Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Oka, A. Relationship between Saliva and Sublingual Immunotherapy. Encyclopedia. Available online: https://encyclopedia.pub/entry/15573 (accessed on 07 February 2026).
Oka A. Relationship between Saliva and Sublingual Immunotherapy. Encyclopedia. Available at: https://encyclopedia.pub/entry/15573. Accessed February 07, 2026.
Oka, Aiko. "Relationship between Saliva and Sublingual Immunotherapy" Encyclopedia, https://encyclopedia.pub/entry/15573 (accessed February 07, 2026).
Oka, A. (2021, October 31). Relationship between Saliva and Sublingual Immunotherapy. In Encyclopedia. https://encyclopedia.pub/entry/15573
Oka, Aiko. "Relationship between Saliva and Sublingual Immunotherapy." Encyclopedia. Web. 31 October, 2021.
Copy Citation
Sublingual immunotherapy (SLIT) can induce the clinical remission of patients from IgE-mediated allergic diseases, but not all patients who undergo SLIT experience substantial improvement in their allergic symptoms. As biomarkers to predict the efficacy of SLIT have been investigated, some of the salivary microorganisms which turn out to be associated with the efficacy of SLIT are detected, and these microorganisms may be used as adjuvants for SLIT in the future.
sublingual immunotherapy
saliva
allergic rhinitis
microbiome
1. Introduction
Allergen-specific immunotherapy (AIT) can not only reduce allergic symptoms but also induce the clinical remission of patients from IgE-mediated allergic diseases including allergic rhinitis (AR), atopic asthma and venom allergy [1][2][3]. AIT consists of several routes of administration such as subcutaneous immunotherapy (SCIT) and sublingual immunotherapy (SLIT), and the demand for SLIT is increasing rapidly because it has fewer adverse effects and offers more convenience for patients than SCIT. The effectiveness of SLIT is reported to be similar to that of SCIT, but not all patients who undergo AIT experience substantial improvement in their allergic symptoms [4]. Biomarkers to predict or monitor the efficacy of SLIT have been investigated. Saliva may be important in SLIT because tablets used in SLIT first come into contact and react with the saliva in the sublingual site. Since a saliva sample is easy to collect without any pain for patients, it would be useful if there was evidence to support its potential to become a biomarker of SLIT. Saliva contains various microorganisms and establishes a local microbiome, and patients with not only local disorders such as dental caries or periodontal diseases but also systemic disorders such as allergic, inflammatory or malignant diseases are reported to have dysbiosis in their saliva [5][6][7]. The salivary microbiome may have some effects on the immune system and also on SLIT. According to the development of research on the salivary microbiome, the discovery of microorganisms which can be used as biomarkers to predict the effectiveness of SLIT and of probiotics to improve the outcome of SLIT is expected.
2. Effects of Salivary pH on Immunity and Sublingual Immunotherapy
Saliva’s chemical properties may affect the immune system in the oral cavity. Saliva’s normal pH range is reported to be between 6.0 and 8.0 in healthy young adults [8]. Saccharolytic bacteria such as certain Streptococcus, Actinomyces and Lactobacillus species generate organic acids [9][10]. Arginine deiminase pathway (ADP)-positive bacteria such as certain species of Streptococcus and Lactobacillus maintain pH homeostasis in the oral cavity through alkali production [11][12]. The low reactivity of salivary IgA with Candida albicans cells grown at acidic pH values has been reported [13]. Another report showed that uric acid, the main antioxidant of saliva, and also salivary pH were significantly lower in HIV-infected individuals [14]. These results suggest that saliva with an acidic pH is associated with immune depression. Protein release from peanut flour at various pH values of saliva-like buffer was investigated, and the result showed the minimum extraction of proteins between pH 3 and pH 6, while protein extractability was high at both lower and higher pH values [15][16][17]. These results suggest that saliva with an acidic pH leads to poor solubility of allergen proteins used in SLIT. However, an association between salivary pH and the efficacy of SLIT has not been reported, and future investigation is desirable.
