
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Gerald Audette | + 4101 word(s) | 4101 | 2021-07-07 06:06:42 | | | |
| 2 | Vivi Li | Meta information modification | 4101 | 2021-08-30 04:57:49 | | |
Video Upload Options
Nanobiotechnology involves the study of structures found in nature to construct nanodevices for biological and medical applications with the ultimate goal of commercialization. Within a cell most biochemical processes are driven by proteins and associated macromolecular complexes. Evolution has optimized these protein-based nanosystems within living organisms over millions of years. Among these are flagellin and pilin-based systems from bacteria, viral-based capsids, and eukaryotic microtubules and amyloids. While carbon nanotubes (CNTs), and protein/peptide-CNT composites, remain one of the most researched nanosystems due to their electrical and mechanical properties, there are many concerns regarding CNT toxicity and biodegradability. Therefore, proteins have emerged as useful biotemplates for nanomaterials due to their assembly under physiologically relevant conditions and ease of manipulation via protein engineering.
1. Introduction
2. Protein Nanofibers and Nanotubes (NTs) from Bacterial Systems
2.1. Flagella-Based Protein Nanofibers and Nanotubes
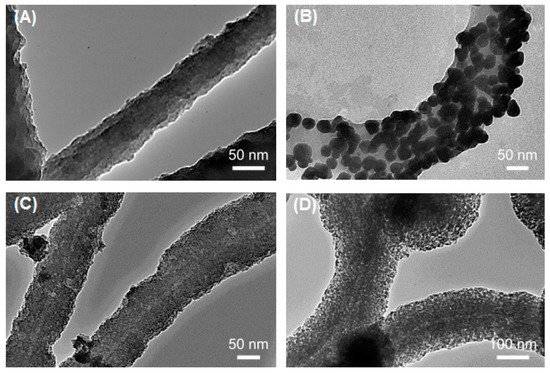
2.2. Pilin-Based Protein Nanotubes
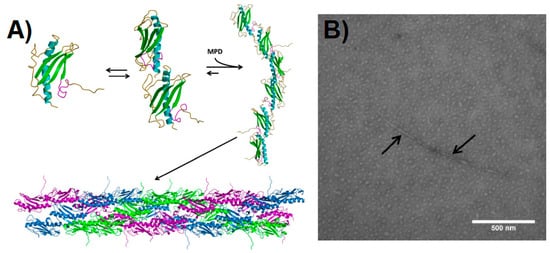
3. Virus-Based Protein Nanotubes (PNTs)
3.1. Tobacco Mosaic Virus (TMV)
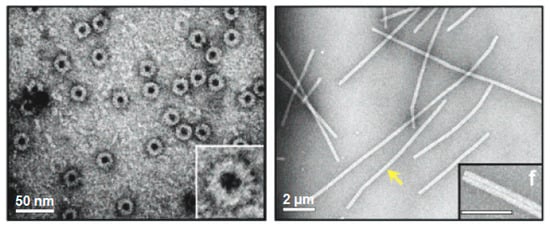
3.2. Cowpea Mosaic Virus (CPMV)
3.3. M13 Bacteriophage
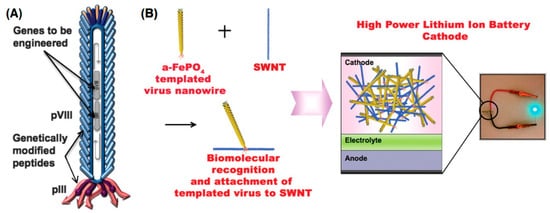
References
- Sanderson, K. Bioengineering: What to make with DNA origami. Nature 2010, 464, 158–159.
- Bell, N.A.W.; Keyser, U.F. Nanopores formed by DNA origami: A review. FEBS Lett. 2014, 588, 3564–3570.
