Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Olivier Namy | + 2853 word(s) | 2853 | 2021-08-30 16:58:42 | | | |
| 2 | Jessie Wu | Meta information modification | 2853 | 2021-09-01 04:57:56 | | | | |
| 3 | Jessie Wu | Meta information modification | 2853 | 2021-09-01 05:03:12 | | | | |
| 4 | Jessie Wu | -43 word(s) | 2810 | 2021-11-19 03:53:35 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Namy, O. RNA Modifications in Translation Fidelity. Encyclopedia. Available online: https://encyclopedia.pub/entry/13762 (accessed on 07 February 2026).
Namy O. RNA Modifications in Translation Fidelity. Encyclopedia. Available at: https://encyclopedia.pub/entry/13762. Accessed February 07, 2026.
Namy, Olivier. "RNA Modifications in Translation Fidelity" Encyclopedia, https://encyclopedia.pub/entry/13762 (accessed February 07, 2026).
Namy, O. (2021, August 31). RNA Modifications in Translation Fidelity. In Encyclopedia. https://encyclopedia.pub/entry/13762
Namy, Olivier. "RNA Modifications in Translation Fidelity." Encyclopedia. Web. 31 August, 2021.
Copy Citation
RNA modifications play an essential role in determining RNA fate. Recent studies have revealed the effects of such modifications on all steps of RNA metabolism. These modifications range from the addition of simple groups, such as methyl groups, to the addition of highly complex structures, such as sugars. Their consequences for translation fidelity are not always well documented. Unlike the well-known m6A modification, they are thought to have direct effects on either the folding of the molecule or the ability of tRNAs to bind their codons.
RNA modifications
translation fidelity
2′-O-methylation
1. Role of rRNA Modifications in Translation Fidelity
Ribosomal RNA is the most abundant non-coding RNA in the cytoplasm. It is the main constituent of the ribosome. In total, 200 modification sites have been mapped on the human ribosome, in which about 2% of the nucleotides are modified [1][2][3][4]. These modifications can modulate all stages in the life of the rRNA, from ribosome biogenesis to translation accuracy [5]. The most frequent modifications observed are pseudo-uridines and 2′-O-methylations, although base methylation and acetylation have been reported [1]. We focus here on the description of the two main modifications of rRNAs known to affect translation fidelity.
1.1. 2′-O-methylation
2′-O-methylation (Nm) is a modification in which a sugar is added to the 2′C hydroxyl group of the nucleotide. The chemical impact of Nm on RNA has been investigated by several studies. It has been reported that Nm biases the sugar pucker equilibrium in favor of the C3′-endo conformation of pyrimidines [6]. Intra-residue steric repulsion occurs between the Nm, the 3′-phosphate, and the 2-carbonyl groups in the C2′-endo conformation, favoring the C3′ form. The Nm modification may, therefore, either stabilize or modulate RNA structures.
In human cells, Nm is mediated by the ribonucleoprotein complex consisting of the methylase fibrillarin (FBL) and the guide RNA (C/D box snoRNA) specific to the methylation site [7]. FBL is an essential protein, but it can be partially inactivated, leading to a decrease of up to 50% in the number of methylation sites in human cells [8][9]. More than 100 2′-O-methylation sites have been mapped on rRNAs, independently of nucleoside identity [10][11].
The role of Nm in miscoding has been explored in human cancer cells [12]. FBL overexpression, leading to hypermethylation of the ribosome, has been shown to trigger an increase in amino-acid incorporation at cognate or near-cognate codons. It is difficult to identify the 2′-O-methylation sites responsible for this phenotype, because site-specific inactivation experiments have not been performed yet on human cells. As FBL methylates all the sites, the only solution would be to inactivate each snoRNA specifically, one-by-one. A study of this type has been performed in yeast, in which knockouts of the various guide C/D box snoRNAs have been performed [13]. The impact of the loss of each snoRNA was evaluated by measuring stop codon readthrough efficiency. Nm-C1639 was identified as the most important of the Nm sites tested. The abolition of Nm at this P-site triggers a slight increase in UAG readthrough. This work revealed a role for Nm-C1639 in the maintenance of ribosome fidelity during termination. There is now a need to reproduce such systematic analyses of Nm sites in humans.
The role of rRNA’s Nm extends beyond miscoding events. The downregulation of FBL has been shown to alter IRES-dependent initiation and frameshifting. A single deletion of Am398 or Gm3745 in the 28S rRNA or of Am163 in the 18S rRNA is embryo-lethal in zebrafish [14]. Moreover, FBL overexpression has been reported during the differentiation of human stem cells, and in several cancer studies, suggesting a central role in these processes [8][12][15][16].
1.2. Pseudouridine and rRNA
With the exception of position 50 in the 5S rRNA that is catalyzed by the enzyme PUS7, the formation of Ψs in rRNA is catalyzed by a ribonucleoprotein complex composed of the pseudo-uridine synthase DKC1 associated with H/ACA box snoRNAs [17][18]. In human rRNAs, Ψs are mapped with a Ψ/U ratio of 5–7%, with a total of about 100 sites [17][19][20][21][22]. DKC1 is as an essential protein, and mutations of its gene have been linked to X-linked dyskeratosis congenita disease. Patients may display alterations to skin color, nail dystrophy, bone marrow failure, and an increase in the risk of developing cancer and pulmonary fibrosis, although it is not clear whether these effects are related to the absence of Ψ from rRNA [18].
