Healthcare electronic records can provide both physicians and healthcare agencies to discover knowledge. This work proposes an overview of the data mining techniques used for knowledge discovery in medical records. Furthermore, based on real healthcare data, this paper also demonstrates a case study of discovering knowledge with the help of three data mining techniques: (1) association analysis; (2) sequential pattern mining; (3) clustering. Particularly, association analysis is used to extract frequent correlations among examinations done by patients with a specific disease, sequential pattern mining allows extracting frequent patterns of medical events and clustering is used to find groups of similar patients. The discovered knowledge may enrich healthcare guidelines, improve their processes and detect anomalous patients’ behavior with respect to the medical guidelines.
1. Introduction
A healthcare system is comprised of institutions, people and resources and aims to deliver health services to meet the health demands of the population. It is, in reality, a complex fusion of human-centered activities, which increasingly depend upon Information Technology (IT) and knowledge
[1]. The digital age has rapidly increased the amount of data storing the medical history of patients, making healthcare systems overwhelmed with data
[1][2][3]. However, the advancement in technology and a few software applications/tools have helped to manage, evaluate and analyze the collected healthcare data
[4]. The analysis of such big data, which are often high-dimensional and sparse, is critical and physicians often do not have adequate tools for extracting useful information
[5].
Data mining helps in finding hidden relationships and global patterns, which exist in large databases
[6]. Among the different data mining techniques, association analysis is used to extract correlations among data in medical records, such as to discover the co-occurrence of treatments or drugs taken by patients
[7][8], to detect the diseases affecting patients depending on the value of specific attributes
[9] or to predict disease risk levels
[10].
Another frequent problem is to extract sequences of medical diagnostic events or examinations done by patients. A diagnostic examination is a medical test, which has been done by the patient for a certain pathology that is normally carried out by the medical team in accordance with the advice of a medical expert. The patterns of such biochemical variables or treatments undergone by patients with specific diseases
[11][12] are used to see if the extracted sequences are coherent with the guidelines or represent anomalies to be further investigated. These issues are addressed by the sequential pattern mining algorithms. A third problem is to discover a group of similar entities (e.g., patients) based on certain similarity measures, such as finding patients with similar gait patterns
[13], similar disease sub-type
[14] or similar examination frequencies
[15]. The data mining technique used to address this problem is clustering.
2. Knowledge Discovery from Real Healthcare Data—A Case Study
This section is devoted to the applications of the data mining techniques previously described. The knowledge discovery from a real healthcare dataset is shown in Figure 1. The knowledge discovery process is comprised of three major steps: data preparation, knowledge discovery and results evaluation. Data preparation steps are prerequisites of the knowledge discovery steps. While results evaluation step is the evaluation of the extracted knowledge. The result evaluation is carried out in two ways: (1) evaluation indices; (2) medical expert. The evaluation index is used according to the data mining technique applied and medical experts evaluated the results in accordance with medical guidelines. The subsequent sections describe the healthcare electronic data used in the study, while the other three subsections report the results of the data mining techniques applied to the dataset to derive medical knowledge. The experiments were performed on a computer with a processor 3 GHz Dual-Core Intel Core i7 and 8 GB RAM. The data mining techniques were implemented in Java programming language due to the fact that the enquired data pre-processing in the transformation of sequence database from the transactional dataset was not supported within existing tools such as WEKA, R and other such tools/application. Besides, the clustering evaluation index also needed to be implemented. However, any other tool/application can also be used in applying data mining algorithms.