3. Salivary Microbiome and Sublingual Immunotherapy
Human monocytes are reported to produce IL-10 in response to saliva, and IL-10 production is associated with signals through TLR2 and TLR4 [18]. These results show that saliva which contains the oral microbiome is highly associated with immunity. On the other hand, salivary IgG4 and IgA are increased in response to SLIT, as shown above. Because the tablet used in SLIT first comes into contact with the oral cavity or saliva, the salivary microbiome may have some effects on SLIT. With the use of the metagenomic sequencing technique, organisms in the salivary microbiome of Japanese cedar-allergic rhinitis patients were shown to be mainly composed of Firmicutes (median: 38.4%), Bacteroidetes (median: 16.9%) and Proteobacteria (median: 14.4%). On the other hand, in non-allergic controls, the composition ratios of Firmicutes, Bacteroidetes and Proteobacteria were 31.8%, 9.6% and 37.3%, respectively. The composition ratio of Bacteroidetes in the saliva was significantly higher compared to the non-allergic controls, and the composition ratio of Bacteroidetes, especially Prevotella species, was also shown to have a positive correlation with IL-10 production in cedar pollen-allergic patients. The composition ratios of Bacteroidetes and Prevotella were also significantly higher in asymptomatic patients showing a visual analog scale (VAS) of 0 after SLIT compared with symptomatic patients showing a VAS above 0 after SLIT [19]. VAS is well validated for the measurement of AR symptoms and correlates well with allergic rhinitis and its impact on asthma (ARIA) severity classification; it also correlates well with the reflective Total Nasal Symptom Score (rTNSS) and rhinoconjunctivitis quality of life questionnaire (RQLQ) [20]. These results suggest that Prevotella in the salivary microbiome may have some effects in inducing IL-10 production and in reducing allergic symptoms (Figure 1), and it may be able to be used as an adjuvant for SLIT. The Lactobacillus genus is commonly reported as probiotics to engage receptors such as TLR2 and TLR4 [21]. Some Lactobacillus species work as adjuvants in SLIT experiments using mice with asthma [22][23]. On the other hand, certain Lactobacillus species increase the antibody responses to an allergen after it is used as a probiotic in SLIT [24]. Since not all allergic patients benefit from SLIT, and also due to the fact that SLIT takes a long time to express the effect of symptom reduction, these microorganisms improving the efficacy of SLIT may potentially be used as adjuvants for SLIT. In addition, SLIT may have the potential to alter the human microbiome, although we found one clinical report showing that the nasal microbiome was not significantly different between AR patients with and without allergen immunotherapy [25]. We would like to investigate the difference between the salivary microbiome in AR patients before and after SLIT in addition to healthy controls. Further research regarding the relationship between immunotherapy and the microbiome is needed.
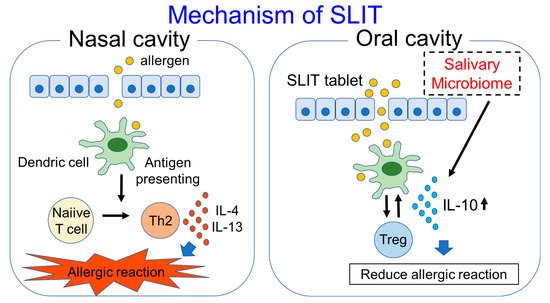
Figure 1. Mechanism of SLIT and participation of salivary microbiome in SLIT are shown in this figure.
The demand for AIT, especially SLIT, is increasing because of its efficacy in inducing clinical remission of allergic diseases and its low risk of side effects. Since not all patients that undergo SLIT demonstrate an improvement in allergic symptoms, biomarkers to predict the outcome and adjuvants for SLIT are expected. Saliva is the first target with which tablets used in SLIT come into contact, and antibodies in saliva such as IgG4 and IgA are increased after SLIT. The salivary microbiome is reported to have some effects on the immune system and is associated with allergic and immune diseases. According to the development of the metagenomic sequencing technique, specific microorganisms which turn out to be associated with the efficacy of SLIT can be detected, and these microorganisms may be used as adjuvants for SLIT in the future
References
- Roberts, G.; Pfaar, O.; Akdis, C.A.; Ansotegui, I.J.; Durham, S.R.; Halken, S.; Larenas-Linnemann, D.; Pawankar, R.; Pitsios, C.; Sheikh, A.; et al. EAACI Guidelines on Allergen Immunotherapy: Allergic rhinoconjunctivitis. Allergy 2018, 73, 765–798.