The role of Ψs in miscoding has been investigated in human cells [23]. SNORA24 (ACA24), a H/ACA box snoRNA guiding the Ψ609 and Ψ863 on the 18S rRNA, has been downregulated in HCC cells [24]. An analysis of ribosomal pre-translocation complex dynamics by sm-FRET indicated changes in tRNA conformation in the A-site in ribosomes lacking Ψ609 and Ψ863 relative to wild-type ribosomes, depending on the tRNA entering the ribosome. It has also been shown that lower levels of SNORA24 expression increase amino-acid misincorporation by 10%–20% and readthrough by 15% at UGA, but not at UAG codons.
The way in which Ψs in rRNAs decrease the accuracy of translation seems to depend on their abundance in the peptidyl transferase and decoding centers of the ribosome [25]. Ψs are known to generate an additional N1 H-bond donor and to stabilize the C3′-endo conformation [26]. This enables Ψs to increase RNA–RNA stability in the fidelity centers of the ribosome [27]. A decrease in the number of Ψ sites is, thus, accompanied by ribosome destabilization, resulting in a decrease in ribosome fidelity.
2. mRNA Modifications Impact the Reading of the Genetic Code
Many studies over the last decade have revealed the importance of mRNA modifications. These modifications are highly dynamic, with eraser proteins able to eradicate the modifications from the mRNA. The dynamic aspect of the modifications allows integration in a very efficient manner of the RNA metabolism and translation to the physiological state of the cell, considering the appearance of possible stresses.
2.1. Inosine on mRNA
The formation of inosine on mRNAs is catalyzed by the adenosine deaminases ADAR1 and 2 [28]. The inosines of mRNAs, like those of tRNAs, play a major role in expansion of the genetic code, with 5072 identified editing sites in human coding sequences [29].
One of the best known examples of the importance of A-to-I editing in mRNA is the modification of the glutamate receptor subunit B (GluRB) precursor messenger RNA: CAG (Gln) → CIG (Arg) in exon 11. This site is modified by ADAR2 and is essential to ensure the impermeability of the glutamate receptor to Ca2+ ions [30]. A defect of this inosine site has, notably, been shown to contribute to neuronal death in amyotrophic lateral sclerosis [31]. The dysregulation of ADAR1 and 2 has also recently been observed in human hepatocellular carcinoma [32]. Patients with an upregulation of ADAR1 and a downregulation of ADAR2 have higher incidences of tumor recurrence and liver cirrhosis, and shorter disease-free survival times. These dysregulations are linked to changes in the number of inosine sites, with, in particular, hyper-editing of the FLNB mRNA and hypo-editing of the COPA mRNA [32]. Finally, ADAR1 seems to act as an oncogene, whereas ADAR2 acts as a tumor suppressor, in hepatocellular carcinoma.
Inosine in mRNAs is known to modulate alternative splicing and stability, but it clearly also plays an essential role as an enhancer of near-cognate tRNA incorporation, ensuring the activity of some proteins [28]. On the other hand, we did not find any significative difference in ribosome profiling between edited and non-edited mRNA in term of translation efficiency in A. Thaliana mitochondria [33]. The conservation of these essential CDS sites, rather than the cognate codon with a G, remains to be evaluated.
2.2. Pseudouridine
Unlike the Ψs found in rRNA, the reaction generating those found in mRNA is catalyzed by pseudo-uridine synthases, which are RNA-independent proteins [21][34][35][36], although the existence of some box H/ACA snoRNAs complementary to mRNAs raises the possibility that RNA-dependent pseudo-urylation of mRNAs also occurs [37][38]. mRNA Ψs are known to be modulated under cellular stress and during development, but no Ψ reader or eraser has yet been described [39]. Within the translated and untranslated regions of mRNAs, pseudo-uridine is present with a Ψ/U ratio of 0.2–0.6%, and 1889 sites have been identified by N3-CMC–enriched pseudo-uridine sequencing [21]. More than 60% of pseudo-uridine residues are located within the coding sequence, suggesting a link with translation [40][41].
In prokaryotes, several studies have described the ability of Ψ to alter base-pairing and induce misincorporation [42][43][44]. However, far fewer studies have been performed on human cells [41]. Amino-acid misincorporation in front of a “U-codon” has been shown to occur at a rate of 1%. The presence of Ψ in mRNA induces the substitution of Ser, Ile or Leu for Phe at UUU/C codons; Cys or His substitution for Tyr at UAU/C codons; and Pro or Gln substitution for Leu at CUA/U/C/G codons (Figure 1).
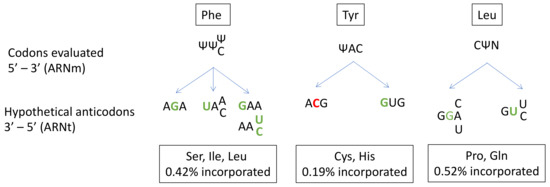
Figure 1. Prediction of anticodon substitution in front of codons with pseudo-uridine (Ψ), based on the amino acid mis-incorporated in the study of Eyler et.al. (2019). Nucleotides in green indicate a mismatch in front of Ψ. Nucleotides in red indicate a mismatch next to Ψ. N: A, U, C or G base. Possibilities of codon-anticodon pairings with more than one mismatch are not represented.
