
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | John Hancock | + 2232 word(s) | 2232 | 2021-07-27 08:11:34 | | | |
| 2 | Lindsay Dong | Meta information modification | 2232 | 2021-08-04 05:14:50 | | |
Video Upload Options
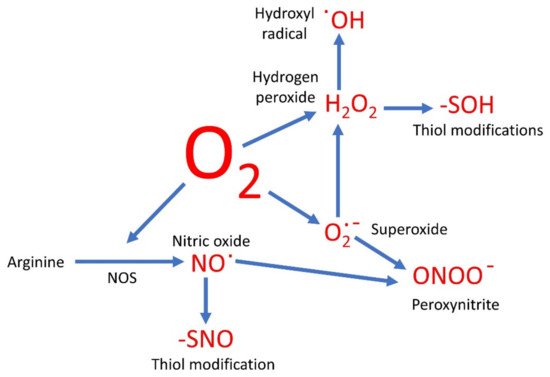
Oxygen-based compounds are an instrumental part of the group of small, relatively reactive molecules which control cellular activities. Traditionally such molecules have been referred to as the reactive oxygen species (ROS) and include hydrogen peroxide (H2O2), superoxide (O2∙−), and hydroxyl radicals (∙OH). However, several other reactive signaling molecules also contain oxygen, although referred to as reactive nitrogen species (RNS). These include nitric oxide (NO) and peroxynitrite (ONOO−), and therefore could be grouped together with the ROS as oxygen-based compounds.
1. Introduction
2. Signaling by ROS
2.1. Superoxide and Its Role
2.2. Hydrogen Peroxide as a Signal
The sequential oxidation of molecule oxygen produces O2∙−, then H2O2, and finally the hydroxyl radical (∙OH) before the four election reduction results in water. Therefore, once the O2∙− anion is formed, a cascade of further products is likely. As discussed below, there are side reactions likely here too. For example, hypochlorous acid can be produced in the presence of the enzyme myeloperoxidase [23].
H2O2 has been the focus of ROS signaling [7][24][25]. One of the ways in which H2O2 is known to alter cell function is by the oxidation of thiol groups in proteins [25], and such modifications can be analyzed by proteomic techniques [26][27]. The -SH group is converted to the sulfenic acid group, -SOH. This is in many ways akin to phosphorylation, and like phosphorylation, the formation of the -SOH group is likely to force a conformational change on the proteins and thus alter its activity. This is not necessarily activation. In tyrosine phosphatase, the interaction with H2O2 leads to the formation of a sulfenyl-amide intermediate and inhibition of the enzyme [28]. This means in the cell that the levels of tyrosine phosphorylation are likely to increase, with the concomitant effects that leads to.
2.3. Hydroxyl Radicals Can Be Signals Too
3. Signaling by RNS
3.1. Nitric Oxide and Working with Other Oxygen-Based Molecules
NO has been found to be involved in the mediation of a wide range of biological functions, from controlled blood flow in humans [45], to controlling stomatal apertures in plants [46]. In animals, the main source is NOS. In humans, there are three isoforms of this enzyme: eNOS, iNOS, and nNOS [47]. However, the existence of such an enzyme in plants has been hotly contested and it is unlikely to exist, at least in the form that would be easily recognizable [48]. It is more likely that in plants the main source of NO is the enzyme nitrate reductase (NR) [49], although as mentioned above there are other sources of NO in biological systems.
A universal mechanism of NO signaling is the modification of protein thiol groups, in what has been dubbed S-nitrosylation [50]. However, this terminology is technically incorrect, and this modification should preferably be called S-nitrosation [51]. Either way, this is the formation of the -SNO group, and like the formation of -SOH by H2O2, this formation of -SNO causes a conformational change on the protein and therefore a modulation of its activity or function. As this is a reversable reaction it is again akin to phosphorylation. However, the thiols are also able to be oxidized, as discussed, so there is likely to be competition for the thiol between the oxidation by ROS and nitrosation by NO. Furthermore, the same thiols may be under attack by H2S, in S-sulfhydration [52][53], as well as being able to be glutathionylated [54]. Which thiol modification actually results depends on the environment of the thiol and the relative concentrations of the molecule trying to attack it. As many of these reactions are reversible, the whole system is likely to be very dynamic, allowing different modifications happening with time and in different locations.
Proteins can also be nitrated on tyrosine. Therefore, NO can mediate the modification of polypeptides in more than one manner [55], and such changes are not mutually exclusive.