- Li, S.; Jiang, Q.; Liu, S.; Zhang, Y.; Tian, Y.; Song, C.; Wang, J.; Zou, Y.; Anderson, G.J.; Han, J.-Y.Y.; et al. A DNA nanorobot functions as a cancer therapeutic in response to a molecular trigger in vivo. Nat. Biotechnol. 2018, 36, 258–264.
- Gerling, T.; Wagenbauer, K.F.; Neuner, A.M.; Dietz, H. Dynamic DNA devices and assemblies formed by shape-complementary, non–base pairing 3D components. Science 2015, 347, 1446–1452.
- Mashaghi, S.; Jadidi, T.; Koenderink, G.; Mashaghi, A. Lipid Nanotechnology. Int. J. Mol. Sci. 2013, 14, 4242–4282.
- Tesauro, D.; Accardo, A.; Diaferia, C.; Milano, V.; Guillon, J.; Ronga, L.; Rossi, F. Peptide-Based Drug-Delivery Systems in Biotechnological Applications: Recent Advances and Perspectives. Molecules 2019, 24, 351.
- De Frates, K.; Markiewicz, T.; Gallo, P.; Rack, A.; Weyhmiller, A.; Jarmusik, B.; Hu, X. Protein polymer-based nanoparticles: Fabrication and medical applications. Int. J. Mol. Sci. 2018, 19, 1717.
- Petrov, A.; Audette, G.F. Peptide and protein-based nanotubes for nanobiotechnology. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2012, 4, 575–585.
- Oliveira, S.F.; Bisker, G.; Bakh, N.A.; Gibbs, S.L.; Landry, M.P.; Strano, M.S. Protein functionalized carbon nanomaterials for biomedical applications. Carbon NY 2015, 95, 767–779.
- Besteman, K.; Lee, J.; Wiertz, F.G.M.; Heering, H.A.; Dekker, C. Enzyme-Coated Carbon Nanotubes as Single-Molecule Biosensors. Nano Lett. 2003, 3, 727–730.
- Unnikrishnan, B.; Palanisamy, S.; Chen, S.-M. A simple electrochemical approach to fabricate a glucose biosensor based on graphene–glucose oxidase biocomposite. Biosens. Bioelectron. 2013, 39, 70–75.
- Barton, S.C.; Gallaway, J.; Atanassov, P. Enzymatic Biofuel Cells for Implantable and Microscale Devices. Chem. Rev. (Washington, DC) 2004, 104, 4867–4886.
- Kannan, A.M.; Renugopalakrishnan, V.; Filipek, S.; Li, P.; Audette, G.F.; Munukutla, L. Bio-Batteries and Bio-Fuel Cells: Leveraging on Electronic Charge Transfer Proteins. J. Nanosci. Nanotechnol. 2009, 9, 1665–1678.
- Dudzik, J.; Chang, W.-C.; Kannan, A.M.; Filipek, S.; Viswanathan, S.; Li, P.; Renugopalakrishnan, V.; Audette, G.F. Cross-linked glucose oxidase clusters for biofuel cell anode catalysts. Biofabrication 2013, 5, 035009.
- Audette, G.F.; Hazes, B. Development of Protein Nanotubes from a Multi-Purpose Biological Structure. J. Nanosci. Nanotechnol. 2007, 7, 2222–2229.
- Miranda, F.F.; Iwasaki, K.; Akashi, S.; Sumitomo, K.; Kobayashi, M.; Yamashita, I.; Tame, J.R.H.; Heddle, J.G. A Self-Assembled Protein Nanotube with High Aspect Ratio. Small 2009, 5, 2077–2084.
- Hopcroft, N.H.; Manfredo, A.; Wendt, A.L.; Brzozowski, A.M.; Gollnick, P.; Antson, A.A. The Interaction of RNA with TRAP: The Role of Triplet Repeats and Separating Spacer Nucleotides. J. Mol. Biol. 2004, 338, 43–53.
