Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Hussein Migdadi | + 1750 word(s) | 1750 | 2021-11-17 07:32:06 | | | |
| 2 | Vivi Li | + 6 word(s) | 1756 | 2021-11-18 02:43:13 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Migdadi, H. Duckweed Species in Saudi Arabia. Encyclopedia. Available online: https://encyclopedia.pub/entry/16114 (accessed on 07 February 2026).
Migdadi H. Duckweed Species in Saudi Arabia. Encyclopedia. Available at: https://encyclopedia.pub/entry/16114. Accessed February 07, 2026.
Migdadi, Hussein. "Duckweed Species in Saudi Arabia" Encyclopedia, https://encyclopedia.pub/entry/16114 (accessed February 07, 2026).
Migdadi, H. (2021, November 17). Duckweed Species in Saudi Arabia. In Encyclopedia. https://encyclopedia.pub/entry/16114
Migdadi, Hussein. "Duckweed Species in Saudi Arabia." Encyclopedia. Web. 17 November, 2021.
Copy Citation
Duckweeds, or Lemnaceae, are widespread aquatic plants. Morphology-based identification of duckweed species is difficult because of their structural complexity. Hence, molecular tools provide significant advantages for characterizing and selecting species or clones for sustainable commercial use.
duckweeds
identification
morphological
DNA barcoding
phylogenetic analysis
1. Introduction
Under the arid conditions of Saudi Arabia, the agricultural sector plays a significant role in food security for humans and animals, as well as economic, social, and sustainable development. Saudi Arabia, with a Vision 2030 strategic plan, aims to enact sustainable development in food production by saving freshwater resources, the environment, and energy. The government supports research on new feed resources for livestock and poultry that maintain high protein value and high water and fertilizer use efficiency. Duckweed is an aquatic plant that contains crude protein up to 45% of its dry weight under optimum conditions and may be used, without further processing, as a complete feed for animals and fish [1]. Duckweed starch content has been reported to be from 5% to 70% of its dry weight. The high level of starch and low lignin and cellulose levels make duckweed a promising feedstock for biofuel production [2]. Duckweed plants float on the surface or are slightly submerged in water. Leaf-like fronds are modified leaves and stem that mainly function in photosynthesis and vegetative reproduction. As a result, the rate of biomass production is high, providing the basis for various practical applications of duckweed in food and feed sustainability [3]. For commercial utilization of duckweed, knowledge of its taxonomy is required. At the morphological level, delineating duckweed species has been shown to be difficult. Many studies have tried to decrease the anatomical complexity based on the chemical composition of flavonoids [4] and to delineate duckweed species using allozyme, which has helped advance morphology techniques [5][6]. The genome size of duckweed varies 13-fold, ranging from 150 Mb in Spirodela polyrhiza (L.), Schleid. to 1881 Mb in Wolffia arrhiza Wimm. With the development of sequencing technology and bioinformatics, five duckweed genomes from Spirodela and Lemna have been sequenced and assembled [7]. Thirty-eight duckweed species have been identified and characterized [8]. With molecular biology tools and techniques (polymerase chain reaction and Q-PCR, DNA barcoding, and high-throughput DNA sequencing tools), a taxonomic analysis at the molecular level offers a more profound understanding than chemotaxonomy. Therefore, among the various DNA analysis methods for taxonomic definition, molecular taxonomy is considered the most suitable method for identifying and evaluating the genetic potential of duckweed species [9]. There are only a few studies in the literature on identifying duckweed genera and species in Saudi Arabia. Masrahi et al. [10] reported the first record of Wolffiella hyalina (Delile) Monod in Saudi Arabia and regarded it as an addition to the flora of Saudi Arabia.