Last, NO can partake in some direct reactions with other important redox molecules. One of the most significant is the generation of S-nitrosoglutathione (GSNO). This not only removes glutathione from its important role as a redox mediator, especially in ROS metabolism, as discussed above [56], but it also creates a new signaling molecule. It has been suggested that GSNO is a buffer for NO, GSNO formation being reversed by S-nitrosoglutathione reductase (GSNOR) [57], but it may also be able to be moved around an organism in the vasculature [58], so allowing long-range NO signaling. NO can also react with H2S in the formation of nitrosothiol, which can act as a signal as well [59].
3.2. Peroxynitrite, as a Signal
4. The Signaling of Carbon Monoxide
Many of the effects of CO are mediated through the action of heme oxygenase [65][66]. This enzyme degrades heme to produce biliverdin, ferrous ions, and CO.
In a similar manner to ROS and RNS, CO is inherently toxic [67]. It can inhibit the activity of Complex IV of the mitochondrial ETC, for example. Even so, as it can inherently interact with metal containing proteins, it is known to modulate the activities of several enzymes, and this can lead to changes to the accumulation of ROS and NO. It can also alter cGMP levels, an instrumental intracellular signaling molecule. Furthermore, CO effects can be mediated by MAPK pathways and by changes in the activity of ion channels [68]. One of the mechanisms of action of H2 is thought to be mediated by heme oxygenase [69], which would then impinge on CO signaling.
5. Conclusions
References
- Beckmann, R.; Flohé, L. The pathogenic role of superoxide radicals in inflammation: Efficacy of exogenous superoxide dismutase. Bull. Eur. Physiopathol. Respir. 1981, 17, 275–286.
- Hohn, D.C.; Lehrer, R.I. NADPH oxidase deficiency in X-linked chronic granulomatous disease. J. Clin. Investig. 1975, 55, 707–713.
- Arnold, D.E.; Heimall, J.R. A review of Chronic Granulomatous Disease. Adv. Ther. 2017, 34, 2543–2557.
- Palmer, R.M.; Ferrige, A.G.; Moncada, S. Nitric oxide release accounts for the biological activity of endothelium-derived relaxing factor. Nature 1987, 327, 524–526.
- Anson, M.L.; Mirsky, A.E. On the combination of nitric oxide with haemoglobin. J. Physiol. 1925, 60, 100–102.
- Brandes, R.P.; Weissmann, N.; Schröder, K. Nox family NADPH oxidases: Molecular mechanisms of activation. Free Radic. Biol. Med. 2014, 76, 208–226.
- Sies, H. Hydrogen peroxide as a central redox signaling molecule in physiological oxidative stress: Oxidative eustress. Redox Biol. 2017, 11, 613–619.
- El-Bahr, S.M. Biochemistry of free radicals and oxidative stress. Sci. Int. 2013, 1, 111–117.
- Speckmann, B.; Steinbrenner, H.; Grune, T.; Klotz, L.O. Peroxynitrite: From interception to signaling. Arch. Biochem. Biophys. 2016, 595, 153–160.
- Morse, D.; Sethi, J.; Choi, A.M. Carbon monoxide-dependent signaling. Crit. Care Med. 2002, 30 Suppl. 1, S12–S17.
- Olas, B. Hydrogen sulfide in signaling pathways. Clin. Chim. Acta. 2015, 439, 212–218.
- Zeng, J.; Zhang, M.; Sun, X. Molecular hydrogen is involved in phytohormone signaling and stress responses in plants. PLoS ONE 2013, 8, e71038.
- Hancock, J.T.; Whiteman, M. Hydrogen sulfide and cell signaling: Team player or referee? Plant Physiol. Biochem. 2014, 78, 37–42.
- Corpas, F.J.; González-Gordo, S.; Cañas, A.; Palma, J.M. Nitric oxide and hydrogen sulfide in plants: Which comes first? J. Exp. Bot. 2019, 70, 4391–4404.
- Monteiro, H.P.; Rodrigues, E.G.; Amorim Reis, A.K.C.; Longo, L.S., Jr.; Ogata, F.T.; Moretti, A.I.S.; da Costa, P.E.; Teodoro, A.C.S.; Toledo, M.S.; Stern, A. Nitric oxide and interactions with reactive oxygen species in the development of melanoma, breast, and colon cancer: A redox signaling perspective. Nitric Oxide 2019, 89, 1–13.
