
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | José Santos | + 4080 word(s) | 4080 | 2020-08-06 10:42:04 | | | |
| 2 | Nicole Yin | -49 word(s) | 4031 | 2020-09-01 04:59:18 | | |
Video Upload Options
Marine bacterial natural products are molecules produced by marine bacteria through secondary metabolism genes, which can provide them with competitive advantages. Certain natural products can have bioactive effects (e.g. antimicrobial properties) and thus can be used by humans as therapeutics or leads for novel therapeutics.
1. History
Nature has long been the most important source of therapeutics. The use of poultices and mixtures of plant material to treat infections goes back to the early bronze age civilizations[1]. Building on the prior knowledge acquired, early medicine, pharmacology and chemistry started to develop therapeutics by studying nature. For example, extracts of willow bark (genus Salix), containing salicylic acid, which was identified to be the bioactive molecule present in the bark in 1828, were already used by Sumerians and Egyptians to treat inflammation and pain. In 1852 acetylsalicylic acid was first synthesised and in 1899, Bayer patented it as aspirin[2]. Ever since then, many terrestrial organisms and in particular plants have been sought for use as natural products. It was only around the 1970s, that attention was first given to the ocean as a source of useful natural products. As the oceans cover most of the Earth’s surface, they are home to a substantial portion of the world’s biodiversity[3] which lives in distinctive and varied conditions and has evolved through a long period of metabolic adaptations. When exploration of the ocean’s biodiversity and metabolic richness began, it resulted in the discovery of thousands of structurally unique bioactive marine natural products[4].
Initially, the exploitation of marine wildlife for natural bioactive products focused on a small number of organisms which included sponges, molluscs, tunicates and macroalgae[5]. These were shown to produce a very diverse range of unique molecular structures, like halogenated terpenes, polyketides and prostaglandins[6][7][8][9]. This diversity of bioactive structures is considered to be part of the defence, survival and predatorial strategies employed by these organisms, such as, for example, sponges, which are sessile, soft-bodied organisms, lacking morphological defences like biological armature or spines[10]. Thus, these organisms appeared to be a great source for the discovery of novel bioactive molecules. However, the bioactive molecules produced by these organisms can be present in quite small amounts. For example, halichondrin B (1) (Figure 1), a macrolide first isolated from Halichondria okadai that has potent anticancer activity, impeding mitotic division by targeting tubulin[11], is present in concentrations as low as 400 µg per kg wet weight of tissue of Lissodendoryx sp.[12]. Due to their low concentrations of bioactive molecules the use of these organisms poses environmental problems, because high quantities of organisms would be needed to produce enough molecules to even begin preclinical trials[13]. Yet, its structure inspired a synthetic analogue, eribulin mesylate, which is now used in breast cancer and lipocarcinoma treatment[14]. As such, sponges, molluscs, tunicates and macroalgae still remain relevant sources of new marine natural products[15].
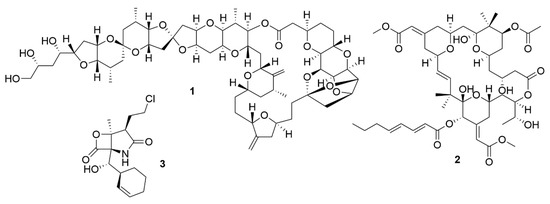
Figure 1. Bioactive metabolites isolated from marine organisms. Halichondrin B (1), a macrolide first isolated from H. okadai but also present in other sponges. Bryostatin 1 (2), which belongs to a family of polyketide macrolides first identified in the marine bryozoan B. neritina. Salinosporamide A (3), is a proteasome inhibitor isolated from bacteria from the genus Salinospora and is in phase III clinical trials for the treatment of multiple myeloma.
The exploitation of other sources of bioactive marine organisms, mainly microorganisms, has also led to the discovery of new promising leads. Indeed, some of the molecules associated with macroorganisms such as sponges, may have their origin in associated microorganisms[16]. This may be the case of bryostatins, found in the marine bryozoan Bugula neritina[17]. The bryostatins, exemplified by bryostatin 1 (2) are polyketide macrolactones with neurological and anticancer properties that work by modulating the activity of the protein kinase C family and they were first isolated from this bryozoan in 1982[18]. Yet, these macrolactones appear to be synthesised by a group of PKS genes of bacterial origin, indicating that bryostatins are produced by a bacterium[19] [56].
Marine microorganisms, like members of Actinobacteria, Proteobacteria, Firmicutes, Cyanobacteria, fungi and dinoflagellates, have shown to be great reservoirs of bioactive molecules[15]. Yet, even though initial predictions pointed to an immediate increase in the number of natural products discovered with the shift in focus from macroorganisms to microorganisms, this did not occur[20]. However, advances in the isolation of novel taxa, provided a boost in the discovery of novel bioactive molecules. Analysis of the literature reveals an increase in the number of new molecules discovered in all microorganisms from 2014 to 2018[15], with a special emphasis on fungi and bacteria. Members of the genus Salinospora (Actinobacteria) are examples of bacteria that lead to the discovery of salinosporamide A (3)[21]. This molecule has a potent cytotoxic activity due to a unique functionalisation of the core-fused γ-lactam-β-lactone bicyclic ring, which contributes significantly to its activity[22]. Salinosporamide A is now in phase III clinical trials for the treatment of multiple myeloma under the brand name Marizomib[23].
Although salinosporamide A is the best example of the potential present in bacteria, many novel molecular structures are discovered each year. In fact, in 2016, 179 new natural products of marine bacterial origin were discovered[24], in 2017, the number rose to 242[25] and in 2018 a total of 240 new molecules were reported[15]. The upwards trend seen in the number of discovered molecules and the remarkable chemical diversity displayed, which ranges from peptides, siderophores and polyketides to esters, macrolactones, quinones and terpenes, shows the bacterial potential for the discovery of novel active principles.
2. Antimicrobial Marine Bacterial Natural Products
Bacteria are promising sources for novel antimicrobial natural product discovery. This is primarily due to two factors. One is their variable and malleable metabolism[26] and the other is their competitive pressure for resources against other microbes. Several recent examples of natural products from marine bacteria are provided below. The novel molecule bacicyclin (4) (Figure 2), which is a cyclic peptide, was isolated from a Bacillus sp. strain BC028 isolated from the common mussel (Mytilus edulis)[27]. It displays antibacterial activity against E. faecalis and S. aureus with minimal inhibitory concentration (MIC) values of 8 and 12 µM, respectively, and can help in the design of analogues with increased antibiotic efficacy[27]. Anthracimycin B (5), a polyketide with powerful anti-Gram-positive bacteria activity that was obtained from a marine-derived Streptomyces cyaneofuscatus M-169, has expanded the knowledge of how the methyl group at C-2 of anthracimycins plays a role in its antibacterial effect[28]. Taromycin B (6), a lipodepsipeptide with potent activity against methicillin-resistant S. aureus and vancomycin-resistant E. faecium which was isolated from the marine actinomycete Saccharomonospora sp. CNQ-490, provides a promising start for the development of novel antibacterial scaffoldings[29]. Janthinopolyenemycin A (7) and B (8) are also polyketides and the first examples of molecules of their structural type. Janthinopolyenemycins were isolated from the proteobacterium Janthinobacterium spp., strains ZZ145 and ZZ148, and have activity against Candida albicans[30]. Streptoseomycin (9), a macrolactone isolated from the actinobacterium Streptomyces seoulensis A01, has specific activity against microaerophilic bacteria, specially the pathogen H. pylori[31]. This restricted activity makes streptoseomycin a good starting point for the discovery of antibiotics for the treatment of H. pylori infections. The polyketides ansalactams B (10), C (11) and D (12) are highly modified ansamycins that show weak and mild anti-methicillin-resistant S. aureus and were identified in cultures of Streptomyces sp. CNH189, isolated from marine sediments. Ansalactams B and D are cyclic polyketides with similarities to ansalactam A. However, ansalactam D shows evidence of an uncommon oxetane ring. Ansalactam C is an open polyketide chain resulting from a Baeyer–Villiger-type oxidation[32]. Micromonohalimanes A (13) and B (14) are rare halimane-type diterpenoids isolated from the actinobacterium Micromonospora sp. WMMC-218. Micromonohalimane A displays a very weak inhibitory effect on methicillin-resistant S. aureus while micromonohalimane B displays moderate bacteriostatic activity against it. Xestostreptin (15) is a modified diketopiperazine isolated from Streptomyces sp. S.4, resulting from the condensation of the aminoacids threonine and alanine[33]. Xestostreptin shows weak activity against the malarial agent P. falciparum. Two macrolides, branimycins B (16) and C (17) were identified from a fermentation of the actinobacterium Pseudonocardia carboxydivorans M-227, isolated from deep sea water[34]. Branimycin B shows moderate antibacterial activity against Gram-positive bacteria, while branimycin C displayed moderate antibacterial activity against Gram-negative bacteria.
