
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Jaspreet Kaur Dhanjal | + 2866 word(s) | 2866 | 2021-10-14 05:49:50 | | | |
| 2 | Camila Xu | Meta information modification | 2866 | 2021-11-03 08:35:37 | | |
Video Upload Options
Cell transdifferentiation and reprogramming refers to a group of approaches that allow researchers to halt/reverse the development of adult cells, or convert them one from one cell type to another. The manipulation of cell fate can be achieved by enrolling exogenous/artificial controls. The chemical/small molecule and regulatory components of transcription machinery serve as potential tools to execute cell transdifferentiation and have thereby uncovered new avenues for disease modeling and drug discovery. At the advanced stage, one can believe these methods can pave the way to develop efficient and sensitive gene therapy and regenerative medicine approaches.
1. Introduction
The quenching of cell stemness as cell progressively proliferates and acquires a differentiated state was initially thought to be an irreversible mechanism [1]. The canonical design of a biological process such as cell differentiation has now largely been disproved in the light of emerging evidence about the cellular reprogramming and transdifferentiation mechanisms that potentiate conversion of a lineage-specific, differentiated cell into another/different lineage/cell type [2]. In the process of differentiation, a pluripotent stem cell systematically proliferates and undergoes the intermediate/progenitor and differentiated progenitor/multipotent stages before losing its plasticity and dividing terminally into the specialized/mature cells which constitute an organ or tissue [3]. Mechanistically, when a differentiated cell reverts to its parental lineage or less-differentiated cell to acquire a proliferative phenotype, the process is generally known as dedifferentiation, while transdifferentiation suggests the direct conversion of a differentiated cell type to another differentiated cell type without entering a pluripotent state. Therefore, transdifferentiation is often called direct cell reprogramming [3][4]. Both differentiation and transdifferentiation events can occur naturally [2]. In contrast, the process of cell reprogramming or induced pluripotency, which refers to the process of reverting specialized/differentiated cells to the induced pluripotent stem cells (iPSCs) state, is largely artificial [5]. The fundamental difference between differentiation, dedifferentiation, cellular reprogramming and transdifferentiation is illustrated in Figure1.
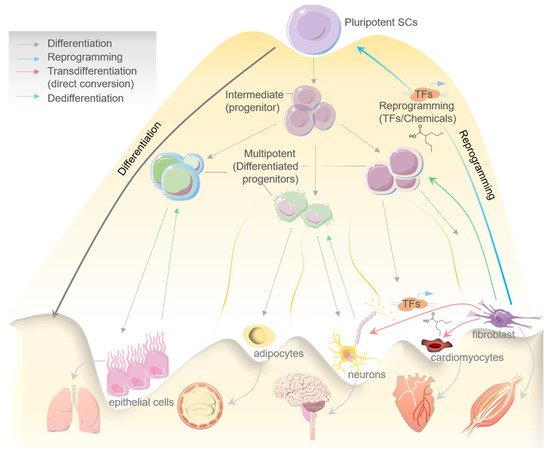
Cell reprogramming can be induced artificially by chemicals/small molecules or by expressing certain transcription factors (TFs), which reprogram a cell to enter an intermediate or pluripotent state [6] (Figure 1). Davies and Weintraub, in the earliest report in 1987, firstly demonstrated the ability of lineage-specific TFs to govern cell fate [7]. Murry et al., later in 1996, brilliantly showed that MyoD expression across different cell lines in vitro stimulates muscle-specific genes’ expression and may further convert these cells into myoblasts [8]. Accumulating evidence in the last three decades has significantly established cellular reprogramming and transdifferentiation in mammals; however, events altering cell fate were also seen to occur naturally [9].
The pathological side of these processes is known in clinical practice, for instance in Barret’s metaplasia, Cdx2 activation transdifferentiates stratified squamous cells into epithelial cells, which potentiates esophagus carcinoma [10]. Earlier reports showed that the transdifferentiation of diverse cell types into myofibroblasts may cause fibrosis in the case of injury or chronic damage to the liver [11], kidney [12], and muscle [13], while a natural transdifferentiation mechanism can be seen in heart [14], liver [15], and in the lens regeneration process in axolotls [16]. Although such remarkable regeneration abilities produced by endogenous transdifferentiation are largely restricted to lower vertebrates, mammals exhibit limited features [9]. For instance, after injury, Lgr5+ led transdifferentiation induces the revival of the hair follicular cells in the inner ear, a rare feature that is exclusive to the neonatal stages. However, in the adults it fails to repair injury significantly [17].
Developments in this field have largely been fueled by investigations into these model organisms and their regenerative abilities, and from the accumulating knowledge on small molecules/chemicals and key cell fate-regulators. The latter includes key transcription factors that can instigate cellular reprogramming and transdifferentiation [18][19][20][21][22], which is largely seen as a promising therapeutic strategy in disease modeling [5][23][24][25][26]. In the following section, we review the role of diverse factors involved in cellular transdifferentiation towards regulating the cell fate in disease modeling.
2. Cell Transdifferentiation: An Overview
A recent development in transdifferentiation or direct lineage-reprogramming— where a cell converts into another cell type without crossing the pluripotent state—offered novel applications to produce functional cells/tissues in disease modeling [18]. Although several functional cell types, including cardiomyocytes, neurons, progenitor/stem cells, hepatic stem cells, hepatocytes, and blood/hematopoietic stem cells have been obtained from fibroblasts/other somatic cells in vitro using the TFs or chemical-mediated transdifferentiation approach, a greater focus of translational research on neural and cardiac cells has been evident.