- Ma, J.; Zhou, H.; Yan, S.; Song, W. Kinetics studies and mechanistic considerations on the reactions of superoxide radical ions with dissolved organic matter. Water Res. 2019, 149, 56–64.
- Winterbourn, C.C.; Kettle, A.J.; Hampton, M.B. Reactive oxygen species and neutrophil function. Annu. Rev. Biochem. 2016, 85, 765–792.
- Wang, Y.; Branicky, R.; Noë, A.; Hekimi, S. Superoxide dismutases: Dual roles in controlling ROS damage and regulating ROS signaling. J. Cell Biol. 2018, 217, 1915–1928.
- Wegerich, F.; Turano, P.; Allegrozzi, M.; Möhwald, H.; Lisdat, F. Cytochrome C mutants for superoxide biosensors. Anal. Chem. 2009, 81, 2976–2984.
- Schröder, K. NADPH oxidases: Current aspects and tools. Redox Biol. 2020.
- Miller, G.; Schlauch, K.; Tam, R.; Cortes, D.; Torres, M.A.; Shulaev, V.; Dangl, J.L.; Mittler, R. The plant NADPH oxidase RBOHD mediates rapid systemic signaling in response to diverse stimuli. Sci. Signal. 2009, 2, ra45.
- Ewald, C.Y. Redox signaling of NADPH oxidases regulates oxidative stress responses, immunity and aging. Antioxidants 2018, 7, 130.
- Aratani, Y. Myeloperoxidase: Its role for host defense, inflammation, and neutrophil function. Arch. Biochem. Biophys. 2018, 640, 47–52.
- Marinho, H.S.; Real, C.; Cyrne, L.; Soares, H.; Antunes, F. Hydrogen peroxide sensing, signaling and regulation of transcription factors. Redox Biol. 2014, 2, 535–562.
- Winterbourn, C.C. Hydrogen peroxide reactivity and specificity in thiol-based cell signalling. Biochem. Soc. Trans. 2020, 48, 745–754.
- Baty, J.W.; Hampton, M.B.; Winterbourn, C.C. Proteomic detection of hydrogen peroxide-sensitive thiol proteins in Jurkat cells. Biochem. J. 2005, 389, 785–795.
- Hawkins, C.L.; Davies, M.J. Detection, identification, and quantification of oxidative protein modifications. J. Biol. Chem. 2019, 294, 19683–19708.
- Salmeen, A.; Andersen, J.N.; Myers, M.P.; Meng, T.C.; Hinks, J.A.; Tonks, N.K.; Barford, D. Redox regulation of protein tyrosine phosphatase 1B involves a sulphenyl-amide intermediate. Nature 2003, 423, 769–773.
- Fischbacher, A.; von Sonntag, C.; Schmidt, T.C. Hydroxyl radical yields in the Fenton process under various pH, ligand concentrations and hydrogen peroxide/Fe(II) ratios. Chemosphere 2017, 182, 738–744.
- Kehrer, J.P. The Haber-Weiss reaction and mechanisms of toxicity. Toxicology 2000, 149, 43–50.
- Fong, K.L.; McCay, P.B.; Poyer, J.L. Evidence for superoxide-dependent reduction of Fe3+ and its role in enzyme-generated hydroxyl radical formation. Chem. Biol. Interact. 1976, 15, 77–89.
- Halliwell, B. Superoxide-dependent formation of hydroxyl radicals in the presence of iron chelates: Is it a mechanism for hydroxyl radical production in biochemical systems? FEBS Lett. 1978, 92, 321–326.
- Richards, S.L.; Wilkins, K.A.; Swarbreck, S.M.; Anderson, A.A.; Habib, N.; Smith, A.G.; McAinsh, M.; Davies, J.M. The hydroxyl radical in plants: From seed to seed. J. Exp. Bot. 2015, 66, 37–46.
- Pottosin, I.; Zepeda-Jazo, I.; Bose, J.; Shabala, S. An anion conductance, the essential component of the hydroxyl-radical-induced ion current in plant roots. Int. J. Mol. Sci. 2018, 19, 897.
- Demidchik, V.; Cuin, T.A.; Svistunenko, D.; Smith, S.J.; Miller, A.J.; Shabala, S.; Sokolik, A.; Yurin, V. Arabidopsis root K+-efflux conductance activated by hydroxyl radicals: Single-channel properties, genetic basis and involvement in stress-induced cell death. J. Cell Sci. 2010, 123, 1468–1479.