- Nagano, S.; Banwell, E.F.; Iwasaki, K.; Michalak, M.; Pałka, R.; Zhang, K.Y.J.; Voet, A.R.D.; Heddle, J.G. Understanding the Assembly of an Artificial Protein Nanotube. Adv. Mater. Interfaces 2016, 3, 1600846.
- Ballister, E.R.; Lai, A.H.; Zuckermann, R.N.; Cheng, Y.; Mougous, J.D. In vitro self-assembly of tailorable nanotubes from a simple protein building block. Proc. Natl. Acad. Sci. USA 2008, 105, 3733–3738.
- Medalsy, I.; Dgany, O.; Sowwan, M.; Cohen, H.; Yukashevska, A.; Wolf, S.G.; Wolf, A.; Koster, A.; Almog, O.; Marton, I.; et al. SP1 protein-based nanostructures and arrays. Nano Lett. 2008, 8, 473–477.
- Uddin, I.; Frank, S.; Warren, M.J.; Pickersgill, R.W. A Generic Self-Assembly Process in Microcompartments and Synthetic Protein Nanotubes. Small 2018, 14, 1704020.
- Yonekura, K.; Maki-Yonekura, S.; Namba, K. Complete atomic model of the bacterial flagellar filament by electron cryomicroscopy. Nature 2003, 424, 643–650.
- Thomson, N.M.; Rossman, F.M.; Ferreira, J.L.; Matthews-Palmer, T.R.; Beeby, M.; Pallen, M.J. Bacterial Flagellins: Does Size Matter? Trends Microbiol. 2017, 1–7.
- Beatson, S.A.; Minamino, T.; Pallen, M.J. Variation in bacterial flagellins: From sequence to structure. Trends Microbiol. 2006, 14, 151–155.
- Yonekura, K.; Maki, S.; Morgan, D.G.; DeRosier, D.J.; Vonderviszt, F.; Imada, K.; Namba, K. The bacterial flagellar cap as the rotary promoter of flagellin self-assembly. Science 2000, 290, 2148–2152.
- Hoshikawa, H.; Kamiya, R. Elastic properties of bacterial flagellar filaments. II. Determination of the modulus of rigidity. Biophys. Chem. 1985, 22, 159–166.
- Kamiya, R.; Asakura, S. Flagella transformations at alkaline pH. J. Mol. Biol. 1977, 108, 513–518.
- Kamiya, R.; Asakura, S. Helical Transformations of Salmonella Flagella in vitro. J. Mol. Biol. 1976, 106, 167–186.
- Kumara, M.T.; Srividya, N.; Muralidharan, S.; Tripp, B.C. Bioengineered flagella protein nanotubes with cysteine loops: Self-assembly and manipulation in an optical trap. Nano Lett. 2006, 6, 2121–2129.
- Kumara, M.T.; Tripp, B.C.; Muralidharan, S. Layer-by-layer assembly of bioengineered flagella protein nanotubes. Biomacromolecules 2007, 8, 3718–3722.
- Kumara, M.T.; Tripp, B.C.; Muralidharan, S. Self-Assembly of Metal Nanoparticles and Nanotubes on Bioengineered Flagella Scaffolds. Chem. Mater. 2007, 19, 2056–2064.
- Jo, W.; Freedman, K.J.; Yi, D.K.; Kim, M.J. Fabrication of tunable silica-mineralized nanotubes using flagella as bio-templates. Nanotechnology 2012, 23, 55601.
- Jo, W.; Darmawan, M.; Kim, J.; Ahn, C.W.; Byun, D.; Baik, S.H.; Kim, M.J. Electrical property measurements of metallized flagella-templated silica nanotube networks. Nanotechnology 2013, 24, 135704.
- Kolappan, S.; Coureuil, M.; Yu, X.; Nassif, X.; Egelman, E.H.; Craig, L. Structure of the neisseria meningitidis type IV pilus. Nat. Commun. 2016, 7, 13015.
