
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Naoyuki Kataoka | + 1361 word(s) | 1361 | 2021-08-01 10:50:17 | | | |
| 2 | Vicky Zhou | Meta information modification | 1361 | 2021-08-12 06:35:06 | | | | |
| 3 | Vicky Zhou | Meta information modification | 1361 | 2021-08-12 06:35:23 | | |
Video Upload Options
Pre-mRNA splicing is an essential process for gene expression in higher eukaryotes, which requires a high order of accuracy. Mutations in splicing factors or regulatory elements in pre-mRNAs often result in many human diseases. Myelodysplastic syndrome (MDS) is a heterogeneous group of chronic myeloid neoplasms characterized by many symptoms and a high risk of progression to acute myeloid leukemia. Recent findings indicate that mutations in splicing factors represent a novel class of driver mutations in human cancers and affect about 50% of Myelodysplastic syndrome (MDS) patients. Somatic mutations in MDS patients are frequently found in genes SF3B1, SRSF2, U2AF1, and ZRSR2. Interestingly, they are involved in the recognition of 3′ splice sites and exons. It has been reported that mutations in these splicing regulators result in aberrant splicing of many genes.
1. Introduction
1.1. Splicing Signals and Splicing Reaction
Pre-mRNA splicing is a critical step for protein expression in higher eukaryotes [1]. For constitutive splicing, all exons are ligated in order without any insertions and deletions of nucleotides. The essential signals for splicing reaction mostly reside at both ends of introns ( Figure 1 ) [1]. At the 5′ end, consensus sequence of GURRGU (R stands for purine) can be found in most of introns in mammals ( Figure 1 ). This site is called 5′ splice site (5′ss). CAG consensus sequence is often discovered at the 3′ end of introns ( Figure 1 ), which is called 3′ splice site (3′ss). In addition to them, pyrimidine (Y) residue stretch precedes to 3′ splice site in order to support recognition of 3′ splice site in mammals ( Figure 1 , (Y)nNCAG). A branch point sequence (BP), at which lariat formation occurs by 2′–5′ phosphodiester bond formation with Guanine residue at 5′ splice site, resides 20–30 nucleotides upstream of the 3′ splice site ( Figure 1 ). Although the sequence for branch point in budding yeast is well-conserved as UACUA A C (underlined A is a branch point) among introns, the conserved sequence around branch point in mammals is YUN A Y (branch point is underlined, Y and N stand for pyrimidine and any nucleotide, respectively), which is more diverse ( Figure 1 ) [2]. Then, pyrimidine residue stretch also supports branch point sequence recognition ( Figure 1 ). The splicing reaction consists of two steps, the first step and the second step. In the first step reaction, cleavage at 5′ ss and formation of lariat structure in intron occur. The second step reaction includes cleavage at 3′ ss and ligation of exons to produce mRNA. Both steps require ATP and divalent cations in vitro. As a divalent cation, magnesium is most efficient in in vitro splicing reaction.