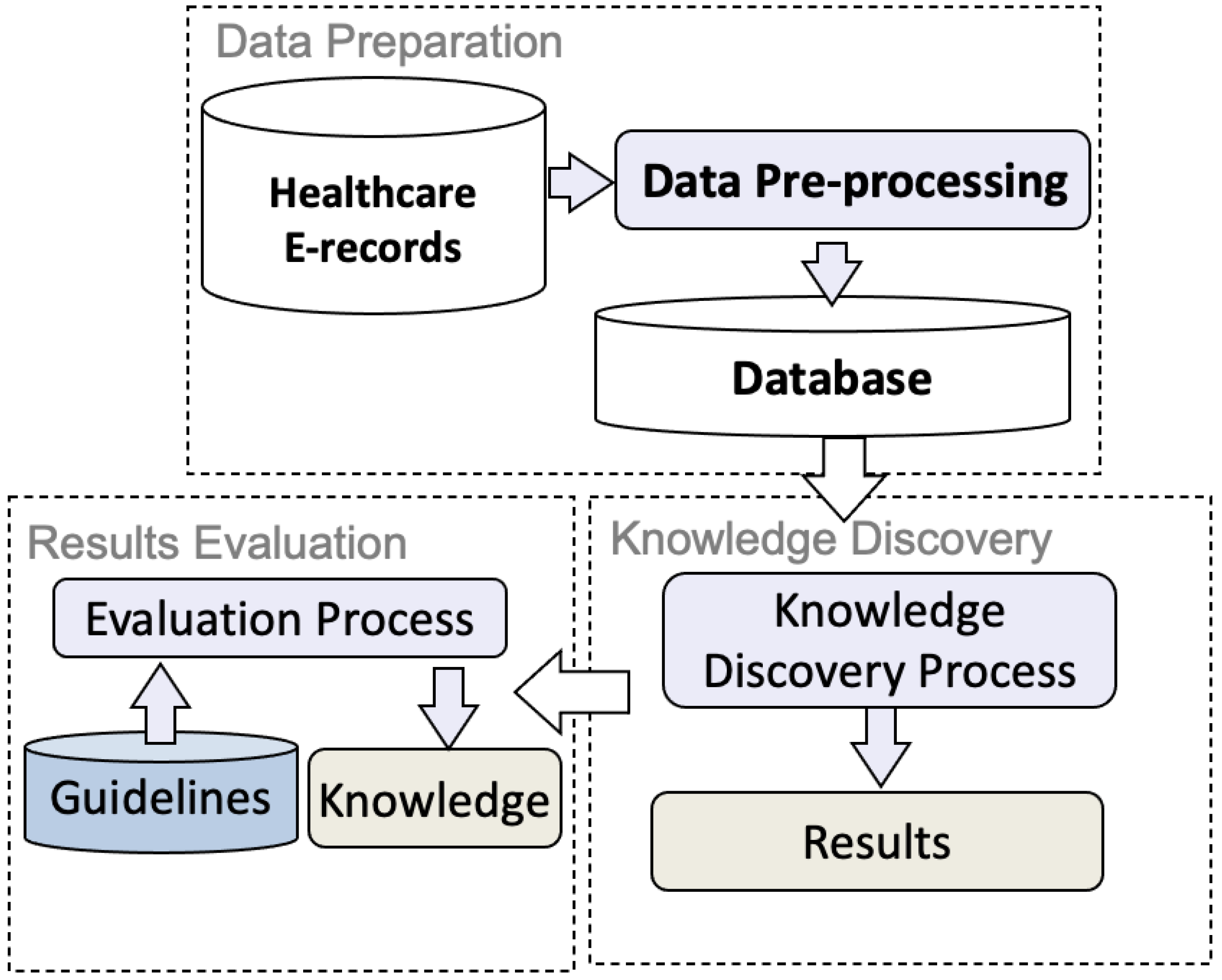
Figure 1. Discovering knowledge from healthcare data.
2.1. Healthcare Electronic Records
Healthcare systems record for each diabetic patient the examinations performed together with their date. From this data, interesting information can be extracted. For example, with the association analysis, the examinations frequently performed on the same day by patients can be detected. This analysis allows healthcare systems to control the examination process because the detected examination sets can deviate from predefined guidelines, for example, if different or additional exams have been carried out, or, if some exams are missing, medical experts may take necessary and timely measures. In addition, the sequential pattern mining can be profitably used to extract sequences of examination sets, and thus compare the medical pathways followed by patients with the theoretical ones. The pathways representing non-compliance may be determined due to patient negligence in strictly following medical treatments, incorrect procedures during record-keeping or the existence of different guidelines for specific cases. Finally, in order to group similar patients, clustering techniques can be adapted to divide the data into smaller and meaningful sets that are easier to analyze.
