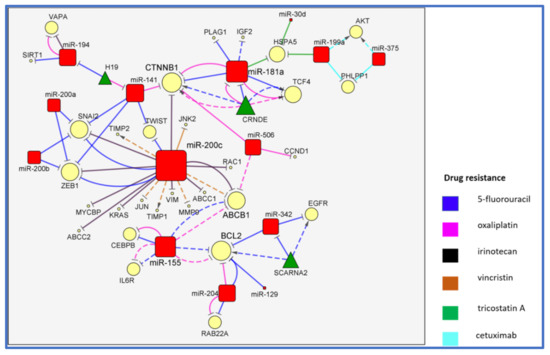
2. The MiR-200/MiR-181/MiR-155 CTNNB1 BCL2 Network
The members of the miR-200 family (miR-200a/b/c and miR-141) and miR-181a play a pivotal role in the multidrug resistance of colorectal carcinoma. These miRNAs were considered as suppressors of cancer growth and metastasis through the regulation of different molecular pathways. MiR-200c and miR-181a are the most-connected miRNAs participating in this network, and both inhibit catenin beta 1 (CTNNB1) expression, a key target associated with three different drug resistances (Figure 1).
Figure 1. MiR-200s/miR-181a and their targets in CRC drug resistance.
The miRNAs in the network are connected with a number of targets (direct or indirect) and are involved in the resistance to vincristine (VCR), irinotecan (CPT11), 5-fluorouracil (5-FU), oxaliplatin (L-OHP), trichostatin A (TSA) and cetuximab (CET). In detail, the overexpression of miR-200c leads to the direct suppression of c-Jun N-terminal kinase 2 (
JNK2) and indirectly to that of
JUN, ATP-binding cassette subfamily B member 1 (
ABCB1) and matrix metallopeptidase 9 (
MMP9), leading, in turn, to the overexpression of TIMP metallopeptidase inhibitor 1 (
TIMP1) and
TIMP2 in HCT8 cells treated with VCR
[1]. The ABCB1 molecular transporter is also an indirect target of miR-506, a negative regulator of CTNNB1 and cyclin D1 (
CCND1), and promotes L-OHP sensitivity in colon cancer after forced expression
[2]. Juang et al. confirmed that miR-200c acted as promoter of CPT11 sensitivity in CRC cells after encapsulation in solid liposomes by suppressing the RAS/CTNNB1/ZEB pathway
[3]. Consistently, the loss of miR-200 and miR-141 were related to the overexpression of the zinc finger E-box-binding homeobox 1 (
ZEB1) and snail family transcriptional repressor 2 (
SNAI2) (targeted by miR-200a, miR-200b and miR-141) and twist family bHLH transcription factors (
TWIST) (targeted by miR-200c and miR-141), all contributing to the epithelial–mesenchymal transition (EMT) in 5-FU-resistant CRC
[4]. Moon et al. investigated the direct correlation between the overexpression of miR-141 and the decrease of the tripartite motif containing 13 (
TRIM13) expression in the 5-FU sensitivity of CRC and the consequent activation of apoptotic pathways
[5]. Ren et al. focused their study on the antagonism between miR-141, which inhibited cancer stemness by the suppression of
CTNNB1, and H19 lncRNA, which promoted cancer growth and L-OHP resistance acting as sponge for miR-141
[6]. Furthermore, miR-194 was reported to be ‘sponged’ by H19 lncRNA, albeit, as in most of these kinds of experiments, the stoichiometry was not reported; the restoration of the miR-194 levels led to the downregulation of sirtuin 1 (
SIRT1), resulting in a decrease of H19/SIRT1-mediated autophagy and in an increase of 5-FU sensitivity
[7].
CTNN1B, one of the most connected proteins of this network, alongside BCL2, was also targeted by miR-181 and CRNDE lncRNA. The repression of miR-181 by CRNDE determined the higher expression of CTNNB1 and transcription factor 4 (
TCF4) miR targets with a promotion of cancer cell growth, 5-FU and L-OHP resistance in CRC cells
[8]. MiR-181a also inhibited the 5-FU resistance directly targeting transcription factor 4 (
PLAG1) and, indirectly, insulin-like growth factor 2 (
IGF2)
[9]. Furthermore, miR-181a cooperated with miR-199a and miR-30d (normally downregulated in colon cancer) to downregulate the endoplasmic reticulum chaperone heat shock protein family A (Hsp70) member 5 (
HSPA5) and increase the TSA sensitivity in CRC cells
[10]. On the other hand, miR-199a, in addition to miR-375, is one of the miRNAs that strengthen the resistance to CET. In details, miR-199a and miR-375 silenced the common target PH domain and leucine-rich repeat protein phosphatase 1 (
PHLPP1), leading to activation of the AKT pathway and increase in CET resistance
[11]. The involvement of miR-199a is the opposite for CET and TSA, since it promotes a resistance to the former (by targeting
PHLPP1 together with miR-375) while it inhibits that to the latter (by targeting
HSPA5 with miR-181a and miR-30d).