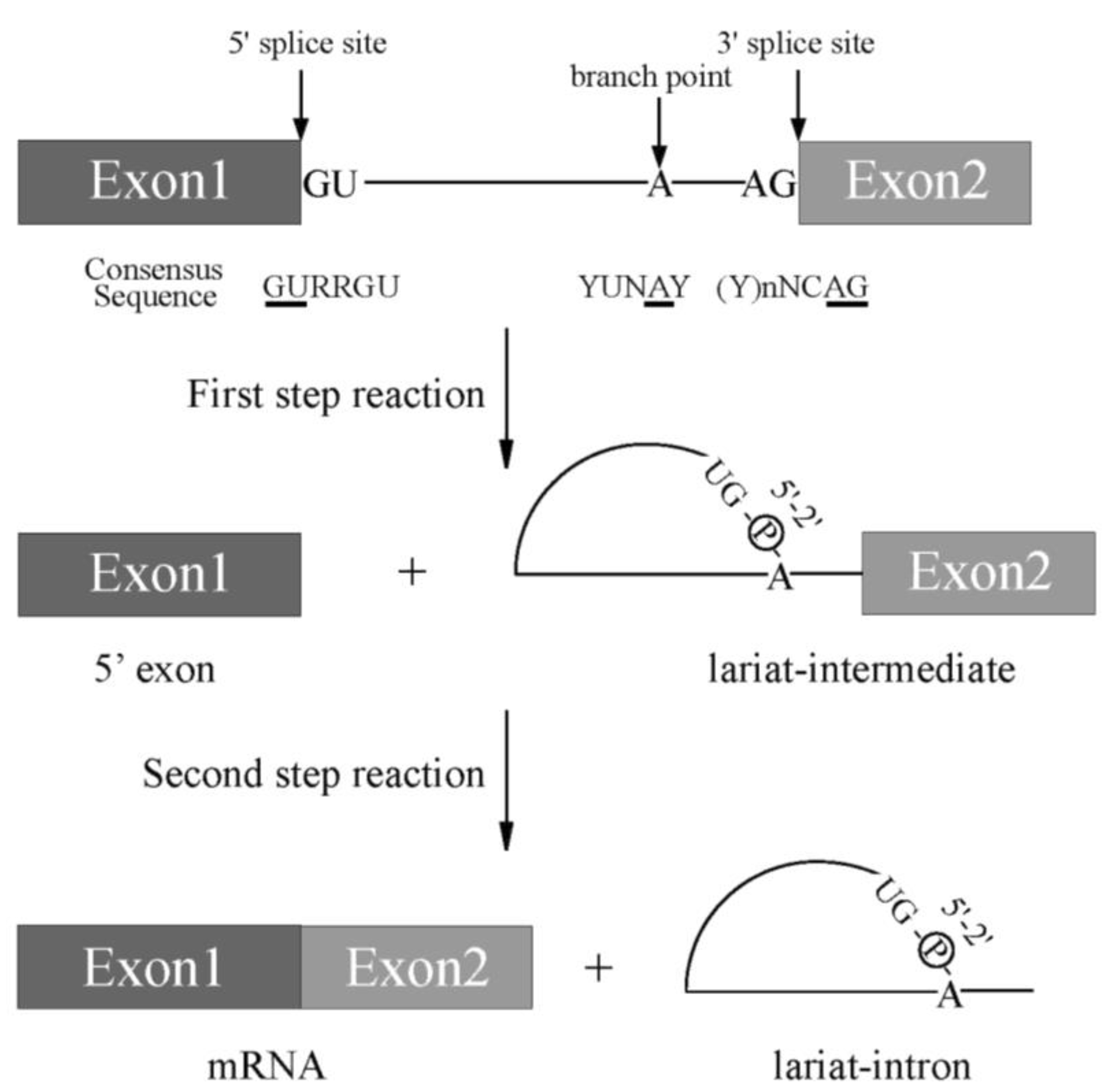 Figure 1. A scheme for splicing reaction with two steps. Schematic representation of sequences required for splicing reaction. Boxes show exons, and lines between boxes represent introns. Conserved sequence elements of metazoan pre-mRNAs. R and Y stand for purine and pyrimidine residues, respectively. N indicates any nucleotides. Conserved 5′ and 3′ splice sites, and Adenosine residue used for branch nucleotide are underlined.
Figure 1. A scheme for splicing reaction with two steps. Schematic representation of sequences required for splicing reaction. Boxes show exons, and lines between boxes represent introns. Conserved sequence elements of metazoan pre-mRNAs. R and Y stand for purine and pyrimidine residues, respectively. N indicates any nucleotides. Conserved 5′ and 3′ splice sites, and Adenosine residue used for branch nucleotide are underlined.
