| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Roberta Prete | + 3341 word(s) | 3341 | 2020-06-19 12:13:39 | | | |
| 2 | Roberta Prete | + 3354 word(s) | 3354 | 2020-06-19 13:09:39 | | | | |
| 3 | Roberta Prete | Meta information modification | 3354 | 2020-06-19 13:23:41 | | | | |
| 4 | Roberta Prete | Meta information modification | 3354 | 2020-06-22 12:18:01 | | | | |
| 5 | Roberta Prete | Meta information modification | 3354 | 2020-06-22 14:25:53 | | | | |
| 6 | Roberta Prete | Meta information modification | 3354 | 2020-06-22 14:34:22 | | | | |
| 7 | Catherine Yang | -1446 word(s) | 1908 | 2020-06-24 06:28:21 | | | | |
| 8 | Nicole Yin | Meta information modification | 1908 | 2020-07-01 04:17:40 | | |
Video Upload Options
Table olives fermentation is the result of a complex set of dynamics involving diverse microbial populations such as lactic acid bacteria, yeasts and moulds. Their metabolic activities determine the characteristics such as flavour, texture and safety of the final product. This chapter offers an overview on the main microbiota characterizing table olives and their role during fermentation.
Introduction
Table olives are the most widespread fermented food in the Mediterranean countries. Their fermentation provides preservation, enhanced nutritional and technological characteristics, as well as health benefits.
The main aim of fermentation is the removal of bitter compounds such as oleuropein. The main technologies are:
- Treated olives (Spanish-style): olives are debittered through an alkaline treatment, followed by washing steps. After washing, olives are brined with an initial concentration of 8-12% NaCl and allowed to ferment for 30-60 days at room temperature;
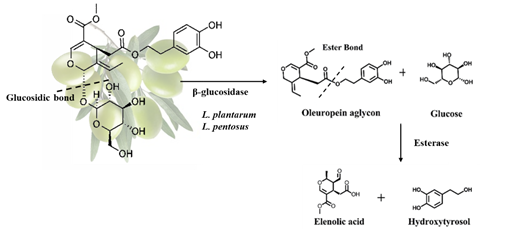
- Natural olives (Greek-style): olives are directly brined and allowed to ferment for 8-12 months and bitterness is removed through the enzymatic activities of indigenous microorganisms especially Lactobacillus plantarum and Lactobacillus pentosus. It is a 2-step process. Initially, oleuropein is converted into oleuropein aglycon and glucose by microbial β-glucosidase and then endogenous esterase can hydrolyse the oleuropein ester bond, forming hydroxytyrosol and elenoic acid as shown in Figure 1 [1][2].
Figure 1. Oleuropein hydrolysis in natural olives.
Olive fermentation is a complex process involving a wide array of microorganisms and mainly lactic acid bacteria (LAB), yeasts and occasionally moulds. These microorganisms and their metabolic activities determine the characteristics (flavour, texture and safety) of the final product [1].
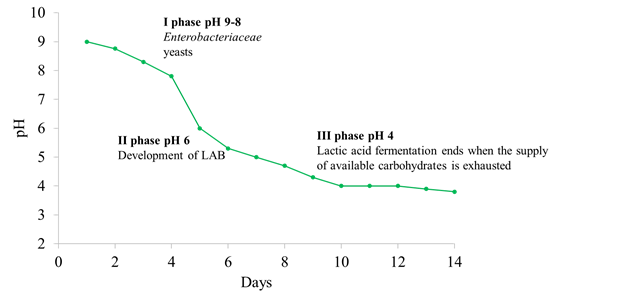
The microbial ecosystem in natural fermentations is quite complex and it is mainly influenced by (a) the indigenous olive microbiota, (b) intrinsic factors prevailing in the brine such as pH, water activity, diffusion of nutrients from the drupe (depending on the structure of the olive skin) and levels of antimicrobial compounds such as oleuropein and other polyphenols and, (c) extrinsic factors such as temperature, oxygen availability and the quantity of salt added in brine. Microbial dynamics are strongly influenced by the technology used. The fermentation process in Spanish-style olives begins at an alkaline pH and the microbial population is mainly composed of Enterobacteriaceae and yeasts, which are able to drop the pH value below 7.0, creating the optimal conditions for LAB growth. The development of LAB induces a further lowering of pH (about 4) (Figure 2). Regarding natural olives, the microbial population starts to grow at an acidic pH because organic acids (acetic, citric, and lactic) are which prevent the growth of Gram-negative bacteria [3].