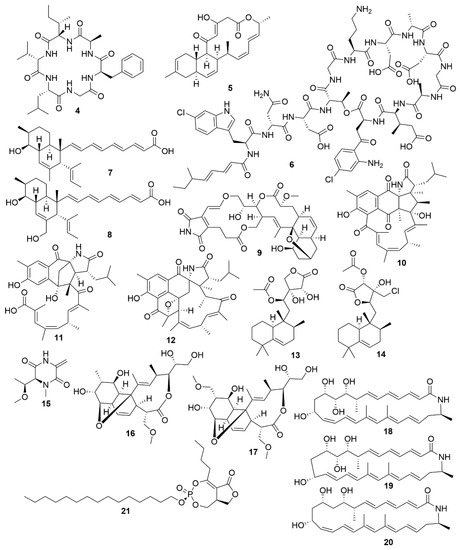
Figure 2. Examples of recently isolated antimicrobial natural products from marine bacteria. Bacicyclin (4), a cyclic peptide isolated from a Bacillus sp. BC028. Anthramicin B (5), a polyketide isolated from S. cyaneofuscatus M-169. Taromycin B (6), a lipopeptide from Saccharomonospora sp. CNQ-490. Janthinopolyenemycin A (7) and Janthinopolyenemycin B (8). The janthinopolyenemycins are polyketides isolated from two strains of the genus Janthinobacterium. Streptoseomycin (9), a macrolactone isolated from S. seoulensis. Ansalactam B (10), a pentacyclic polyketide. Ansalactam C (11), an open polyketide unlike ansalactam B. Ansalactam D (12), a hexacyclic polyketide. Ansalactams B, C and D were isolated from Streptomyces sp. CNH189. Micromonohalimane A (13). Micromonohalimane B (14). Micromonohalimanes A and B are terpenes isolated from Micromonospora sp. WMMC-218. Xestostreptin (15), a diketopiperazine isolated from Streptomyces sp. S.4. Branimycin B (16) and Branimycin C (17) C are macrolides isolated from the deep-sea bacterium P. carboxydivorans M-227. Lobosamide A (18), Lobosamide B (19) and Lobosamide C (20) are macrolactams isolated from Micromonospora sp. RL09-050-HVF-A. Salinipostin A (21), a bicyclic phosphotriester isolated from Salinispora sp. RLUS08-036-SPS-B.
Utilizing a genome-assisted discovery strategy, three macrolactams, lobosamides A (18), B (19) and C (20), were isolated from Micromonospora sp. RL09-050-HVF-A[35]. Lobosamides A and B showed bioactivity against the microbial agent of African trypanosomiasis, Trypanosoma brucei in low concentrations. However, lobosamide C was not bioactive. Schulze and colleagues[36] also identified salinipostins A-K bicyclic phosphotriesters isolated from the actinobacterium Salinispora sp. RLUS08-036-SPS-B with potent and selective activity against P. falciparum. The salinipostin scaffold considerably differs from any of the known antimalarial compounds, representing a novel lead structure in the development of therapeutics for malaria. Experiments with salinipostin A (21), the most bioactive of the 11 salinipostins, indicate that it exhibits growth stage-specific effects and no apparent resistance could be identified in parasite populations. A hybrid peptide-polyketide, mollemycin A (22) (Figure 3) was isolated from the marine bacterium Streptomyces sp. CMB-M0244 and shows potent antimalarial and broad antibacterial activities[37]. Actinosporin A (23) is a glycosilated polyketide which shows antiparasitic activity against T. brucei and was isolated from a marine sponge associated Actinokineospora sp. EG49[38]. Likewise, actinosporin B, was also isolated but showed no bioactivity, suggesting that actinosporin A is acting selectively against the parasite. The linear lipopetides, gageopeptides A-D (24–27), gageotetrins A–C (28–30) and gageostatins A–C (31–33) were isolated from the marine Bacillus subtilis 109GGC020. These lipopetides showed a range of different antimicrobial bioactivities, with gageostatins A, B and C all showing good antimicrobial activity and moderate cytotoxic activity to lung cancer cell line NCI-H23[39]. Gageotetrins A, B and C showed potent antimicrobial bioactivities but not cytotoxic effect on human myeloid leukaemia K-562[40]. Furthermore, gageopeptides A, B, C and D all showed good antifungal and moderate broad antibacterial activity, while not showing cytotoxicity to human myeloid leukaemia K-562 and mouse leukemic macrophage RAW 264.7 cell lines[41].
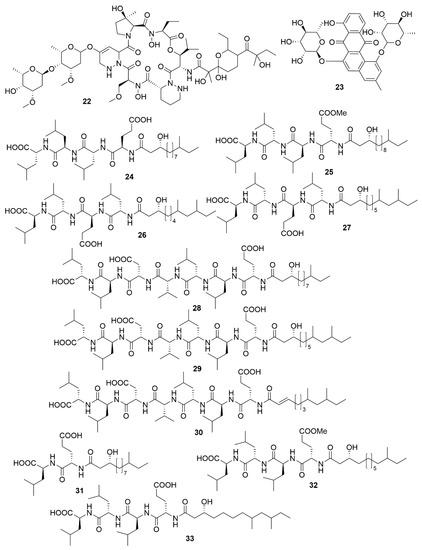
Figure 3. Examples of recently isolated antimicrobial natural products from marine bacteria. Mollemycin A (22) is a hybrid peptide-polyketide isolated from Streptomyces sp. CMB-M0244. Actinosporin A (23), a polyketide isolated from Actinokineospora sp. EG49. Gageopeptide A (24). Gageopeptide B (25). Gageopeptide C (26), Gageopeptide D (27), Gageotetrin A (28). Gageotetrin B (29). Gageotetrin C (30). Gageostatin A (31). Gageostatin B (32). Gageostatin C (33). The gageopeptides, gageotetrins and gageostatins are linear lipopeptides isolated from B. subtilis 109GGC020.
