
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Stanislav Naryzhny | + 4179 word(s) | 4179 | 2021-07-09 11:37:12 | | | |
| 2 | Ron Wang | -1 word(s) | 4178 | 2021-08-02 08:03:56 | | | | |
| 3 | Ron Wang | -1 word(s) | 4178 | 2021-08-02 08:05:35 | | |
Video Upload Options
Haptoglobin (Hp) is a blood plasma glycoprotein that plays a critical role in tissue protection and the prevention of oxidative damage. Haptoglobin is an acute-phase protein, its concentration in plasma changes in pathology, and the test for its concentration is part of normal clinical practice. Haptoglobin is a conservative protein and is the subject of research as a potential biomarker of many diseases, including malignant neoplasms. The Human Hp gene is polymorphic and controls the synthesis of three major phenotypes—homozygous Hp1-1 and Hp2-2, and heterozygous Hp2-1, determined by a combination of allelic variants that are inherited. Numerous studies indicate that the phenotype of haptoglobin can be used to judge the individual’s predisposition to various diseases. In addition, Hp undergoes various post-translational modifications (PTMs). Glioblastoma multiform (GBM) is the most malignant primary brain tumor. In our study, we have analyzed the state of Hp proteoforms in plasma and cells using 1D (SDS-PAGE) and 2D electrophoresis (2DE) with the following mass spectrometry (LC ES-MS/MS) or Western blotting.
1. Introduction
2. Results
To obtain more information about plasma proteomes in connection to glioblastoma we have carried out a comparative proteomic analysis of plasma samples from GBM patients and other donors. A gel-based proteomic analysis using SDS-PAGE (1DE), 2DE, and sectional 2DE techniques was performed in combination with MS and immunodetection (Western blotting).
2.1. Detection of differentially expressed proteins in GBM plasma samples versus non-tumorous plasma samples based on SDS-PAGE
Initially, we have performed a comparative analysis of different groups of plasma samples using SDS-PAGE. These groups were: a) GBM (n=22), b) other cancers (n=7), c) acute-phase, non-related with cancer (n=6), d) healthy group (n=16). The equal amounts of proteins from each sample were loaded, and after electrophoresis, the gels were stained with Coomassie R350. It should be mentioned that among all samples, it was four samples (two from the healthy group, one from other cancers, and one from the GBM group) where α2-chain was absent. That means these persons have an Hp1-1 phenotype, and for statistical analysis of α2-chain abundancy, these samples were not included. Talking about Hp phenotypes in all samples, the main one was Hp2-2, a few Hp2-1, and only 2 – Hp1-1. The representative gels of such analysis are shown in Figure 1(a,b,c). Densitometry of each line was performed, and the intensity of every band was measured. Among different bands, the bands that correspond to Mw ~35000 and ~18000 were overexpressed in GBM samples. Western blot analysis and two-dimensional gel electrophoresis (2DE) revealed that these bands correspond to Hp β-chain and α2-chain (Figure 2(a,b,c)). According to densitometry-based statistical analysis, a level of α2- and β-chains is increased in acute-phase samples. It is no wonder as Hp is an acute-phase protein. In the cancer-specific samples, the Hp level is even higher and is highest in GBM samples (Figure 1(d,e)). Interestingly, comparative analysis based on densitometry of Coomassie-stained gels or immune-stained membranes gave similar results (Figure 1(d,e) and Figure 2(d,e)). It means that for α2-chain and β-chain we can detect their amounts in SDS-PAGE just after Coomassie staining. According to the known amount of loaded protein, we can calculate concentrations of α2- and β-chain of Hp in the samples. This data shows that using just Coomassie staining of SDS-PAGE gels we can evaluate the level of Hp α2- chain and β-chain in a sample.
A more detailed evaluation of Hp proteoforms was performed based on 2DE followed by shot-gun mass-spectrometry (ESI LC-MS/MS).