Figure 2. Different steps and microbial evolution during Spanish-style olives fermentation.
Yeasts associated to table olives
Several yeast species have been reported in different table olives cultivars and the main are reported in Table 1. Some species e.g. Saccharomyces cerevisiae, Wickerhamomyces anomalus, Candida boidinii, Pichia manshurica (ex Pichia galeiformis), and Pichia membranifaciens are associated to different cultivars, suggesting that they can be considered a part of the usual microbiota of table olives. The number of yeasts present on olive drupe is generally limited and the dominant species are not the same ones prevailing in brines. In addition, the yeast biota originating from olive surface is not necessarily involved in the fermentation process. In fact, at the end of fermentation the most common recovered species are S. cerevisiae, W. anomalus, P. kuyveri and P. membranifaciens.
Yeasts can have both a positive and a negative role during olive fermentation. Concerning the negative aspects, some authors reported high pH value and low free acidity during table olive fermentations dominated by yeasts [4][5]. These microorganisms are responsible of the so called “Alambrado” alteration inducing the formation of gas pockets because of CO2 production [6]. The growth of yeasts may also favour the clouding of brines in packed olives. Finally, they possess a wide array of enzymatic activities e.g. proteases, xylanases and pectinases causing the softening of drupes (for a review see Arroyo-Lopez et al. [7]). Despite these negative aspects, yeasts exert a lot of positive actions during table olive fermentation and the main are: i) release of aroma compounds such as esters, higher alcohols which impart a desirable flavour to the final product; ii) degradation of polyphenols thanks to their β-glucosidase activity; iii) release of anti-microbial compounds active against fungi and spoilage yeasts; iv) enhancement of LAB growth through the release of vitamins, amino acids and purines; v) antioxidant action. In this context, it is important to remark that some yeast species isolated from table olives are able to degrade phytate complexes and mycotoxins, which have a negative impact on human health, and to produce bioactive compounds such as carotenoids, tocopherols, and glutathione. Finally, K. lactis and S. cerevisiae isolated from table olives are able to reduce cholesterol content [7].
Table 1. Main yeast species associated to table olives.
|
Olive cultivar |
Yeast species |
References |
|
Gemlik |
Candida mycetangi, Candida hellenica, Candida membranaefaciens, Candida famata, Candida pelliculosa, S. cerevisiae, Zygosaccharomyces mrakii, Aureobasidium pullulans, Kloeckera apiculate, Cryptococcus saitoi, Candida boidinii, W. anomalus, Candida aaseri, Meyerozyma sp., Zygoascus hellenicus, Pichia kudriavzevii, Schwanniomyces etchellsii, Candida atlantica |
|
|
Bosana |
Wickerhamomyces anomalus Nakazawaea molendini-olei, Candida boidinii, Zygotorulaspora mrakii, Saccharomyces cerevisiae |
[10] |
|
Spanish-style green table olives |
Candida thaimueangensis, Saccharomyces cerevisiae and Hanseniaspora sp. |
[10] |
|
Itrana bianca, Itrana nera, Peranzana, Nocellara del Belice, Cellina di Nardò and Bella di Cerignola |
S. cerevisiae, P. manshurica, W. anomalus, Candida boidinii, Candida ishiwadae |
[11] |
|