3. Antiviral Marine Bacterial Natural Products
It is estimated that as many as 1031 viruses inhabit the oceans[42], with concentrations ranging from 3 × 106 viruses mL−1 in deep sea waters to 108 viruses mL−1 in coastal waters[43], many of which are bacteriophages. Consequently, marine bacteria are subjected to evolutionary pressure to develop defences against viral attacks. As such, marine bacteria may be great reservoirs of antiviral leads.
A number of examples of marine bacterial natural products with antiviral bioactivities have been recently reported. Three novel abyssomicin monomers, neoabyssomicins D (34) (Figure 4), E and A2 and two dimers—neoabyssomicins F (35) and G (36)—were isolated from the marine Streptomyces koyangensis SCSIO 5802, with neoabyssomicin D showing moderate anti-herpes simplex virus activity, and neoabyssomicins F and G showing low activity against vesicular stomatitis virus[44]. Streptomyces sp. OUCMDZ-3434, isolated from the marine alga Enteromorpha prolifera, was shown to produce five new phenolic polyketides[45]. Of these molecules, wailupemycin J (37) and (R)-wailupemycin K (38) proved to be bioactive against the influenza A virus (H1N1). The indolosesquiterpenoids xiamycins C (39), D (40) and E (41) were isolated from the marine-derived Streptomyces sp. #HK18, and showed strong inhibitory effect against the coronavirus porcine epidemic diarrhoea virus[46]. As such, xiamycins may provide useful leads in the development of antivirals with broader spectrum activity against other coronaviruses.
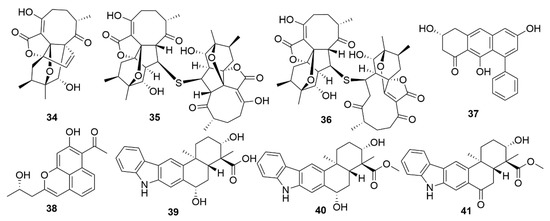
Figure 4. Examples of recently isolated natural products from marine bacteria with antiviral activity. Neoabyssomicin D (34). Neoabyssomicin F (35) and Neoabyssomicin E (36) are polycyclic polyketides isolated from S. koyangensis SCSIO 5802. Wailupemycin J (37) and (R)-wailupemycin K (38) are phenolic polyketides isolated from Streptomyces sp. OUCMDZ-3434. Xiamycin C (39), Xiamycin D (40) and Xiamycin E (41) were isolated from Streptomyces sp. #HK18.
4. Anticancer Marine Bacterial Natural Products
As with antimicrobials, it is ascertained that marine bacteria are great reservoirs for cytotoxic natural products. While salinosporamide A, already mentioned above, is a great example of marine cytotoxic drug discovery [58], every year, novel anticancer bioactive structures are discovered. Actinobacteria, especially those of the genera Streptomyces and Micromonospora, have been a very prolific source of cytotoxic compounds. There are several recent examples of structures isolated from these bacteria. Dentigerumycin E (42) (Figure 5), is a cyclic hexapeptide bearing three piperazic acids and a pyran-bearing polyketide acyl chain, isolated from the marine actinoabcterium Streptomyces albogriseolus JB5[47]. It showed moderate cytotoxicity against lung carcinoma A549, colorectal cancer HCT116, breast cancer MDA-MB-231, liver cancer SK-HEP-1 and stomach cancer SNU638 cell lines, while not being cytotoxic to the normal human breast epithelial cell line MCF-10A [84]. Likewise, neothioviridamide (43) is a polythioamide cyclic peptide with strong cytotoxicity against human ovarian adenocarcinoma (SKOV-3), malignant pleural mesothelioma (Meso-1) and immortalized human T lymphocyte (Jurkat) cell lines[48]. It was isolated after discovery of a novel biosynthetic cluster (thioviridamide-like biosynthetic gene) in Streptomyces sp. MSB090213SC12 by genome mining and heterologous expression of a bacterial artificial chromosome in Streptomyces avermitilis SUKA. Three cyclic depsipeptides, rakicidins G-I (44–46), isolated from the marine actinobacterium Micromonospora chalcea FIM 02–523, have potent cytotoxic activity against the human pancreatic cancer cell line PANC-1 and human colon carcinoma cell line HCT-8[49]. Rakicidins G-I differ in the length of their β-hydroxy fatty acid moiety. The 26-membered polyene macrolactam, FW05328-1 (47), isolated from Micromonospora sp. FIM05328, has potent bioactivity against three cells lines of human oesophageal squamous cell carcinoma (KYSE30, KYSE180 and EC109)[49]. The integration of genomic data in association with nuclear magnetic resonance (NMR) analysis allowed the determination of the stereostructure of neaumycin B, a cytovaricin-ossamycin-oligomycin macrolide, that was isolated from Micromonospora sp. CNY-010[50]. Neaumycin B (48) has shown potent anti-human glioblastoma cell line U87 activity but it is unstable. Through genetic manipulation of promoters, six new polyketides, pactamides A–F, were isolated from Streptomyces pactum SCSIO 02999. Pactamides B-F showed low to moderate cytotoxic activity against human glioblastoma cell line (SF-268), human breast cancer cell (MCF-7), human large-cell lung carcinoma (NCI-H460) and human liver cancer cell (HepG2) while pactamide A (49) showed potent activity[51]. Two cyclodepsipeptides, streptodepsipeptides P11A (50) and P11B (51), were isolated from Streptomyces sp. P11-23B and displayed potent anti-proliferative bioactivity in four cell lines of human glioblastoma (U251, U87-MG, SHG-44 and C6)[52]. Research with Streptomyces sp. strain THS-55 yielded four new antimycin alkaloids, antimycins E-H (52–55), which showed potent cytotoxic effect on HPV-transformed human cervix adenocarcinoma (HeLa) cells and moderate anti-proliferative activity in human cervical cancer cell SiHa, human myelogenous leukaemia cell line K562, and human leukaemia cell line HL-60[53]. However, cytotoxic effects were shown in healthy human embryonic kidney cells 293T. In a knockout mutant of Streptomyces sp. CHQ-64, two new alkaloids, geranylpyrrol A and piericidin F (56), were discovered[54]. Of these, piericidin F showed potent anti-proliferative activity against several cancer cell lines, including HeLa, human acute promyelocytic leukaemia (NB4) and human lung carcinoma (A549 and H1975).
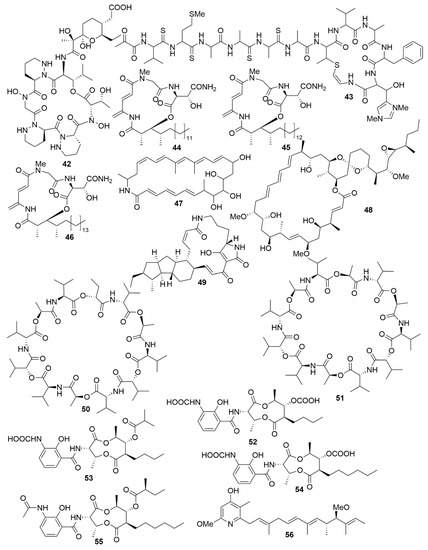
Figure 5. Examples of recently isolated cytotoxic natural products from marine bacteria. Dentigerumycin E (42), a cyclic hexapeptide isolated from S. albogriseolus JB5. Neothioviridamide (43), a cyclic peptide from Streptomyces sp. MSB090213SC12. Rakicidin G (44), Rakicidin H (45) and Rakicidin I (46) are cyclic depsipeptides isolated from M. chalcea FIM 02–523. FW05328-1 (47), a polyene macrolactam isolated from Micromonospora sp FIM05328. Neaumycin B (48), a macrolide from Micromonospora sp. CNY-010. Pactamide A (49), a polyketide isolated from S. pactum SCSIO 02999. Streptodepsipeptide P11A (50) and Streptodepsipeptide P11B (51) are cyclodepsipeptides from Streptomyces sp. P11-23B. Antimycin E (52), Antimycin F (53), Antimycin G (54) and Antimycin H (55) are alkaloids isolated from Streptomyces sp. THS-55. Piericidin F (56) is an alkaloid isolated Streptomyces sp. CHQ-64.
