobic glycolysis, are known as the Warburg effect [3][4]. The function of the Warburg effect has been simply understood as a metabolic switch, but a breakthrough to explain the Warburg effect regarding cancer metabolism in vivo has taken place recently [5][6][7][8].
Tumor hypoxia is known to be the most important factor to account for biological aggressiveness and resistance to chemotherapy and radiotherapy through the expression of multidrug resistance 1 (MDR1) and cell cycle arrest [8][9]. Accelerated proliferation and metabolism of cancer cells lead to an imbalance in the form of insufficient oxygen supply in relation to oxygen demand in solid tumors [10][11]. Anti-neoplastic drugs and ionizing radiation have effects on oxygen to generate reactive oxygen species (ROS) in cancer cells, causing oxidative stress, which results in apoptosis. However, cancer cells can survive in the hypoxic area, which is usually seen at 100 μm from tumor vessels, because of the decreased generation of ROS [12]. In the area of hypoxia, a transcription factor, hypoxia-inducible factor 1 (HIF-1), is activated to induce the expression of various genes responsible for adaption to hypoxic metabolism from oxidative phosphorylation to glycolytic ATP production, explained by the Warburg effect, as mentioned above [13][14], invasion and metastases of cancer cells through the formation of pre-metabolic niche and epithelial–mesenchymal transition (EMT) to escape from hypoxia [15][16], increased erythropoiesis through upregulation of erythropoietin, and angiogenesis to reoxygenation of hypoxic area [17]. An α-subunit of HIF-1 (HIF-1α) induces expression of glucose transporter 1 and glycolytic enzymes to increase glucose uptake and anaerobic glycolysis to compensate for ATP production [18][19]. FDG-PET can therefore evaluate tumor aggressiveness and resistance to chemotherapy and radiotherapy by detecting increased glucose metabolism and is a possible therapeutic marker to monitor responses.
Figure 1 shows FDG-PET/CT images of a patient with NSCLC. Two foci of increased nodular FDG uptake are seen in the upper lobe of the right lung and the upper mediastinum on the left side. These lesions show SUVmax of 7.2 and 4.8, respectively. Do these images provide the radiologist with sufficient information for correct interpretation? Radiologists cannot diagnose correctly without additional information about clinical history, because every radiologist knows that the most common sites of metastasis of lung cancer are ipsilateral hilar and mediastinal lymph nodes and that metastasis to the contralateral mediastinal lymph node usually occurs after ipsilateral mediastinal lymph nodes metastases [20]. Information about past history of left lung cancer with T3N1M0, stage IIIA, for which left upper lobe segmentectomy and lymph node dissection was performed 2 years before, is a clue for correct diagnosis. The patient had undergone surgery followed by chemotherapy on the basis of correct diagnosis. How accurately can FDG-PET images estimate the efficacy of chemotherapy and prognosis of this patient? Do quantitative parameters help to predict response to chemotherapy and prognosis?
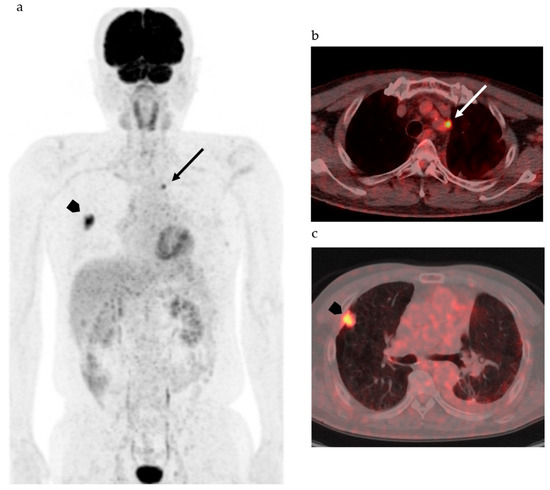
Figure 1. 2-[18F] fluoro-2-deoxy-d-glucose positron emission tomography/computed tomography (FDG-PET/CT) of patients with non-small cell lung cancer. (a) Maximum intense projection image shows abnormal FDG uptake in the left upper mediastinum (arrow) and the right lung (arrowhead). (b) PET/CT shows increased FDG uptake in the lymph node adjacent to the left subclavian artery at the level of the upper mediastinum (arrow). (c) High FDG uptake is seen in the nodule at the right upper lobe (arrow head).
