
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Anna Sroka-Bartnicka | + 2067 word(s) | 2067 | 2020-06-08 10:50:22 | | | |
| 2 | Nora Tang | + 14 word(s) | 2081 | 2020-11-03 12:35:58 | | |
Video Upload Options
Fourier transform infrared (FT-IR) and Raman spectroscopy and mapping were applied to
the analysis of biofilms produced by bacteria of the genus Streptococcus. Bacterial biofilm, also called
dental plaque, is the main cause of periodontal disease and tooth decay. It consists of a complex
microbial community embedded in an extracellular matrix composed of highly hydrated extracellular
polymeric substances and is a combination of salivary and bacterial proteins, lipids, polysaccharides,
nucleic acids, and inorganic ions. This study confirms the value of Raman and FT-IR spectroscopies
in biology, medicine, and pharmacy as effective tools for bacterial product characterization.
1. Introduction
Dental caries is an infectious disease associated with the accumulation of bacterial plaque on the tooth surface [1]. Dental plaque, the biofilm formed on the tooth surface, consists of a complex microbial community (less than 10% of biofilm dry weight) embedded in a bacterial and salivary-origin matrix of highly hydrated extracellular polymeric substances (EPS, more than 90% of biofilm dry weight). Dental plaque formation is a multistep process, which involves acquired pellicle formation, initial sucrose-independent and subsequent polysaccharide-mediated attachment of cells to the tooth surface, biofilm maturation, and dispersion of biofilm cells [2]. Mutans streptococci (MS, mainly Streptococcus mutans and S. sobrinus) have been isolated from human dental plaque and have been implicated as a primary causative agent of dental caries [3]. Adhesion, acidogenicity, and acid tolerance are the main virulence factors of the bacteria. MS secrete constitutive glucosyltransferases (Gtfs) that cooperatively synthesize polysaccharide components of EPS from ingested sucrose. S. mutans produce three glucosyltransferases. GtfD synthesizes water-soluble (1→6)-α-D-glucans (its activity is primer-dependent), GtfB and GtfC synthesize water-insoluble (1→3)-α-D-glucans, and the second produces a mixture of water-soluble and water-insoluble glucans, respectively [4][5][6]. It has been found that simultaneous synthesis of glucans by GtfB and GtfC is essential for formation of high-density biofilm with high adhesion, which promotes their binding to an apatite surface [7]. S. sobrinus strains extracellularly produce at least four kinds of Gtfs: (1→3)-α-D-glucan synthase (GtfI) and (1→6)-α-D-glucan synthase (GtfU), (1→6)-α-D-glucan synthase (GtfT), and an oligo-isomaltosaccharide synthase (GtfS) [8]. Glucosyltransferases and their polysaccharide products have been shown to be fundamental virulence factors in the pathogenesis of dental caries because they are responsible for close adhesion to the tooth surface in the presence of sucrose. Additional virulence factors of mutans streptococci are glucan-binding proteins (Gbps). S. mutans produces at least four Gbps: GbpA, GbpB, GbpC, and GbpD. The importance of these proteins is to maintain biofilm architecture by linking bacteria and extracellular molecules of glucan [9]. Another factor that is associated with the virulence of S. mutans is the cell surface protein antigen c (PAc). PAc participates in sucrose-independent bacterial adherence to the tooth surface via interaction with the salivary pellicle [7].
Fourier transform infrared (FT-IR) and Raman spectroscopies are powerful techniques for generating direct information about the molecular and chemical composition of biological samples [10][11][12]. Compared with conventional histological and/or microscopic methods, the microspectroscopic approach is considered advantageous because it is fast, non-invasive, staining- and labeling-free, and less susceptible to human subjective analysis. The combination of these complementary spectroscopic techniques can offer a more comprehensive approach to the analysis of intact samples and ensures more detailed chemical information [13]. The coupling of FT-IR or Raman vibrational spectrometers with a microscope can provide useful information on molecular differences and spatial distributions within and between various healthy and pathological cells and tissues at a microscopic level [13][14][15]. A recent Raman spectroscopic study on bacterial biofilms demonstrated that this technique enables the identification and analysis of nucleic acids, carbohydrates, proteins, and extracellular polymeric substances in biofilms created by a Pseudomonas sp. strain [16]. It has also been applied to characterization of the typology and matrix composition of biofilm produced by Pseudoalteromonas haloplanktis TAC125 in the context of environmental and cold adaptation [17]. In the field of dentistry, it can be used to assess the mineral properties of calcified tissue [18], examine the hydroxyapatite single crystallites [19], compare dental tissues, including enamel and dentin [20], and characterize hydroxylated phosphates [21]. It can also be applied to dental material research, [22][23] and finally utilized for tooth caries diagnosis [24][25].