Two new chromodepsipeptides, neo-actinomycin A (57) (Figure 6) and neo-actinomycin B (58) were isolated from a marine-derived Streptomyces sp. IMB094[55]. They showed strong cytotoxic effect on human colorectal carcinoma cell line HCT116 and human lung carcinoma cell line A549.
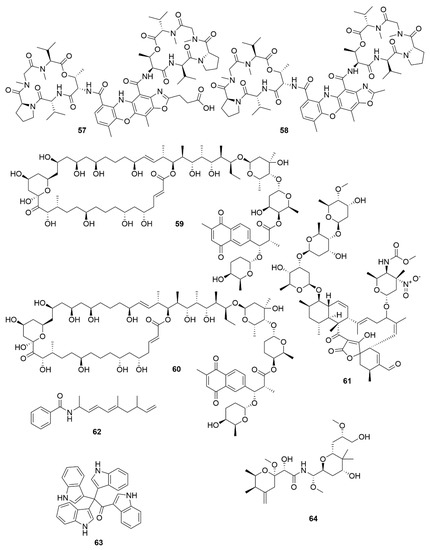
Figure 6. Examples of recently isolated cytotoxic natural products from marine bacteria. Neo-actinomycin A (57) and Neo-actinomycin B (58) are chromopeptides from Streptomyces sp. IMB094. PM100117 (59) and PM100118 (60) are macrolides isolated from S. caniferus GUA-06-05-006A. Lobophorin I (61), a spirotetronate isolated from Streptomyces sp. 1053U.I.1a.3b. Haliamide (62), a hybrid of a polyketide synthase from H. ochraceum SMP-2. Tetra(indol-3-yl)ethenone (63), an indole isolated from P. denitrificans BBCC725. O-Demethylpederin (64), a polyketide with a tetrahydropyran-core from Labrenzia sp. PHM005.
Two new macrolides, PM100117 (59) and PM100118 (60) were isolated from a marine Streptomyces caniferus GUA-06-05-006A[56]. Both PM100117 and PM100118 show potent cytotoxic effect on human breast adenocarcinoma (MDA-MB-231), human lung carcinoma (A549) and human colorectal carcinoma (HT-29) cell lines. The study of a symbiotic Streptomyces sp. (strain 1053U.I.1a.3b) of cone snails lead to the isolation of two lobophorins, H and I[57]. Lobophorins are a large family of spirotetronates with antimicrobial and cytotoxic bioactivities. Of these, lobophorin I (61) showed potent cytotoxic activity against human T-cell leukaemia cell line CEM-TART.
Besides Actinobacteria, other marine bacterial phyla are also proving to be relevant for the isolation of novel bioactive molecules. As a result of a hybrid polyketide synthase (PKS) and non-ribosomal peptide synthetase (NRPS) biosynthesis, haliamide (62) was isolated from a marine myxobacterium, Haliangium ochraceum SMP-2, and shows moderate cytotoxicity towards the HeLa cell line[58]. A novel cytotoxic indole, tetra(indol-3-yl)ethenone (63) was isolated from the marine proteobacterium Pseudovibrio denitrificans BBCC725[59]. Tetra(indol-3-yl)ethenone has moderate cytotoxicity to human lung carcinoma cell line A549 and the mouse fibroblasts cell line L929. Another marine proteobacterium, Labrenzia sp. PHM005 produced a new tetrahydropyran-core polyketide and analogue of pederin[60]. This novel pederin, 18-O-demethylpederin (64) shows potent anti-proliferative activity against four cell lines: human lung carcinoma cell line A549, human colon adenocarcinoma cell line HT-29, human breast adenocarcinoma cell line MDA-MB-231 and human pancreas adenocarcinoma cell line PSN-1.
All these examples show the great potential displayed by marine bacteria which reveals the extraordinary chemical diversity of their metabolism. When analysing the bioactive bacteria and their taxonomic groups, Actinobacteria proved to be the most prolific and diverse producers (Tables 1–3). However, ignoring other less studied phyla denies access to valuable chemical diversity, which is essential in the drug discovery process. Moreover, marine bacterial metabolites have shown to have potential as treatment in both human and animal pathologies. Thus, marine bacterial natural products have great significance under the “One Health” framework.
Table 1. Recently isolated bioactive molecules from marine bacteria with antimicrobial properties.
|
Molecule |
Bacterial Origin |
Chemical Structure |
Bioactivity |
||
|
Strain Identification |
Phyla |
Effect |
Target |
||
|
Actinosporin A |
Actinokineospora sp. EG49. |
Actinobacteria |
Polyketide |
AP |
TBB |
|
Lobosamide A |
Micromonospora sp. RL09-050-HVF-A |
Actinobacteria |
Polyketide |
AP |
TBB |
|
Lobosamide B |
Micromonospora sp. RL09-050-HVF-A |
Actinobacteria |
Polyketide |
AP |
TBB |
|
Micromonohalimane A |
Micromonospora sp. WMMC-218 |
Actinobacteria |
Polyketide |
AB |
MRSA |
|
Micromonohalimane B |
Micromonospora sp. WMMC-218 |
Actinobacteria |
Polyketide |
AB |
MRSA |
|
Branimycin B |
Pseudonocardia carboxydivorans M-227 |
Actinobacteria |
Polyketide |
AB |
G+ |
|
Branimycin C |
Pseudonocardia carboxydivorans M-227 |
Actinobacteria |
Polyketide |
AB |
G- |
|
Taromycin B |
Saccharomonospora sp. CNQ-490 |
Actinobacteria |
Peptide |
AB |
MRSA; VRE |
|
Salinipostin A |
Salinispora sp. RLUS08-036-SPS-B |
Actinobacteria |
Polyketide |
AP |
PF |
|
Anthracimycin B |
Streptomyces cyaneofuscatus M-169 |
Actinobacteria |
Polyketide |
AB |
G+ |
|
Streptoseomycin |
Streptomyces seoulensis A01 |
Actinobacteria |
Polyketide |
AB |
H. pylori |
|
Mollemycin A |
Streptomyces sp. CMB-M0244 |
Actinobacteria |
Peptide-polyketide |
AB/AP |
Broad spectrum/PF |
|
Ansalactam B |
Streptomyces sp. CNH189 |
Actinobacteria |
Polyketide |
AB |
MRSA |
|
Ansalactam C |
Streptomyces sp. CNH189 |
Actinobacteria |
Polyketide |
AB |
MRSA |
|
Ansalactam D |
Streptomyces sp. CNH189 |
Actinobacteria |
Polyketide |
AB |
MRSA |
|
Xestostreptin |
Streptomyces sp. S.4 |
Actinobacteria |
Peptide |
AP |
PF |
|
Bacicyclin |
Bacillus sp. BC028 |
Firmicutes |
Peptide |
AB |
G+ |
|
Gageopeptide A |
Bacillus subtillis strain 109GGC020 |
Firmicutes |
Peptide |
AB/AF |
Broad spectrum |
|
Gageopeptide B |
Bacillus subtillis strain 109GGC020 |
Firmicutes |
Peptide |
AB/AF |
Broad spectrum |
|
Gageopeptide C |
Bacillus subtillis strain 109GGC020 |
Firmicutes |
Peptide |
AB/AF |
Broad spectrum |
|
Gageopeptide D |
Bacillus subtillis strain 109GGC020 |
Firmicutes |
Peptide |
AB/AF |
Broad spectrum |
|
Gageotetrin A |
Bacillus subtillis strain 109GGC020 |
Firmicutes |
Peptide |
AF |
Broad spectrum |
|
Gageotetrin B |
Bacillus subtillis strain 109GGC020 |
Firmicutes |
Peptide |
AF |
Broad spectrum |
|
Gageotetrin C |
Bacillus subtillis strain 109GGC020 |
Firmicutes |
Peptide |
AF |
Broad spectrum |
|
Gageostatin A |
Bacillus subtillis strain 109GGC020 |
Firmicutes |
Peptide |
AB/AF |
Broad spectrum |
|
Gageostatin B |
Bacillus subtillis strain 109GGC020 |
Firmicutes |
Peptide |
AB/AF |
Broad spectrum |
|
Gageostatin C |
Bacillus subtillis strain 109GGC020 |
Firmicutes |
Peptide |
AB/AF |
Broad spectrum |
|
Janthinopolyenemycin A |
Janthinobacterium spp. ZZ145 and ZZ148 |
Proteobacteria |
Polyketide |
AF |
CA |
|
Janthinopolyenemycin B |
Janthinobacterium spp. ZZ145 and ZZ148 |
Proteobacteria |
Polyketide |
AF |
CA |
|
AB = Antibacterial, AF = Antifungal AP = Antiparasitic; CA = Candida albicans; G+ = Gram-positive bacteria; G- = Gram-negative bacteria; MRSA = Methicillin-resistant S. aureus; PF = Plasmodium falciparum; TBB = Trypanosoma brucei; VRE = Vancomycin-resistant E. faecium |
|||||
Table 2. Recently isolated bioactive molecules from marine bacteria with antiviral properties.