Recently, Qin et al. demonstrated the transdifferentiation of human fibroblast cells into DA-neuron-like cells by using a combination of protein factors and small molecules [27]. Their method exhibited efficient direct conversion, as 95% of yielded cells were TUJ1-positive, and the process did not include an intermediate neural stem/progenitor stage. In another recent report, Song et al., by using a doxycycline-inducible TFs system (carrying Ngn2, Ascl1, and Dlx2) in human pluripotent stem cells, performed the successful transdifferentiation of these cells into excitatory and inhibitory neurons, exhibiting an equivalent phenotype and molecular signature [28].
Although these studies still do not qualify directly for therapeutic applications or diseases modeling, they do demonstrate proof-of-principal that neurons with post-mitotic state can be transdifferentiated from different cell types, or cell-to-cell conversion can be programmed. These reports decisively affirmed that TFs-mediated neural stem/progenitor cells’ transdifferentiation can critically shape its therapeutic applications, and utility in disease modeling, more specifically in neurodegenerative and age-related neuronal diseases.
Obtaining human cells and stem cells is a practical impediment. Given the non-invasive source of multiple types of cells, urine can be obtained from patients of any age. Urine cell-derived competent cells have emerged as a major tool for research given its therapeutic importance [29]. Xu et al. demonstrated direct transdifferentiation of human urine cells to neurons using a seven small molecule cocktail (CHIR99021, A8301, Y-27632, TTNPB, Forskolin, VPA, NaB) [30]. The transdifferentiated neurons exhibited a mature neuron-like phenotype and molecular signature as validated by the expression of neuronal markers. Further, Qin et al. used a combination of small molecules and protein factors and successfully performed transdifferentiation of human fibroblasts into neuron-like cells without passing through a neural stem/progenitor intermediate stage [27]. Although these reports showed efficient neuronal transdifferentiation from various cell types, the underlying molecular mechanism of these processes warrants further investigation.
3. Cellular Reprogramming for Generating Induced Pluripotent Stem Cells
Cellular reprogramming refers to a group of approaches that allow researchers to halt or reverse the development of adult cells. The validation of cellular reprogramming in human cells has paved the way for a slew of new stem cell biology, disease modeling, drug development, and regenerative medicine applications [19]. The presence of pluripotent stem cells in a population that gives rise to all cells is one of the most defining elements of early mammalian development [31]. Due to a shortage of primary cells from the human central nervous system (CNS) and peripheral nervous system, human-induced pluripotent stem cells (hiPSCs) can also be studied for neurogenerative disease [32]. However, researchers have been able to conduct studies on the recapitulation of physiological and pathological pathways in patient-derived lines. This has resulted in more realistic disease modeling platforms [33]. These are widely utilized in drug discovery and safety investigations, for instance in the development of AD drugs with the goal of identifying chemicals that can inhibit or lower amyloid-beta levels [34]. Some of the recent studies where chemical/small molecules or transcription factors have been employed for inducing cellular reprogramming to study neuronal and cardiac systems have been listed in Table 1 and Table 2, respectively.
Table 1. Chemicals/small molecules-induced cellular reprogramming and their molecular activity/function(s) in neuronal and cardiac model systems.
| Chemicals/Small Molecules |
Molecular Activity/Induced Mechanism(s) |
Cellular Reprogramming Function(s) |
References |
|---|---|---|---|
| RepSox (E-616452) | TGF-βRI (ALK5) inhibitor | CiNPC, CiN, CiCM | [35][36][37] |
| TTNPB | RAR ligand | CiCM, CiN | [38][37] |
| Forskolin | Adenylyl cyclase activator | CiN, CiCM | [39][36][37] |
| CHIR99021 | GSK3 inhibitor | CiNPC, CiNSLCe, CiNf, CiCM | [39][36][38][37][40][41] |
| VPA | HDAC inhibitor | CiPSCa, CiNPCb, CiNc, CiCMd | [35][36][38][37] |
| LiCl and Li2CO3 | GSK3 inhibitor | CiNPC | [35] |
| SB431542 | TGF-βRI inhibitor | CiPSC, CiNPC, CiN, hiEndoPC | [35][38] |
| NaB | HDAC inhibitor | CiNPC | [35] |
| Tranilast | Inhibit TGF-β1 secretion | CiNPC | [35] |
| TSA (Trichostatin A) | HDAC inhibitor | CiNPC | [35] |
| RG108 | DNA methyltransferase inhibitor | CiNSLC | [42] |
| A-83-01 | TGF-βRI (ALK4/5/7) inhibitor | CiNSLC, CiCM | [42][40] |