The aim of the present study was to investigate the composition (the molecular structure and distribution of particular chemical components) of bacterial biofilms produced by various cariogenic strains of Streptococcus spp. with the use of FT-IR and Raman spectroscopic imaging. This vibrational spectroscopic approach proves to be useful in determining the structure of biological samples, which could potentially reduce the cost of experiments and shorten the time of analysis.
2. Discussion
Generally, biofilms are described as “complex communities of bacteria residing within an exopolysaccharide matrix that adheres to a surface” [26]. Biofilm production by bacterial strains is a significant medical and clinical problem because it may be the cause of chronic disease or infections from hospitals, and may be related to infections from implantable medical devices (i.e., dialysis catheters, artificial heart valves, heart Pacemakers, drainage tubes, orthopedic prostheses) [27]. In dentistry, biofilms contribute mainly to dental plaque formation, which, in turn, leads to tooth caries and chronic gingivitis. Biofilms also contribute to infection in the para-nasal sinuses and adhere to dental prostheses and implants, constituting a particular risk for patients with impaired immunity [28]. The phenomenon of quorum sensing—the way bacterial cells communicate with each other, determining surface adhesion, EPS, and virulence factor production—is involved in the formation of bacterial biofilms [29]. EPSs are mainly polysaccharides and, in the matrix of the dental plaque, mostly occur as glucose homopolymers, such as (1→3)-α-, (1→4)-α-, (1→6)-α-D-glucans, while (1→3),(1→6)-α-D-glucan remains crucial for dental caries ethology [2]. The spectral analysis of glucans in biofilms is difficult due to the fact that the presence of these components is attributed to several wavenumbers in both FT-IR and Raman spectra. Furthermore, carbohydrate bands overlap with those of other compounds, such as DNA/RNA, phosphorylated lipids, and proteins [30]. The specificity of glucans can be confirmed by other methods, e.g., polysaccharide-specific monoclonal anti-bodies [31]. FT-IR spectra revealed that a greater amount of (1→3),(1→6)-α-D-glucan is contained in biofilms formed by S. sobrinus DSMZ 20381 and S. sobrinus CAPM 6070. They also exhibit a similar profile to that of other polysaccharides, and may have a higher caries-forming potential than other tested streptococci strains. Raman spectra confirmed the differences in glucans and xylose content and the lower glucose quantity in S. sobrinus/downei CCUG 21020 and the mixture of strains. The FT-IR and Raman chemical images presented in Figure 1 and Figure 2 show the specific component distribution within the measured area. The distributions of the Amide I and (1→3)-α-D-glucan band vary between the different samples, indicating different cariogenic potential.
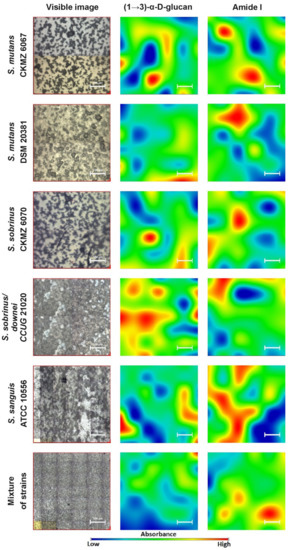
Figure 1. The FT-IR chemical maps of compound distributions in bacterial biofilms. The white bar corresponds to 100 µm
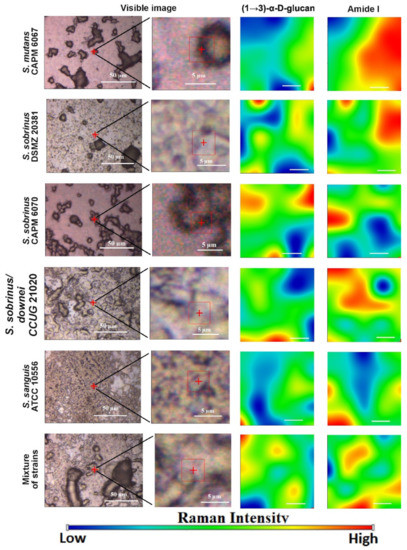
Figure 2. The Raman chemical maps of compound distributions in bacterial biofilms. The red cross and frame in the optical images indicate the mapping area. The white bar in the chemical maps corresponds to 1 µm.