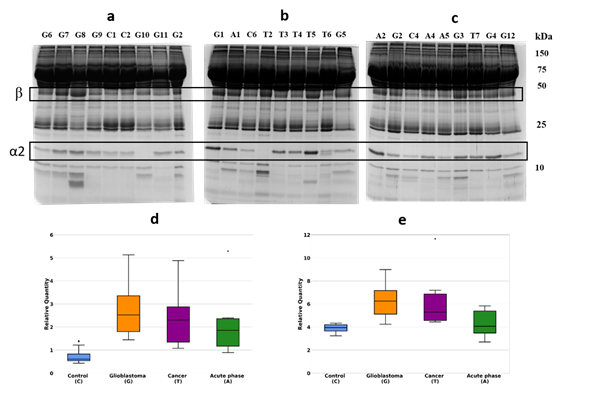
Figure 1. SDS-PAGE of plasma samples, Coomassie staining. The representative gels stained with Coomassie (a,b,c). Statistical analysis of the band intensity of α2-chain (d).). Statistical analysis of the band intensity of β-chain e). GBM samples are marked as G, control – as C, cancer – as T, acute phase – as A.
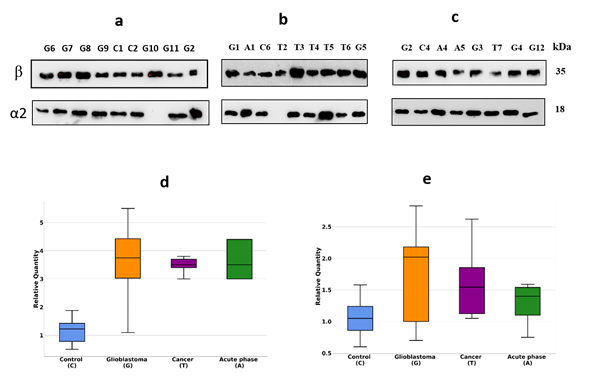
Figure 2. Western blotting of plasma samples. The duplicates of the gels shown in Figure 1 were tested using Ab against α2-chain and β-chain (a,b,c). Intensity statistical analysis of the bands of α2-chain and β-chain (d,e). GBM samples are marked as G, control – as C, cancer – as T, acute phase – as A.
2.2. An analysis based on spots in 2DE gels
To obtain profiles (patterns) of Hp proteoforms we performed 2DE separation of plasma proteins. Proteins after 2DE separation were stained with Coomassie and the produced spots, were analyzed by mass spectrometry (ESI LC-MS/MS analysis). Proteins located in many spots were identified. The obtained data are mainly following the 2DE plasma map deposited at the SWISS-2DPAGE database http://world-2dpage.expasy.org/swiss-2dpage/ [29]. Additionally, we found that many spots contain several different proteins (Supplementary Table 1). We have paid special attention to spots, where Hp was detected (Hp spots). The proteoform clusters (patterns) of α2-chain and β-chain are well-represented in plasma (Figure 3).
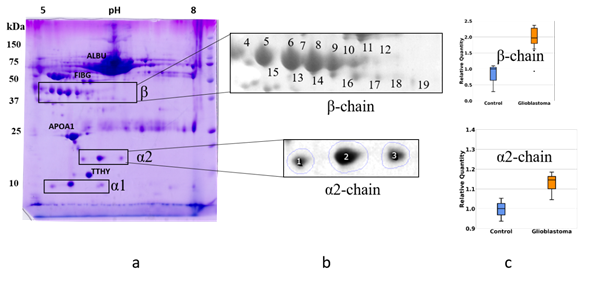

Figure 3. The comparative 2DE analysis of the abundance of Hp α2- and β-chain proteoforms in plasma of control and GBM patients. A representative Coomassie-stained 2DE gel of plasma proteins (GBM) with identified positions of α- and β-chains (a). The enlarged areas with spots containing α2- and β-chain proteoforms (b). MS data about proteins detected in different spots can be found in the Supplementary Table. A comparative statistical analysis of sum spot intensities in control and GBM samples (c).