|
Molecule |
Bacterial Origin |
Chemical Structure |
Bioactivity |
|
|
Strain Identification |
Phyla |
Target |
||
|
Neoabyssomicin D |
S. koyangensis SCSIO 5802 |
Actinobacteria |
Polyketide |
HSV |
|
Neoabyssomicin F |
S. koyangensis SCSIO 5802 |
Actinobacteria |
Polyketide |
VSV |
|
Neoabyssomicin G |
S. koyangensis SCSIO 5802 |
Actinobacteria |
Polyketide |
VSV |
|
Wailupemycin J |
Streptomyces sp. OUCMDZ-3434 |
Actinobacteria |
Polyketide |
H1N1 |
|
R-wailupemycin K |
Streptomyces sp. OUCMDZ-3435 |
Actinobacteria |
Polyketide |
H1N1 |
|
Xiamycin C |
Streptomyces sp. #HK18 |
Actinobacteria |
Polyketide |
PEDV |
|
Xiamycin D |
Streptomyces sp. #HK18 |
Actinobacteria |
Polyketide |
PEDV |
|
Xiamycin E |
Streptomyces sp. #HK18 |
Actinobacteria |
Polyketide |
PEDV |
HSV = Herpes simplex virus; VSV = vesicular stomatitis virus; H1N1 = influenza A virus; PEDV = porcine epidemic diarrhea virus
Table 3. Recently isolated bioactive molecules from marine bacteria with cytotoxic properties.
|
Molecule |
Bacterial Origin |
Chemical Structure |
Bioactivity |
|
|
Strain Identification |
Phyla |
Target |
||
|
Dentigerumycin E |
Streptomyces albogriseolus JB5 |
Actinobacteria |
Peptide |
HCT116; A549; MDA-MB-231; SK-HEP-1; SNU638 |
|
Neothioviridamide |
Streptomyces sp. MSB090213SC12 |
Actinobacteria |
Peptide |
SKOV-3; Meso-1; Jurkat |
|
Rakicidin G |
Micromonospora chalcea FIM 02-523 |
Actinobacteria |
Peptide |
PANC-1; HCT-8 |
|
Rakicidin H |
Micromonospora chalcea FIM 02-523 |
Actinobacteria |
Peptide |
PANC-1; HCT-8 |
|
Rakicidin I |
Micromonospora chalcea FIM 02-523 |
Actinobacteria |
Peptide |
PANC-1; HCT-8 |
|
FW05328-1 |
Micromonospora sp. FIM05328 |
Actinobacteria |
Polyketide |
KYSE30; KYSE180; EC109 |
|
Neaumycin B |
Micromonospora sp. CNY-010 |
Actinobacteria |
Polyketide |
U87 |
|
Pactamide A |
Streptomyces pactum SCSIO 02999 |
Actinobacteria |
Polyketide |
SF-268; MCF-7; NCI-H460; Hep-G2 |
|
Pactamide B |
Streptomyces pactum SCSIO 02999 |
Actinobacteria |
Polyketide |
SF-268; MCF-7; NCI-H460; Hep-G2 |
|
Pactamide C |
Streptoyces pactum SCSIO 02999 |
Actinobacteria |
Polyketide |
SF-268; MCF-7; NCI-H460; Hep-G2 |
|
Pactamide D |
Streptomyces pactum SCSIO 02999 |
Actinobacteria |
Polyketide |
SF-268; MCF-7; NCI-H460; Hep-G2 |
|
Pactamide E |
Streptomyces pactum SCSIO 02999 |
Actinobacteria |
Polyketide |
SF-268; MCF-7; NCI-H460; Hep-G2 |
|
Pactamide F |
Streptomyces pactum SCSIO 02999 |
Actinobacteria |
Polyketide |
SF-268; MCF-7; NCI-H460; Hep-G2 |
|
Streptodepsipeptide P11A |
Streptomyces sp. P11-23B |
Actinobacteria |
Peptide |
U251; U87; SHG-44; C6 |
|
Streptodepsipeptide P11B |
Streptomyces sp. P11-23B |
Actinobacteria |
Peptide |
U251; U87; SHG-44; C6 |
|
Antimycin E |
Streptomyces sp. THS-55 |
Actinobacteria |
Polyketide |
HeLa; SiHa; K562; HL-60; 293T |
|
Antimycin F |
Streptomyces sp. THS-55 |
Actinobacteria |
Polyketide |
HeLa; SiHa; K562; HL-60; 293T |
|
Antimycin G |
Streptomyces sp. THS-55 |
Actinobacteria |
Polyketide |
HeLa; SiHa; K562; HL-60; 293T |
|
Antimycin H |
Streptomyces sp. THS-55 |
Actinobacteria |
Polyketide |
HeLa; SiHa; K562; HL-60; 293T |
|
Piericidin F |
Streptomyces sp. CHQ-64 |
Actinobacteria |
Polyketide |
HeLa; NB4; A549; H1975 |
|
Neo-actinomycin A |
Streptomyces sp. IMB094 |
Actinobacteria |
Peptide |
HCT116; A549 |
|
Neo-actinomycin B |
Streptomyces sp. IMB094 |
Actinobacteria |
Peptide |
HCT116; A549 |
|
PM100117 |
Streptomyces caniferus GUA-06-05-006A |
Actinobacteria |
Polyketide |
A549; MDA-MB-231; HT-29 |
|
PM100118 |
Streptomyces caniferus GUA-06-05-006A |
Actinobacteria |
Polyketide |
A549; MDA-MB-231; HT-29 |
|
Lobophorin I |
Streptomyces sp. 1053U.I.1a.3b |
Actinobacteria |
Polyketide |
CEM-TART |
|
Haliamide |
Haliangium ochraceum SMP-2 |
Myxobacteria |
Polyketide |
HeLa |
|
Tetra(indol-3-yl)ethanone |
Pseudovibrio denitrificans BBCC725 |
Proteobacteria |
Polyketide |
L929; A549 |
|
18-O-demethylpederin |
Labrenzia sp. PHM005 |
Proteobacteria |
Polyketide |
A549; HT-29; MDA-MB-231; PSN-1; |
HCT116 = HCT-8 = HT-29 = human colorectal carcinoma; A549 = H1975 = human lung carcinoma; MDA-MB-231 = MCF-7 = human breast adenocarcinoma; SK-HEP-1 = Hep-G2 = human hepatic adenocarcinoma; SNU638 = human gastric carcinoma; SKOV-3 = human ovarian adenocarcinoma; Meso-1 = malignant pleural mesothelioma; Jurkat = immortalized human T lymphocyte; PANC-1 = PSN-1 = human pancreas adenocarcinoma; KYSE30 = KYSE180 = EC109 = human oesophageal squamous cell carcinoma; U251 = U87 = SHG-44 = C6 = SF-268 = human glioblastoma cell line; HeLa = SiHa = human cervix adenocarcinoma; K562 = HL-60 = NB4 = human leukaemia cell line; 293T = human embryonic kidney cells; L929 = mouse fibroblasts cell; CEM-TART = human T-cell leukaemia
References
- R D Forrest; Early history of wound treatment.. Journal of the Royal Society of Medicine 1982, 75, 198-205.