Given the high frequency of Ψ in mRNA and its role in near-cognate tRNA recognition, Ψ modifications probably make a major contribution to translation fidelity. A closer look at codon/anticodon base-pairing in the case of the misincorporation of Cys at a Tyr codon reveals a central mismatch between an A and a C. This unfavorable interaction is probably compensated for by the strong ability of Ψ to stabilize the codon/anticodon structure by stacking interactions. Indeed, Ψ is known to enhance RNA structure stability. Despite its ability to form a supplementary N1-hydrogen bond, Ψ has the same Watson-Crick base-pairing properties as U [26].
2.3. N6-methyladenosine (m6A)
The N6-methyladenosine (m6A) modification involves the addition of a methyl group to the N atom linked to the C6 of adenosine. Chemical predictions of the impact of m6A on RNA–RNA base-pairing suggest a disruption of this interaction due to the methyl group [45]. Indeed, this group must adopt an anti-conformation in the context of A–U pairing. This conformation is less energetically favorable than the syn conformation, leading to destabilization of the RNA–RNA accommodation.
In humans, more than 12000 m6A sites are estimated to be present on 7000 mRNAs [46][47]. About 35% of m6A sites are located within the coding region [48]. m6A is a dynamic modification that has been reported to interact with several enzymes called readers [49][50]. A heterodimeric methylase complex (METTL3-METTL14) is responsible for adding the methyl group. Once modified, the site can be recognized by reader proteins, most of which belong to the YTH-domain protein family (YTHDC and YTHDF), or eraser proteins, which are demethylases (such as FTO and ALKBH5).
The impact of m6A at the first or second position of the codon has been measured by quench flow techniques [51]. This modification delays tRNA incorporation, by slowing tRNA accommodation at site A of the ribosome. However, it has been reported that m6A at the middle position of the codon has a lesser effect on pairing for near-cognate than for cognate tRNAs [52]. This difference in kinetics suggests that tRNA misincorporation rates are likely to be higher in the presence of m6A at the middle position. However, contrary to these findings for prokaryotic systems, mass spectrometry assays in eukaryotes (wheat germ and HEK293T) identified no miscoding effect of the m6A modification [53][54]. The method used for eukaryote systems may be insufficiently sensitive to detect misincorporation in the context of cognate/near-cognate competition. Indeed, the same study found no miscoding effect of Ψ modification, contradicting the findings of another team published in the same year [41]. In the face of these conflicting data, further studies are required to clarify the impact of m6A on miscoding events.
m6A is one of the most commonly studied RNA modifications because of its broad influence on RNA maturation and degradation, RNA-protein interactions and translation efficiency, implicating this modification in a number of different biological processes [48][55][56][57][58][59]. Focusing on human health, altered m6A levels have been implicated in the regulation of the expression of genes relating to cancer pathogenesis and development [60].
2.4. 5-methylcytosine
5-methylcytosine (m5C) is a cytosine with an additional methyl group on C5. Like m6A, m5C is a dynamic modification, with writer, reader and eraser proteins. NSUN2 and the Aly/REF export factor are the principal m5C mRNA writer and reader proteins, respectively [61]. m5C has been mapped on several transcriptomes in humans [62][63][64][65]. Although NSUN2 and NSUN6 are well-known tRNA-modification enzymes, they also appear to modify mRNA. The number of m5C sites in mRNA has been estimated at about a thousand by bisulfite RNA sequencing [66]. Interestingly, viral RNAs are particularly rich in m5C modifications, suggesting that it could play a role in the discrimination of endogenous and exogenous RNAs.
The question of the impact of m5C on translation has been addressed by ribosome profiling in Hues9 human embryonic stem cells with a knockout of NSUN6 gene [67]. No global translational defect was observed, but the absence of NSUN6 was found to trigger stop codon enrichment at the P-site of the ribosome, possibly after readthrough, and an increase in ribosomes bound to the 3′UTR of mRNAs modified by NSUN6. These data suggest that m5C sites in the 3′UTR of mRNA enhance translation termination efficiency by decreasing the readthrough rate. It remains unclear how m5C in the 3′UTR affects termination. Another study in HEK293T cells assessed the impact of m5C at the three codon positions by mass spectrometry [54]. None of the three positions was found to modulate the misincorporation of amino acids.
m5C is linked to human health. Indeed, NSUN2 mutations are associated with growth retardation, neurodevelopmental defects, and have been identified as a possible treatment target for tumors [63][68][69][70][71]. Moreover, the m5C reader and eraser proteins cited above are known to display altered expression levels in various types of cancer [61].
3. Utilization of RNA Modifications to Treat Human Diseases
The field of RNA modifications is undoubtedly a very promising area in human therapy. Synthetic modified mRNAs can be used in diverse therapeutic contexts, including cardiac regeneration, asthma, cystic fibrosis or lung diseases [72][73][74][75]. The best-known application is probably the current COVID-19 vaccines of Pfizer/BioNtech and Moderna. In these mRNA-based vaccines, all the uridine residues are replaced by N1-methyl-pseudouridines to prevent the recognition of the vaccine mRNA by host RNA sensors and to stimulate translation initiation by attenuating eIF2α phosphorylation [76][77]. For those interested in eiF2α stress response and translational regulations, please see the following review [78]. It is also possible to target mRNAs directly, through the use of artificial snoRNAs to replace a U residue with a Ψ at a specific position [42]. In this example, changing the U to a Ψ at the first position of a premature termination codon leads to the incorporation of several amino acids rather than a stopping of translation. This could restore production of the full-length protein, thereby correcting the genetic defect.