| Hh-Ag 1.5 | Smoothened agonist | CiNSLC | [42] |
| SMER28 | Autophagy modulator | CiNSLC | [42] |
| Retinoic acid | RAR ligand | CiNSLC | [42] |
| LDN193189 | BMP type I receptor (ALK2/3) inhibitor | CiNSLC | [42] |
| GO6983 | PKC inhibitor | CiN | [36] |
| ISX9 | neurogenesis inducer | CiN | [39] |
| Dorsomorphin | AMPK and BMP I receptor inhibitor | CiN | [36] |
| I-BET151 | BET inhibitor | CiN | [39] |
| SP600125 | JNK inhibitor | CiN | [36] |
| SAG | Smoothened agonist | CiN | [38] |
| Y-27632 | ROCK inhibitor | CiN, CiCM | [36][40] |
| Purmorphamine | Smoothened agonist | CiN | [38] |
| DAPT | Gamma-secretase inhibitor | CiN | [38] |
| SC1 | ERK1 and RasGAP inhibitor | CiCM | [40] |
| Thiazovivin | ROCK inhibitor | CiN | [38] |
| OAC2 | Epigenetic modulation | CiCM | [40] |
| AS8351 | Epigenetic modulator | CiCM | [40] |
| SU16F | PDGFR-β inhibitor | CiCM | [40] |
| JNJ10198409 | PDGFR-α and PDGFR-β inhibitor | CiCM | [40] |
| Bix01294 | Histone methyl transferase inhibitor | CiCM | [40] |
| Reprogramming Factors (TFs) | Species/Model/Cell Type | Obtained Cell Types | Efficiency | Results/Functional Outcome | References | |
|---|---|---|---|---|---|---|
| Neuronal | Brn2, Myt1l, Zic1, Olig2, and Ascl1 | Mouse embryonic and postnatal fibroblast cells | iN (mostly GABAergic and glutamatergic neurons) | ∼50% | Synaptic maturation, functional electrophysiology | [41] |
| Ascl1, Brn2 and Myt1l | iN (mostly excitatory neurons) | 19.50% | Synaptic maturation, functional electrophysiology | [41][43] | ||
| Forskolin, ISX9, CHIR99021 and SB431542 | Mouse fibroblast cells | iN | >90% | Functional electrophysiology | [39] | |
| Ascl1, Brn2, Myt1l | Mouse hepatocytes | iN | >90% | Functional electrophysiology | [44] | |
| Mash1, Nurr1 and Lmx1a | Mouse and human cells/fibroblast cells | iN (mostly dopaminergic neurons) | High | - | [45] | |
| Ascl1, Brn2 and Myt1l | neurons | 20% | Functional | [46] | ||
| Sox2 and Mash1 | Pericyte-derived cells of the adult human cerebral cortex | GABAergic neurons | ∼50% | Obtained iN acquire the ability of action potential firing, synaptic targets for neurons | [47] | |
| LDN193189, SB431542, TTNPB, Tzv, CHIR99021, VPA, DAPT, SAG, Purmo | Human astrocytes | Functional neurons (mainly glutamatergic neurons) | >90% | Functional | [38] | |
| ASCL1, NGN2, SOX2, NURR1 and PITX3 | Human fibroblast cells | iN (mostly dopaminergic neurons) | ∼80% | Functional electrophysiology | [48] | |
| NeuroD1, Ascl1, Brn2, and Mytl1 | iN | ∼60% | Functional neurons | [43] | ||
| Ascl1, Lmx1a, FoxA2, and FEV | serotonergic (i5HT) neurons | ∼25% | Showed spontaneous electrophysiological activity, Active synaptic transmission observed | [49] | ||
| Cardiac | GATA4, MEF2C, TBX5, HAND2 | Mouse | iCMs from MEFs | ~70–80% | Spontaneous beating, Ca2+ transients | [50] |
| GATA4, MYOD-MEF2C, TBX5, HAND2 | iCMs from embryonic head fibroblasts | 10-20% | Spontaneous beating, Ca2+ transients | [51] | ||
| GATA4, MEF2C, TBX5, HAND2, NKX2.5, SB431542 | iCMs from MEFs | 17% | Spontaneous beating, Ca2+ transients | [52] | ||
| MEF2C, GATA4, TBX5 | iCMs from CFs | ~10% | Action potentials, spontaneous beating, Ca2+ transients | [53] | ||
| GATA4, MEF2C, TBX5, HAND2, miR-1, miR-133, A83-01, Y-27632 | iCMs from MEFs | 60% | Action potentials, spontaneous beating, Ca2+ transients | [54] | ||
| GATA4, MEF2C, TBX5, (HAND2), Bmi1 shRNA | iCMs from CFs | 22% | Spontaneous beating, Ca2+ transients | [55] | ||
| GATA4, MEF2C, TBX5, SB431542, XAV939 | iCMs from CFs | ~30% | Spontaneous beating, Ca2+ transients | [56] | ||
| GATA4, MEF2C, TBX5, HAND2, DAPT | iCMs from MEFs | ~38% | Ca2+ transients, spontaneous beating | [57] | ||
| GATA4, MEF2C, TBX5, MESP1, MYOCD | Human | iCMs from HCFs | 5.90% | Ca2+ transients, action potentials | [58] | |
| GATA4, MEF2C, TBX5, ESRGG, MESP1, MYOCD, ZFPM2 | iCMs from hESC-derived fibroblasts | 13% | Ca2+ transients, action potentials | [59] | ||
| GATA4, MEF2C, TBX5 (+ MESP1, MYOCD) with miR-133 | iCMs from HCFs | 27.80% | Ca2+ transients | [60] | ||
| GATA4, MEF2C, TBX5, (HAND2, MYOCD or miR-590) | Human, rat, porcine | iCMs from adult HCFs | ~40% | No spontaneous beating in human iCMs | [61] |
Importantly, iPSCs and ESCs have a high degree of similarity, providing new promise for the use of pluripotent stem cells for regenerative therapies with fewer ethical problems and potentially improved patient specificity [62]. The development of innovative stem cell-based models to investigate the underlying processes of lineage differentiation and embryonic morphogenesis has been aided by the availability of embryo-derived stem cells that capture the lineage propensity [63].