2. Morphological Characterization and DNA Barcoding of Duckweed Species in Saudi Arabia
The identification of duckweed clones, collected from ten locations in Saudi Arabia, was made through morphological and molecular characterization (Figure 1). Morphological variation was recorded based on the different measurements (Table 1 and Table 2). All clones showed obovate fronds shape and rounded frond apex, and the turions were present. The frond shape of the clones from the Riyadh, Taif, Tohama, Tanomah, Qassim, and Madinah-1 regions was asymmetric, while those from the remaining regions were symmetric. The frond color of clones from five regions (Riyadh, Tohama, Al-Baha, Qassim, and Madinah-1) were dark green, and, in five regions (Taif, Jazan, Tanomah, Dhahran, and Madinah-2), the frond color was recorded as light green. All the studied duckweed clones showed frond apexes and turions. The analysis of variance for frond and root traits showed highly significant differences among the clones (Table 2). The number of roots ranged from 1.67 to 9.22; moreover, the Madinah-2 and Al-Qassim clones had the highest number of roots and differed significantly from other clones. The maximum number of roots, i.e., 9.22, was recorded for the Madinah-2 clones, while the Al-Baha clones had the lowest number of roots, i.e., 1.67. The Tanomah clones had the longest roots (3.76 mm), and the Madinah-2 clones had the shortest roots (0.74 mm). Regarding the characteristics of fronds, clones collected from the Al-Qassim region had the highest number of fronds (6.0), while clones from the Riyadh region had the longest fronds (7 mm). The widest and thickest clones were 8.5 mm from the Madinah-1 region and 3.2 mm from the Al-Taif region, respectively.
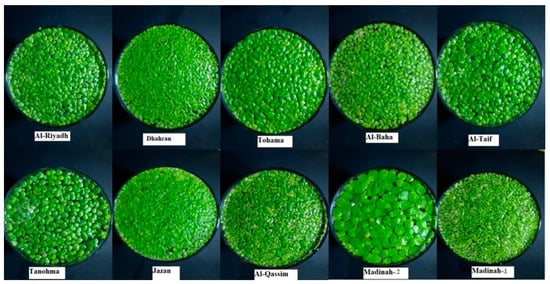

Figure 1. Duckweed clones collected from Saudi Arabia were used for morphological characterization and molecular identification.
Table 1. Qualitative morphological traits for studied duckweed clones of the ten locations of Saudi Arabia (n = 10 samples from each clone averaged and replicated three times, total samples = 30 reads). Frond shape—FSh; frond symmetry—FS; frond color—FC; frond apex—FA; turions—T.
| FSh | FS | FC | FA | T | |
|---|---|---|---|---|---|
| Riyadh | Obovate | Asymmetry | Dark Green | Rounded | Present |
| Dhahran | Obovate | Symmetry | Light Green | Rounded | Present |
| Tohama | Obovate | Asymmetry | Dark Green | Rounded | Present |
| Al-Baha | Obovate | Symmetry | Dark Green | Rounded | Present |
| Al-Taif | Obovate | Asymmetry | Light Green | Rounded | Present |
| Tanomah | Obovate | Asymmetry | Light Green | Rounded | Present |
| Jazan | Obovate | Symmetry | Light Green | Rounded | Present |
| Al-Qassim | Obovate | Asymmetry | Dark Green | Rounded | Present |
| Madinah-1 | Obovate | symmetry | Light Green | Rounded | Present |
| Madinah-2 | Obovate | Asymmetry | Dark Green | Rounded | Present |
Table 2. Analysis of variance (ANOVA) for the quantitative morphological traits of the studied duckweed clones. Mean and standard deviation of the evaluated traits (n = 10 samples and three replications, total = 30).