We performed a comparative 2DE quantitation analysis of α2-chain and β-chain patterns in plasma of healthy people and GBM patients. Also, to obtain more reliable data about the presence of different proteoforms of Hp in plasma from healthy people or GBM patients, we applied Western blotting using Ab against α2-chain (Figure 4).
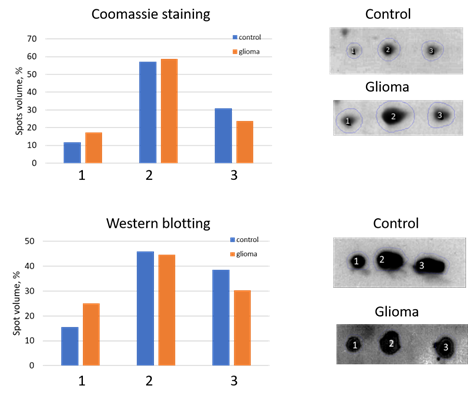
Figure 4. An analysis of three main spots of α2-chain. The relative intensity of each spot in control or GBM samples was estimated after Coomassie staining (top) or Western blotting (down). The spot numbers are the same as in Figure 3.
The 2DE-based analysis confirmed the results obtained using SDS-PAGE. The levels of proteoforms of Hp α2-chain and β-chain are elevated in the plasma of GBM patients. As Hp α2-chain has several proteoforms it is necessary to figure out which proteoform(s) could be responsible for Hp elevated level in plasma of GBM patients. According to data obtained, it looks like there is no preference between different proteoforms in this upregulation. In the case of α2-chain, three major proteoforms can be observed in Coomassie-stained 2DE gel (Figure 3). Comparative analysis using Coomassie spot absorbance, or immunostaining (Western blotting) did not show preference of any spot for the elevated level of Hp α2-chain in plasma of GBM patients. What is more, the ratios in intensities of three spots (proteoforms) obtained using Coomassie staining, or immunostaining (Western blotting) were practically the same. It shows that the impurity of Hp spots by other proteins does not affect the assessment of Hp representation in these spots. A similar situation was observed in the case of major proteoforms of β-chain (data not shown). But it should be mentioned that the spot-based MS-analysis also revealed more locations of Hp proteoforms than it was detected before (Supplementary Table 1). Especially many proteoforms the β-chain has. This data pointed to the possibility of detecting more Hp proteoforms if the whole gel will be analyzed. So, we applied our 2DE sectional (pixel) analysis to plasma samples.
2.3. A sectional 2DE analysis
As haptoglobin was detected not only in “Hp spots” but also in some spots that “are occupied” by other plasma proteins (Supplementary Table 1), it means that there is a chance of detecting more Hp proteoforms when a whole gel analysis is performed. It can be done using our sectional (pixel) 2DE analysis [30]. In with case, a whole 2DE gel is cut into 96 sections with determining coordinates (pI/Mw), and each section is treated according to the protocol for shot-gun mass-spectrometry (ESI LC-MS/MS analysis). Finally, after analysis of mass-spectra, a list of proteins located in each section is obtained. If a protein is detected in different sections, it is considered that this protein has different proteoforms.
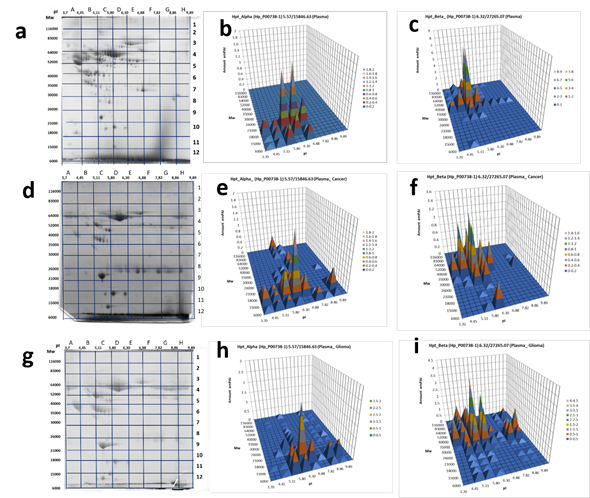
Figure5. 2DE sectional analysis of Hp proteoforms in plasma. (From left to right) Sections in 2DE gel selected for the following ESI LC−MS/MS analysis, Hp α-chain or β-chain detection in sections: (a, b, c) control, а pooled plasma of the mix of samples from 54 healthy donors [31] , (d, e, f) a patient with colon cancer, (g, h, i) a GBM patient.