- Folkhard, W.; Marvin, D.A.; Watts, T.H.; Paranchych, W. Structure of polar pili from Pseudomonas aeruginosa strains K and O. J. Mol. Biol. 1981, 149, 79–93.
- Craig, L.; Taylor, R.K.; Pique, M.E.; Adair, B.D.; Arvai, A.S.; Singh, M.; Lloyd, S.J.; Shin, D.S.; Getzoff, E.D.; Yeager, M.; et al. Type IV pilin structure and assembly: X-ray and EM analyses of Vibrio cholerae toxin-coregulated pilus and Pseudomonas aeruginosa PAK pilin. Mol. Cell 2003, 11, 1139–1150.
- Burrows, L.L. Pseudomonas aeruginosa Twitching Motility: Type IV Pili in Action. Annu. Rev. Microbiol. 2012, 66, 493–520.
- Harvey, H.; Habash, M.; Aidoo, F.; Burrows, L.L. Single-Residue Changes in the C-Terminal Disulfide-Bonded Loop of the Pseudomonas aeruginosa Type IV Pilin Influence Pilus Assembly and Twitching Motility. J. Bacteriol. 2009, 191, 6513–6524.
- Craig, L.; Li, J. Type IV pili: Paradoxes in form and function. Curr. Opin. Struct. Biol. 2008, 18, 267–277.
- Burrows, L.L. Weapons of mass retraction. Mol. Microbiol. 2005, 57, 878–888.
- Craig, L.; Pique, M.E.; Tainer, J.A. Type IV pilus structure and bacterial pathogenicity. Nat. Rev. Microbiol. 2004, 2, 363–378.
- Hazes, B.; Sastry, P.A.; Hayakawa, K.; Read, R.J.; Irvin, R.T. Crystal structure of Pseudomonas aeruginosa PAK pilin suggests a main-chain-dominated mode of receptor binding. J. Mol. Biol. 2000, 299, 1005–1017.
- Dunlop, K.V.; Irvin, R.T.; Hazes, B. Pros and cons of cryocrystallography: Should we also collect a room-temperature data set? Acta Crystallogr. Sect. D Biol. Crystallogr. 2005, 61, 80–87.
- Keizer, D.W.; Slupsky, C.M.; Kalisiak, M.; Campbell, A.P.; Crump, M.P.; Sastry, P.A.; Hazes, B.; Irvin, R.T.; Sykes, B.D. Structure of a Pilin Monomer from Pseudomonas aeruginosa: Implications for the assembly of pili. J. Biol. Chem. 2001, 276, 24186–24193.
- Audette, G.F.; Irvin, R.T.; Hazes, B. Crystallographic analysis of the Pseudomonas aeruginosa strain K122-4 monomeric pilin reveals a conserved receptor-binding architecture. Biochemistry 2004, 43, 11427–11435.
- Kao, D.J.; Churchill, M.E.A.; Irvin, R.T.; Hodges, R.S. Animal Protection and Structural Studies of a Consensus Sequence Vaccine Targeting the Receptor Binding Domain of the Type IV Pilus of Pseudomonas aeruginosa. J. Mol. Biol. 2007, 374, 426–442.
- Nguyen, Y.; Jackson, S.G.; Aidoo, F.; Junop, M.; Burrows, L.L. Structural Characterization of Novel Pseudomonas aeruginosa Type IV Pilins. J. Mol. Biol. 2010, 395, 491–503.
- Parge, H.E.; Forest, K.T.; Hickey, M.J.; Christensen, D.A.; Getzoff, E.D.; Tainer, J.A. Structure of the fibre-forming protein pilin at 2.6 A resolution. Nature 1995, 378, 32–38.