Aloreña de Málaga |
S. cerevisiae, I. occidentalis, G. candidum, C. apicola, C. diddensiae, P. manshurica, P. membranifaciens |
|
|
Gordal and Manzanilla |
C. tropicalis, P. galeiformis, W. anomalus |
[14] |
|
Arbequina |
C. boidinii, C. sorbosa, C. diddensiae, K. lactis, P. membranifaciens, W. anomalus, P. kluyveri, R. glutinis |
|
|
Hojiblanca and Manzanilla |
C. boidinii, H. guilliermondii, S. cerevisiae, P. manshurica, P. membranifaciens |
|
|
Kalamata |
D. hansenii, W. anomalus, P. membranifaciens, S. cerevisiae |
Lactic acid bacteria associated to table olives
Olive fermentation represents a stressful environment for microorganisms because of the high content of polyphenols and salt as well as the low temperature characterizing the process. Therefore, only few adapted strains are able to grow up. The main LAB genera isolated from table olives are: Lactobacillus, Enterococcus, Pediococcus, Leuconostoc, and Lactococcus. In particular, Lb. brevis, Lb. casei, Lb. coryniformis, Lb. fermentum, Lb. helveticus, Lb. paraplantarum, Lb. plantarum, Lb. pentosus, E. casseliflavus, E. durans, E. faecalis, P. acidilactici, P. ethanolidurans, P. parvolus P. pentosaceus, Leu. cremoris, Leu. mesenteroides, Leu. paramesenteroides, Leu. pseudomesenteroides, L. lactis are the dominant species in most fermentations (for a review see Hurtado et al. [20]). Lactic acid bacteria are able to face these conditions and are considered the main microbial groups involved in table fermentation. In fact, they play important roles during table olive fermentations. First of all, they produce lactic acid which favour pH decrease and free acidity increase. The release of this acid is essential to inhibit the development of pathogens and spoilage microbes. Moreover, LAB produce volatile compounds involved in the definition of the organoleptic characteristics of the final product, and finally yet importantly, they are the main responsible of oleuropein degradation [2]. The development of LAB during table olive fermentation is influenced by several factors including: a) the pH of the fruit, the residual NaOH in treated olives and the washing process of the olives; b) the amount of NaCl in the brine and its effect on water activity; c) the temperature of the fermentation; d) the availability of nutrients and their diffusion through the olive skin; e) the polyphenol content of the fruit; f) the aeration of the vats; g) the size and kind of vats where the fermentations take place [20].
Recent studies revealed that some LAB strains isolated from table olives present probiotic traits. Lb. pentosus isolated from different table olives cultivars (i.e. Nocellara del Belice and Aloreña green table olives) adhere to human intestinal epithelial Caco-2 and vaginal cells and auto-aggregate and/or co-aggregate with pathogenic bacteria (for a review see Perpetuini et al. [2]). Benítez-Cabello et al. [21] showed that different strains of Lb. pentosus and Lb. plantarum isolated from table olives, favored the release of pro-inflammatory (IL-6) and anti-inflammatory (IL-10) interleukins on macrophages, suppressed the secretion of IL-8 and showed anti-proliferative activity on the HT-29 cell line. A probiotic Lb. pentosus strain survived for 200 days in packed olives, confirming the possibility to incorporate probiotic strains and thus produce functional table olives [22].