1.2. Spliceosome Formation with Splicing Factors
Splicing reaction takes place in a large ribonucleoprotein complex, termed the spliceosome [1]. The assembly of the spliceosome on pre-mRNA occurs with stepwise association of the uridine (U)-rich small nuclear RNPs (snRNPs) (U1, U2, U4, U5, and U6) (Figure 2) and a multitude of non-snRNP splicing factors [1]. U snRNPs consist of short RNA, Sm proteins, and a few specific proteins of each U snRNPs. As the first step of the reaction, 5′ splice site is recognized by U1 snRNP by RNA-RNA pairing. U2 snRNP then come to associate with a branch point sequence with the help of U2 snRNP auxiliary factor (U2AF) complex that consists of U2AF1 and U2AF2 heterodimer. The RNA component of U2 snRNP also hybridizes with pre-mRNA to recognize BP. The tri-snRNP, U4/U5/U6, then becomes joining to the spliceosome. Two U snRNPs, U4 and U6, form a heterodimer by pairing their RNA components. The spliceosome is activated by removal of U1 and U4 snRNPs to remodel pre-mRNA-U snRNPs and U snRNP-U snRNP interactions, and the first step reaction, the cleavage at the 5′ splice site and formation of a lariat structure, takes place. Then, the cleavage at the 3′ splice site and ligation of two exons occur as the second step reaction. Several lines of evidence suggested that U6 snRNA has catalytic activity for the splicing reaction.
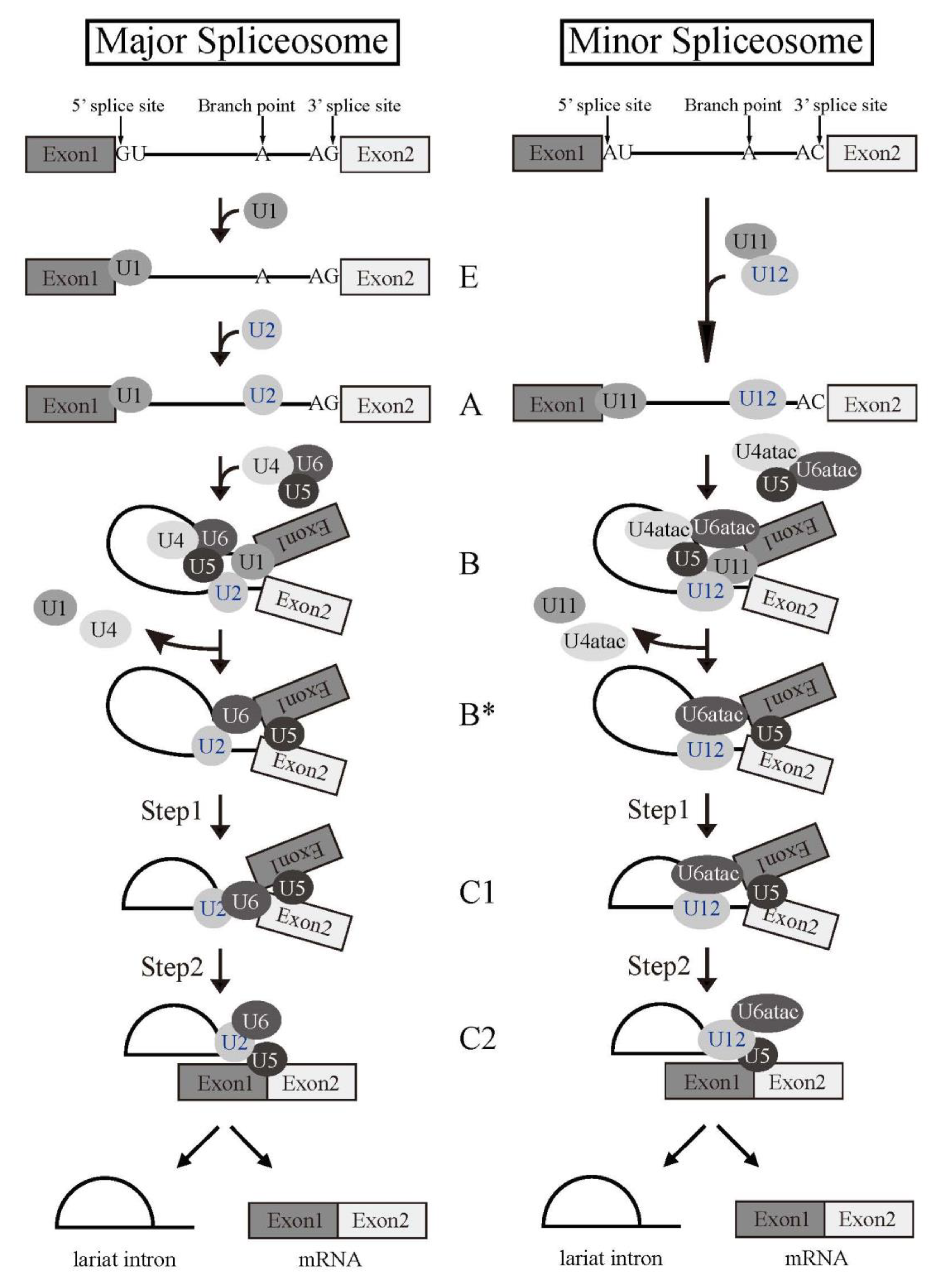 Figure 2. Splicing reaction and formation of spliceosome. Schematic representations of major and minor spliceosome formations. Both splicing reactions take place stepwise in a spliceosome. Spliceosomal Uridine-rich small nuclear ribonucleoproteins (U snRNPs) are indicated with their names. The name of each spliceosome intermediate complex is shown in the middle.
Figure 2. Splicing reaction and formation of spliceosome. Schematic representations of major and minor spliceosome formations. Both splicing reactions take place stepwise in a spliceosome. Spliceosomal Uridine-rich small nuclear ribonucleoproteins (U snRNPs) are indicated with their names. The name of each spliceosome intermediate complex is shown in the middle.
2. Splicing Factors Mutated in Myelodysplastic Syndrome