- Lu, W.J.; Lin, K.H.; Hsu, M.J.; Chou, D.S.; Hsiao, G.; Sheu, J.R. Suppression of NF-κB signaling by andrographolide with a novel mechanism in human platelets: Regulatory roles of the p38 MAPK-hydroxyl radical-ERK2 cascade. Biochem. Pharmacol. 2012, 84, 914–924.
- Chen, W.; Ding, S.; Wu, J.; Shi, G.; Zhu, A. In situ detection of hydroxyl radicals in mitochondrial oxidative stress with a nanopipette electrode. Chem. Commun. 2020, 56, 13225–13228.
- Sakai, T.; Imai, J.; Ito, T.; Takagaki, H.; Ui, M.; Hatta, S. The novel antioxidant TA293 reveals the role of cytoplasmic hydroxyl radicals in oxidative stress-induced senescence and inflammation. Biochem. Biophys. Res. Commun. 2017, 482, 1183–1189.
- Xu, G.; Chance, M.R. Hydroxyl radical-mediated modification of proteins as probes for structural proteomics. Chem. Rev. 2007, 107, 3514–3543.
- Tejero, I.; Gonzalez-Lafont, A.; Lluch, J.M.; Eriksson, L.A. Theoretical modeling of hydroxyl-radical-induced lipid peroxidation reactions. J. Phys. Chem. B 2007, 111, 5684–5693.
- Gilbert, B.C.; King, D.M.; Thomas, C.B. The oxidation of some polysaccharides by the hydroxyl radical: An e.s.r. investigation. Carbohydr. Res. 1984, 125, 217–235.
- Kumar, A.; Prasad, A.; Sedlářová, M.; Pospíšil, P. Data on detection of singlet oxygen, hydroxyl radical and organic radical in Arabidopsis thaliana. Data Brief. 2018, 21, 2246–2252.
- Shen, B.; Jensen, R.G.; Bohnert, H.J. Mannitol protects against oxidation by hydroxyl radicals. Plant Physiol. 1997, 115, 527–532.
- Lipinski, B. Hydroxyl radical and its scavengers in health and disease. Oxid. Med. Cell Longev. 2011, 2011, 809696.
- Toda, N.; Ayajiki, K.; Okamura, T. Cerebral blood flow regulation by nitric oxide: Recent advances. Pharmacol. Rev. 2009, 61, 62–97.
- Sun, L.R.; Yue, C.M.; Hao, F.S. Update on roles of nitric oxide in regulating stomatal closure. Plant Signal. Behav. 2019, 14, 1649569.
- Stuehr, D.J.; Vasquez-Vivar, J. Nitric oxide synthases-from genes to function. Nitric Oxide 2017, 63, 29.
- Astier, J.; Gross, I.; Durner, J. Nitric oxide production in plants: An update. J. Exp. Bot. 2018, 69, 3401–3411.
- Chamizo-Ampudia, A.; Sanz-Luque, E.; Llamas, A.; Galvan, A.; Fernandez, E. Nitrate reductase regulates plant nitric oxide homeostasis. Trends Plant Sci. 2017, 22, 163–174.
- Feng, J.; Chen, L.; Zuo, J. Protein S-nitrosylation in plants: Current progresses and challenges. J. Integr. Plant Biol. 2019, 61, 1206–1223.
- Gupta, K.J.; Hancock, J.T.; Petrivalsky, M.; Kolbert, Z.; Lindermayr, C.; Durner, J.; Barroso, J.B.; Palma, J.M.; Brouquisse, R.; Wendehenne, D.; et al. Recommendations on terminology and experimental best practice associated with plant nitric oxide research. New Phytol. 2020, 225, 1828–1834.
- Mustafa, A.K.; Gadalla, M.M.; Sen, N.; Kim, S.; Mu, W.; Gazi, S.K.; Barrow, R.K.; Yang, G.; Wang, R.; Snyder, S.H. H2S signals through protein S-sulfhydration. Sci. Signal. 2009, 2, ra72.
- Aroca, Á.; Serna, A.; Gotor, C.; Romero, L.C. S-sulfhydration: A cysteine posttranslational modification in plant systems. Plant Physiol. 2015, 168, 334–342.
- Checconi, P.; Limongi, D.; Baldelli, S.; Ciriolo, M.R.; Nencioni, L.; Palamara, A.T. Role of glutathionylation in infection and inflammation. Nutrients 2019, 11, 1952.