Moulds associated to table olives
Few data are available concerning the occurrence of moulds during olive fermentation. The main species associated to this niche are: Aspergillus fumigatus, Penicillium commune, Galactomyces geotrichum, Rhizomucor variabilis var. regularior, Penicillium chrysogenum, Penicillium citrinum, Penicillium verrucosum, Mucor fragilis, Rhizopus oryzae, Mycocladus corymbiferus, Penicillium crustosum, Penicillium expansum, Penicillium roqueforti, Mucor circinelloides, Aspergillus niger, and Lichtheimia corymbifera [23][24][25][26]. They can develop on the brine surface and produce a thick layer on the top during fermentation. Their role is not completely understood and they are usually considered as spoilage microbes responsible of flesh softening, production of mouldy odour and taste and release of toxic compounds. Regarding this last aspect, moulds are associated to mycotoxins production. Moreover, the occurrence of P. citrinum and P. verrucosum during fermentation is associated to the accumulation of ochratoxin A and citrinin, while the contamination by aflatoxin B1 is mainly related to the development of A. flavus [23][24][27][28]. However, some strains of A. fumigatus, A. niger, R. variabilis, R. oryzae and M. fragilis have β-glucosidase activity. Further studies are necessary to better investigate the role of moulds during olive fermentation and eventually select suitable strains to be used as starter cultures.
Microbial biofilm communities on olive surface
Biofilms are defined as functional consortia of microorganisms attached to a surface which are embedded in the extracellular polymeric substances (EPS) produced by the microorganisms [29]. The presence of biofilms on olive surface were firstly described by Nychas et al. [30] who revealed the presence of LAB and yeasts on the epidermis of fermented Greek black olives. Starting from this point several studies demonstrated the presence of polymicrobial communities adhered to both biotic (olive skin) and abiotic (glass slides, vessels) surfaces during olive fermentation process. The main species detected on olive surface were P. manshurica (ex P. galeiformis), P. membranifaciens, Candida sorbosa, Geotrichum candidum, and Lb. pentosus [7][31][32][33][34]. Abiotic surfaces were mainlycolonized by species belonging to Candida genus, W. anomalus, D. hansenii and P. guilliermondii, while among LAB, Lb. pentosus was the most abundant species detected [35]. This feature is particularly interesting since it can turn olives into a carrier for delivering different probiotic strains into the human gastro intestinal tract. Lavermicocca et al. [36] used table olives as carrier for the probiotic Lb. paracasei strain IMPC2.1 and the strain was recovered in human faeces after fermented olive intake. The formation of complex microbial communities on olive surface could be due to a high availability and concentration of nutrients at the olive/brine interface. However, the mechanisms regulating this process are poorly understood. Some studies revealed a peculiar genetic structure of Lb. pentosus strains – isolated from table olives – which could favour its adhesion on olive epidermidis. Some of them are involved in the glycolytic pathway (e.g. enolase 1, glucose-6-phosphate isomerase, phosphoglycerate mutase and glucosamine-6-phosphate deaminase) while others in stress response (small heat shock protein) and transcription (transcription elongation factor GreA) [37][38]. However, further research is needed to investigate the development and significance of the microbial association on and within both green and black olives during fermentation.
*Corresponding author: Dr. Giorgia Perpetuini, Faculty of BioScience and Technology for Food, Agriculture and Environment, University of Teramo, Via R Balzarini, 1 - 64100 Teramo, Italy. E-mail address: gperpetuini@unite.it
References
- Corsetti, A.; Perpetuini, G.; Schirone, M.; Tofalo, R.; Suzzi, G. Application of starter cultures to table olive fermentation: an overview on the experimental studies. Front Microbiol 2012, 3, 248, doi:10.3389/fmicb.2012.00248.
- Perpetuini, G.; Prete, R.; Garcia-Gonzalez, N.; Khairul Alam, M.; Corsetti, A. Table Olives More than a Fermented Food. Foods 2020, 9, doi:10.3390/foods9020178.
- Garrido-Fernández, A.; Fernández-Díez, M.J.; Adams, R.M. Table Olives: Production and Processing, 1 ed.; London Chapman & Hall 1997; pp. XII, 496.
- Tassou, C.C.; Panagou, E.Z.; Katsaboxakis, K.Z. Microbiological and physicochemical changes of naturally black olives fermented at different temperatures and NaCl levels in the brines. Food Microbiology 2002, 19, 605-615, doi:10.1006/fmic.2002.0480.