- Maria Rosa Montinari; Sergio Minelli; Raffaele De Caterina; The first 3500 years of aspirin history from its roots – A concise summary. Vascular Pharmacology 2019, 113, 1-8, 10.1016/j.vph.2018.10.008.
- Burkhard Haefner; Drugs from the deep: marine natural products as drug candidates.. Drug Discovery Today 2003, 8, 536-544, 10.1016/s1359-6446(03)02713-2.
- Daniel A. Dias; Sylvia Urban; Ute Roessner; A Historical Overview of Natural Products in Drug Discovery. Metabolites 2012, 2, 303-336, 10.3390/metabo2020303.
- William H Gerwick; Bradley S. Moore; Lessons from the Past and Charting the Future of Marine Natural Products Drug Discovery and Chemical Biology. Chemistry & Biology 2012, 19, 85-98, 10.1016/j.chembiol.2011.12.014.
- Weinheimer, A.J.; Spraggins, R.L. The occurrence of two new prostaglandin derivatives (15-epi-PGA2 and its acetate, methyl ester) in the gorgonian Plexaura homomalla chemistry of coelenterates. XV. Tetrahedron Lett. 1969, 10, 5185–5188.
- Masuda, M.; Abe, T.; Suzuki, T.; Suzuki, M. Morphological and chemotaxonomic studies on Laurencia composita and L. okamurae (Ceramiales, Rhodophyta). Phycologia 1996, 35, 550–562.
- Taylor, R.E. Tedanolide and the evolution of polyketide inhibitors of eukaryotic protein synthesis. Nat. Prod. Rep. 2008, 25, 854–861.
- Selvin, J.; Ninawe, A.S.; Kiran, G.; Arya, S.; Priyadharshini, S. Bioactive Marine Natural Products: Insights into Marine Microbes, Seaweeds, and Marine Sponges as Potential Sources of Drug Discovery. In Marine Microorganisms Extraction and Analysis of Bioactive Compounds, 1st ed.; Nollet, L.M.L., Ed.; CRC Press: Boca Raton, FL, USA, 2016
- M S Laport; O C S Santos; G. Muricy; Marine sponges: potential sources of new antimicrobial drugs.. Current Pharmaceutical Biotechnology 2009, 10, 86-105, 10.2174/138920109787048625.
- R L Bai; K D Paull; C L Herald; L Malspeis; G R Pettit; E Hamel; Halichondrin B and homohalichondrin B, marine natural products binding in the vinca domain of tubulin. Discovery of tubulin-based mechanism of action by analysis of differential cytotoxicity data.. Journal of Biological Chemistry 1991, 266, 15882-15889.
- Detmer Sipkema; Ronald Osinga; Wolfgang Schatton; Dominick Mendola; Johannes Tramper; Large-scale production of pharmaceuticals by marine sponges: Sea, cell, or synthesis?. Biotechnology and Bioengineering 2005, 90, 201-222, 10.1002/bit.20404.
- Murray H. G. Munro; John W. Blunt; Eric J. Dumdei; Sarah J.H. Hickford; Rachel E. Lill; Shangxiao Li; Christopher N. Battershill; Alan R Duckworth; The discovery and development of marine compounds with pharmaceutical potential.. Journal of Biotechnology 1999, 70, 15-25, 10.1016/s0168-1656(99)00052-8.
- Sudeep Gupta; Nishitha Shetty; Eribulin drug review. South Asian Journal of Cancer 2014, 3, 57-59, 10.4103/2278-330X.126527.
- Anthony R. Carroll; Brent R. Copp; Rohan A. Davis; Robert A. Keyzers; Michele R. Prinsep; Marine natural products. Natural Product Reports 2020, 37, 175-223, 10.1039/c9np00069k.
- Margo G. Haygood; E W Schmidt; S K Davidson; D J Faulkner; Microbial symbionts of marine invertebrates: opportunities for microbial biotechnology.. Journal of Molecular Microbiology and Biotechnology 1999, 1, 33–43.
- G. R. Pettit; The Bryostatins. Fortschritte der Chemie organischer Naturstoffe / Progress in the Chemistry of Organic Natural Products 1991, 57, 153-195, 10.1007/978-3-7091-9119-4_3.
- George R. Pettit; Cherry L. Herald; Dennis L. Doubek; Delbert L. Herald; Edward Arnold; Jon Clardy; Isolation and structure of bryostatin 1. Journal of the American Chemical Society 1982, 104, 6846-6848, 10.1021/ja00388a092.
- Sebastian Sudek; Nicole B. Lopanik; Laura E. Waggoner; Mark Hildebrand; Christine Anderson; Haibin Liu; Amrish Patel; David H. Sherman; Margo G. Haygood; Christine M. Bassis; et al. Identification of the Putative Bryostatin Polyketide Synthase Gene Cluster from “CandidatusEndobugula sertula”, the Uncultivated Microbial Symbiont of the Marine BryozoanBugula neritina. Journal of Natural Products 2007, 70, 67-74, 10.1021/np060361d.
- John Faulkner, D.; Marine natural products. Nat. Prod. Rep. 2000, 17, 7–55.
- Robert H. Feling; Greg O. Buchanan; Tracy J. Mincer; Christopher A. Kauffman; Paul R. Jensen; William Fenical; Salinosporamide A: A Highly Cytotoxic Proteasome Inhibitor from a Novel Microbial Source, a Marine Bacterium of the New Genus Salinospora. Angewandte Chemie International Edition 2003, 42, 355-357, 10.1002/anie.200390115.
- John W. Blunt; Brent R. Copp; Murray H. G. Munro; Peter T. Northcote; Marine natural products. Natural Product Reports 2005, 22, 15-61, 10.1039/b415080p.
- Sergey A. Dyshlovoy; Friedemann Honecker; Marine Compounds and Cancer: The First Two Decades of XXI Century. Marine Drugs 2019, 18, 20, 10.3390/md18010020.