In the case of plasma proteins, it happened that Hp α-chain was detected at least in 10 sections and β-chain – at least in 30 sections of the 2DE gel (Figure 5, Supplementary Table 2). Keeping in mind that sections are big enough to accommodate several different proteoforms, the real number of proteoforms is even bigger than the number of sections. For instance, in the case of β-chain, section C6 with coordinates: pI 5.11-5.8/Mw 35 000−40 000 accommodates at least ten spots representing the β-chain of Hp that shown in Figure 2. This data shows that plasma Hp can exist in much more proteoforms than was reported previously [29]. What is especially interesting, this number is increased in the case of cancer (Figure 5).
Another aspect demanding attention is colocalization in some cases of α- and β-chains. Mainly, as expected, the patterns of Hp α- and β-chain proteoforms occupy different sections because of the big difference in Mw (theoretical Mw for α1-chain – 9192, α2-chain - 15847, β-chain – 27265) of these polypeptides. Also, we see something special, when both α- and β-chains are detected in the same section. In this case, if a section corresponds to a polypeptide with Mw bigger than 40000 and contains both α2- and β-chains it means that located here α2- and β-chains are covalently linked together. In other words, the polypeptide detected in this section represents an unprocessed Pre-Hp precursor (zonulin). The theoretical biophysical parameters of zonulin are pI 6.3 / Mw 43000. Section D6 (pI 5.80-6.30 / Mw 35000-40000) is close to these parameters. What is more interesting, the similar situation we observed also in sections F6, G5, G6, but only for the plasma of GBM patients (not in the control sample, or colon cancer sample) (Figure 5). As the pH range of these sections (8-10) is much higher than the theoretical pI 6.3, it means that zonulin detected in these sections is bearing some PTMs that are responsible for this shift.
As this form of zonulin was detected only in the plasma of GBM patients we also have checked the Hp pattern in glioblastoma cells (Figure 6 (a,c)). Interestingly, two proteoforms were detected: one in section B2 (pI 4.45-5.11 / Mw 83000-116000) and another in section G5 (pI 7.82-8.86 / Mw 40000-52000).
What is interesting, a similar Hp pattern was obtained with extract of LEH (human lung embryonic fibroblasts) (Figure 6 (b,d), Supplementary Table 3). It means that only full-length preHp2-2 (zonulin) presents in these cells. This information is in full concord with known data about the involvement of zonulin not only in intestinal permeability but also in a wide range of extraintestinal epithelia as well as the ubiquitous vascular endothelium, including the BBB [32]. Zonulin can cross BBB, enter the bloodstream, and be detected in plasma.
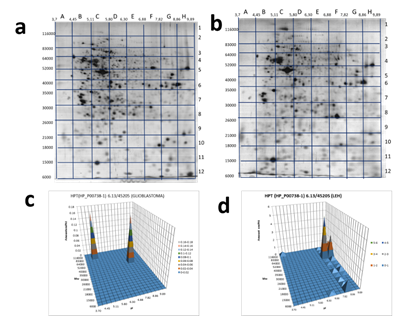
Figure 6. 2DE sectional analysis of Hp proteoforms in cells. Glioblastoma 2DE gel (a). A 2DE pattern of Hp proteoforms in glioblastoma cells (b). LEH 2DE gel (c). A 2DE pattern of Hp proteoforms in LEH cells (d).