- Piepenbrink, K.H.; Maldarelli, G.A.; Martinez de la Peña, C.F.; Dingle, T.C.; Mulvey, G.L.; Lee, A.; von Rosenvinge, E.; Armstrong, G.D.; Donnenberg, M.S.; Sundberg, E.J. Structural and Evolutionary Analyses Show Unique Stabilization Strategies in the Type IV Pili of Clostridium difficile. Structure 2015, 23, 385–396.
- Piepenbrink, K.H.; Maldarelli, G.A.; Martinez de la Peña, C.F.; Mulvey, G.L.; Snyder, G.A.; De Masi, L.; von Rosenvinge, E.C.; Günther, S.; Armstrong, G.D.; Donnenberg, M.S.; et al. Structure of Clostridium difficile PilJ Exhibits Unprecedented Divergence from Known Type IV Pilins. J. Biol. Chem. 2014, 289, 4334–4345.
- Wang, F.; Coureuil, M.; Osinski, T.; Orlova, A.; Altindal, T.; Gesbert, G.; Nassif, X.; Egelman, E.H.; Craig, L. Cryoelectron Microscopy Reconstructions of the Pseudomonas aeruginosa and Neisseria gonorrhoeae Type IV Pili at Sub-nanometer Resolution. Structure 2017, 25, 1423–1435.
- Audette, G.F.; Van Schaik, E.J.; Hazes, B.; Irvin, R.T. DNA-binding protein nanotubes: Learning from nature’s nanotech examples. Nano Lett. 2004, 4, 1897–1902.
- Lombardo, S.; Jasbi, S.Z.; Jeung, S.; Morin, S.; Audette, G.F. Initial Studies of Protein Nanotube Oligomerization from a Modified Gold Surface. J. Bionanosci. 2009, 3, 61–65.
- Petrov, A.; Lombardo, S.; Audette, G.F. Fibril-mediated oligomerization of pilin-derived protein nanotubes. J. Nanobiotechnol. 2013, 11, 24.
- Lento, C.; Wilson, D.J.; Audette, G.F. Dimerization of the type IV pilin from Pseudomonas aeruginosa strain K122-4 results in increased helix stability as measured by time-resolved hydrogen-deuterium exchange. Struct. Dyn. 2016, 3, 012001.
- Yu, B.; Giltner, C.L.; Van Schaik, E.J.; Bautista, D.L.; Hodges, R.S.; Audette, G.F.; Li, D.Y.; Irvin, R.T. A Novel Biometallic Interface: High Affinity Tip-AssociatedBinding by Pilin-Derived Protein Nanotubes. J. Bionanosci. 2007, 1, 73–83.
- Reguera, G.; McCarthy, K.D.; Mehta, T.; Nicoll, J.S.; Tuominen, M.T.; Lovley, D.R. Extracellular electron transfer via microbial nanowires. Nature 2005, 435, 1098–1101.
- Reguera, G. Harnessing the power of microbial nanowires. Microb. Biotechnol. 2018, 1–16.
- Yi, H.; Nevin, K.P.; Kim, B.-C.C.; Franks, A.E.; Klimes, A.; Tender, L.M.; Lovley, D.R. Selection of a variant of Geobacter sulfurreducens with enhanced capacity for current production in microbial fuel cells. Biosens. Bioelectron. 2009, 24, 3498–3503.
- Malvankar, N.S.; Vargas, M.; Nevin, K.P.; Franks, A.E.; Leang, C.; Kim, B.-C.C.; Inoue, K.; Mester, T.; Covalla, S.F.; Johnson, J.P.; et al. Tunable metallic-like conductivity in microbial nanowire networks. Nat. Nanotechnol. 2011, 6, 573–579.
- Malvankar, N.S.; Mester, T.; Tuominen, M.T.; Lovley, D.R. Supercapacitors Based on C-Type Cytochromes Using Conductive Nanostructured Networks of Living Bacteria. Chem. Phys. Chem. 2012, 13, 463–468.