Reprogramming the adult somatic cells into induced pluripotent stem cells (iPSCs) is another effective model that has a bright future as regenerative medicine. Therefore, disease models are critical for revealing the molecular basis of a variety of diseases, enabling the development of new treatments.
Pluripotent stem cells (PSCs), which include embryonic stem cells (ESCs) and induced pluripotent stem cells (iPSCs), have a limitless ability to self-renew and proliferate. This feature allows them to generate a therapeutically relevant number of cells for regenerative therapy [24]. This would help the researchers to better understand the mechanisms driving a variety of human genetic, malignant, and non-malignant disorders. Genome editing techniques have also been utilized to fix disease-specific iPSC mutations, resulting in gene-corrected iPSCs that can be employed for autologous cell-based treatment [64]. The number and kind of cells, their efficiency, footprint, and long-term translational goal influences all its reprogramming approaches. However, fibroblasts and peripheral blood mononuclear cells remain the gold standard, despite the usage of diverse cell types. When compared to iPSCs produced from other parental tissues, blood cells were less likely to develop aberrant DNA methylation, and these cells exhibited stronger hematopoietic differentiation ability [65][66]. Therefore, the generation of patient-specific iPSCs provides a safer alternative for clinical applications.
4. Therapeutic Applications of Transdifferentiation and Cellular Reprogramming
Cellular reprogramming and transdifferentiation have diverse therapeutic applications that include gene therapy/correction, cellular therapy, tissue engineering, and disease modeling (Figure 2).
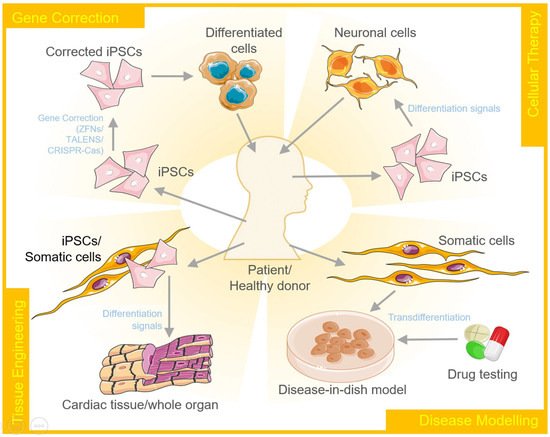
A disease model represents the abnormal state of cells that occur in a specific disease. Therefore, it allows researchers to investigate and understand the intricate mechanisms that lead to the onset and further progression of the disease. These models can further be explored for developing and testing therapeutics. Cellular reprogramming of stem cells to create disease-in-a-dish models has gained a lot of attention over the past few years. These disease models are capable of self-renewal and also differentiate into desired cellular types to capture the disease pathogenesis [67]. Using iPSCs, one of the earliest disease models developed was to study spinal muscular atrophy. The motor neurons produced by diseased iPSCs carried the histological markers of the disease and degenerated at a rate faster than the wildtype control neurons [68]. On similar lines, disease models for many other neurological and cardiac disorders have been developed to date that include Down syndrome, Parkinson's disease, Alzheimer's disease, long QT syndrome, catecholaminergic polymorphic ventricular tachycardia, arrhythmogenic right ventricular cardiomyopathy/dysplasia, dilated cardiomyopathy, left ventricular non-compaction, hypertrophic cardiomyopathy, Barth syndrome, fatty acid oxidation disorders and Pompe diseases to name a few.
The concept of regenerative medicine involves the switching of stem cells or dedifferentiating somatic cells into stem cell-like multipotent cells. These cells can proliferate and then re-differentiate into the desired lineage to repopulate the damaged or degenerated tissue with functional cells. The reprogramming of the cells can be conducted in vitro, in vivo, or ex vivo to regain their regenerative properties. The use of a single transcription factor, such as FOXN1, has been shown to regenerate the thymus in aged mice. Though a lot of efforts are being made to explore and understand mammalian stem cell biology, the knowledge regarding the regenerative capacity of the mammalian system is still limited. However, it is known that the cellular environment, including the modulators present in the extracellular matrix, cytokines, and growth factors, plays a crucial role in this process [69].
An alternative to the natural regenerative potential of mammalian stem cells is to induce transdifferentiation in somatic cells. Differentiated cells, such as neurons derived from iPSCs, have been observed to represent an embryo-like stage. The epigenetic changes that a cell undergoes as it ages or becomes diseased are therefore not reflected by the matured cells. This results in the importance of the transdifferentiation process, whereby the phenotype of one somatic cell type can be converted into another without an intermediate progenitor stage [37]. For instance, Ieda and his colleagues used a combination of Gata4, Mef2c, and Tbx5 developmental transcription factors to transdifferentiate postnatal cardiac or dermal fibroblasts into cardiomyocyte-like cells. The gene expression profile and function of the differentiated cells was also found to be similar to the adult cardiomyocytes [70]. Similar studies enhancing the in vivo efficiency of cardiac cell reprogramming [71] and the use of small molecules for the same have also been reported [40].