- Aponte, M.; Ventorino, V.; Blaiotta, G.; Volpe, G.; Farina, V.; Avellone, G.; Lanza, C.M.; Moschetti, G. Study of green Sicilian table olive fermentations through microbiological, chemical and sensory analyses. Food Microbiol 2010, 27, 162-170, doi:10.1016/j.fm.2009.09.010.
- Arroyo-López, F.N.; Querol, A.; Bautista-Gallego, J.; Garrido-Fernández, A. Role of yeasts in table olive production. Int J Food Microbiol 2008, 128, 189-196, doi:10.1016/j.ijfoodmicro.2008.08.018.
- Arroyo-López, F.N.; Romero-Gil, V.; Bautista-Gallego, J.; Rodríguez-Gómez, F.; Jiménez-Díaz, R.; García-García, P.; Querol, A.; Garrido-Fernández, A. Potential benefits of the application of yeast starters in table olive processing. Front Microbiol 2012, 5, 34, doi:10.3389/fmicb.2012.00161.
- Mujdeci, G.; Arévalo-Villena, M.; Ozbas, Z.Y.; Briones Pérez, A. Yeast Identification During Fermentation of Turkish Gemlik Olives. J Food Sci 2018, 83, 1321-1325, doi:10.1111/1750-3841.14124.
- Leventdurur, S.; Sert-Aydın, S.; Boyaci-Gunduz, C.P.; Agirman, B.; Ben Ghorbal, A.; Francesca, N.; Martorana, A.; Erten, H. Yeast biota of naturally fermented black olives in different brines made from cv. Gemlik grown in various districts of the Cukurova region of Turkey. Yeast 2016, 33, 289-301, doi:10.1002/yea.3170.
- Porru, C.; Rodríguez-Gómez, F.; Benítez-Cabello, A.; Jiménez-Díaz, R.; Zara, G.; Budroni, M.; Mannazzu, I.; Arroyo-López, F.N. Genotyping, identification and multifunctional features of yeasts associated to Bosana naturally black table olive fermentations. Food Microbiol 2018, 69, 33-42, doi:10.1016/j.fm.2017.07.010.
- Tofalo, R.; Perpetuini, G.; Schirone, M.; Suzzi, G.; Corsetti, A. Yeast biota associated to naturally fermented table olives from different Italian cultivars. Int J Food Microbiol 2013, 161, 203-208, doi:10.1016/j.ijfoodmicro.2012.12.011.
- Arroyo-López, F.N.; Durán-Quintana, M.C.; Ruiz-Barba, J.L.; Querol, A.; Garrido-Fernández, A. Use of molecular methods for the identification of yeast associated with table olives. Food Microbiol 2006, 23, 791-796, doi:10.1016/j.fm.2006.02.008.
- Abriouel, H.; Benomar, N.; Lucas, R.; Gálvez, A. Culture-independent study of the diversity of microbial populations in brines during fermentation of naturally-fermented Aloreña green table olives. Int J Food Microbiol 2011, 144, 487-496, doi:10.1016/j.ijfoodmicro.2010.11.006.
- Bautista-Gallego, J.; Rodríguez-Gómez, F.; Barrio, E.; Querol, A.; Garrido-Fernández, A.; Arroyo-López, F.N. Exploring the yeast biodiversity of green table olive industrial fermentations for technological applications. Int J Food Microbiol 2011, 147, 89-96, doi:10.1016/j.ijfoodmicro.2011.03.013.
- Hurtado, A.; Reguant, C.; Esteve-Zarzoso, B.; Bordons, A.; Rozès, N. Microbial population dynamics during the processing of Arbequina table olives. Food Research International 2008, 41, 738-744, doi:10.1016/j.foodres.2008.05.007.
- Hurtado, A.; Reguant, C.; Bordons, A.; Rozès, N. Influence of fruit ripeness and salt concentration on the microbial processing of Arbequina table olives. Food Microbiol 2009, 26, 827-833, doi:10.1016/j.fm.2009.05.010.