Considering the heterogeneity of tumor tissue, more sophisticated indexes surpassing SUV and other parameters as well as diagnostic algorithms are needed to accurately classify the tumor on the basis of biological malignancy, effectiveness of various types of therapy, and prognosis of patients. Recent advancements in computer science and artificial intelligence (AI) have shown the possibility for machine learning systems to take on the practice of radiology, which was previously thought to be limited to human radiologists. AI including machine learning technologies has the potential to transform radiological imaging by using the vast amount of clinical data including pathologic and genetic examinations to automate the integrated diagnostic radiology workflow and diagnosis. Machine learning algorithms, such as random forests, support vector machines, and artificial neural networks, have been used for classification of images by training with input data set and knowledge, and then the best model is applied for the prediction of pathophysiology. Due to deep learning and convolutional neural networks (CNN), the capability to learn and master given tasks to perform computer-aided diagnosis (CAD) has made remarkable advances in clinical radiology in the past decade [21][22]. Wang et al. have suggested that the performance of CNN from FDG PET/CT images is comparable to the best classical machine learning and human radiologists and that CNN is more convenient and objective than the classical methods, because it does not need tumor segmentation, feature selection, or texture features for classifying mediastinal lymph node metastasis in patients with NSCLC [21]. They also suggested that the performance of the CNN would be improved by incorporating diagnostic features like SUV and tumor size [21]. For example, in mediastinal lymph node metastasis in patients with NSCLC, accurate diagnosis is a challenge, as indicated in Figure 1; however, lymph node metastasis evaluated by FDG uptake has been reported to be prognostic as compared with pathological lymph node metastasis [23]. Therefore, diagnosis of lymph node status during diagnostic work up is of the utmost importance.
Machine learning is already being applied in the practice of radiology, including in the field of mammography. There have been many papers describing a performance level in lesion detection similar to that of experienced radiologists [24][25][26]. CAD was approved by the Food and Drug Administration (FDA) and has been used for mammography in radiology practices [27]; however, improvement of the diagnostic ability has not been satisfactory, and the majority of radiologists have rarely changed their reports as a result of findings generated by CAD [28][29][30]. Machine learning has been reported to be unlikely to replace radiologists but will provide quantitative tools to increase the value of imaging as a biomarker including therapeutic response evaluation [31]. Recently, radiology professionals have reminded that AI algorithms must be as safe and effective as the physician by rigorous testing, longitudinal surveillance, and investigation of oversight mechanisms to ensure generalizability across patients as well as variable imaging and imaging protocols [32]. However, radiologists cannot disregard autonomous radiology AI, because AI can tirelessly improve the image reading capacity and may drastically acquire interpretation capabilities if AI can incorporate available medical information and contextual integration of data that would typically be identified during physician interpretation in order to render a medical judgement.
On the basis of considering tumor heterogeneity, texture analysis has been explored, especially in the field of nuclear medicine [32][33][34][35][36]. The most exciting part of machine learning in medical imaging would be to extract patterns that are beyond human perception and classification due to the application of deep learning for diagnostic algorithms [37][38]. Radiologists should seek to work alongside AI in the future.
Malignant cells survive in a complex balance in the immune system. Both CTLA-4 and PD-1 suppress T cell activities. Therefore, agents that block CTLA-4, PD-1, and PD-L1 are able to produce an anti-tumor response through immune activation. Inhibition of CXCR4 exaggerates the anti-tumor immune response and CXCR4-targeted therapy is a possible therapeutic option to eradicate CSCs. Recent studies have indicated that dual blockade of PD-1–PD-L1 and CXCL-12–CXCR4 pathways reduces specific cellular and functional elements within the immunosuppressive tumor microenvironment and augments tumor-specific cell-mediated immune responses. The complexity of these interactions and heterogeneity of immune cells in the tumor microenvironment are challenges in the development and the evaluation of the therapeutic efficacy of new immune therapies in vivo. Imaging of immune cells that are major players in anti-cancer therapy is challenging because many subtypes of cells exist and play different roles in the tumor microenvironment.