3. Current Insights
4. Conclusions
The collected up-to-date information shows that the concentration of certain proteins in plasma including Hp may indicate the presence of GBM. Despite these findings, none of these proteins alone was sufficiently specific and sensitive to serve as a diagnostic marker. So far no single molecular marker was found to follow the dynamics of glioblastoma and so do not have clinical value [35]. The lack of consistency is due to the disease and patient heterogeneity and involvement of these proteins in various physiological and pathological processes. Another important limitation is a wide range of Hp levels in normal serum. In the future, a signature of multiple biomarkers including Hp may prove to be an especially useful tool in glioma subtypes diagnosis that will improve information about prediction and prognosis. Additionally, the characterization of a Hp phenotype may increase the sensitivity and reliability of the screening biomarkers.
References
[1] Furnari, F.B., Fenton, T., Bachoo, R.M., Mukasa, A., et al., Malignant astrocytic glioma: Genetics, biology, and paths to treatment. Genes Dev. 2007, 21, 2683–2710.
[2] Louis, D.N., Ohgaki, H., Wiestler, O.D., Cavenee, W.K., et al., The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol. 2007, 114, 97–109.
[3] Stupp, R., Mason, W.P., Van Den Bent, M.J., Weller, M., et al., Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N. Engl. J. Med. 2005, 352, 987–996.
[4] Szopa W, Burley TA, Kramer-Marek G, K.W., in:, Diagnostic Ther. Biomarkers Glioblastoma Curr. Status Futur. Perspect., 2017, p. 13.
[5] Alix-Panabieres, C.& P., Challenges in circulating tumour cell research. Nat. Rev. Cancer 2014, 14, 523–631.
[6] Miller, A. M., Shah, R. H., Pentsova, E. I., Pourmaleki, M., Briggs, S., Distefano, N. et al, Tracking tumour evolution in glioma through liquid biopsies of cerebrospinal fluid. Nature 2019, 565, 654–658.
[7] Müller Bark, J., Kulasinghe, A., Chua, B., Day, B.W., Punyadeera, C., Circulating biomarkers in patients with glioblastoma. Br. J. Cancer 2020, 122, 295–305.
[8] Chen, Z. & Hambardzumyan, D., Immune microenvironment in glioblastoma subtypes. Front. Immunol 2018, 9, 1004.
[9] Allard, W. J., Matera, J., Miller, M. C., Repollet, M., Connelly, M. C., Rao, C. et al., Tumor cells circulate in the peripheral blood of all major carcinomas but not in healthy subjects or patients with nonmalignant diseases. Clin. Cancer Res 2004, 10, 6897–6904.
[10] Zhang, R., Tremblay, T.L., McDermid, A., Thibault, P., Stanimirovic, D., Identification of differentially expressed proteins in human glioblastoma cell lines and tumors. Glia 2003, 42, 194–208.
[11] Cohen, A.L., Colman, H., Glioma biology and molecular markers. Cancer Treat. Res. 2015, 163, 15–30.
[12] Naryzhny, S.N., Ronzhina, N.L., Mainskova, M.A., Belyakova, N. V., et al., Development of barcode and proteome profiling of glioblastoma. Biochem. Suppl. Ser. B Biomed. Chem. 2014, 8, 243–251.
[13] Petrenko, E.S., Kopylov, A.T., Kleist, O.A., Legina, O.K., et al., Searching for Specific Markers of Glioblastoma: Analysis of Glioblastoma Cell Proteoforms. Cell tissue biol. 2018, 12, 455–459.
[14] Naryzhny, Proteomic Profiling of High-grade Glioblastoma Using Virtual-experimental 2DE. J Proteomics Bioinform 2016, 9, 158–165.
[15] Naryzhny, S., Volnitskiy, A., Kopylov, A., Zorina, E., et al., Proteome of Glioblastoma-Derived Exosomes as a Source of Biomarkers. Biomedicines 2020, 8, 216.
[16] Zhang, S., Shang, S., Li, W., Qin, X., Liu, Y., Insights on N-glycosylation of human haptoglobin and its association with cancers. Glycobiology 2016, 26, 684–692.