- Rodríguez-Gómez, F.; Arroyo-López, F.N.; López-López, A.; Bautista-Gallego, J.; Garrido-Fernández, A. Lipolytic activity of the yeast species associated with the fermentation/storage phase of ripe olive processing. Food Microbiol 2010, 27, 604-612, doi:10.1016/j.fm.2010.02.003.
- Bleve, G.; Tufariello, M.; Durante, M.; Perbellini, E.; Ramires, F.A.; Grieco, F.; Cappello, M.S.; De Domenico, S.; Mita, G.; Tasioula-Margari, M., et al. Physico-chemical and microbiological characterization of spontaneous fermentation of Cellina di Nardò and Leccino table olives. Front Microbiol 2014, 5, 570, doi:10.3389/fmicb.2014.00570.
- Bleve, G.; Tufariello, M.; Durante, M.; Grieco, F.; Ramires, F.A.; Mita, G.; Tasioula-Margari, M.; Logrieco, A.F. Physico-chemical characterization of natural fermentation process of Conservolea and Kalamàta table olives and developement of a protocol for the pre-selection of fermentation starters. Food Microbiol 2015, 46, 368-382, doi:10.1016/j.fm.2014.08.021.
- Hurtado, A.; Reguant, C.; Bordons, A.; Rozès, N. Lactic acid bacteria from fermented table olives. Food Microbiol 2012, 31, 1-8, doi:10.1016/j.fm.2012.01.006.
- Benítez-Cabello, A.; Torres-Maravilla, E.; Bermúdez-Humarán, L.; Langella, P.; Martín, R.; Jiménez-Díaz, R.; Arroyo-López, F.N. Probiotic Properties of Lactobacillus Strains Isolated from Table Olive Biofilms. Probiotics Antimicrob Proteins 2019, 10.1007/s12602-019-09604-y, doi:10.1007/s12602-019-09604-y.
- Rodríguez-Gómez, F.; Romero-Gil, V.; Bautista-Gallego, J.; García-García, P.; Garrido-Fernández, A.; Arroyo-López, F.N. Production of potential probiotic Spanish-style green table olives at pilot plant scale using multifunctional starters. Food Microbiol 2014, 44, 278-287, doi:10.1016/j.fm.2014.03.023.
- Ghitakou, S.; Koutras, K.; Kanellou, E.; Markaki, P. Study of aflatoxin B1 and ochratoxin A production by natural microflora and Aspergillus parasiticus in black and green olives of Greek origin. Food Microbiol 2006, 23, 612-621, doi:10.1016/j.fm.2005.12.008.
- Heperkan, D.; Meric, B.E.; Sismanoglu, G.; Dalkiliç, G.; Güler, F.K. Mycobiota, mycotoxigenic fungi, and citrinin production in black olives. Adv Exp Med Biol 2006, 571, 203-210, doi:10.1007/0-387-28391-9_13.
- Baffi, M.A.; Romo-Sánchez, S.; Ubeda-Iranzo, J.; Briones-Pérez, A.I. Fungi isolated from olive ecosystems and screening of their potential biotechnological use. N Biotechnol 2012, 29, 451-456, doi:10.1016/j.nbt.2011.05.004.
- Heperkan, D. Microbiota of table olive fermentations and criteria of selection for their use as starters. Front Microbiol 2013, 4, 143, doi:10.3389/fmicb.2013.00143.
- El Adlouni, C.; Tozlovanu, M.; Naman, F.; Faid, M.; Pfohl-Leszkowicz, A. Preliminary data on the presence of mycotoxins (ochratoxin A, citrinin and aflatoxin B1) in black table olives "Greek style" of Moroccan origin. Mol Nutr Food Res 2006, 50, 507-512, doi:10.1002/mnfr.200600055.