So far, it was confirmed that mixed bacterial species’ biofilms produce considerably more biomass compared with biofilms of one bacterial species, with no need to provide additional external nutrients [32][33]. Moreover, it was stated that these complex bacterial communities have unique properties, e.g., greater resistance to antimicrobial agents and chemical stress/substances, as well as superior expansiveness [32][34]. In our study, the biofilm formed by the mixture of strains has a very similar composition to those of S. mutans CAPM 6067, S. sanguis ATCC 10556, and S. sanguis/downei CCUG 21020 biofilms, while it was characterized by a different polysaccharide content, what can contribute to its different features.
Lipids represent only circa 1.8% of the biofilm matrix [35], but can lead to binding to metals (e.g., in dental prostheses), enhancing virulence, and increasing microbial adherence together with lipopolysaccharides [36]. It was proven that strongly adherent microbial cells increased the production of saturated membrane lipids [37]. They may also play a role as biosurfactants, like viscosin, surfactant, and emulsan, which enable the bioavailability of dispersed hydrophobic substances [38]. Due to the low content of lipids in bacterial biofilms, we did not study their spatial distribution by FT-IR and Raman microspectroscopies, but we observed that lipid composition varied in the studied strains, especially in the ~2920 cm−1 FT-IR band, influencing the cell membrane saturation. In combination with the sugar composition, it may indicate different adhesion potential. S. mutans CAPM 6067, S. sanguis ATCC 10556, and the mixture of strains had the most similar lipid profile, but also sugar profile, and thus probably have congenial adhesive properties.
Protein in the extracellular matrix mainly has two different functions, depending on the location—as enzymes and virulence factors [39]. Enzymatic proteins are involved in the degradation of water-soluble (proteins, nucleic acids, and polysaccharides) and insoluble (lipids) organic components prevalent in biofilms. They promote cell dispersion and, therefore, colonization of new areas [35]. Virulence agents participate in infection processes, including within the oral cavity [40]. Moreover, it was described that S. mutans CAPM 6067 produces glucan-binding proteins, like lectins, leading to formation and stabilization of the matrix [9]. In our research, in terms of protein content, biofilm produced by S. sanguis ATCC 10556 significantly stands out. In the FT-IR spectrum, a higher absorbance intensity was recorded, a different secondary structure of proteins was detected, and a much higher content of Amide I distribution in spectral mapping was found. In the Raman studies, alteration in the β-sheet secondary structure was also revealed. This may testify for the higher colonization potential of this strain.
The quantification of various cellular structures like lipids, proteins, and sugars evidences not only the modifications in the metabolism of bacterial cells and bacterial products, but can also serve as a potential marker for cariogenic processes [41]. FT-IR and Raman spectroscopies are very efficient tools applied for the detection, characterization, and analysis of the above-mentioned molecules. These techniques remain an attractive approach because they are cheap and do not require additional reagents, high-grade solvents, or expensive internal standards and equipment. Moreover, they are widely accessible in standard basic laboratories, and these techniques are potent and adequate for routine studies [42].
Although FT-IR and Raman microspectroscopies have been commonly used as potential techniques for analysis of metabolic profiles of cells and their products in biomedical science [43], they have some limitations. Due to their high complexity connected with overlapping or broadened signals from different simultaneously absorbing cellular components, sometimes, it becomes a problem to ascribe variations in absorbance at a particular wavenumber to a specific molecule [30][44]. To overcome this difficulty, advanced mathematical methodologies for spectral data analyses [45][46], such as second-order derivative determination, Gaussian and/or Lorentzian curve fitting, and Voight deconvolution, can be applied [47]. In our case, the second-order derivative function proved to be sufficient because we compared the composition of the biofilms produced by various streptococci strains, not the changes in biofilm created by one bacterial strain during different conditions (such as drug treatment or alternating ion composition, pH, or temperature).