- Giltner, C.L.; van Schaik, E.J.; Audette, G.F.; Kao, D.; Hodges, R.S.; Hassett, D.J.; Irvin, R.T. The Pseudomonas aeruginosa type IV pilin receptor binding domain functions as an adhesin for both biotic and abiotic surfaces. Mol. Microbiol. 2006, 59, 1083–1096.
- Young, M.; Willits, D.; Uchida, M.; Douglas, T. Plant viruses as biotemplates for materials and their use in nanotechnology. Annu. Rev. Phytopathol. 2008, 46, 361–384.
- Fraenkel-Conrat, H.; Williams, R.C. Reconstitution of Active Tobacco Mosaic Virus from Its Inactive Protein and Nucleic Acid Components. Pro.c Natl. Acad. Sci. USA 1955, 41, 690–698.
- Perham, R.N.; Wilson, T.M. The characterization of intermediates formed during the disassembly of tobacco mosaic virus at alkaline pH. Virology 1978, 84, 293–302.
- Atabekov, J.; Nikitin, N.; Arkhipenko, M.; Chirkov, S.; Karpova, O. Thermal transition of native tobacco mosaic virus and RNA-free viral proteins into spherical nanoparticles. J. Gen. Virol. 2011, 92, 453–456.
- Balci, S.; Bittner, A.M.; Hahn, K.; Scheu, C.; Knez, M.; Kadri, A.; Wege, C.; Jeske, H.; Kern, K. Copper nanowires within the central channel of tobacco mosaic virus particles. Electrochim. Acta 2006, 51, 6251–6257.
- Balci, S.; Bittner, A.M.; Schirra, M.; Thonke, K.; Sauer, R.; Hahn, K.; Kadri, A.; Wege, C.; Jeske, H.; Kern, K. Catalytic coating of virus particles with zinc oxide. Electrochim. Acta 2009, 54, 5149–5154.
- Miller, R.A.; Presley, A.D.; Francis, M.B. Self-assembling light-harvesting systems from synthetically modified tobacco mosaic virus coat proteins. J. Am. Chem. Soc. 2007, 129, 3104–3109.
- Endo, M.; Fujitsuka, M.; Majima, T. Porphyrin light-harvesting arrays constructed in the recombinant tobacco mosaic virus scaffold. Chem. A Eur. J. 2007, 13, 8660–8666.
- Lin, T.; Chen, Z.; Usha, R.; Stauffacher, C.V.; Dai, J.B.; Schmidt, T.; Johnson, J.E. The refined crystal structure of cowpea mosaic virus at 2.8 A resolution. Virology 1999, 265, 20–34.
- Steinmetz, N.F.; Shah, S.N.; Barclay, J.E.; Rallapalli, G.; Lomonossoff, G.P.; Evans, D.J. Virus-templated silica nanoparticles. Small 2009, 5, 813–816.
- Porta, C.; Spall, V.E.; Findlay, K.C.; Gergerich, R.C.; Farrance, C.E.; Lomonossoff, G.P. Cowpea mosaic virus-based chimaeras. Effects of inserted peptides on the phenotype, host range, and transmissibility of the modified viruses. Virology 2003, 310, 50–63.
- Lewis, J.D.; Destito, G.; Zijlstra, A.; Gonzalez, M.J.; Quigley, J.P.; Manchester, M.; Stuhlmann, H. Viral nanoparticles as tools for intravital vascular imaging. Nat. Med. 2006, 12, 354–360.
- Brunel, F.M.; Lewis, J.D.; Destito, G.; Steinmetz, N.F.; Manchester, M.; Stuhlmann, H.; Dawson, P.E. Hydrazone ligation strategy to assemble multifunctional viral nanoparticles for cell imaging and tumor targeting. Nano Lett. 2010, 10, 1093–1097.