The miR-200c/
ZEB1 and miR-200c/
ABCB1 relations are confirmed in two different papers, with the first couple involved in 5-FU resistance
[3][4] and the second one involved directly with CPT11 and indirectly with VCR
[1][3]. Furthermore, the influence of CTNNB1 on L-OHP is confirmed by two different papers, although via different miRNAs: miR-141 or miR-181a
[6][8]. Finally,
PHLPP1,
HSPA5 and
CTNN1B are first-order targets of several miRNA families in the context of drugs resistance. The lower and left portions of this network have genes and miRNAs that likely arise from the tumor microenvironment and are not expressed in the cancer cells themselves. MiR-204 and miR-129, acting as onco-suppressors, directly affect 5-FU resistance by targeting
BCL2, an antiapoptotic oncoprotein, which was also downregulated by miR-204/miR-155 in L-OHP resistance. MiR-204 and miR-155 were both downregulated in tumor-associated macrophages (TAMs), due to the inhibitory role of the activated interleukin 6 (IL6)/signal transducer and activator of the transcription 3 (STAT3) pathway, with a consequent upregulation of CCAAT enhancer-binding protein beta (
CEBPB), IL6 receptor (
IL6R),
ABCB1 (by miR-155),
RAB22A (by miR-204) and the shared
BCL2 target
[12]. This molecular mechanism, possibly involving exosomes and validated by a coculture of TAMs and CRC cells in vitro, conferred L-OHP and 5-FU resistance to CRCs. The miR-204 activity on RAB22A, a member of the RAS oncogene family, and the promotion of chemosensitivity after miRNA’s ectopic expression was confirmed in L-OHP-resistant CRCs
[13]. The resistance to 5-FU was also associated with a low expression of miR-129. After an ectopic expression of miR-129 and the consequent targeting of
BCL2, CRC apoptosis and 5-FU sensitivity were, in fact, promoted
[14]. Furthermore, miR-342 was competitively bound by SCARNA2, a non-coding RNA highly expressed in CRC tissues, thus leading to a secondary upregulation of both the epidermal growth factor receptor (EGFR) and BCL2 oncoproteins and to a sustained 5-FU resistance
[15]. BCL2 is one of the most-connected proteins (together with CTNNB1) and one of the most affected by miRNA activity, as reported by a number of studies on 5-FU resistance. Nevertheless, the implications of miR-204/RAB22A on the resistance to L-OHP were reported by two independent research groups
[12][13]. To understand the functional involvement of the genes in this network, we looked for the most-represented cellular pathways using Fisher’s exact test (
Table S3). The false detection rate was additionally computed to control for multiple testing
[16]. As expected, RAS and PI3K are among the most-represented pathways (FDR <0.05), although the signaling by the cholecystokinin (CCK) receptor is the one spanning the most members (
n = 7) in the network (fold enrichment of 29.9 and FDR 5.6 × 10
−7). In both gastric and colon cancer cells transfected with the cholecystokinin 2 receptor (
CCK2R), gastrin has been shown to enhance cyclooxygenase-2 (
COX-2) gene expression. This key enzyme is known to play an important role in inflammation and carcinogenesis. COX-2 has been involved in hyperproliferation, transformation, invasion, and angiogenesis. In CRC, the extracellular signal-regulated kinase 1/2 (ERK1/2) and PI3-kinase pathways are also involved in gastrin-induced COX-2 expression
[17].
The largest miRNA network in CRC drug resistance spanned miR-200s/miR-181a, among others, and was implicated in the action of six different anticancer treatments (
Figure 1). In this network, CTNNB1 plays a pivotal role, and it is at the interface of two miRNA subnetworks. CTNNB1 is part of a complex of proteins forming adherens junctions, which are important for the establishment and maintenance of epithelial cell layers by regulating cell growth and adhesion between adjacent cells
[18].
CTNNB1 is altered in 4.81% of colorectal carcinoma patients mutations, which are commonly homo- or hemizygous, indicating a higher threshold of CTNNB1 stabilization to be required for transformation in the colon as compared to extracolonic sites
[19]. Moreover, different mutational hotspots in
CTNNB1 for MSI-H and MSS CRCs suggest different effects on CTNNB1 stabilization. Reduced E-cadherin may also contribute to higher levels of transcriptionally active CTNNB1, and it is not directly linked to the
CTNNB1 mutational status. Another target shared by both miR-181a and miR-200s is ABCB1, a membrane transport involved in multidrug resistance. ABCB1 links the larger portion of this network to the miR-155 lobe. MiR-155 is expressed both in CRC cells and in the tumor immune infiltrates, with the presence of CEBPB pointing to tumor-associated macrophages as additional actors in drug resistance. The potency of miR-155 indirectly regulates IL6R, which also suggests the inclusion of granulocytes in the relevant immune cells. Finally, there is a higher-order downregulation of the
BCL2 and
EGFR oncogenes by both miR-155 and miR-342. The molecular mechanisms underlying multiple drug resistance are revealed here as crossing different types of cells and some of them appearing to be exosome-mediated.
Another network that stands out, albeit a smaller one, is highly concentrated around miR-34a
[20] and comprises heavy-weight cancer genes, namely TP53 and MYC, together with some other outstanding oncoproteins, such as MYCN, NOTCH1, WNT1, CSF1R, CDC34 and the stem cell regulator NANOG. The notorious onco-miR-17, which is transcribed by MYC
[20], seems to have an opposite influence when compared to miR-34a. This small network has been reported in the resistance to five different cancer drugs.
A small number of microRNAs and proteins in the networks and clusters that we defined through our work are critically involved in major anticancer treatments for colon cancer. In particular, the family of miR-200, miR-34a, miR-155 and miR-17 appear among the key microRNAs. Thus, the regulation of these miRNAs and their downstream targets or effectors might help to interfere with several drug resistance mechanisms in CRC. As evidenced by our study, few miRNAs seem to have pleiotropic effects on different anticancer drugs. These miRNAs and their partners might also be used in predictive hybrid coding/non-coding gene signatures to address patients to the most effective therapy.