From another standpoint, RNA modifications affect diverse biological processes, and the correct incorporation of many of these modifications, at the correct sites, is required for normal development. Alterations to these modifications have been implicated in several diseases, including cancers and resistance to therapy of melanoma cells [79]. The role of m6A in cancer is very well documented, and m5C has also emerged as a major player in cancer development [80][81][82]. Given the crucial roles of writer, reader and eraser proteins in cell homeostasis, these proteins have naturally emerged as potential treatment targets [83]. Ribosome modifications are also of potential interest in this context, and DKC1 and FBL may serve as potential anticancer targets, as shown by the changes in their levels of expression in many cancers [12][84].
As discussed above, it is possible to target mRNA with an H/ACA snoRNA for the incorporation of a Ψ at a specific position. This approach could be used in genetic diseases caused by the presence of a premature termination codon (PTC). Proof-of-concept has been obtained through the demonstration that replacing the U of the stop codon with Ψ converts the stop codon into a sense codon [42]. Indeed, serine and threonine were found at ΨAA and ΨAG codons, whereas tyrosine and phenylalanine were found at ΨGA codons. In principle, it should be possible to change the modification status of tRNAs to modulate translation fidelity. This would be particularly useful in diseases linked to the appearance of a premature stop codon, which are treated with readthrough-inducing molecules. These molecules, such as aminoglycosides, target the ribosome, enabling it to read through the stop codon, but it should be possible to improve the incorporation of specific tRNAs by altering their modifications [85]. However, in this case, a delicate balance must be found between promoting high levels of readthrough without compromising normal tRNA usage. The recent publication describing the stimulation of UGA readthrough by inhibiting the Cm34 modification on tRNATrp with 2,6-diaminopurine (DAP) paves the way for the development of such therapeutic approaches [86]. We are still at the dawning of the epi-transcriptomic era, particularly as concerns human treatments, but this field promises to yield extraordinary advances.
References
- Sloan, K.E.; Warda, A.S.; Sharma, S.; Entian, K.D.; Lafontaine, D.L.J.; Bohnsack, M.T. Tuning the ribosome: The influence of rRNA modification on eukaryotic ribosome biogenesis and function. RNA Biol. 2017, 14, 1138–1152.
- Shubina, M.Y.; Musinova, Y.R.; Sheval, E.V. Proliferation, cancer, and aging-novel functions of the nucleolar methyltransferase fibrillarin? Cell Biol. Int. 2018, 42, 1463–1466.
- Natchiar, S.K.; Myasnikov, A.G.; Kratzat, H.; Hazemann, I.; Klaholz, B.P. Visualization of chemical modifications in the human 80S ribosome structure. Nature 2017, 551, 472–477.
- Taoka, M.; Nobe, Y.; Yamaki, Y.; Sato, K.; Ishikawa, H.; Izumikawa, K.; Yamauchi, Y.; Hirota, K.; Nakayama, H.; Takahashi, N.; et al. Landscape of the complete RNA chemical modifications in the human 80S ribosome. Nucleic Acids Res. 2018, 46, 9289–9298.
- Baudin-Baillieu, A.; Namy, O. Saccharomyces cerevisiae, a powerful model for studying rRNA modifications and their effects on translation fidelity. Int. J. Mol. Sci. 2021, 22, 7419.
- Assi, H.A.; Shi, H.; Liu, B.; Clay, M.; Erharter, K.; Kreutz, C.; Holley, C.; Al-Hashimi, H. 2′-O-methylation alters the RNA secondary structural ensemble. bioRxiv 2020, 2020, 121996.
- Watkins, N.J.; Bohnsack, M.T. The box C/D and H/ACA snoRNPs: Key players in the modification, processing and the dynamic folding of ribosomal RNA. Wiley Interdiscip. Rev. RNA 2012, 3, 397–414.
- Erales, J.; Marchand, V.; Panthu, B.; Gillot, S.; Belin, S.; Ghayad, S.E.; Garcia, M.; Laforêts, F.; Marcel, V.; Baudin-Baillieu, A.; et al. Evidence for rRNA 2′-O-methylation plasticity: Control of intrinsic translational capabilities of human ribosomes. Proc. Natl. Acad. Sci. USA 2017, 114, 12934–12939.
- Sharma, S.; Marchand, V.; Motorin, Y.; Lafontaine, D.L.J. Identification of sites of 2′-O-methylation vulnerability in human ribosomal RNAs by systematic mapping. Sci. Rep. 2017, 7, 1–15.
- Krogh, N.; Jansson, M.D.; Häfner, S.J.; Tehler, D.; Birkedal, U.; Christensen-Dalsgaard, M.; Lund, A.H.; Nielsen, H. Profiling of 2′-O-Me in human rRNA reveals a subset of fractionally modified positions and provides evidence for ribosome heterogeneity. Nucleic Acids Res. 2016, 44, 7884–7895.
- Incarnato, D.; Anselmi, F.; Morandi, E.; Neri, F.; Maldotti, M.; Rapelli, S.; Parlato, C.; Basile, G.; Oliviero, S. High-throughput single-base resolution mapping of RNA 2-O-methylated residues. Nucleic Acids Res. 2017, 45, 1433–1441.