SRSF2 is involved in exon recognition through ESE binding. Other factors, SF3B1, U2Af1 and ZRSR2 are also involved in branch point and 3' splice site recognition. Taken together, it is likely that a splicing mode called exon recognition ( Figure 3 ) [3] participates in aberrant splicing in MDS. In higher eukaryotes, the average length of introns is much longer than that of lower eukaryotes. In fact, the average length of human introns is 5849 nucleotides, while that of nematodes is 335 nucleotides [4]. In contrast, the average length of internal exons, which is no longer than 300 nucleotides, does not differ between vertebrates and lower eukaryotes. Therefore, exon recognition is likely a major mode for splicing in vertebrates whose intron size is large, while intron recognition is dominant in lower eukaryotes in which introns are relatively short ( Figure 3 ). The facts that 5′ splice site mutations result in skipping of adjacent exons and cause human diseases also support the exon recognition model. For 5′ terminal exons, it is assumed that the cap structure serves as a substitute of the 3′ splice site. The cap structure is recognized by a nuclear cap binding protein complex that consists of NCBP1/2 proteins in the nucleus [5][6][7]. As NCBP1 was demonstrated to associate with U2 snRNP [7] , it is possible that NCBP1-U2 snRNP interacts with U1 snRNP at the 5′ splice site to define the first exon. As for 3′ terminal exons, poly(A) addition signal and poly(A) addition machinery are assumed to serve as interactors with U2 snRNP on the branch point in the last intron. As supporting evidence, mutation of the 3′ splice sites inhibits the polyadenylation cleavage reaction in vitro [8]. In the exon recognition model, definition of the 3′ splice site region highly likely takes place first, and this step is critical for exon recognition. Although many excellent works have been performed and provide information for the mechanism of vertebrate exon recognition, it remains unclear whether different factors/mechanism are involved in different exons. It is expected that precise analyses of the aberrant splicing mechanism in MDS with mutant splicing factors also contribute to uncovering the regulation of alternative splicing through exon recognition in vertebrates.