Even though FT-IR and Raman spectroscopies are complementary techniques that measure the vibrational energies of molecules, both methods are based on different selection rules—an absorption process and an inelastic scattering effect of electromagnetic radiation, respectively. Therefore, the combination of these complementary spectroscopies, as we did in this research, can offer a more comprehensive approach for analyzing intact samples, and can ensure more detailed chemical information [13]. The differences in the results obtained using both techniques are due to the different sensitivities to detection of particular chemical groups and types of vibrations; therefore, some overlapping or low-intensity bands can be distinguished only by one of these methods. For instance, by means of Raman spectroscopy, the most intensive bands are recorded from the symmetric, non-polar groups, e.g., C–C, C=C, C–S, and S–S, but, generally, vibrational spectroscopies are receptive to the anomeric configuration of glycosidic bonds. Additionally, the Raman scatter from water is relatively weak. [48]. In order to avoid water contribution problems, other techniques should be applied in the study of the biochemical composition of bacterial biofilms, which include, among others, the combination of IR and Raman with confocal scanning light microscopy (CSLM), small-angle x-ray scattering (SAXS), surface plasmon resonance imaging (SPRi), electrochemical surface plasmon resonance (EC-SPR), and microscopic approaches: Scanning electron microscopy (SEM) and atomic force microscopy [49].
In summary, our data demonstrate that FT-IR and Raman vibrational spectroscopies coupled with a microscopic approach can be utilized in combination with other biochemical techniques as additional determination and confirmation of the cariogenic potential of bacteria of the genus Streptococcus. The general technique we employed for biochemical analysis is applicable for investigating the bacterial cells’ inherence and proliferation, as well as their extracellular polymeric substance (EPS) production. This opens the possibility of applying non-invasive spectral optical techniques to monitor bacterial adhesion and biofilm production directly on tooth enamel, providing a valuable tool for measuring dental pathologies, such as caries, in vivo.
References
- Marsh, P.D. Dental diseases—Are these examples of ecological catastrophes? Int. J. Dent. Hyg. 2006, 4 (Suppl. 1), 3–10.
- Pleszczyńska, M.; Wiater, A.; Janczarek, M.; Szczodrak, J. (1→3)-α-d-Glucan hydrolases in dental biofilm prevention and control: A review. Int. J. Biol. Macromol. 2015, 79, 761–778.
- Loesche, W.J. Role of Streptococcus mutans in human dental decay. Microbiol. Rev. 1986, 50, 353–380.
- Aoki, H.; Shiroza, T.; Hayakawa, M.; Sato, S.; Kuramitsu, H.K. Cloning of a streptococcus mutans glucosyltransferase gene coding for insoluble glucan synthesis. Infect. Immun. 1986, 53, 587–594.
- Hanada, N.; Kuramitsu, H.K. Isolation and characterization of the Streptococcus mutans gtfD gene, coding for primer-dependent soluble glucan synthesis. Infect. Immun. 1989, 57, 2079–2085.
- Hanada, N.; Kuramitsu, H.K. Isolation and characterization of the streptococcus mutans gtfc gene, coding for synthesis of both soluble and insoluble glucans. Infect. Immun. 2005, 56, 1999–2005.
- Matsumoto-nakano, M. Role of Streptococcus mutans surface proteins for biofilm formation. Jpn. Dent. Sci. Rev. 2018, 54, 22–29.
- Nanbu, A.; Hayakawa, M.; Takada, K.; Shinozaki, N. Production, characterization, and application of monoclonal antibodies which distinguish four glucosyltransferases from Streptococcus sobrinus. FEMS Immunol. Med. Microbiol. 2000, 27, 9–15.
- Lynch, D.J.; Fountain, T.L.; Mazurkiewicz, J.E.; Banas, J.A. Glucan-binding proteins are essential for shaping Streptococcus mutans biofilm architecture. FEMS Microbiol. Lett. 2007, 268, 158–165.
- Carter, E.A.; Tam, K.K.; Armstrong, R.S.; Lay, P.A. Vibrational spectroscopic mapping and imaging of tissues and cells. Biophys. Rev. 2009, 1, 95–103.
- Song, C.L.; Kazarian, S.G. Three-dimensional depth profiling of prostate tissue by micro ATR-FTIR spectroscopic imaging with variable angles of incidence. Analyst 2019, 144, 2954–2964.