Gene editing, in combination with stem cell technology, has the potential to revolutionize the field of medicine, especially for the corrective therapy of monogenic diseases such as sickle cell anemia. In the most simplistic representation, it works by generating patient-specific iPSCs, correcting the genetic defect, ex vivo/in vivo differentiation of the modified cells, followed by the transplantation of the corrected cells/tissue into the patient. The discovery of gene-editing tools, such as zinc finger nucleases, TALENS, and CRISPR/Cas9 has relatively simplified the process of gene editing, making the dream come true. One of the encouraging examples is the replacement of CAG expansion by normal repeats in the huntingtin gene in iPSCs derived from fibroblast cells of Huntington patient. The correction was sustained by the differentiation DARPP-32-positive neurons as well under both in vitro and in vivo conditions [72].
References
- Graf, T. Historical Origins of Transdifferentiation and Reprogramming. Cell Stem Cell 2011, 9, 504–516.
- Slack, J.M.W. Metaplasia and transdifferentiation: From pure biology to the clinic. Nat. Rev. Mol. Cell Biol. 2007, 8, 369–378.
- Jopling, C.; Boue, S.; Belmonte, J.C.I. Dedifferentiation, transdifferentiation and reprogramming: Three routes to regeneration. Nat. Rev. Mol. Cell Biol. 2011, 12, 79–89.
- Roccio, M. Directed differentiation and direct reprogramming: Applying stem cell technologies to hearing research. Stem Cells 2021, 39, 375–388.
- Guo, J.; Wang, H.; Hu, X. Reprogramming and Transdifferentiation Shift the Landscape of Regenerative Medicine. DNA Cell Biol. 2013, 32, 565–572.
- Zhao, Z.; Xu, M.; Wu, M.; Tian, X.; Zhang, C.; Fu, X. Transdifferentiation of Fibroblasts by Defined Factors. Cell. Reprogramm. 2015, 17, 151–159.
- Davis, R.L.; Weintraub, H.; Lassar, A.B. Expression of a single transfected cDNA converts fibroblasts to myoblasts. Cell 1987, 51, 987–1000.
- Murry, C.E.; Kay, M.A.; Bartosek, T.; Hauschka, S.D.; Schwartz, S.M. Muscle differentiation during repair of myocardial necrosis in rats via gene transfer with MyoD. J. Clin. Investig. 1996, 98, 2209–2217.
- Mollinari, C.; Zhao, J.; Lupacchini, L.; Garaci, E.; Merlo, D.; Pei, G. Transdifferentiation: A new promise for neurodegenerative diseases. Cell Death Dis. 2018, 9, 830.
- Barrett, N.R. The lower esophagus lined by columnar epithelium. Surgery 1957, 41, 881–894.
- Tsukamoto, H.; She, H.; Hazra, S.; Cheng, J.; Miyahara, T. Anti-adipogenic regulation underlies hepatic stellate cell transdifferentiation. J. Gastroenterol. Hepatol. 2006, 21 (Suppl. 3), S102–S105.
- Hay, E.D.; Zuk, A. Transformations between epithelium and mesenchyme: Normal, pathological, and experimentally induced. Am. J. Kidney Dis. 1995, 26, 678–690.
- Li, Y.; Huard, J. Differentiation of Muscle-Derived Cells into Myofibroblasts in Injured Skeletal Muscle. Am. J. Pathol. 2002, 161, 895–907.
- Zhang, R.; Han, P.; Yang, H.; Ouyang, K.; Lee, D.; Lin, Y.-F.; Ocorr, K.; Kang, G.; Chen, J.; Stainier, D.Y.R.; et al. In vivo cardiac reprogramming contributes to zebrafish heart regeneration. Nature 2013, 498, 497–501.
- He, J.; Lu, H.; Zou, Q.; Luo, L. Regeneration of Liver After Extreme Hepatocyte Loss Occurs Mainly via Biliary Transdifferentiation in Zebrafish. Gastroenterology 2014, 146, 789–800.
- Suetsugu-Maki, R.; Maki, N.; Nakamura, K.; Sumanas, S.; Zhu, J.; Del Rio-Tsonis, K.; Tsonis, P.A. Lens regeneration in axolotl: New evidence of developmental plasticity. BMC Biol. 2012, 10, 103.
- Wang, T.; Chai, R.; Kim, G.S.; Pham, N.; Jansson, L.; Nguyen, D.-H.; Kuo, B.; May, L.A.; Zuo, J.; Cunningham, L.; et al. Lgr5+ cells regenerate hair cells via proliferation and direct transdifferentiation in damaged neonatal mouse utricle. Nat. Commun. 2015, 6, 6613.
- Chambers, S.M.; Studer, L. Cell Fate Plug and Play: Direct Reprogramming and Induced Pluripotency. Cell 2011, 145, 827–830.
- Cherry, A.B.; Daley, G.Q. Reprogramming Cellular Identity for Regenerative Medicine. Cell 2012, 148, 1110–1122.
- Baranek, M.; Belter, A.; Naskręt-Barciszewska, M.Z.; Stobiecki, M.; Markiewicz, W.T.; Barciszewski, J. Effect of small molecules on cell reprogramming. Mol. BioSyst. 2016, 13, 277–313.
- Julian, L.M.; McDonald, A.C.; Stanford, W.L. Direct reprogramming with SOX factors: Masters of cell fate. Curr. Opin. Genet. Dev. 2017, 46, 24–36.