- Heperkan, D.; Dazkir, G.S.; Kansu, D.Z.; Güler, F.K. Influence of temperature on citrinin accumulation by Penicillium citrinum and Penicillium verrucosum in black table olives. Toxin Reviews 2009, 28, 180-186, doi:10.1080/15569540903084982.
- Monds, R.D.; O'Toole, G.A. The developmental model of microbial biofilms: ten years of a paradigm up for review. Trends Microbiol 2009, 17, 73-87, doi:10.1016/j.tim.2008.11.001.
- Nychas, G.J.; Panagou, E.Z.; Parker, M.L.; Waldron, K.W.; Tassou, C.C. Microbial colonization of naturally black olives during fermentation and associated biochemical activities in the cover brine. Lett Appl Microbiol 2002, 34, 173-177, doi:10.1046/j.1472-765x.2002.01077.x.
- Domínguez-Manzano, J.; León-Romero, Á.; Olmo-Ruiz, C.; Bautista-Gallego, J.; Arroyo-López, F.N.; Garrido-Fernández, A.; Jiménez-Díaz, R. Biofilm formation on abiotic and biotic surfaces during Spanish style green table olive fermentation. Int J Food Microbiol 2012, 157, 230-238, doi:10.1016/j.ijfoodmicro.2012.05.011.
- Grounta, A.; Panagou, E.Z. Mono and dual species biofilm formation between Lactobacillus pentosus and Pichia membranifaciens on the surface of black olives under different sterile brine conditions. Annals of Microbiology 2014, 64, 1757 – 1767.
- Benítez-Cabello, A.; Romero-Gil, V.; Rodríguez-Gómez, F.; Garrido-Fernández, A.; Jiménez-Díaz, R.; Arroyo-López, F.N. Evaluation and identification of poly-microbial biofilms on natural green Gordal table olives. Antonie Van Leeuwenhoek 2015, 108, 597-610, doi:10.1007/s10482-015-0515-2.
- León-Romero, Á.; Domínguez-Manzano, J.; Garrido-Fernández, A.; Arroyo-López, F.N.; Jiménez-Díaz, R. Formation of In Vitro Mixed-Species Biofilms by Lactobacillus pentosus and Yeasts Isolated from Spanish-Style Green Table Olive Fermentations. Appl Environ Microbiol 2016, 82, 689-695, doi:10.1128/AEM.02727-15.
- Grounta, A.; Doulgeraki, A.I.; Panagou, E.Z. Quantification and characterization of microbial biofilm community attached on the surface of fermentation vessels used in green table olive processing. Int J Food Microbiol 2015, 203, 41-48, doi:10.1016/j.ijfoodmicro.2015.03.001.
- Lavermicocca, P.; Valerio, F.; Lonigro, S.L.; De Angelis, M.; Morelli, L.; Callegari, M.L.; Rizzello, C.G.; Visconti, A. Study of adhesion and survival of lactobacilli and bifidobacteria on table olives with the aim of formulating a new probiotic food. Appl Environ Microbiol 2005, 71, 4233-4240, doi:10.1128/AEM.71.8.4233-4240.2005.
- Perpetuini, G.; Pham-Hoang, B.N.; Scornec, H.; Tofalo, R.; Schirone, M.; Suzzi, G.; Cavin, J.F.; Waché, Y.; Corsetti, A.; Licandro-Seraut, H. In Lactobacillus pentosus, the olive brine adaptation genes are required for biofilm formation. Int J Food Microbiol 2016, 216, 104-109, doi:10.1016/j.ijfoodmicro.2015.10.002.
- Pérez Montoro, B.; Benomar, N.; Caballero Gómez, N.; Ennahar, S.; Horvatovich, P.; Knapp, C.W.; Alonso, E.; Gálvez, A.; Abriouel, H. Proteomic analysis of Lactobacillus pentosus for the identification of potential markers of adhesion and other probiotic features. Food Res Int 2018, 111, 58-66, doi:10.1016/j.foodres.2018.04.072.