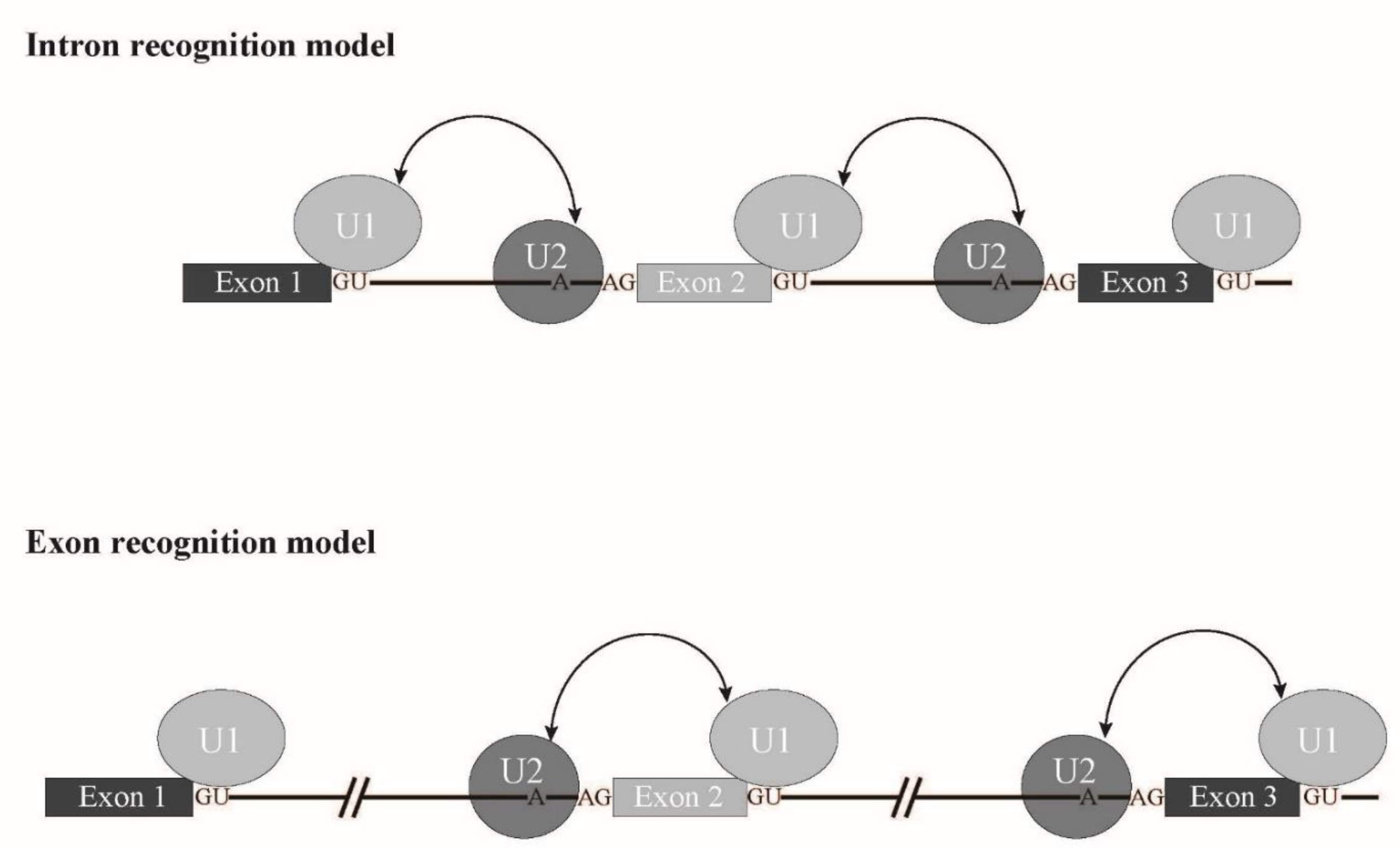 Figure 3. Schematic representation of exon recognition and intron recognition models during splicing. In lower eukaryotes, whose intron size is small, intron recognition is a dominant mode for splicing (upper panel). Introns are recognized by crosstalk between U1 snRNP and U2 snRNP that bind to the 5′ splice site and a branch point, respectively. On the other hand, exon recognition is major for splicing in vertebrates, in which introns are long. In this type, exons are recognized via interaction over exons between U2 snRNP and U1 snRNP that bind to a branch point and the 5′ splice site, respectively (lower panel).