[17] Kumar, D.M., Thota, B., Shinde, S.V., Prasanna, K. V., et al., Proteomic identification of haptoglobin α2 as a glioblastoma serum biomarker: Implications in cancer cell migration and tumor growth. J. Proteome Res. 2010, 9, 5557–5567.
[18] Gollapalli, K., Ray, S., Srivastava, R., Renu, D., et al., Investigation of serum proteome alterations in human glioblastoma multiforme. Proteomics 2012, 12, 2378–2390.
[19] Garibay-Cerdenares, O.L., Hernández-Ramírez, V.I., Osorio-Trujillo, J.C., Hernández-Ortíz, M., et al., Proteomic identification of fucosylated haptoglobin alpha isoforms in ascitic fluids and its localization in ovarian carcinoma tissues from Mexican patients. J. Ovarian Res. 2014, 7.
[20] Ye, B., Cramer, D.W., Skates, S.J., Gygi, S.P., et al., Haptoglobin-α subunit as potential serum biomarker in ovarian cancer: Identification and characterization using proteomic profiling and mass spectrometry. Clin. Cancer Res. 2003, 9, 2904–2911.
[21] Naryzhny, S.N., Legina, O.K., Haptoglobin as a biomarker. Biomeditsinskaya Khimiya 2021, 67, 105–118.
[22] Shih, A.W.Y., Mcfarlane, A., Verhovsek, M., Haptoglobin testing in hemolysis: Measurement and interpretation. Am. J. Hematol. 2014.
[23] Clerc, F., Reiding, K.R., Jansen, B.C., Kammeijer, G.S.M., et al., Human plasma protein N-glycosylation. Glycoconj. J. 2016, 33, 309–343.
[24] Pompach, P., Brnakova, Z., Sanda, M., Wu, J., et al., Site-specific glycoforms of haptoglobin in liver cirrhosis and hepatocellular carcinoma. Mol. Cell. Proteomics 2013, 12, 1281–1293.
[25] Mikkat, S., Koy, C., Ulbrich, M., Ringel, B., Glocker, M.O., Mass spectrometric protein structure characterization reveals cause of migration differences of haptoglobin α chains in two-dimensional gel electrophoresis. Proteomics 2004, 4, 3921–3932.
[26] Fasano, A., Zonulin and its regulation of intestinal barrier function: The biological door to inflammation, autoimmunity, and cancer. Physiol. Rev. 2011, 91, 151–175.
[27] Tripathi, A., Lammers, K.M., Goldblum, S., Shea-Donohue, T., et al., Identification of human zonulin, a physiological modulator of tight junctions, as prehaptoglobin-2. Proc. Natl. Acad. Sci. U. S. A. 2009.
[28] Ohlsson B, Orho-Melander M, N.P., Higher Levels of Serum Zonulin May Rather Be Associated with Increased Risk of Obesity and Hyperlipidemia, Than with Gastrointestinal Symptoms or Disease Manifestations. Int J Mol Sci . 2017, 18, 582.
[29] Hoogland, C., Mostaguir, K., Sanchez, J.C., Hochstrasser, D.F., Appel, R.D., SWISS-2DPAGE, ten years later. Proteomics 2004, 4, 2352–2356.
[30] Naryzhny, S.N., Maynskova, M.A., Zgoda, V.G., Ronzhina, N.L., et al., Virtual-Experimental 2DE Approach in Chromosome-Centric Human Proteome Project. J. Proteome Res. 2016, 15, 525–530.
[31] Kopylov A.T, Ilgisonis E.V, Moysa A.A, Tikhonova O.V, et al., Targeted Quantitative Screening of Chromosome 18 Encoded Proteome in Plasma Samples of Astronaut Candidates. J. Proteome Res. 2016, 15, 4039-4046.
[32] Skardelly, M., Armbruster, F.P., Meixensberger, J., Hilbig, H., Expression of zonulin, c-kit, and glial fibrillary acidic protein in human gliomas. Transl. Oncol. 2009, 2, 117–120.