- John W. Blunt; Anthony R. Carroll; Brent R. Copp; Rohan A. Davis; Robert A. Keyzers; Michele R. Prinsep; Marine natural products. Natural Product Reports 2018, 35, 8-53, 10.1039/c7np00052a.
- Anthony R. Carroll; Brent R. Copp; Rohan A. Davis; Robert A. Keyzers; Michele R. Prinsep; Marine natural products. Natural Product Reports 2019, 36, 122-173, 10.1039/c8np00092a.
- Marnix H. Medema; Michael A. Fischbach; Computational approaches to natural product discovery. Nature Chemical Biology 2015, 11, 639-648, 10.1038/nchembio.1884.
- Jutta Wiese; Usama Ramadan Abdelmohsen; Asa Motiei; Ute Hentschel Humeida; Johannes F. Imhoff; Bacicyclin, a new antibacterial cyclic hexapeptide from Bacillus sp. strain BC028 isolated from Mytilus edulis. Bioorganic & Medicinal Chemistry Letters 2018, 28, 558-561, 10.1016/j.bmcl.2018.01.062.
- Víctor Rodríguez; Jesús Martín; Aida Sarmiento-Vizcaíno; Mercedes De La Cruz; Luis A. García; Gloria Blanco; Fernando Reyes; Anthracimycin B, a Potent Antibiotic against Gram-Positive Bacteria Isolated from Cultures of the Deep-Sea Actinomycete Streptomyces cyaneofuscatus M-169. Marine Drugs 2018, 16, 406, 10.3390/md16110406.
- Kirk A Reynolds; Hanna Luhavaya; Jie Li; Samira Dahesh; Victor Nizet; Kazuya Yamanaka; Bradley S. Moore; Isolation and structure elucidation of lipopeptide antibiotic taromycin B from the activated taromycin biosynthetic gene cluster.. The Journal of Antibiotics 2017, 71, 333-338, 10.1038/ja.2017.146.
- Komal Anjum; Izhar Sadiq; Lei Chen; Sidra Kaleem; Xing-Cong Li; Zhizhen Zhang; Xiao-Yuan Lian; Novel antifungal janthinopolyenemycins A and B from a co-culture of marine-associated Janthinobacterium spp. ZZ145 and ZZ148. Tetrahedron Letters 2018, 59, 3490-3494, 10.1016/j.tetlet.2018.08.022.
- Bo Zhang; Kai Biao Wang; Wen Wang; Shu Feng Bi; Ya Ning Mei; Xin Zhao Deng; Rui Hua Jiao; Ren-Xiang Tan; Hui Ming Ge; Discovery, Biosynthesis, and Heterologous Production of Streptoseomycin, an Anti-Microaerophilic Bacteria Macrodilactone. Organic Letters 2018, 20, 2967-2971, 10.1021/acs.orglett.8b01006.
- Tu Cam Le; Inho Yang; Yeo Joon Yoon; Sang-Jip Nam; William Fenical; Ansalactams B–D Illustrate Further Biosynthetic Plasticity within the Ansamycin Pathway. Organic Letters 2016, 18, 2256-2259, 10.1021/acs.orglett.6b00892.
- L. Harinantenaina Rakotondraibe; Rado Rasolomampianina; Hyun-Young Park; Jie Li; Carla Slebodnik; Peggy J. Brodie; Leah C. Blasiak; Russel Hill; Karen TenDyke; Yongchun Shen; et al.María B. CasseraFelicite RejoDavid G. I. Kingston Antiproliferative and antiplasmodial compounds from selected Streptomyces species.. Bioorganic & Medicinal Chemistry Letters 2015, 25, 5646-5649, 10.1016/j.bmcl.2015.07.103.
- Alfredo F. Braña; Aida Sarmiento-Vizcaíno; Ignacio Pérez-Victoria; Luis Otero; Jonathan Fernández; Juan José Palacios; Jesús Martín; Mercedes De La Cruz; Caridad Díaz; Francisca Vicente; et al.Fernando ReyesLuis A. GarcíaGloria Blanco Branimycins B and C, Antibiotics Produced by the Abyssal Actinobacterium Pseudonocardia carboxydivorans M-227. Journal of Natural Products 2017, 80, 569-573, 10.1021/acs.jnatprod.6b01107.
- Christopher J. Schulze; Mohamed S. Donia; Jair Lage Siqueira-Neto; Debalina Ray; Jevgenij A. Raskatov; Richard E. Green; James H. McKerrow; Michael A. Fischbach; Roger G Linington; Genome-Directed Lead Discovery: Biosynthesis, Structure Elucidation, and Biological Evaluation of Two Families of Polyene Macrolactams against Trypanosoma brucei. ACS Chemical Biology 2015, 10, 2373-2381, 10.1021/acschembio.5b00308.
- Christopher J. Schulze; Gabriel Navarro; Daniel Ebert; Joseph DeRisi; Roger G Linington; Salinipostins A–K, Long-Chain Bicyclic Phosphotriesters as a Potent and Selective Antimalarial Chemotype. The Journal of Organic Chemistry 2015, 80, 1312-1320, 10.1021/jo5024409.
- Ritesh Raju; Zeinab G. Khalil; Andrew M. Piggott; Antje Blumenthal; Nald L. Gardiner; Tina Skinner-Adams; Robert J. Capon; Robert J. Capon; Mollemycin A: An Antimalarial and Antibacterial Glyco-hexadepsipeptide-polyketide from an Australian Marine-Derived Streptomyces sp. (CMB-M0244). Organic Letters 2014, 16, 1716-1719, 10.1021/ol5003913.
- Usama Ramadan Abdelmohsen; Cheng Cheng; Christina Viegelmann; Tong Zhang; Tanja Grkovic; Safwat A. Ahmed; Ronald J. Quinn; Ute Hentschel; RuAngelie Edrada-Ebel; Dereplication Strategies for Targeted Isolation of New Antitrypanosomal Actinosporins A and B from a Marine Sponge Associated-Actinokineospora sp. EG49. Marine Drugs 2014, 12, 1220-1244, 10.3390/md12031220.
- Fakir Shahidullah Tareq; Min Ah Lee; Hyi-Seung Lee; Jong Seok Lee; Yeon-Ju Lee; Hee Jae Shin; Gageostatins A–C, Antimicrobial Linear Lipopeptides from a Marine Bacillus subtilis. Marine Drugs 2014, 12, 871-885, 10.3390/md12020871.
- Fakir Shahidullah Tareq; Min Ah Lee; Hyi-Seung Lee; Yeon-Ju Lee; Jong Seok Lee; Choudhury M. Hasan; Tofazzal Islam; Hee Jae Shin; Gageotetrins A–C, Noncytotoxic Antimicrobial Linear Lipopeptides from a Marine BacteriumBacillus subtilis. Organic Letters 2014, 16, 928-931, 10.1021/ol403657r.
- Fakir Shahidullah Tareq; Min Ah Lee; Hyi-Seung Lee; Yeon-Ju Lee; Jong Seok Lee; Choudhury M. Hasan; Tofazzal Islam; Hee Jae Shin; Non-cytotoxic Antifungal Agents: Isolation and Structures of Gageopeptides A–D from a Bacillus Strain 109GGC020. Journal of Agricultural and Food Chemistry 2014, 62, 5565-5572, 10.1021/jf502436r.
- Mya Breitbart; Forest Rohwer; Here a virus, there a virus, everywhere the same virus?. Trends in Microbiology 2005, 13, 278-284, 10.1016/j.tim.2005.04.003.