- Cheung, C.T.; Singh, R.; Kalra, R.S.; Kaul, S.C.; Wadhwa, R. Collaborator of ARF (CARF) Regulates Proliferative Fate of Human Cells by Dose-dependent Regulation of DNA Damage Signaling. J. Biol. Chem. 2014, 289, 18258–18269.
- De Lázaro, I.; Kostarelos, K. Engineering Cell Fate for Tissue Regeneration by In Vivo Transdifferentiation. Stem Cell Rev. Rep. 2015, 12, 129–139.
- Suman, S.; Domingues, A.; Ratajczak, J.; Ratajczak, M.Z. Potential Clinical Applications of Stem Cells in Regenerative Medicine. Adv. Exp. Med. Biol. 2019, 1201, 1–22.
- Kalra, R.S.; Chaudhary, A.; Omar, A.; Cheung, C.T.; Garg, S.; Kaul, S.C.; Wadhwa, R. Stress-induced changes in CARF expression determine cell fate to death, survival, or malignant transformation. Cell Stress Chaperones 2020, 25, 481–494.
- Wadhwa, R.; Kalra, R.S.; Kaul, S.C. CARF is a multi-module regulator of cell proliferation and a molecular bridge between cellular senescence and carcinogenesis. Mech. Ageing Dev. 2017, 166, 64–68.
- Qin, H.; Zhao, A.-D.; Sun, M.-L.; Ma, K.; Fu, X.-B. Direct conversion of human fibroblasts into dopaminergic neuron-like cells using small molecules and protein factors. Mil. Med. Res. 2020, 7, 52.
- Song, S.; Ashok, A.; Williams, D.; Kaufman, M.; Duff, K.; Sproul, A. Efficient Derivation of Excitatory and Inhibitory Neurons from Human Pluripotent Stem Cells Stably Expressing Direct Reprogramming Factors. Curr. Protoc. 2021, 1, e141.
- Gautam, S.; Biswas, S.; Singh, B.; Guo, Y.; Deng, P.; Deng, W. Urine Cells-derived iPSCs: An Upcoming Frontier in Regenerative Medicine. Curr. Med. Chem. 2021, 28, 1.
- Xu, G.; Wu, F.; Gu, X.; Zhang, J.; You, K.; Chen, Y.; Getachew, A.; Zhuang, Y.; Zhong, X.; Lin, Z.; et al. Direct Conversion of Human Urine Cells to Neurons by Small Molecules. Sci. Rep. 2019, 9, 16707.
- Baillie-Benson, P.; Moris, N.; Arias, A.M. Pluripotent stem cell models of early mammalian development. Curr. Opin. Cell Biol. 2020, 66, 89–96.
- Ford, E.; Pearlman, J.; Ruan, T.; Manion, J.; Waller, M.; Neely, G.G.; Caron, L. Human Pluripotent Stem Cells-Based Therapies for Neurodegenerative Diseases: Current Status and Challenges. Cells 2020, 9, 2517.
- Kim, S.-Y.; Kim, Y.-R.; Park, W.-J.; Kim, H.S.; Jung, S.-C.; Woo, S.-Y.; Jo, I.; Ryu, K.-H.; Park, J.-W. Characterisation of insulin-producing cells differentiated from tonsil derived mesenchymal stem cells. Differentiation 2015, 90, 27–39.
- González, J.F.; Alcántara, A.R.; Doadrio, A.L.; Sánchez-Montero, J.M. Developments with multi-target drugs for Alzheimer’s disease: An overview of the current discovery approaches. Expert Opin. Drug Discov. 2019, 14, 879–891.
- Cheng, L.; Hu, W.; Qiu, B.; Zhao, J.; Yu, Y.; Guan, W.; Wang, M.; Yang, W.; Pei, G. Generation of neural progenitor cells by chemical cocktails and hypoxia. Cell Res. 2014, 24, 665–679.
- Hu, W.; Qiu, B.; Guan, W.; Wang, Q.; Wang, M.; Li, W.; Gao, L.; Shen, L.; Huang, Y.; Xie, G.; et al. Direct Conversion of Normal and Alzheimer’s Disease Human Fibroblasts into Neuronal Cells by Small Molecules. Cell Stem Cell 2015, 17, 204–212.
- Fu, Y.; Huang, C.; Xu, X.; Gu, H.; Ye, Y.; Jiang, C.; Qiu, Z.; Xie, X. Direct reprogramming of mouse fibroblasts into cardiomyocytes with chemical cocktails. Cell Res. 2015, 25, 1013–1024.
- Zhang, L.; Yin, J.; Yeh, H.; Ma, N.-X.; Lee, G.; Chen, X.A.; Wang, Y.; Lin, L.; Chen, L.; Jin, P.; et al. Small Molecules Efficiently Reprogram Human Astroglial Cells into Functional Neurons. Cell Stem Cell 2015, 17, 735–747.
- Li, X.; Zuo, X.; Jing, J.; Ma, Y.; Wang, J.; Liu, D.; Zhu, J.; Du, X.; Xiong, L.; Du, Y.; et al. Small-Molecule-Driven Direct Reprogramming of Mouse Fibroblasts into Functional Neurons. Cell Stem Cell 2015, 17, 195–203.
- Cao, N.; Huang, Y.; Zheng, J.; Spencer, C.I.; Zhang, Y.; Fu, J.-D.; Nie, B.; Xie, M.; Zhang, M.; Wang, H.; et al. Conversion of human fibroblasts into functional cardiomyocytes by small molecules. Science 2016, 352, 1216–1220.