- Marcel, V.; Ghayad, S.E.; Belin, S.; Therizols, G.; Morel, A.P.; Solano-Gonzàlez, E.; Vendrell, J.A.; Hacot, S.; Mertani, H.C.; Albaret, M.A.; et al. P53 Acts as a Safeguard of Translational Control by Regulating Fibrillarin and rRNA Methylation in Cancer. Cancer Cell 2013, 24, 318–330.
- Baudin-baillieu, A.; Fabret, C.; Liang, X.H.; Piekna-Przybylska, D.; Fournier, M.J.; Rousset, J.P. Nucleotide modifications in three functionally important regions of the Saccharomyces cerevisiae ribosome affect translation accuracy. Nucleic Acids Res. 2009, 37, 7665–7677.
- Higa-Nakamine, S.; Suzuki, T.; Uechi, T.; Chakraborty, A.; Nakajima, Y.; Nakamura, M.; Hirano, N.; Suzuki, T.; Kenmochi, N. Loss of ribosomal RNA modification causes developmental defects in zebrafish. Nucleic Acids Res. 2012, 40, 391–398.
- Watanabe-Susaki, K.; Takada, H.; Enomoto, K.; Miwata, K.; Ishimine, H.; Intoh, A.; Ohtaka, M.; Nakanishi, M.; Sugino, H.; Asashima, M.; et al. Biosynthesis of Ribosomal RNA in Nucleoli Regulates Pluripotency and Differentiation Ability of Pluripotent Stem Cells. Stem Cells 2014, 32, 3099–3111.
- Su, H.; Xu, T.; Ganapathy, S.; Shadfan, M.; Long, M.; Huang, T.H.M.; Thompson, I.; Yuan, Z.M. Elevated snoRNA biogenesis is essential in breast cancer. Oncogene 2014, 33, 1348–1358.
- Schattner, P.; Barberan-Soler, S.; Lowe, T.M. A computational screen for mammalian pseudouridylation guide H/ACA RNAs. RNA 2006, 12, 15–25.
- Jack, K.; Bellodi, C.; Landry, D.M.; Niederer, R.O.; Meskauskas, A.; Musalgaonkar, S.; Kopmar, N.; Krasnykh, O.; Dean, A.M.; Thompson, S.R.; et al. RRNA Pseudouridylation Defects Affect Ribosomal Ligand Binding and Translational Fidelity from Yeast to Human Cells. Mol. Cell 2011, 44, 660–666.
- Hughes, D.G.; Maden, E.H. The pseudouridine contents of the ribosomal ribonucleic acids of three vertebrate species. Numerical correspondence between pseudouridine residues and 2′-O-methyl groups is not always conserved. Biochem. J. 1978, 171, 781–786.
- Maden, B.E.H. The Numerous Modified Nucleotides in Eukaryotic Ribosomal RNA. Prog. Nucleic Acid Res. Mol. Biol. 1990, 39, 241–303.
- Li, X.; Zhu, P.; Ma, S.; Song, J.; Bai, J.; Sun, F.; Yi, C. Chemical pulldown reveals dynamic pseudouridylation of the mammalian transcriptome. Nat. Chem. Biol. 2015, 11, 592–597.
- Bakin, A.V.; Ofengand, J. Mapping of pseudouridine residues in RNA to nucleotide resolution. Methods Mol. Biol. 1998, 77, 297–309.
- McMahon, M.; Contreras, A.; Holm, M.; Uechi, T.; Forester, C.M.; Pang, X.; Jackson, C.; Calvert, M.E.; Chen, B.; Quigley, D.A.; et al. A single H/ACA small nucleolar RNA mediates tumor suppression downstream of oncogenic RAS. Elife 2019, 8, e48847.
- Kiss, A.M.; Jády, B.E.; Bertrand, E.; Kiss, T. Human Box H/ACA Pseudouridylation Guide RNA Machinery. Mol. Cell. Biol. 2004, 24, 5797–5807.
- De Zoysa, M.D.; Yu, Y.T. Posttranscriptional RNA Pseudouridylation. In Enzymes; Academic Press: Cambridge, MA, USA, 2017; Volume 41, pp. 151–167.
- Deb, I.; Popenda, Ł.; Sarzyńska, J.; Małgowska, M.; Lahiri, A.; Gdaniec, Z.; Kierzek, R. Computational and NMR studies of RNA duplexes with an internal pseudouridine-adenosine base pair. Sci. Rep. 2019, 9, 1–13.
- Liang, X.H.; Liu, Q.; Fournier, M.J. rRNA Modifications in an Intersubunit Bridge of the Ribosome Strongly Affect Both Ribosome Biogenesis and Activity. Mol. Cell 2007, 28, 965–977.
- Nishikura, K. Functions and regulation of RNA editing by ADAR deaminases. Annu. Rev. Biochem. 2010, 79, 321–349.
- Sakurai, M.; Yano, T.; Kawabata, H.; Ueda, H.; Suzuki, T. Inosine cyanoethylation identifies A-to-I RNA editing sites in the human transcriptome. Nat. Chem. Biol. 2010, 6, 733–740.
- Sommer, B.; Köhler, M.; Sprengel, R.; Seeburg, P.H. RNA editing in brain controls a determinant of ion flow in glutamate-gated channels. Cell 1991, 67, 11–19.