| Number of Roots | Frond Width (mm) | Frond Thickness (mm) | Frond Length (mm) | NCF Number of Contiguous Fronds |
Root Length (mm) | Number of Fronds | |
|---|---|---|---|---|---|---|---|
| Riyadh | 2.00 ± 0.01 b | 5.7 ± 0.12 b | 2.3 ± 0.17 c | 7.0 ± 0.02 a | 4.0 ± 0.01 b | 2.35 ± 0.13 b | 4.00 ± 0.01 b |
| Al-Taif | 2.00 ± 0.01 b | 5.8 ± 0.06 b | 3.2 ± 0.06 a | 6.5 ± 0.06 b | 5.0 ± 0.01 a | 2.02 ± 0.32 b | 3.67 ± 0.50 b |
| Tohama | 3.00 ± 0.01 b | 5.8 ± 0.06 b | 3.0 ± 0.00 b | 6.5 ± 0.04 b | 3.0 ± 0.01 c | 2.29 ± 0.37 b | 4.00 ± 0.01 b |
| Jazan | 2.44 ± 0.53 b | 3.5 ± 0.10 d | 0.5 ± 0.06 f | 4.6 ± 0.04 e | 3.0 ± 0.01 c | 0.84 ± 0.05 c | 3.67 ± 0.50 b |
| Tanomah | 1.89 ± 0.33 b | 3.4 ± 0.06 d | 0.5 ± 0.0 f | 4.5 ± 0.09 e | 4.0 ± 0.01 b | 3.76 ± 0.49 a | 3.56 ± 0.53 b |
| Al-Baha | 1.67 ± 0.50 b | 3.5 ± 0.06 d | 1.0 ± 0.0 d | 4.7 ± 0.09 e | 4.0 ± 0.01 b | 1.98 ± 0.45 b | 3.00 ± 0.01 bc |
| Dhahran | 2.00 ± 0.01 b | 3.0 ± 0.01 e | 0.4 ± 0.0 f | 5.0 ± 0.07 d | 3.0 ± 0.01 c | 1.54 ± 0.28 b | 3.22 ± 0.44 bc |
| Al-Qassim | 8.11 ± 0.78 a | 4.0 ± 0.01 c | 0.7 ± 0.0 e | 6.0 ± 0.06 c | 5.0 ± 0.01 a | 2.14 ± 0.09 b | 6.00 ± 0.01 a |
| Madinah-1 | 2.00 ± 0.01 b | 8.5 ± 0.01 a | 0.9 ± 0.0 d | 1.0 ± 0.06 g | 4.0 ± 0.01 b | 0.84 ± 0.09 c | 2.89 ± 0.33 bc |
| Madinah-2 | 9.22 ± 2.39 a | 4.0 ± 0.06 c | 0.4 ± 0.0 f | 4.0 ± 0.05 f | 3.0 ± 0.01 c | 0.74 ± 0.09 c | 2.44 ± 0.53 bc |
| Tukey HSD | 1.60 | 0.38 | 0.18 | 0.45 | 0.42 | 0.60 | 1.10 |
Means followed with the same letter in the column were not significantly different based on Tukey HSD, p < 0.05.
To facilitate interpretation of the correlation matrix, produced from the quantitative and qualitative morphological traits for ten duckweed clones, a principal component analysis (PCA) was applied. The PCA plots provided a visual overview of how different traits influenced the distribution of the duckweed clones. The first three principal components explained 88.27% of the total variance (i.e., PC1 46.4%, PC2 24.3%, and PC3 17.51%). The plot for the PC1 and PC2 showed that some traits described the same variation among the clones (Figure 2). The number of roots formed the underlying dimension for PC1, with a value of 0.95 positive loadings; whereas, frond width (−0.2) was negatively loaded and placed on the left-hand side of the plot. PC2 explained 24.3% of the variance and showed a positive correlation with frond traits, such as frond length (0.79), number of fronds (0.26), and frond thickness (0.22), besides root length (0.26). PC3 was loaded with frond width with positive loading (0.76). The Al-Qassim and Madina-2 clones showed the most positive loading in PC1, with 4.9 and 5.8, respectively, while Madina-1 clones had the most negative loading on PC1 and PC2. Those clones were considered to be the most divergent clones.
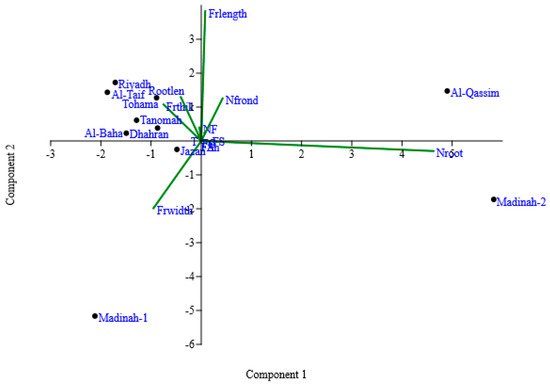
Figure 2. Two-dimensional biplot ordination of the duckweed clones on principal component axes according to qualitative and rescaled quantitative data traits. Principal component 1 (PC1) represented 46.4% of the total variance, and PC2 represented 24.3% of the total variance. Frond shape (FSh); frond symmetry (FS); frond color (FC); frond apex (FA); turions (T); the number of roots—Nroot; frond width—Frwidth; frond thickness (Frthick); frond length (Frlength); the number of contiguous fronds (NCF); root length (Rootlen); the number of fronds (Nfrond).