Non-invasive evaluation procedures for therapy outcomes, such as biomarkers and molecular imaging, are expected to represent precise strategies of cancer therapy. FDG-PET can play an important role in fulfilling this purpose, as mentioned earlier [39][40][41]. Uptake of FDG reflects the viability of cancer cells and all other players of the immune system in the microenvironment. No uptake of FDG means complete remission of the tumor; however, increased uptake does not always indicate progression of the tumor, because of the pseudoprogression phenomenon and increased anaerobic glycolysis in the therapy-induced hypoxia, as mentioned above [18][39][42]. Cancer cell-specific imaging has the potential to evaluate quantitatively the residual cancer cells that had been able to evade anti-cancer agent of immune response. However, phenotypic changes due to genetic alteration, such as therapy resistant mutation and de novo mutation after therapy, may decrease specificity to the specific imaging agent. Metabolism-based PET tracers other than FDG can be used to evaluate therapeutic efficacy [43][44]. However, metabolic diversity and instability, especially those acquired on the progression course or after therapy, of cancer cells would be sources of inaccuracy in evaluating the response.
Prediction and evaluation of therapeutic efficacy would be possible with a tumor-specific PET tracer. Prostate-specific membrane antigen (PSMA) ligand labeled with gallium-68 (68Ga-PSMA) is a PET tracer used to determine the eligibility for PSMA-targeted radionuclide therapy with 177Lu-PSMA or 235Ac-PSMA (Table 1). Peptide receptor radionuclide therapy for neuroendocrine carcinoma with 90Y- and 177Lu-dodecane-tetraacetic acid-Tyr3-octreotate (DOTA-TATE) is another radionuclide therapy performed successfully for solid tumors. 68Ga-DOTA-TATE is a diagnostic counterpart of therapeutics. These examples are representative theranostics in nuclear medicine practice that will be followed by the future radionuclide therapy. A major role of specific imaging in the theranostics is to confirm the indication of therapy. Another role would be dosimetry analysis to determine the therapeutic dose by calculating absorbed doses in the tumor for efficacy and target organs for toxicity. It may be possible for PET imaging with specific tracers to evaluate therapeutic efficacy by measuring the amount of target molecules; however, the expression of the target molecules may change after the therapy—then, accurate response evaluation would be difficult with these target-specific PET studies.
Table 1. Representative Pair of Radiopharmaceuticals for Theranostics.
| Radiopharmaceutical for Therapy | Radiation | Half-Life | Radiopharmaceutical for Diagnosis |
|---|---|---|---|
| 177Lu-DOTA-TATE | Beta ray (β− particle) | 78 h | 68Ga-DOTA-TATE |
| 213Bi-DOTA-TOC | Alpha ray (He2+ particle) | 0.76 h | 68Ga-DOTA-TOC |
| 177Lu-PSMA | Beta ray (β− particle) | 78 h | 68Ga-PSMA |
| 225Ac-PSMA | Alpha ray (He2+ particle) | 10 d | 68Ga-PSMA |
DOTA-TATE: dodecane-tetraacetic acid-Tyr3-octreotate; DOTA-TOC: dodecane-tetraacetic acid-D-Phe1-Tyr3-octreotide; PSMA: prostate-specific membrane antigen.
Considering the present availability and required standardization, FDG-PET may be favorable for response evaluation in solid tumors. Since there is a variety of therapeutics that have effects on both cancer cells and the immune system, individualized evaluation criteria based on therapeutic agents and clinicopathologic information may be appropriate. Clinicopathologic data include therapeutic regimen and time from administration, immune function status, temporal changes in size and attenuation of tumor on CT, and pathological parameters, such as proliferation, invasion, differentiation, vascularity, and interstitial findings. These data as well as image features and quantitative indices like SUV and MTV of PET are subjected to artificial intelligence (AI) for radiomics analysis. Other available data such as MRI and contrast enhancement are welcome by AI for more detailed analyses.