To perform such analyses, we collected electronic records of diabetic patients from an agency (the name is kept secret due to privacy), and we arranged them in the form of a transactional patient dataset, reporting examinations done for each patient with time, as shown in Table 1. Table 2 shows the sequence dataset corresponding to the transactional patient dataset in Table 1.
Table 1. Diabetic patient dataset.
| Patient ID |
Date |
Examination |
| 1 |
17 February 2007 |
Glucose |
| 1 |
17 February 2007 |
Capillary blood |
| 2 |
25 July 2007 |
Glucose |
| 3 |
8 January 2007 |
Urine Test |
| 3 |
15 February 2007 |
Venous blood |
| 3 |
15 February 2007 |
Glucose |
| . |
. |
. |
Table 2. Sequence dataset.
| Patient ID |
Examination Sequence |
| 1 |
{Glucose, Capillary blood} |
| 2 |
{Glucose} |
| 3 |
{Urine Test} {Venous blood, Glucose} |
| . |
. |
2.2. Association Analysis
We applied the association analysis to extract the frequent examination sets and frequent correlation between examination sets. To this aim, closed association rules
[16] were generated from the considered dataset. The Java implementation of the closed association rules was provided by Philippe Fournier-Vigier
[17], and it was properly modified to include the lift computation.
By setting a minimum support threshold of 10%, the total number of extracted item sets (i.e., closed item sets) is 2231. From the analysis of frequent closed item sets, it emerged that the extracted sets are generally in accordance with the medical guidelines, even though some anomalies were detected. The diagnostic examinations that are the base tests and routinely repeated by diabetic patients remained at the top extraction. These results, as shown in the top of Table 3, help in monitoring the concentration of sugar in the blood.
Table 3. Extracted closed examination sets and closed association rules.
| Closed Frequent Item sets |
Support |
|
|
| {Glucose} |
84.8% |
|
|
| {Venous blood} |
79.2% |
|
|
| {Capillary blood} |
75.0% |
|
|
| {Urinalysis} |
74.9% |
|
|
| {Glucose, Urinalysis} |
74.8% |
|
|
| {Glucose, Capillary blood} |
74.4% |
|
|
| {Glucose, Urinalysis, Capillary blood} |
73.9% |
|
|
| {Glucose, Venous blood} |
71.0% |
|
|
| {Glucose, Urinalysis, Capillary blood, Venous blood} |
57.4% |
|
|
| {Hemoglobin} |
46.4% |
|
|
| {Hemoglobin, Venous blood} |
43.0% |
|
|
| {Cholesterol} |
36.0% |
|
|
| {Triglycerides} |
35.7% |
|
|
| {HDL cholesterol} |
35.4% |
|
|
| {Cholesterol, Triglycerides, HDL cholesterol} |
33.7% |
|
|
| Closed Association Rules |
Support |
Confidence |
Lift |
| {Venous blood, AST, HDL cholesterol} ⇒ {ALT, Total cholesterol} |
25% |
98% |
3.66 |
| {Glucose, Venous blood, ALT, Hemoglobin} ⇒ {AST} |
26% |
98% |
3.27 |
| {Venous blood, Hemoglobin, HDL cholesterol} ⇒ {Triglycerides, Total cholesterol} |
33% |
97% |
2.72 |
| {Venous blood, Triglycerides, Total cholesterol, HDL cholesterol} ⇒ {Hemoglobin} |
33% |
96% |
2.03 |
| {Gluocse, Capillary blood, Venous blood} ⇒ {Urine Test} |
64% |
100% |
1.32 |
| {Capillary blood, Venous blood} ⇒ {Urine Test} |
65% |
99% |
1.32 |
| {Glucose, Urine Test} ⇒ {Capillary blood} |
73% |
98% |
1.32 |
| {Urine Test, Venous blood} ⇒ {Glucose} |
65% |
99% |
1.17 |
2.3. Sequential Pattern Mining
Table 4 reports the frequent examination sequences, which were detected (26,838 sequences) using BIDE algorithm
[18] with a minimum support threshold of 25%. It was analyzed that glucose level has been examined twice for 58% of patients, thrice for 32% of patients and four times for 15% of patients during a one-year period. These sequence patterns are in line with medical knowledge, although the frequency of the sequences remained less than expected. This can highlight the fact that many patients do not access the public service but prefer to perform examinations privately.