- Vierbuchen, T.; Ostermeier, A.; Pang, Z.; Kokubu, Y.; Südhof, T.C.; Wernig, M. Direct conversion of fibroblasts to functional neurons by defined factors. Nat. Cell Biol. 2010, 463, 1035–1041.
- Zhang, M.; Lin, Y.-H.; Sun, Y.J.; Zhu, S.; Zheng, J.; Liu, K.; Cao, N.; Li, K.; Huang, Y.; Ding, S. Pharmacological Reprogramming of Fibroblasts into Neural Stem Cells by Signaling-Directed Transcriptional Activation. Cell Stem Cell 2016, 18, 653–667.
- Pang, Z.; Yang, N.; Vierbuchen, T.; Ostermeier, A.; Fuentes, D.; Yang, T.Q.; Citri, A.; Sebastiano, V.; Marro, S.; Südhof, T.C.; et al. Induction of human neuronal cells by defined transcription factors. Nat. Cell Biol. 2011, 476, 220–223.
- Marro, S.; Pang, Z.; Yang, N.; Tsai, M.-C.; Qu, K.; Chang, H.Y.; Südhof, T.C.; Wernig, M. Direct Lineage Conversion of Terminally Differentiated Hepatocytes to Functional Neurons. Cell Stem Cell 2011, 9, 374–382.
- Caiazzo, M.; Dell’Anno, M.T.; Dvoretskova, E.; Lazarevic, D.; Taverna, S.; Leo, D.; Sotnikova, T.D.; Menegon, A.; Roncaglia, P.; Colciago, G.; et al. Direct generation of functional dopaminergic neurons from mouse and human fibroblasts. Nat. Cell Biol. 2011, 476, 224–227.
- Torper, O.; Pfisterer, U.; Wolf, D.A.; Pereira, M.; Lau, S.; Jakobsson, J.; Björklund, A.; Grealish, S.; Parmar, M. Generation of induced neurons via direct conversion in vivo. Proc. Natl. Acad. Sci. USA 2013, 110, 7038–7043.
- Karow, M.; Sánchez, R.; Schichor, C.; Masserdotti, G.; Ortega, F.; Heinrich, C.; Gascón, S.; Khan, M.A.; Lie, D.C.; Dellavalle, A.; et al. Reprogramming of Pericyte-Derived Cells of the Adult Human Brain into Induced Neuronal Cells. Cell Stem Cell 2012, 11, 471–476.
- Liu, X.; Li, F.; Stubblefield, E.A.; Blanchard, B.; Richards, T.L.; Larson, G.A.; He, Y.; Huang, Q.; Tan, A.C.; Zhang, D.; et al. Direct reprogramming of human fibroblasts into dopaminergic neuron-like cells. Cell Res. 2011, 22, 321–332.
- Xu, Z.; Jiang, H.; Zhong, P.; Yan, Z.; Chen, S.; Feng, J. Direct conversion of human fibroblasts to induced serotonergic neurons. Mol. Psychiatry 2015, 21, 62–70.
- Zhang, Z.; Zhang, W.; Nam, Y.-J. Stoichiometric optimization of Gata4, Hand2, Mef2c, and Tbx5 expression for contractile cardiomyocyte reprogramming. Sci. Rep. 2019, 9, 14970.
- Hirai, H.; Katoku-Kikyo, N.; Keirstead, S.A.; Kikyo, N. Accelerated direct reprogramming of fibroblasts into cardiomyocyte-like cells with the MyoD transactivation domain. Cardiovasc. Res. 2013, 100, 105–113.
- Ifkovits, J.L.; Addis, R.C.; Epstein, J.A.; Gearhart, J.D. Inhibition of TGFβ Signaling Increases Direct Conversion of Fibroblasts to Induced Cardiomyocytes. PLoS ONE 2014, 9, e89678.
- Inagawa, K.; Miyamoto, K.; Yamakawa, H.; Muraoka, N.; Sadahiro, T.; Umei, T.; Wada, R.; Katsumata, Y.; Kaneda, R.; Nakade, K.; et al. Induction of Cardiomyocyte-Like Cells in Infarct Hearts by Gene Transfer of Gata4, Mef2c, and Tbx5. Circ. Res. 2012, 111, 1147–1156.
- Zhao, Y.; Londono, P.; Cao, Y.; Sharpe, E.J.; Proenza, C.; O’Rourke, R.; Jones, K.L.; Jeong, M.Y.; Walker, L.; Buttrick, P.M.; et al. High-efficiency reprogramming of fibroblasts into cardiomyocytes requires suppression of pro-fibrotic signalling. Nat. Commun. 2015, 6, 8243.
- Zhou, Y.; Wang, L.; Vaseghi, H.R.; Liu, Z.; Lu, R.; Alimohamadi, S.; Yin, C.; Fu, J.; Wang, G.G.; Liu, J.; et al. Bmi1 Is a Key Epigenetic Barrier to Direct Cardiac Reprogramming. Cell Stem Cell 2016, 18, 382–395.
- Mohamed, T.M.A.; Stone, N.R.; Berry, E.C.; Radzinsky, E.; Huang, Y.; Pratt, K.; Ang, Y.-S.; Yu, P.; Wang, H.; Tang, S.; et al. Chemical Enhancement of In Vitro and In Vivo Direct Cardiac Reprogramming. Circulation 2017, 135, 978–995.