- Kawahara, Y.; Ito, K.; Sun, H.; Aizawa, H.; Kanazawa, I.; Kwak, S. RNA editing and death of motor neurons: There is a glutamate-receptor defect in patients with amyotrophic lateral sclerosis. Nature 2004, 427, 801.
- Chan, T.H.M.; Lin, C.H.; Qi, L.; Fei, J.; Li, Y.; Yong, K.J.; Liu, M.; Song, Y.; Chow, R.K.K.; Ng, V.H.E.; et al. A disrupted RNA editing balance mediated by ADARs (Adenosine DeAminases that act on RNA) in human hepatocellular carcinoma. Gut 2014, 63, 832–843.
- Planchard, N.; Bertin, P.; Quadrado, M.; Dargel-Graffin, C.; Hatin, I.; Namy, O.; Mireau, H. The translational landscape of Arabidopsis mitochondria. Nucleic Acids Res. 2018, 46, 6218.
- Carlile, T.M.; Rojas-Duran, M.F.; Zinshteyn, B.; Shin, H.; Bartoli, K.M.; Gilbert, W.V. Pseudouridine profiling reveals regulated mRNA pseudouridylation in yeast and human cells. Nature 2014, 515, 143–146.
- Carlile, T.M.; Martinez, N.M.; Schaening, C.; Su, A.; Bell, T.A.; Zinshteyn, B.; Gilbert, W.V. mRNA structure determines modification by pseudouridine synthase 1. Nat. Chem. Biol. 2019, 15, 966–974.
- Safra, M.; Nir, R.; Farouq, D.; Slutzkin, I.V.; Schwartz, S. TRUB1 is the predominant pseudouridine synthase acting on mammalian mRNA via a predictable and conserved code. Genome Res. 2017, 27, 393–406.
- Hüttenhofer, A.; Brosius, J.; Bachellerie, J.P. RNomics: Identification and function of small, non-messenger RNAs. Curr. Opin. Chem. Biol. 2002, 6, 835–843.
- Cavaillé, J.; Buiting, K.; Kiefmann, M.; Lalande, M.; Brannan, C.I.; Horsthemke, B.; Bachellerie, J.P.; Brosius, J.; Hüttenhofer, A. Identification of brain-specific and imprinted small nucleolar RNA genes exhibiting an unusual genomic organization. Proc. Natl. Acad. Sci. USA 2000, 97, 14311–14316.
- Vandivier, L.E.; Gregory, B.D. Reading the Epitranscriptome: New Techniques and Perspectives. In Enzymes; Academic Press: Cambridge, MA, USA, 2017; Volume 41, pp. 269–298.
- Gilbert, W.V.; Bell, T.A.; Schaening, C. Messenger RNA modifications: Form, distribution, and function. Science 2016, 352, 1408–1412.
- Eyler, D.E.; Franco, M.K.; Batool, Z.; Wu, M.Z.; Dubuke, M.L.; Dobosz-Bartoszek, M.; Jones, J.D.; Polikanov, Y.S.; Roy, B.; Koutmou, K.S. Pseudouridinylation of mRNA coding sequences alters translation. Proc. Natl. Acad. Sci. USA 2019, 116, 23068–23074.
- Karijolich, J.; Yu, Y.T. Converting nonsense codons into sense codons by targeted pseudouridylation. Nature 2011, 474, 395–399.
- Fernández, I.S.; Ng, C.L.; Kelley, A.C.; Wu, G.; Yu, Y.T.; Ramakrishnan, V. Unusual base pairing during the decoding of a stop codon by the ribosome. Nature 2013, 500, 107–110.
- Svidritskiy, E.; Madireddy, R.; Korostelev, A.A. Structural Basis for Translation Termination on a Pseudouridylated Stop Codon. J. Mol. Biol. 2016, 428, 2228–2236.
- Jordan Ontiveros, R.; Stoute, J.; Liu, K.F. The chemical diversity of RNA modifications. Biochem. J. 2019, 476, 1227–1245.
- Meyer, K.D.; Saletore, Y.; Zumbo, P.; Elemento, O.; Mason, C.E.; Jaffrey, S.R. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 2012, 149, 1635–1646.
- Dominissini, D.; Moshitch-Moshkovitz, S.; Schwartz, S.; Salmon-Divon, M.; Ungar, L.; Osenberg, S.; Cesarkas, K.; Jacob-Hirsch, J.; Amariglio, N.; Kupiec, M.; et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 2012, 485, 201–206.
- Mao, Y.; Dong, L.; Liu, X.M.; Guo, J.; Ma, H.; Shen, B.; Qian, S.B. m6A in mRNA coding regions promotes translation via the RNA helicase-containing YTHDC2. Nat. Commun. 2019, 10, 1–11.
- Meyer, K.D.; Jaffrey, S.R. Rethinking m6A readers, writers, and erasers. Annu. Rev. Cell Dev. Biol. 2017, 33, 319–342.
- Balacco, D.L.; Soller, M. The m6A Writer: Rise of a Machine for Growing Tasks. Biochemistry 2019, 58, 363–378.
- Choi, J.; Ieong, K.W.; Demirci, H.; Chen, J.; Petrov, A.; Prabhakar, A.; O’Leary, S.E.; Dominissini, D.; Rechavi, G.; Soltis, S.M.; et al. N6-methyladenosine in mRNA disrupts tRNA selection and translation-elongation dynamics. Nat. Struct. Mol. Biol. 2016, 23, 110–115.