- Langeveld, J.P.; Brennan, F.R.; Martinez-Torrecuadrada, J.L.; Jones, T.D.; Boshuizen, R.S.; Vela, C.; Casal, J.I.; Kamstrup, S.; Dalsgaard, K.; Meloen, R.H.; et al. Inactivated recombinant plant virus protects dogs from a lethal challenge with canine parvovirus. Vaccine 2001, 19, 3661–3670.
- Huang, Y.; Chiang, C.Y.; Lee, S.K.; Gao, Y.; Hu, E.L.; De Yoreo, J.; Belcher, A.M. Programmable assembly of nanoarchitectures using genetically engineered viruses. Nano Lett. 2005, 5, 1429–1434.
- Smith, G.P.; Petrenko, V.A. Phage Display. Chem. Rev. 1997, 97, 391–410.
- Wu, C.H.; Liu, I.J.; Lu, R.M.; Wu, H.C. Advancement and applications of peptide phage display technology in biomedical science. J. Biomed. Sci. 2016, 23, 1–14.
- Rami, A.; Behdani, M.; Yardehnavi, N.; Habibi-Anbouhi, M.; Kazemi-Lomedasht, F. An overview on application of phage display technique in immunological studies. Asian Pac. J. Trop. Biomed. 2017, 7, 599–602.
- Ledsgaard, L.; Kilstrup, M.; Karatt-Vellatt, A.; McCafferty, J.; Laustsen, A.H. Basics of antibody phage display technology. Toxins 2018, 10, 236.
- Henry, K.A.; Arbabi-Ghahroudi, M.; Scott, J.K. Beyond phage display: Non-traditional applications of the filamentous bacteriophage as a vaccine carrier, therapeutic biologic, and bioconjugation scaffold. Front. Microbiol. 2015, 6, 1–18.
- Mao, C.; Solis, D.J.; Reiss, B.D.; Kottmann, S.T.; Sweeney, R.Y.; Hayhurst, A.; Georgiou, G.; Iverson, B.; Belcher, A.M. Virus-based toolkit for the directed synthesis of magnetic and semiconducting nanowires. Science 2004, 303, 213–217.
- Sarikaya, M.; Tamerler, C.; Jen, A.K.; Schulten, K.; Baneyx, F. Molecular biomimetics: Nanotechnology through biology. Nat. Mater. 2003, 2, 577–585.
- Nam, K.T.; Kim, D.W.; Yoo, P.J.; Chiang, C.Y.; Meethong, N.; Hammond, P.T.; Chiang, Y.M.; Belcher, A.M. Virus-enabled synthesis and assembly of nanowires for lithium ion battery electrodes. Science 2006, 312, 885–888.
- Lee, Y.J.; Li, H.; Kim, W.-J.; Kang, K.; Yun, D.S.; Strano, M.S.; Ceder, G.; Belcher, A.M. Fabricating genetically engineered high-power lithium-ion batteries using multiple virus genes. Science 2009, 324, 1051–1055.
- Wang, Q.; Li, K.; Chen, Y.; Li, S.; Nguyen, H.G.; Niu, Z.; You, S.; Mello, C.M.; Lu, X.; Wang, Q. Chemical modification of M13 bacteriophage and its application in cancer cell imaging. Bioconjug. Chem. 2010, 21, 1369–1377.
- Frenkel, D.; Solomon, B. Filamentous phage as vector-mediated antibody delivery to the brain. Proc. Natl. Acad. Sci. USA 2002, 99, 5675–5679.
- Dickerson, T.J.; Kaufmann, G.F.; Janda, K.D. Bacteriophage-mediated protein delivery into the central nervous system and its application in immunopharmacotherapy. Expert Opin. Biol. Ther. 2005, 5, 773–781.
- Carrera, M.R.; Kaufmann, G.F.; Mee, J.M.; Meijler, M.M.; Koob, G.F.; Janda, K.D. Treating cocaine addiction with viruses. Proc. Natl. Acad. Sci. USA 2004, 101, 10416–10421.