Figure 3. Schematic representation of exon recognition and intron recognition models during splicing. In lower eukaryotes, whose intron size is small, intron recognition is a dominant mode for splicing (upper panel). Introns are recognized by crosstalk between U1 snRNP and U2 snRNP that bind to the 5′ splice site and a branch point, respectively. On the other hand, exon recognition is major for splicing in vertebrates, in which introns are long. In this type, exons are recognized via interaction over exons between U2 snRNP and U1 snRNP that bind to a branch point and the 5′ splice site, respectively (lower panel).
Although some aberrant splicing patterns in dysregulated genes have been identified to be involved in MDS onset as described above, it is still under investigation how different mutations in different splicing factors cause different MDS phenotypes. To date, there seems to be no common gene(s) whose aberrant splicing is responsible for MDS onset caused by mutations in four main splicing factors SF3B1, SRSF2, U2AF1, and ZRSR2. It is assumed that hot spot mutations among them in SF3B1, SRSF2, and U2AF1 do not cause reduction of the encoded proteins, whereas mutations in ZRSR2 reduce functional protein amount. Splicing pattern analyses implicate that common pathways affected by mutations of those factors are epigenetics and signal transduction pathways. These points have to be addressed in future analyses. The approaches from mechanistic analyses of aberrant splicing caused by mutated splicing factors should shed light on research for therapies of MDS by identifying drug targets.
References
- Wahl, M.C.; Will, C.L.; Lührmann, R. The Spliceosome: Design Principles of a Dynamic RNP Machine. Cell 2009, 136, 701–718.
- Gao, K.; Masuda, A.; Matsuura, T.; Ohno, K. Human branch point consensus sequence is yUnAy. Nucleic Acids Res. 2008, 36, 2257–2267.
- Berget, S.M. Exon Recognition in Vertebrate Splicing. J. Biol. Chem. 1995, 270, 2411–2414.
- Zhu, L.; Zhang, Y.; Zhang, W.; Yang, S.; Chen, J.-Q.; Tian, D. Patterns of exon-intron architecture variation of genes in eukaryotic genomes. BMC Genom. 2009, 10, 47.
- Izaurralde, E.; Lewis, J.; McGuigan, C.; Jankowska, M.; Darzynkiewicz, E.; Mattaj, I.W. A nuclear cap binding protein complex involved in pre-mRNA splicing. Cell 1994, 78, 657–668.
- Kataoka, N.; Ohno, M.; Moda, I.; Shimura, Y. Identification of the factors that interact with NCBP, an 80 kDa nuclear cap binding protein. Nucleic Acids Res. 1995, 23, 3638–3641.
- Ohno, M.; Kataoka, N.; Shimura, Y. A nuclear cap binding protein from HeLa cells. Nucleic Acids Res. 1990, 18, 6989–6995.
- Niwa, M.; Rose, S.D.; Berget, S.M. In vitro polyadenylation is stimulated by the presence of an upstream intron. Genes Dev. 1990, 4, 1552–1559.