- Ieong, K.-W.; Indrisiunaite, G.; Prabhakar, A.; Puglisi, J.D.; Ehrenberg, M. N 6-Methyladenosines in mRNAs reduce the accuracy of codon reading by transfer RNAs and peptide release factors. Nucleic Acids Res. 2021, 49, 2684–2699.
- You, C.; Dai, X.; Wang, Y. Position-dependent effects of regioisomeric methylated adenine and guanine ribonucleosides on translation. Nucleic Acids Res. 2017, 45, 9059–9067.
- Hoernes, T.P.; Heimdörfer, D.; Köstner, D.; Faserl, K.; Nußbaumer, F.; Plangger, R.; Kreutz, C.; Lindner, H.; Erlacher, M.D. Eukaryotic translation elongation is modulated by single natural nucleotide derivatives in the coding sequences of mRNAs. Genes 2019, 10, 84.
- Wang, X.; Lu, Z.; Gomez, A.; Hon, G.C.; Yue, Y.; Han, D.; Fu, Y.; Parisien, M.; Dai, Q.; Jia, G.; et al. N 6-methyladenosine-dependent regulation of messenger RNA stability. Nature 2014, 505, 117–120.
- Wang, X.; Zhao, B.S.; Roundtree, I.A.; Lu, Z.; Han, D.; Ma, H.; Weng, X.; Chen, K.; Shi, H.; He, C. N6-methyladenosine modulates messenger RNA translation efficiency. Cell 2015, 161, 1388–1399.
- Liu, N.; Dai, Q.; Zheng, G.; He, C.; Parisien, M.; Pan, T. N6 -methyladenosine-dependent RNA structural switches regulate RNA-protein interactions. Nature 2015, 518, 560–564.
- Tang, Y.; Chen, K.; Song, B.; Ma, J.; Wu, X.; Xu, Q.; Wei, Z.; Su, J.; Liu, G.; Rong, R.; et al. M6A-Atlas: A comprehensive knowledgebase for unraveling the N6-methyladenosine (m6A) epitranscriptome. Nucleic Acids Res. 2021, 49, D134–D143.
- Meyer, K.D.; Patil, D.P.; Zhou, J.; Zinoviev, A.; Skabkin, M.A.; Elemento, O.; Pestova, T.V.; Qian, S.B.; Jaffrey, S.R. 5′ UTR m6A Promotes Cap-Independent Translation. Cell 2015, 163, 999–1010.
- He, L.; Li, H.; Wu, A.; Peng, Y.; Shu, G.; Yin, G. Functions of N6-methyladenosine and its role in cancer. Mol. Cancer 2019, 18, 1–15.
- García-Vílchez, R.; Sevilla, A.; Blanco, S. Post-transcriptional regulation by cytosine-5 methylation of RNA. Biochim. Biophys. Acta Gene Regul. Mech. 2019, 1862, 240–252.
- Squires, J.E.; Patel, H.R.; Nousch, M.; Sibbritt, T.; Humphreys, D.T.; Parker, B.J.; Suter, C.M.; Preiss, T. Widespread occurrence of 5-methylcytosine in human coding and non-coding RNA. Nucleic Acids Res. 2012, 40, 5023–5033.
- Blanco, S.; Dietmann, S.; Flores, J.V.; Hussain, S.; Kutter, C.; Humphreys, P.; Lukk, M.; Lombard, P.; Treps, L.; Popis, M.; et al. Aberrant methylation of t RNA s links cellular stress to neuro-developmental disorders. EMBO J. 2014, 33, 2020–2039.
- Hussain, S.; Sajini, A.A.; Blanco, S.; Dietmann, S.; Lombard, P.; Sugimoto, Y.; Paramor, M.; Gleeson, J.G.; Odom, D.T.; Ule, J.; et al. NSun2-mediated cytosine-5 methylation of vault noncoding RNA determines its processing into regulatory small RNAs. Cell Rep. 2013, 4, 255–261.
- Huang, T.; Chen, W.; Liu, J.; Gu, N.; Zhang, R. Genome-wide identification of mRNA 5-methylcytosine in mammals. Nat. Struct. Mol. Biol. 2019, 26, 380–388.
- Schumann, U.; Zhang, H.N.; Sibbritt, T.; Pan, A.; Horvath, A.; Gross, S.; Clark, S.J.; Yang, L.; Preiss, T. Multiple links between 5-methylcytosine content of mRNA and translation. BMC Biol. 2020, 18, 40.
- Selmi, T.; Hussain, S.; Dietmann, S.; Heiß, M.; Borland, K.; Flad, S.; Carter, J.M.; Dennison, R.; Huang, Y.L.; Kellner, S.; et al. Sequence- and structure-specific cytosine-5 mRNA methylation by NSUN6. Nucleic Acids Res. 2021, 49, 1006–1022.
- Khan, M.A.; Rafiq, M.A.; Noor, A.; Hussain, S.; Flores, J.V.; Rupp, V.; Vincent, A.K.; Malli, R.; Ali, G.; Khan, F.S.; et al. Mutation in NSUN2, which encodes an RNA methyltransferase, causes autosomal-recessive intellectual disability. Am. J. Hum. Genet. 2012, 90, 856–863.