- Song, C.L.; Vardaki, M.Z.; Goldin, R.D.; Kazarian, S.G. Fourier transform infrared spectroscopic imaging of colon tissues: Evaluating the significance of amide I and C–H stretching bands in diagnostic applications with machine learning. Anal. Bioanal. Chem. 2019, 411, 6969–6981.
- Lin, S.; Li, M.; Cheng, W. FT-IR and Raman vibrational microspectroscopies used for spectral biodiagnosis of human tissues. Spectroscopy 2007, 21, 1–30.
- Kazarian, S.G.; Chan, K.L.A. Applications of ATR-FTIR spectroscopic imaging to biomedical samples. Biochim. Biophys. Acta 2006, 1758, 858–867.
- Prince, R.C.; Potma, E.O. Going visible: High-resolution coherent Raman imaging of cells and tissues. Light Sci. Appl. 2019, 8, 8–9.
- Henry, V.A.; Jessop, J.L.P.; Peeples, T.L. Differentiating Pseudomonas sp. strain ADP cells in suspensions and biofilms using Raman spectroscopy and scanning electron microscopy. Anal. Bioanal. Chem. 2017, 409, 1441–1449.
- Ricciardelli, A.; Casillo, A.; Vergara, A.; Balasco, N.; Corsaro, M.M.; Tutino, M.L.; Parrilli, E. Environmental conditions shape the biofilm of the Antarctic bacterium Pseudoalteromonas haloplanktis TAC125. Microbiol. Res. 2019, 218, 66–75.
- Imbert, L.; Gourion-Arsiquaud, S.; Villarreal-Ramirez, E.; Spevak, L.; Taleb, H.; van der Meulen, M.C.; Boskey, A.L. Dynamic structure and composition of bone investigated by nanoscale infrared spectroscopy. PLoS ONE 2018, 13, e0202833.
- Fu, B.; Sun, X.; Qian, W.; Shen, Y.; Chen, R.; Hannig, M. Evidence of chemical bonding to hydroxyapatite by phosphoric acid esters. Biomaterials 2005, 26, 5104–5110.
- Tsuda, H.; Ruben, J.; Arends, J. Raman spectra of human dentin mineral. Eur. J. Oral. Sci. 1996, 104, 123–131.
- Bista, R.K.; Bruch, R.F. Near-infrared spectroscopic studies of self-forming lipids and nanovesicles. In Nanoscale Imaging, Sensing, and Actuation for Biomedical Applications VI; SPIE—International Society for Optics and Photonics: San Francisco, CA, USA, 2009; p. 718809.
- Liu, Y.; Yao, X.; Liu, Y.W.; Wang, Y. A Fourier transform infrared spectroscopy analysis of carious dentin from transparent zone to normal zone. Caries Res. 2014, 48, 320–329.
- Hȩdzelek, W.; Wachowiak, R.; Marcinkowska, A.; Domka, L. Infrared spectroscopic identification of chosen dental materials and natural teeth. Acta Phys. Pol. A 2008, 114, 471–484.
- El-Sharkawyi, Y.H. Detection and characterization of human teeth caries using 2D correlation raman spectroscopy. J. Biomed. Phys. Eng. 2019, 9, 167–178.
- Buchwald, T.; Buchwald, Z. Assessment of the Raman spectroscopy effectiveness in determining the early changes in human enamel caused by artificial caries. Analyst 2019, 144, 1409–1419.
- Khatoon, Z.; Mctiernan, C.D.; Suuronen, E.J.; Mah, T. Bacterial bio fi lm formation on implantable devices and approaches to its treatment and prevention. Heliyon 2018, 4, e01067.
- Jabbouri, S.; Sadovskaya, I. Characteristics of the biofilm matrix and its role as a possible target for the detection and eradication of Staphylococcus epidermidis associated with medical implant infections. FEMS Immunol. Med. Microbiol. 2010, 59, 280–291.
- Wroblewska, M.; Struzycka, I.; Mierzwinska-Nastalska, E. Significance of biofilms in dentistry. Przegl. Epidemiol. 2015, 69, 739–744.
- Nadell, C.D.; Xavier, J.B.; Levin, S.A.; Foster, K.R. The evolution of quorum sensing in bacterial biofilms. PLoS Biol. 2008, 6, 0171–0179.