[33] Oh, M.K., Park, H.J., Lee, J.H., Bae, H.M., Kim, I.S., Single chain precursor prohaptoglobin promotes angiogenesis by upregulating expression of vascular endothelial growth factor (VEGF) and VEGF receptor2. FEBS Lett. 2015, 589, 1009–1017.
[34] Díaz-Coránguez, M., Segovia, J., López-Ornelas, A., Puerta-Guardo, H., et al., Transmigration of Neural Stem Cells across the Blood Brain Barrier Induced by Glioma Cells. PLoS One 2013, 8.
[35] Linhares, P., Carvalho, B., Vaz, R., Costa, B.M., Glioblastoma: Is there any blood biomarker with true clinical relevance? Int. J. Mol. Sci. 2020, 21, 1–16.
References
- Furnari, F.B.; Fenton, T.; Bachoo, R.M.; Mukasa, A.; Stommel, J.M.; Stegh, A.; Hahn, W.C.; Ligon, K.L.; Louis, D.N.; Brennan, C.; et al. Malignant astrocytic glioma: Genetics, biology, and paths to treatment. Genes Dev. 2007, 21, 2683–2710.
- Louis, D.N.; Ohgaki, H.; Wiestler, O.D.; Cavenee, W.K.; Burger, P.C.; Jouvet, A.; Scheithauer, B.W.; Kleihues, P. The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol. 2007, 114, 97–109.
- Stupp, R.; Mason, W.P.; Van Den Bent, M.J.; Weller, M.; Fisher, B.; Taphoorn, M.J.; Belanger, K.; Brandes, A.A.; Marosi, C.; Bogdahn, U.; et al. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N. Engl. J. Med. 2005, 352, 987–996.
- Szopa, W.; Burley, T.A.; Kramer-Marek, G.; Kaspera, W. Diagnostic and Therapeutic Biomarkers in Glioblastoma: Current Status and Future Perspectives. Biomed. Res. Int. 2017, 2017, 8013575.
- Alix-Panabieres, C.P. Challenges in circulating tumour cell research. Nat. Rev. Cancer 2014, 14, 523–631.
- Miller, A.M.; Shah, R.H.; Pentsova, E.I.; Pourmaleki, M.; Briggs, S.; Distefano, N.; Zheng, Y.; Skakodub, A.; Mehta, S.A.; Campos, C.; et al. Tracking tumour evolution in glioma through liquid biopsies of cerebrospinal fluid. Nature 2019, 565, 654–658.
- Müller Bark, J.; Kulasinghe, A.; Chua, B.; Day, B.W.; Punyadeera, C. Circulating biomarkers in patients with glioblastoma. Br. J. Cancer 2020, 122, 295–305.
- Chen, Z.; Hambardzumyan, D. Immune microenvironment in glioblastoma subtypes. Front. Immunol. 2018, 9, 1004.
- Allard, W.J.; Matera, J.; Miller, M.C.; Repollet, M.; Connelly, M.C.; Rao, C.; Tibbe, A.G.; Uhr, J.W.; Terstappen, L.W. Tumor cells circulate in the peripheral blood of all major carcinomas but not in healthy subjects or patients with nonmalignant diseases. Clin. Cancer Res. 2004, 10, 6897–6904.
- Zhang, R.; Tremblay, T.L.; McDermid, A.; Thibault, P.; Stanimirovic, D. Identification of differentially expressed proteins in human glioblastoma cell lines and tumors. Glia 2003, 42, 194–208.
- Cohen, A.L.; Colman, H. Glioma biology and molecular markers. Cancer Treat. Res. 2015, 163, 15–30.
- Naryzhny, S.N.; Ronzhina, N.L.; Mainskova, M.A.; Belyakova, N.V.; Pantina, R.A.; Filatov, M.V. Development of barcode and proteome profiling of glioblastoma. Biochem. Suppl. Ser. B Biomed. Chem. 2014, 8, 243–251.