Table 4. Frequent examination sequences.
| Examination Sequences |
Frequency (%) |
| {Glucose}{Capillary blood} |
53 |
| {Venous blood}{Glucose} |
53 |
| {Venous blood}{Capillary blood} |
48 |
| {Venous blood}{Glucose, Capillary blood, Urine} |
48 |
| {Capillary blood}{Glucose}{Glucose} |
26 |
| {Glucose, Urine}{Glucose}{Glucose, Capillary blood} |
25 |
| {Venous blood} {Glucose, Capillary blood}{Venous blood} |
20 |
| {Capillary blood}{Glucose, Venous blood}{Glucose} |
20 |
| {Capillary blood, Venous blood}{Glucose}{Glucose} |
17 |
| {Haemoglobin}{Glucose}{Venous blood} |
16 |
2.4. Clustering
Instead of manually selecting a subset of data on which to perform the analysis, clustering algorithms can be exploited to automatically divide patients into groups, based on the similarity of their performed examinations. Among clustering techniques, DBSCAN demonstrated noteworthy characteristics for clustering patients, because it is less sensitive towards noisy data and outliers. In addition, prior information about a number of clusters is not required for DBSCAN. The discovery of patient clusters using DBSCAN from the healthcare diabetic dataset was implemented with the help of RapidMiner
[19].
To apply the clustering, the dataset was represented according to the Vector Space Model (VSM)