- Curtis A. Suttle; Viruses in the sea. Nature 2005, 437, 356-361, 10.1038/nature04160.
- Hongbo Huang; Yongxiang Song; Xin Li; Xin Wang; Chunyao Ling; Xiangjing Qin; Zhenbin Zhou; Qinglian Li; Xiaoyi Wei; Jianhua Ju; et al. Abyssomicin Monomers and Dimers from the Marine-Derived Streptomyces koyangensis SCSIO 5802. Journal of Natural Products 2018, 81, 1892-1898, 10.1021/acs.jnatprod.8b00448.
- Haishan Liu; Zhengbo Chen; Guoliang Zhu; Liping Wang; Yuqi Du; Yi Wang; Wei-Ming Zhu; Phenolic polyketides from the marine alga-derived Streptomyces sp. OUCMDZ-3434. Tetrahedron 2017, 73, 5451-5455, 10.1016/j.tet.2017.07.052.
- Seong-Hwan Kim; Thi-Kim-Quy Ha; Won Keun Oh; Jongheon Shin; Dong-Chan Oh; Antiviral Indolosesquiterpenoid Xiamycins C–E from a Halophilic Actinomycete. Journal of Natural Products 2015, 79, 51-58, 10.1021/acs.jnatprod.5b00634.
- Daniel Shin; Woong Sub Byun; Kyuho Moon; Yun Kwon; Munhyung Bae; Soohyun Um; Sang Kook Lee; Dong-Chan Oh; Coculture of Marine Streptomyces sp. With Bacillus sp. Produces a New Piperazic Acid-Bearing Cyclic Peptide. Frontiers in Chemistry 2018, 6, 498, 10.3389/fchem.2018.00498.
- Teppei Kawahara; Miho Izumikawa; Ikuko Kozone; Junko Hashimoto; Noritaka Kagaya; Hanae Koiwai; Mamoru Komatsu; Manabu Fujie; Noriyuki Sato; Haruo Ikeda; et al.Kazuo Shin-Ya Neothioviridamide, a Polythioamide Compound Produced by Heterologous Expression of a Streptomyces sp. Cryptic RiPP Biosynthetic Gene Cluster. Journal of Natural Products 2018, 81, 264-269, 10.1021/acs.jnatprod.7b00607.
- Yi-Lei Nie; Yun-Dan Wu; Chuanxi Wang; Ru Lin; Yang Xie; Dong-Sheng Fang; Hong Jiang; Yun-Yang Lian; Structure elucidation and antitumour activity of a new macrolactam produced by marine-derived actinomycete Micromonospora sp. FIM05328. Natural Product Research 2017, 32, 2133-2138, 10.1080/14786419.2017.1366479.
- Min Cheol Kim; Henrique Machado; Kyoung Hwa Jang; Lynnie Trzoss; Paul R. Jensen; William Fenical; Integration of Genomic Data with NMR Analysis Enables Assignment of the Full Stereostructure of Neaumycin B, a Potent Inhibitor of Glioblastoma from a Marine-Derived Micromonospora. Journal of the American Chemical Society 2018, 140, 10775-10784, 10.1021/jacs.8b04848.
- Subhasish Saha; Wenjun Zhang; Guangtao Zhang; Yiguang Zhu; Yuchan Chen; Wei Liu; Chengshan Yuan; Qingbo Zhang; Haibo Zhang; Liping Zhang; et al.Weimin ZhangChangsheng Zhang Activation and characterization of a cryptic gene cluster reveals a cyclization cascade for polycyclic tetramate macrolactams. Chem. Sci. 2017, 8, 1607-1612, 10.1039/c6sc03875a.
- Xuewei Ye; Komal Anjum; Tengfei Song; Wenling Wang; Ying Liang; Mengxuan Chen; Haocai Huang; Xiao-Yuan Lian; Zhizhen Zhang; Antiproliferative cyclodepsipeptides from the marine actinomycete Streptomyces sp. P11-23B downregulating the tumor metabolic enzymes of glycolysis, glutaminolysis, and lipogenesis. Phytochemistry 2017, 135, 151-159, 10.1016/j.phytochem.2016.12.010.
- Weiyi Zhang; Qian Che; Hongsheng Tan; Xin Qi; Jing Li; Dehai Li; QianQun Gu; Tianjiao Zhu; Ming Liu; Marine Streptomyces sp. derived antimycin analogues suppress HeLa cells via depletion HPV E6/E7 mediated by ROS-dependent ubiquitin–proteasome system. Scientific Reports 2017, 7, 42180, 10.1038/srep42180.
- Xiaoning Han; Zengzhi Liu; Zhenzhen Zhang; Xiaomin Zhang; Tianjiao Zhu; QianQun Gu; Wenli Li; Qian Che; Dehai Li; Geranylpyrrol A and Piericidin F from Streptomyces sp. CHQ-64 ΔrdmF. Journal of Natural Products 2017, 80, 1684-1687, 10.1021/acs.jnatprod.7b00016.
- Qiang Wang; Yixuan Zhang; Mian Wang; Yi Tan; Xinxin Hu; Hongwei He; Chunling Xiao; Xuefu You; Yiguang Wang; Maoluo Gan; et al. Neo-actinomycins A and B, natural actinomycins bearing the 5H-oxazolo[4,5-b]phenoxazine chromophore, from the marine-derived Streptomyces sp. IMB094. Scientific Reports 2017, 7, 3591, 10.1038/s41598-017-03769-8.
- Marta Pérez; Carmen Schleissner; Rogelio Fernandez; Pilar Rodriguez; Fernando Reyes; Paz Zuniga; Fernando De La Calle; Carmen Cuevas; PM100117 and PM100118, new antitumor macrolides produced by a marine Streptomyces caniferus GUA-06-05-006A. The Journal of Antibiotics 2015, 69, 388-394, 10.1038/ja.2015.121.
- Zhenjian Lin; Michael Koch; Christopher D. Pond; Gaiselle Mabeza; Romell A. Seronay; Gisela P. Concepcion; Louis R. Barrows; Baldomero M. Olivera; Eric W. Schmidt; Structure and activity of lobophorins from a turrid mollusk-associated Streptomyces sp. The Journal of Antibiotics 2013, 67, 121-126, 10.1038/ja.2013.115.
- Yuwei Sun; Tomohiko Tomura; Junichi Sato; Takashi Iizuka; Ryosuke Fudou; Makoto Ojika; Isolation and Biosynthetic Analysis of Haliamide, a New PKS-NRPS Hybrid Metabolite from the Marine Myxobacterium Haliangium ochraceum. Molecules 2016, 21, 59, 10.3390/molecules21010059.
- Alice Ms Rodrigues; Clémence Rohée; Thibaut Fabre; Nicole Batailler; François Sautel; Isabelle Carletti; Stéphanie Nogues; Marcelino T. Suzuki; Didier Stien; Cytotoxic indole alkaloids from Pseudovibrio denitrificans BBCC725. Tetrahedron Letters 2017, 58, 3172-3173, 10.1016/j.tetlet.2017.07.005.
- Carmen Schleissner; Librada M. Canedo; Pilar Rodríguez; Cristina Crespo; Paz Zúñiga; Ana Peñalver; Fernando De La Calle; Carmen Cuevas; Bacterial Production of a Pederin Analogue by a Free-Living Marine Alphaproteobacterium. Journal of Natural Products 2017, 80, 2170-2173, 10.1021/acs.jnatprod.7b00408.