- Sahu, R.K.; Salman, A.; Mordechai, S. Tracing overlapping biological signals in mid-infrared using colonic tissues as a model system. World J. Gastroenterol. 2017, 23, 286–296.
- Mitchell, K.F.; Zarnowski, R.; Sanchez, H.; Edward, J.A.; Reinicke, E.L.; Nett, J.E.; Andes, D.R. Community participation in biofilm matrix assembly and function. Proc. Natl. Acad. Sci. USA 2015, 112, 4092–4097.
- Burmølle, M.; Webb, J.S.; Rao, D.; Hansen, L.H.; Sørensen, S.J.; Kjelleberg, S. Enhanced biofilm formation and increased resistance to antimicrobial agents and bacterial invasion are caused by synergistic interactions in multispecies biofilms. Appl. Environ. Microbiol. 2006, 72, 3916–3923.
- Ren, D.; Madsen, J.S.; Sørensen, S.J.; Burmølle, M. High prevalence of biofilm synergy among bacterial soil isolates in cocultures indicates bacterial interspecific cooperation. ISME J. 2015, 9, 81–89.
- Rice, S.A.; Wuertz, S.; Kjelleberg, S. Next-generation studies of microbial biofilm communities. Microb. Biotechnol. 2016, 9, 677–680.
- Flemming, H.C.; Wingender, J. The biofilm matrix. Nat. Rev. Microbiol. 2010, 8, 623–633.
- Flemming, H.C.; Neu, T.R.; Wozniak, D.J. The EPS matrix: The “House of Biofilm Cells”. J. Bacteriol. 2007, 189, 7945–7947.
- Matsuyama, T.; Nakagawa, Y. Surface-active exolipids: Analysis of absolute chemical structures and biological functions. J. Microbiol. Methods 1996, 25, 165–175.
- Ward, O.P. Microbial Biosurfactants and Biodegradation. Adv. Exp. Med. Biol. 2010, 672, 65–74.
- Baughn, A.D.; Rhee, K.Y. Biofilm matix proteins. Microbiol. Spectr. 2014, 2, 201–222.
- Arciola, R.C.; Campoccia, D.; Speziale, P.; Montanaro, L.; William, J. Biomaterials Bio fi lm formation in Staphylococcus implant infections. A review of molecular mechanisms and implications for bio fi lm-resistant materials. Biomaterials 2012, 33, 5967–5982.
- Nishikawara, F.; Nomura, Y.; Imai, S.; Senda, A.; Hanada, N. Evaluation of cariogenic bacteria. Eur. J. Dent. 2007, 01, 031–039.
- Derenne, A.; Vandersleyen, O.; Goormaghtigh, E. Lipid quantification method using FTIR spectroscopy applied on cancer cell extracts. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2014, 1841, 1200–1209.
- Diem, M.; Romeo, M.; Boydston-White, S.; Miljkovic, M.; Matthaus, M. A decade of vibrational micro-spectroscopy of human cells and tissue (1994–2004). Analyst 2004, 129, 880–885.
- Zwielly, A.; Gopas, J.; Brkic, G.; Mordechai, S. Discrimination between drug-resistant and non-resistant human melanoma cell lines by FTIR spectroscopy. Analyst 2009, 134, 294–300.
- Guo, W.; Piao, S.; Yang, T.C.; Guo, J.; Iqbal, K. High-resolution power spectral estimation method using deconvolution. IEEE J. Ocean. Eng. 2020, 45, 489–499.
- Morhac, M.; Matousek, V. High-resolution boosted deconvolution of spectroscopic data. J. Comput. Appl. Math. 2011, 235, 1629–1640.
- Váczi, T. A new, simple approximation for the deconvolution of instrumental broadening in spectroscopic band profiles. Appl. Spectrosc. 2014, 68, 1274–1278.
- Synytsya, A.; Novak, M. Structural analysis of glucans. Ann. Transl. Med. 2014, 2, 17.
- Wilson, C.; Lukowicz, R.; Merchant, S.; Valquier-Flynn, H.; Caballero, J.; Sandoval, J.; Clement, B. Quantitative and qualitative assessment methods for biofilm growth: A mini-review. Res. Rev. J. Eng. Technol. 2017, 6, 1–8.