- Kolbert, Z.; Feigl, G.; Bordé, Á.; Molnár, Á.; Erdei, L. Protein tyrosine nitration in plants: Present knowledge, computational prediction and future perspectives. Plant Physiol. Biochem. 2017, 113, 56–63.
- Schafer, F.Q.; Buettner, G.R. Redox environment of the cell as viewed through the redox state of the glutathione disulfide/glutathione couple. Free Radic. Biol. Med. 2001, 30, 1191–1212.
- Ventimiglia, L.; Mutus, B. The physiological implications of S-nitrosoglutathione reductase (GSNOR) activity mediating NO signalling in plant root structures. Antioxidants 2020, 9, 1206.
- Rassaf, T.; Preik, M.; Kleinbongard, P.; Lauer, T.; Heiss, C.; Strauer, B.E.; Feelisch, M.; Kelm, M. Evidence for in vivo transport of bioactive nitric oxide in human plasma. J. Clin. Investig. 2002, 109, 1241–1248.
- Whiteman, M.; Li, L.; Kostetski, I.; Chu, S.H.; Siau, J.L.; Bhatia, M.; Moore, P.K. Evidence for the formation of a novel nitrosothiol from the gaseous mediators nitric oxide and hydrogen sulphide. Biochem. Biophys. Res. Commun. 2006, 343, 303–310.
- Vandelle, E.; Delledonne, M. Peroxynitrite formation and function in plants. Plant Sci. 2011, 181, 534–539.
- Alvarez, B.; Radi, R. Peroxynitrite reactivity with amino acids and proteins. Amino Acids 2003, 25, 295–311.
- Staszek, P.; Gniazdowska, A. Peroxynitrite induced signaling pathways in plant response to non-proteinogenic amino acids. Planta 2020, 252, 5.
- Klotz, L.O.; Schroeder, P.; Sies, H. Peroxynitrite signaling: Receptor tyrosine kinases and activation of stress-responsive pathways. Free Radic. Biol. Med. 2002, 33, 737–743.
- Shacka, J.J.; Sahawneh, M.A.; Gonzalez, J.D.; Ye, Y.Z.; D’alessandro, T.L.; Estevez, A.G. Two distinct signaling pathways regulate peroxynitrite-induced apoptosis in PC12 cells. Cell Death Differ. 2006, 13, 1506–1514.
- Wilks, A. Heme oxygenase: Evolution, structure, and mechanism. Antioxid. Redox Signal. 2002, 4, 603–614.
- Wegiel, B.; Nemeth, Z.; Correa-Costa, M.; Bulmer, A.C.; Otterbein, L.E. Heme oxygenase-1: A metabolic nike. Antioxid. Redox Signal. 2014, 20, 1709–1722.
- Goldbaum, L.R.; Ramirez, R.G.; Absalon, K.B. What is the mechanism of carbon monoxide toxicity? Aviat. Space Environ. Med. 1975, 46, 1289–1291.
- Peers, C.; Boyle, J.P.; Scragg, J.L.; Dallas, M.L.; Al-Owais, M.M.; Hettiarachichi, N.T.; Elies, J.; Johnson, E.; Gamper, N.; Steele, D.S. Diverse mechanisms underlying the regulation of ion channels by carbon monoxide. Br. J. Pharmacol. 2015, 172, 1546–1556.
- Lin, Y.; Zhang, W.; Qi, F.; Cui, W.; Xie, Y.; Shen, W. Hydrogen-rich water regulates cucumber adventitious root development in a heme oxygenase-1/carbon monoxide-dependent manner. J. Plant Physiol. 2014, 171, 1–8.
- Hancock, J.T. Harnessing evolutionary toxins for signaling: Reactive oxygen species, nitric oxide and hydrogen sulfide in plant cell regulation. Front. Plant Sci. 2017, 8, 189.
- Hancock, J.T. Considerations of the importance of redox state for reactive nitrogen species action. J. Exp. Bot. 2019, 70, 4323–4331.
- Hillion, M.; Antelmann, H. Thiol-based redox switches in prokaryotes. Biol. Chem. 2015, 396, 415–444.
- Hildebrandt, T.; Knuesting, J.; Berndt, C.; Morgan, B.; Scheibe, R. Cytosolic thiol switches regulating basic cellular functions: GAPDH as an information hub? Biol. Chem. 2015, 396, 523–537.