[20]. A patient is mapped as a vector in the examination space, and each vector element correlates with a different examination. The value of a vector element is the number of times an examination is done by a patient, divided by the number of patients who have undergone that examination. This representation is known as The Term Frequency (TF)—Inverse Document Frequency (IDF) in text mining
[6].
To measure the similarity between patients, we exploited the cosine similarity measure between the weighted examination frequency vectors. To select the value of the DBSCAN parameters (the minimum number of elements in each cluster (MinPts) and the maximum distance among elements of the same cluster (Eps)), we performed a set of experiments varying the parameters and measuring the average (Avg.) silhouette and standard deviation (STD) of silhouette values, as shown in Figure 2.
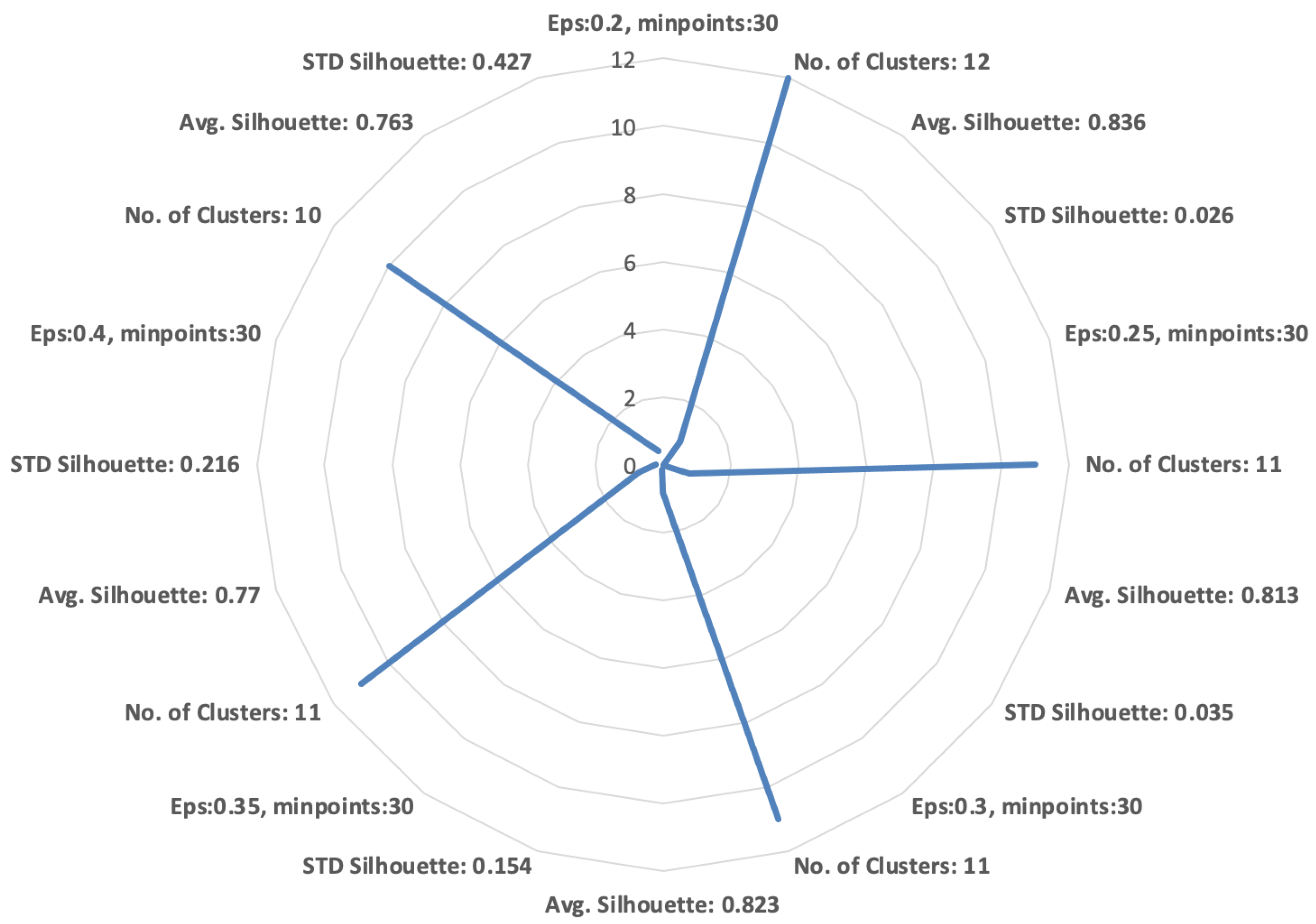 Figure 2.
Figure 2. Cluster evaluation index—silhouette.
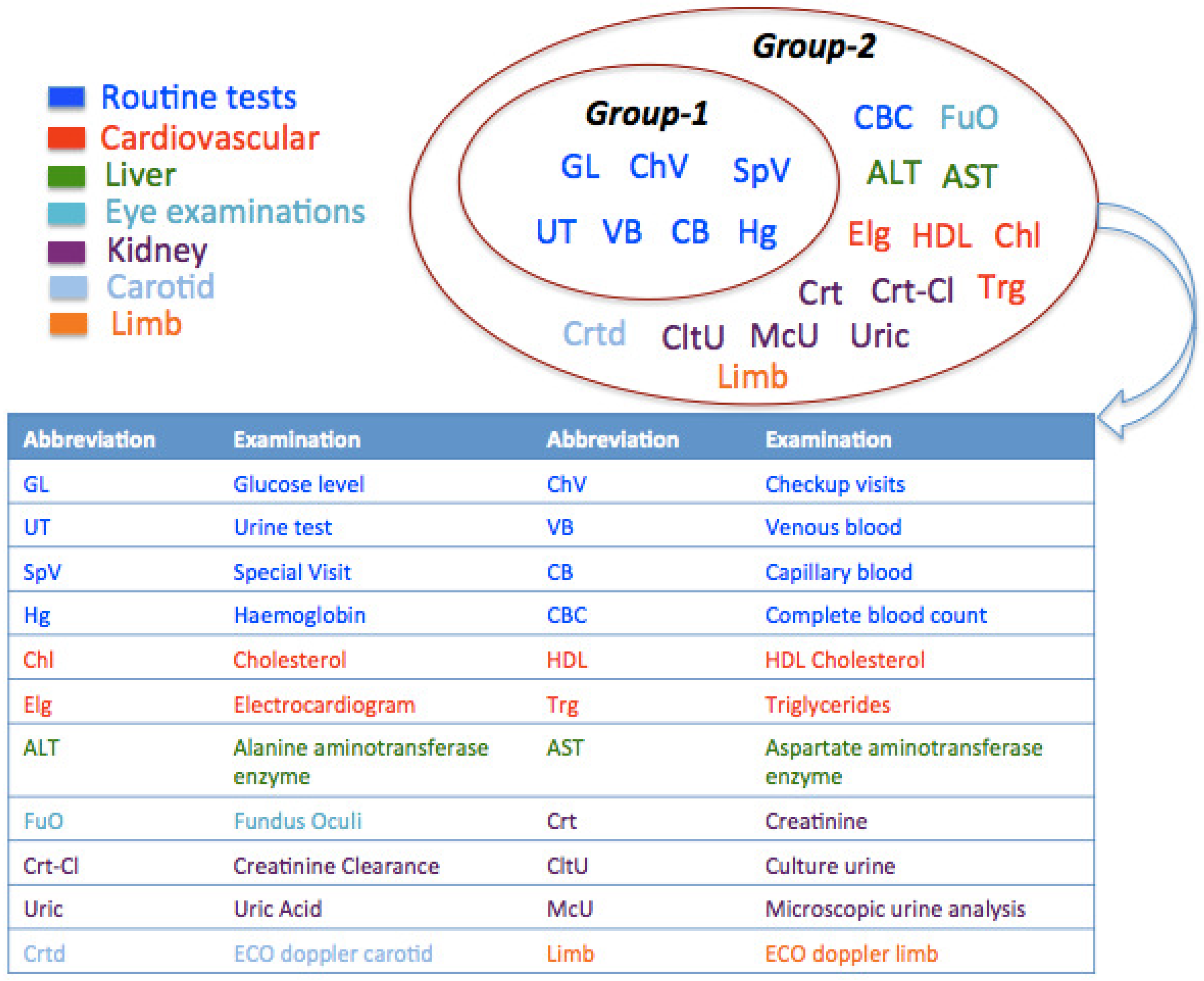
Figure 3 reports, for each group of clusters, the set of examinations done by patients in these clusters. In the following, the main characteristics of each cluster are reported.
Figure 3. Clustering results representing different subgroups of patients.
3. Conclusions and Future Work
The different data mining techniques are applied in knowledge discovery from real healthcare electronic records of diabetic patients. The experimental results show the effectiveness of the three data mining techniques (i.e., association rule mining, sequential pattern mining and clustering) exploited in this study. In particular, sequential pattern mining rebuilt the treatment procedures from the considered healthcare data. Association rule mining potentially uncovered the associations between diagnostic examinations and their interdependency on each other. Moreover, the clustering technique deeply investigated the dataset and detected several subgroups with homogeneous medical treatments present in the dataset. The detected subgroups highlighted the different complications being seen within diabetic treatments. The discovered information may be applied in existing guidelines to enrich them, and, at the same time, it may help healthcare management to utilize their resources efficiently since experimental results define the dependency of examinations and possible severe conditions of the diabetic patients. This work is only limited to diagnostic examinations and three techniques were applied. As future work, we intend to work on time series of sequential patterns, i.e., what patterns are observed with respect to time and age factors that must be considered for further analysis.


 Figure 2. Cluster evaluation index—silhouette.
Figure 2. Cluster evaluation index—silhouette.