- Dhami, S.; Kakourou, A.; Asamoah, F.; Agache, I.; Lau, S.; Jutel, M.; Muraro, A.; Roberts, G.; Akdis, C.A.; Bonini, M.; et al. Allergen immunotherapy for allergic asthma: A systematic review and meta-analysis. Allergy 2017, 72, 1825–1848.
- Kucuksezer, U.C.; Ozdemir, C.; Cevhertas, L.; Ogulur, I.; Akdis, M.; Akdis, C.A. Mechanisms of allergen-specific immu-notherapy and allergen tolerance. Allergol. Int. 2020, 69, 549560.
- Dhami, S.; Nurmatov, U.; Arasi, S.; Khan, T.; Asaria, M.; Zaman, H.; Agarwal, A.; Netuveli, G.; Roberts, G.; Pfaar, O.; et al. Allergen immunotherapy for allergic rhinoconjunctivitis: A systematic review and meta-analysis. Allergy 2017, 72, 1597–1631.
- Lundmark, A.; Hu, Y.O.O.; Huss, M.; Johannsen, G.; Andersson, A.F.; Yucel-Lindberg, T. Identification of Salivary Mi-crobiota and Its Association with Host Inflammatory Mediators in Periodontitis. Front. Cell. Infect. Microbiol. 2019, 9, 216.
- Qi, Y.; Zang, S.-Q.; Wei, J.; Yu, H.-C.; Yang, Z.; Wu, H.-M.; Kang, Y.; Tao, H.; Yang, M.-F.; Jin, L.; et al. High-throughput sequencing provides insights into oral microbiota dysbiosis in association with inflammatory bowel disease. Genomics 2021, 113, 664–676.
- Chattopadhyay, I.; Verma, M.; Panda, M. Role of Oral Microbiome Signatures in Diagnosis and Prognosis of Oral Cancer. Technol. Cancer Res. Treat. 2019, 18.
- Foglio-Bonda, P.L.; Migliario, M.; Rocchetti, V.; Pattarino, F.; Foglio-Bonda, A. Daily and annually variation of unstimu-lated whole saliva flow rate and pH and their relation with body profile in healthy young adults. Eur. Rev. Med. Pharmacol. Sci. 2013, 17, 2538–2545.
- Takahashi, N. Oral Microbiome Metabolism: From “Who Are They?” to “What Are They Doing?” . J. Dent. Res. 2015, 94, 1628–1637.
- Abranches, J.; Zeng, L.; Kajfasz, J.K.; Palmer, S.R.; Chakraborty, B.; Wen, Z.T.; Richards, V.P.; Brady, L.J.; Lemos, J. Biology of Oral Streptococci. Microbiol. Spectr. 2018, 6.
- Huang, X.; Zhang, K.; Deng, M.; Exterkate, R.A.M.; Liu, C.; Zhou, X.; Cheng, L.; Cate, J.M.T. Effect of arginine on the growth and biofilm formation of oral bacteria. Arch. Oral Biol. 2017, 82, 256–262.
- Nascimento, M.M. Potential Uses of Arginine in Dentistry. Adv. Dent. Res. 2018, 29, 98–103.
- Bikandi, J.; Moragues, M.D.; Quindós, G.; Polonelli, L.; Pontón, J. Influence of environmental pH on the reactivity of Candida albicans with salivary IgA. J. Dent. Res. 2000, 79, 1439–1442.
- Motamayel, F.A.; Amjad, S.V.; Goodarzi, M.T.; Poorolajal, J. Evaluation of Salivary Uric Acid and pH in Human Immunodeficiency Virus Infected Patients: A Historical Cohort Study. Infect. Disord. Drug Targets 2018, 18, 35–40.