- Haratizadeh, S.; Bojnordi, M.N.; Darabi, S.; Karimi, N.; Naghikhani, M.; Hamidabadi, H.G.; Seifi, M. Condition medium of cerebrospinal fluid and retinoic acid induces the transdifferentiation of human dental pulp stem cells into neuroglia and neural like cells. Anat. Cell Biol. 2017, 50, 107–114.
- Wada, R.; Muraoka, N.; Inagawa, K.; Yamakawa, H.; Miyamoto, K.; Sadahiro, T.; Umei, T.; Kaneda, R.; Suzuki, T.; Kamiya, K.; et al. Induction of human cardiomyocyte-like cells from fibroblasts by defined factors. Proc. Natl. Acad. Sci. USA 2013, 110, 12667–12672.
- Fu, J.; Stone, N.R.; Liu, L.; Spencer, C.I.; Qian, L.; Hayashi, Y.; Olguín, P.D.; Ding, S.; Bruneau, B.G.; Srivastava, D. Direct Reprogramming of Human Fibroblasts toward a Cardiomyocyte-like State. Stem Cell Rep. 2013, 1, 235–247.
- Muraoka, N.; Yamakawa, H.; Miyamoto, K.; Sadahiro, T.; Umei, T.; Isomi, M.; Nakashima, H.; Akiyama, M.; Wada, R.; Inagawa, K.; et al. MiR-133 promotes cardiac reprogramming by directly repressing Snai1 and silencing fibroblast signatures. EMBO J. 2014, 33, 1565–1581.
- Singh, V.P.; Mathison, M.; Patel, V.; Sanagasetti, D.; Gibson, B.W.; Yang, J.; Rosengart, T.K. MiR-590 Promotes Transdifferentiation of Porcine and Human Fibroblasts Toward a Cardiomyocyte-Like Fate by Directly Repressing Specificity Protein 1. J. Am. Heart Assoc. 2016, 5, e003922.
- Doss, M.X.; Sachinidis, A. Current Challenges of iPSC-Based Disease Modeling and Therapeutic Implications. Cells 2019, 8, 403.
- Kingham, E.; Oreffo, R.O. Embryonic and Induced Pluripotent Stem Cells: Understanding, Creating, and Exploiting the Nano-Niche for Regenerative Medicine. ACS Nano 2013, 7, 1867–1881.
- Ben Jehuda, R.; Shemer, Y.; Binah, O. Genome Editing in Induced Pluripotent Stem Cells using CRISPR/Cas9. Stem Cell Rev. Rep. 2018, 14, 323–336.
- Wattanapanitch, M. Recent Updates on Induced Pluripotent Stem Cells in Hematological Disorders. Stem Cells Int. 2019, 2019, 5171032.
- Nishizawa, M.; Chonabayashi, K.; Nomura, M.; Tanaka, A.; Nakamura, M.; Inagaki, A.; Nishikawa, M.; Takei, I.; Oishi, A.; Tanabe, K.; et al. Epigenetic Variation between Human Induced Pluripotent Stem Cell Lines Is an Indicator of Differentiation Capacity. Cell Stem Cell 2016, 19, 341–354.
- Sterneckert, J.L.; Reinhardt, P.; Scholer, H.R. Investigating human disease using stem cell models. Nat. Rev. Genet. 2014, 15, 625–639.
- Ebert, A.D.; Yu, J.; Rose, F.F., Jr.; Mattis, V.B.; Lorson, C.L.; Thomson, J.A.; Svendsen, C.N. Induced pluripotent stem cells from a spinal muscular atrophy patient. Nature 2009, 457, 277-280.
- Colwell, A.S.; Longaker, M.T.; Lorenz, H.P. Mammalian Fetal Organ Regeneration. Blue Biotechnol. 2005, 93, 83–100.
- Ieda, M.; Fu, J.; Olguín, P.D.; Vedantham, V.; Hayashi, Y.; Bruneau, B.; Srivastava, D. Direct Reprogramming of Fibroblasts into Functional Cardiomyocytes by Defined Factors. Cell 2010, 142, 375–386.
- Mathison, M.; Gersch, R.P.; Nasser, A.; Lilo, S.; Korman, M.; Fourman, M.; Hackett, N.; Shroyer, K.; Yang, J.; Ma, Y.; et al. In Vivo Cardiac Cellular Reprogramming Efficacy Is Enhanced by Angiogenic Preconditioning of the Infarcted Myocardium with Vascular Endothelial Growth Factor. J. Am. Heart Assoc. 2012, 1, e005652.
- An, M.C.; Zhang, N.; Scott, G.; Montoro, D.; Wittkop, T.; Mooney, S.; Melov, S.; Ellerby, L.M. Genetic correction of Huntington's disease phenotypes in induced pluripotent stem cells. Cell Stem Cell 2012, 11, 253-263.
- An, M.C.; Zhang, N.; Scott, G.; Montoro, D.; Wittkop, T.; Mooney, S.; Melov, S.; Ellerby, L.M. Genetic correction of Huntington's disease phenotypes in induced pluripotent stem cells. Cell Stem Cell 2012, 11, 253-263.
- Ebert, A.D.; Yu, J.; Rose, F.F., Jr.; Mattis, V.B.; Lorson, C.L.; Thomson, J.A.; Svendsen, C.N. Induced pluripotent stem cells from a spinal muscular atrophy patient. Nature 2009, 457, 277-280.