- Abbasi-Moheb, L.; Mertel, S.; Gonsior, M.; Nouri-Vahid, L.; Kahrizi, K.; Cirak, S.; Wieczorek, D.; Motazacker, M.M.; Esmaeeli-Nieh, S.; Cremer, K.; et al. Mutations in NSUN2 cause autosomal- Recessive intellectual disability. Am. J. Hum. Genet. 2012, 90, 847–855.
- Martinez, F.J.; Lee, J.H.; Lee, J.E.; Blanco, S.; Nickerson, E.; Gabrie, S.; Frye, M.; Al-Gazali, L.; Gleeson, J.G. Whole exome sequencing identifies a splicing mutation in NSUN2 as a cause of a Dubowitz-like syndrome. J. Med. Genet. 2012, 49, 380–385.
- Blanco, S.; Bandiera, R.; Popis, M.; Hussain, S.; Lombard, P.; Aleksic, J.; Sajini, A.; Tanna, H.; Cortés-Garrido, R.; Gkatza, N.; et al. Stem cell function and stress response are controlled by protein synthesis. Nature 2016, 534, 335–340.
- Kaur, K.; Zangi, L. Modified mRNA as a Therapeutic Tool for the Heart. Cardiovasc. Drugs Ther. 2020, 34, 871–880.
- Mays, L.E.; Ammon-Treiber, S.; Mothes, B.; Alkhaled, M.; Rottenberger, J.; Müller-Hermelink, E.S.; Grimm, M.; Mezger, M.; Beer-Hammer, S.; Von Stebut, E.; et al. Modified Foxp3 mRNA protects against asthma through an IL-10-dependent mechanism. J. Clin. Investig. 2013, 123, 1216–1228.
- Haque, A.K.M.A.; Dewerth, A.; Antony, J.S.; Riethmüller, J.; Schweizer, G.R.; Weinmann, P.; Latifi, N.; Yasar, H.; Pedemonte, N.; Sondo, E.; et al. Chemically modified hCFTR mRNAs recuperate lung function in a mouse model of cystic fibrosis. Sci. Rep. 2018, 8, 1–14.
- Sahu, I.; Haque, A.K.M.A.; Weidensee, B.; Weinmann, P.; Kormann, M.S.D. Recent Developments in mRNA-Based Protein Supplementation Therapy to Target Lung Diseases. Mol. Ther. 2019, 27, 803–823.
- Nance, K.D.; Meier, J.L. Modifications in an Emergency: The Role of N1-Methylpseudouridine in COVID-19 Vaccines. ACS Cent. Sci. 2021, 7, 748–756.
- Svitkin, Y.V.; Cheng, Y.M.; Chakraborty, T.; Presnyak, V.; John, M.; Sonenberg, N. N1-methyl-pseudouridine in mRNA enhances translation through eIF2α-dependent and independent mechanisms by increasing ribosome density. Nucleic Acids Res. 2017, 45, 6023–6036.
- Wek, R.C. Role of eIF2α kinases in translational control and adaptation to cellular stress. Cold Spring Harb. Perspect. Biol. 2018, 10, a032870.
- Rapino, F.; Delaunay, S.; Rambow, F.; Zhou, Z.; Tharun, L.; De Tullio, P.; Sin, O.; Shostak, K.; Schmitz, S.; Piepers, J.; et al. Codon-specific translation reprogramming promotes resistance to targeted therapy. Nature 2018, 558, 605–609.
- Dong, S.; Wu, Y.; Liu, Y.; Weng, H.; Huang, H. N6-methyladenosine Steers RNA Metabolism and Regulation in Cancer. Cancer Commun. 2021, cac2.12161.
- Huang, J.; Chen, Z.; Chen, X.; Chen, J.; Cheng, Z.; Wang, Z. The role of RNA N6-methyladenosine methyltransferase in cancers. Mol. Ther. Nucleic Acids 2021, 23, 887–896.
- Chen, X.; Li, A.; Sun, B.F.; Yang, Y.; Han, Y.N.; Yuan, X.; Chen, R.X.; Wei, W.S.; Liu, Y.; Gao, C.C.; et al. 5-methylcytosine promotes pathogenesis of bladder cancer through stabilizing mRNAs. Nat. Cell Biol. 2019, 21, 978–990.
- Nombela, P.; Miguel-López, B.; Blanco, S. The role of m6A, m5C and Ψ RNA modifications in cancer: Novel therapeutic opportunities. Mol. Cancer 2021, 20, 1–30.
- Ruggero, D.; Grisendi, S.; Piazza, F.; Rego, E.; Mari, F.; Rao, P.H.; Cordon-Cardo, C.; Pandolfi, P.P. Dyskeratosis congenita and cancer in mice deficient in ribosomal RNA modification. Science 2003, 299, 259–262.
- Bidou, L.; Bugaud, O.; Belakhov, V.; Baasov, T.; Namy, O. Characterization of new-generation aminoglycoside promoting premature termination codon readthrough in cancer cells. RNA Biol. 2017, 14, 378–388.
- Trzaska, C.; Amand, S.; Bailly, C.; Leroy, C.; Marchand, V.; Duvernois-Berthet, E.; Saliou, J.M.; Benhabiles, H.; Werkmeister, E.; Chassat, T.; et al. 2,6-Diaminopurine as a highly potent corrector of UGA nonsense mutations. Nat. Commun. 2020, 11, 1–12.
More
Information
Subjects:
Cell Biology
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.3K
Revisions:
4 times
(View History)
Update Date:
19 Nov 2021
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No