- Koppelman, S.J.; Smits, M.; Tomassen, M.; de Jong, G.A.H.; Baumert, J.; Taylor, S.L.; Witkamp, R.; Veldman, R.J.; Pieters, R.; Wichers, H. Release of Major Peanut Allergens from Their Matrix under Various pH and Simulated Saliva Condi-tions-Ara h2 and Ara h6 Are Readily Bio-Accessible. Nutrients 2018, 10, 1281.
- Sathe, S.K.; Venkatachalam, M.; Sharma, G.M.; Kshirsagar, H.H.; Teuber, S.S.; Roux, K.H. Solubilization and elec-trophoretic characterization of select edible nut seed proteins. J. Agric. Food Chem. 2009, 57, 7846–7856.
- Walczyk, N.E.; Smith, P.M.C.; Tovey, E.R.; Roberts, T.H. Peanut protein extraction conditions strongly influence yield of allergens Ara h1and 2 and sensitivity of immunoassays. Food Chem. 2017, 221, 335–344.
- Haruna, T.; Kariya, S.; Fujiwara, T.; Yuta, A.; Higaki, T.; Zhao, P.; Ogawa, Y.; Kanai, K.; Hirata, Y.; Oka, A.; et al. Role of whole saliva in the efficacy of sublingual immunotherapy in seasonal allergic rhinitis. Allergol. Int. 2019, 68, 82–89.
- Oka, A.; Kidoguchi, M.; Kariya, S.; Fujiwara, T.; Yuta, A.; Miyashita, H.; Higaki, T.; Ogawa, Y.; Kanai, K.; Makihara, S.; et al. Role of salivary microbiome in IL-10 production and efficacy of sublingual immunotherapy. Allergy 2021, 76, 2617–2620.
- Klimek, L.; Bergmann, K.C.; Biedermann, T.; Bousquet, J.; Hellings, P.; Jung, K.; Merk, H.; Olze, H.; Schlenter, W.; Stock, P.; et al. Visual analogue scales (VAS): Measuring instruments for the documentation of symptoms and therapy monitoring in cases of allergic rhinitis in everyday health care. Allergo J. Int. 2017, 26, 16–24.
- Hisbergues, M.; Magi, M.; Rigaux, P.; Steuve, J.; Garcia, L.; Goudercourt, D.; Pot, B.; Pestel, J.; Jacquet, A. In vivo and in vitro immunomodulation of Der p 1 allergen-specific response by Lactobacillus plantarum bacteria. Clin. Exp. Allergy 2007, 37, 1286–1295.
- Van Overtvelt, L.; Moussu, H.; Horiot, S.; Samson, S.; Lombardi, V.; Mascarell, L.; van de Moer, A.; Bourdet-Sicard, R.; Moingeon, P. Lactic acid bacteria as adjuvants for sublingual allergy vaccines. Vaccine 2010, 28, 2986–2992.
- Moussu, H.; Van Overtvelt, L.; Horiot, S.; Tourdot, S.; Airouche, S.; Zuercher, A.; Holvoet, S.; Prioult, G.; Nutten, S.; Mercenier, A.; et al. Bifidobacterium bifidum NCC453 promotes tolerance induction in murine models of sublingual immunotherapy. Int. Arch. Allergy Immunol. 2012, 158, 35–42.
- Brisbin, J.T.; Gong, J.; Orouji, S.; Esufali, J.; Mallick, A.I.; Parvizi, P.; Shewen, P.E.; Sharif, S. Oral treatment of chickens with lactobacilli influences elicitation of immune responses. Clin. Vaccine Immunol. 2011, 18, 1447–1455.
- Bender, M.E.; Read, T.D.; Edwards, T.S.; Hargita, M.; Cutler, A.J.; Wissel, E.F.; Wise, S.K. A Comparison of the Bacterial Nasal Microbiome in Allergic Rhinitis Patients Before and After Immunotherapy. Laryngoscope 2020, 130, E882–E888.
More
Information
Subjects:
Allergy
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
940
Revisions:
3 times
(View History)
Update Date:
29 Mar 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No