Modalities used in the clinical setting include PET and single photon emission computed tomography, as well as MRI and ultrasonography. Optical imaging, such as fluorescence and bioluminescence imaging, plays an important role in preclinical settings; however, penetration of these signals is too shallow to detect labeled immune cells in clinical situations, and currently used contrast materials, such as gadolinium based agents, super paramagnetic iron oxide, and perfluorocarbon labeled with fluorine-19, for MRI are non-specific for immune cells. Therefore, nuclear medicine imaging is a possible procedure to elucidate anti-cancer immune responses [45][46]. Cell tracking of particular cell subsets would be done by radiolabeling in vitro prior to re-administration or by injecting a radiopharmaceutical that binds to a specific membrane antigen in vivo [47][48]. There have been many radiopharmaceuticals for cell tracking; however, none of these have been successfully used in clinical practice so far (Table 2).
Table 2. Potential Radiopharmaceuticals to Image Immune Cells and Cell Tracking.
| Target | Radiolabeling Agent | Application/Mechanism | References |
|---|---|---|---|
| T lymphocytes | 111In-oxine, 89Zr-oxine | Tumor infiltration | [46][49][50] |
| 18F-FDG | Cytokine production | ||
| SPIO | |||
| NK cells | 111In-oxine, 89Zr-oxine | Tumor infiltration | [46][51][52] |
| 18F-FDG, 11C-methyl iodide | NK cell homing | ||
| SPIO | |||
| Macrophages | 111In-oxine, 89Zr-nanoparticles | Tumor infiltration | [46][53][54][55][56][57] |
| 18F-FDG | Tumor-associated macrophages | ||
| SPIO, 19F-perfluorocarbon | |||
| Interleukin-2 | Iodine-123, Technetium-99 m, Fluorine-18 | Interleukin-2 receptors on T cells | [58] |
| Anti-CD8 cys-diabody | Zirconium-89, Copper-64 | CD8+ T cells | [59][60][61] |
| Anti-CD8 mAb | |||
| PK11195 | Carbon-11 | Tumor-associated macrophages, Translocator protein | [62] |
| Anti-TCR mAb | Copper-64 | Tumor infiltration of T cells | [63] |
| Anti-CD56 mAb | Technetium-99 m | NK cells | [64] |
18F-FDG: 2-[18F] fluoro-2-deoxy-d-glucose; SPIO: super paramagnetic iron oxide; TCR: T cell receptor; mAb: monoclonal antibody.
Quantitative parameters of FDG-PET, such as SUV and TLG, have been used to evaluate therapeutic responses. Metabolic changes and temporal enlargement due to immune cell infiltration seen after immune checkpoint inhibitors, anti-PD-1, and anti-PD-L1 antibodies facilitate the modification of FDG-PET response evaluation criteria as well as conventional RECIST. Dynamic interaction between cancer and immune cells, CSCs, and metabolism of cancer cells in the tumor microenvironment are promising targets to eradicate cancer. Accumulation of FDG reflects glucose metabolism of both cancer cells and immunologically competent cells in the tumor. Considering inter- and intra-patient tumor heterogeneity, immunological reaction to the therapy differs among patients according to the individual immune function and tumor heterogeneity. This limits the use of current response evaluation criteria and the revised ones may not be relevant enough for use in the clinical setting. Then, imaging of immune cells tracking may be crucial but is still a challenge, due to the fact that radiopharmaceuticals or MRI probes which are highly specific for biomarkers expressed in different immune cells are not likely to be determined. A radiomics approach which combines quantitative image features and deep learning algorithms has the potential to improve response assessment on the basis of elucidating pathologic mechanisms in more personalized treatments in the era of precision nuclear medicine. Multimodal imaging to highlight new therapeutic biomarkers in the complexed tumor response may be required to improve the management of cancer patients.