The results generated from the six plastid markers (psbK-psbI, trnH-psbA, atpF-atpH, matK, rpoC1, and rbcL) for species identification, and their accessions numbers, are presented in Table 3, and the sequences generated from each sample are shown in Supplementary Table S1 (https://www.mdpi.com/2223-7747/10/11/2438#supplementary). The clones collected from the Riyadh, Dhahran, Tohama, Al-Baha, Al-Taif, and Tanomah regions represented the same species (Table 3). However, the duckweed clones collected from the Jazan region represented L. aequinoctialis, and all the sequence data markers confirmed its exact identification, except that rbcL identified L. minor. Similarly, clones collected from the Al-Qassim and Madinah (Madinah-1 and Madinah-2) regions were identified as L. punctata, L. perpusilla, and S. polyryiza, respectively.
Table 3. DNA barcoding-based identifications and accessions number among 10 duckweeds clones.
| Region | psbK-psbI | trnH-psbA | matK | atpF-atpH | rpoC1 | rbcL |
|---|---|---|---|---|---|---|
| Riyadh | L. japonica (OK546023) |
L. gibba (OK103562) |
L. gibba (OK095301) |
L. gibba (OK095300) |
L. minor (OK376426) |
L. gibba (OK571367) |
| Dhahran | L. japonica (OK571365) |
L. gibba (OK383787) |
L. gibba (OK375248) |
L.gibba (OK598953) |
L. gibba (Ok546019) |
L. gibba (OK598946) |
| Tohama | L. japonica (OK350360) |
L. gibba (OK383788) |
L. gibba (OK546020) |
L. gibba (OK598959) |
L. minor (OK376427) |
L. gibba (OK598947) |
| Al-Baha | L. japonica (OK350359) |
L. gibba (OK103563) |
L. gibba (OK571364) |
----- | L. minor (OK493446) |
L. gibba (OK598949) |
| Jazan | L. perpusilla (OK350358) |
L. aequinoctialis (OK383786) |
L. aequinoctialis (OK375249) |
L. aequinoctialis (OK598956) |
L. aequinoctialis (OK493447) |
L. minor (OK598948) |
| Al-Taif | L. japonica (OK546024) |
L. gibba (OK383789) |
L. gibba (OK546021) |
L. gibba (OK598958) |
L. gibba (OK493448) |
L. gibba (OK598951) |
| Tanomah | L. japonica (OK350361) |
L. gibba (OK383790) |
L. gibba (OK546026) |
L. gibba (OK598957) |
L. minor (OK493449) |
L. gibba (OK598952) |
| Al-Qassim | L. punctata (OK350363) |
L. punctata (OK247674) |
L. punctata (OK375250) |
L. punctata (OK598955) |
L. punctata (OK493450) |
L. punctata (OK571368) |
| Madinah-1 | L. perpusilla (OK350362) |
L.aequinoctialis (OK571366) |
L.aequinoctialis (OK546022) |
L. aequinoctialis (OK598954) |
L. aequinoctialis (OK493451) |
L. minor (OK571369) |
| Madinah-2 | S. polyryiza (OK546025) |
S. polyryiza (OK247675) |
----- | S. polyryiza (OK247676) |
S. polyryiza (OK493452) |
S. polyryiza (OK598950) |
After sequencing, the data were cleaned and aligned using the Bioedit software, and the species-level phylogenetic tree was constructed. According to the six markers, the duckweed clones, isolated from the Riyadh region, were identified as L. gibba by four markers (trnH-psbA, mat K, atpF-atpH, and rbcL), while L. japonica was recognized by trnH-psbA, and L. minor was recognized by rpoC1. Similar results were recorded for the Dhahran, Tohama, Al-Baha, Al-Taif, and Tanomah regions. Most markers identified the Jazan isolate as L. aequinoctialis, but it was identified as L. minor only by the rbcL marker. However, the duckweed clones isolated from Al-Qassim and Madinah-2 were identified as L. punctata and S. polyryiza by all markers, respectively. The clones from Madinah-1 were identified as L. perpusilla with three markers (psbK-psbI, trnH-psbA, and matK). In comparison, two markers (atpF-atpH and rpoC1) identified the isolate as L. aequinoctialis, and rbcL identified it as L. minor. Based on multiple nucleotide sequence alignment (Figure S1), divergence score matrix (Figure S2), and the phylogenetic tree (Figure 3), four species were mostly distributed in Saudi Arabia, which were L. gibba, L. aequinoctialis, L. punctata, and S. polyryiza (Figure 3).
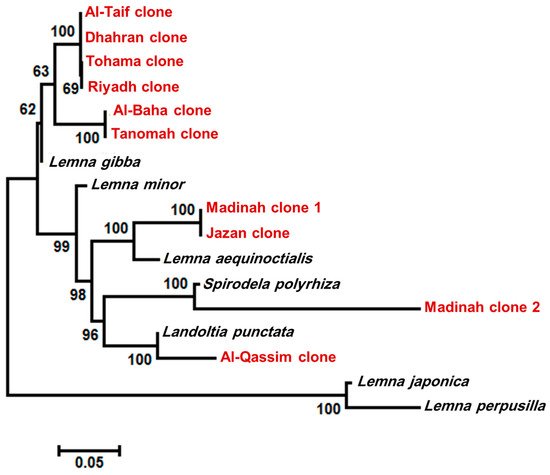
Figure 3. Phylogenetic tree based on maximum likelihood methods with 1000 bootstraps constructed in MEGA 10.0 using the concatenated sequences of psbK-psbI, trnH-psbA, and atpF-atpH intergenic spacer region, and matK, rpoC1, and rbcL gene. The numbers on the nodes represent the percentage of bootstrap values from 1000 replicates. The genetic distances are indicated by the horizontal bar.
References
- Gupta, C.; Prakash, D. Duckweed: An effective tool for phytoremediation. Toxicol. Environ. Chem. 2013, 95, 1256–1266.
- Cheng, J.J.; Stomp, A.M. Growing Duckweed to recover nutrients from Wastewaters and for production of fuel ethanol and animal feed. Clean Soil Air Water 2009, 37, 17–26.
- Sońta, M.; Rekiel, A.; Batorska, M. Use of Duckweed (Lemna L.) in sustainable livestock production and aquaculture—A review. Ann. Anim. Sci. 2019, 19, 257–271.
- Les, D.H.; Landolt, E.; Crawford, D.J. Systematics of the Lemnaceae (duckweeds): Inferences from micromolecular and morphological data. Plant Syst. Evol. 1997, 204, 161–177.
- Crawford, D.J.; Landolt, E.; Les, D.H. An allozyme study of two sibling species of Lemna (Lemnaceae) with comments on their morphology, ecology and distribution. Bull. Torrey Bot. Club 1996, 123, 1–6.
- Crawford, D.J.; Landolt, E.; Les, D.H.; Archibald, J.K.; Kimball, R.T. Allozyme variation within and divergence between Lemna gibba and L. disperma: Systematic and biogeographic implications. Aquat. Bot. 2005, 83, 119–128.
- An, D.; Li, C.; Zhou, Y.; Wu, Y.; Wang, W. Genomes and transcriptomes of duckweeds. Front. Chem. 2018, 6, 230.
- Wang, W.; Wu, Y.; Yan, Y.; Ermakova, M.; Kerstetter, R.; Messing, J. DNA barcoding of the Lemnaceae, a family of aquatic monocots. BMC Plant Biol. 2010, 10, 205.
- Bog, M.; Appenroth, K.-J.; Sree, K.S. Duckweed (Lemnaceae): Its molecular taxonomy. Front. Sustain. Food Syst. 2019, 3, 117.
- Masrahi, Y.S.; Al-Turki, T.A.; Sayed, O.H. Wolffiella hyalina (Delile) Monod (Lemnaceae)—A new record for the flora of Saudi Arabia. Feddes Rep. 2010, 121, 189–193.
More
Information
Subjects:
Plant Sciences; Genetics & Heredity
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
868
Revisions:
2 times
(View History)
Update Date:
18 Nov 2021
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No