- Petrenko, E.S.; Kopylov, A.T.; Kleist, O.A.; Legina, O.K.; Belyakova, N.; Pantina, R.; Naryzhny, S. Searching for Specific Markers of Glioblastoma: Analysis of Glioblastoma Cell Proteoforms. Cell Tissue Biol. 2018, 12, 455–459.
- Naryzhny, S.N.; Maynskova, M.A.; Zgoda, V.G.; Ronzhina, N.L.; Novikova, S.E.; Belyakova, N.V.; Kleyst, O.A.; Legina, O.K.; Pantina, R.A.; Filatov, M. Proteomic Profiling of High-grade Glioblastoma Using Virtual-experimental 2DE. J. Proteom. Bioinform. 2016, 9, 158–165.
- Naryzhny, S.; Volnitskiy, A.; Kopylov, A.; Zorina, E.; Kamyshinsky, R.; Bairamukov, V.; Garaeva, L.; Shlikht, A.; Shtam, T. Proteome of Glioblastoma-Derived Exosomes as a Source of Biomarkers. Biomedicines 2020, 8, 216.
- Zhang, S.; Shang, S.; Li, W.; Qin, X.; Liu, Y. Insights on N-glycosylation of human haptoglobin and its association with cancers. Glycobiology 2016, 26, 684–692.
- Kumar, D.M.; Thota, B.; Shinde, S.V.; Prasanna, K.V.; Hegde, A.S.; Arivazhagan, A.; Chandramouli, B.A.; Santosh, V.; Somasundaram, K. Proteomic identification of haptoglobin α2 as a glioblastoma serum biomarker: Implications in cancer cell migration and tumor growth. J. Proteome Res. 2010, 9, 5557–5567.
- Gollapalli, K.; Ray, S.; Srivastava, R.; Renu, D.; Singh, P.; Dhali, S.; Bajpai Dikshit, J.; Srikanth, R.; Moiyadi, A.; Srivastava, S. Investigation of serum proteome alterations in human glioblastoma multiforme. Proteomics 2012, 12, 2378–2390.
- Garibay-Cerdenares, O.L.; Hernández-Ramírez, V.I.; Osorio-Trujillo, J.C.; Hernández-Ortíz, M.; Gallardo-Rincón, D.; De León, D.C.; Encarnación-Guevara, S.; Villegas-Pineda, J.; Talamás-Rohana, P. Proteomic identification of fucosylated haptoglobin alpha isoforms in ascitic fluids and its localization in ovarian carcinoma tissues from Mexican patients. J. Ovarian Res. 2014, 7, 27.
- Ye, B.; Cramer, D.W.; Skates, S.J.; Gygi, S.P.; Pratomo, V.; Fu, L.; Horick, N.K.; Licklider, L.J.; Schorge, J.O.; Berkowitz, R.S.; et al. Haptoglobin-α subunit as potential serum biomarker in ovarian cancer: Identification and characterization using proteomic profiling and mass spectrometry. Clin. Cancer Res. 2003, 9, 2904–2911.
- Naryzhny, S.N.; Legina, O.K. Haptoglobin as a biomarker. Biomed. Khimiya 2021, 67, 105–118.
- Shih, A.W.Y.; Mcfarlane, A.; Verhovsek, M. Haptoglobin testing in hemolysis: Measurement and interpretation. Am. J. Hematol. 2014, 89, 443–447.
- Clerc, F.; Reiding, K.R.; Jansen, B.C.; Kammeijer, G.S.; Bondt, A.; Wuhrer, M. Human plasma protein N-glycosylation. Glycoconj. J. 2016, 33, 309–343.
- Pompach, P.; Brnakova, Z.; Sanda, M.; Wu, J.; Edwards, N.; Goldman, R. Site-specific glycoforms of haptoglobin in liver cirrhosis and hepatocellular carcinoma. Mol. Cell. Proteom. 2013, 12, 1281–1293.
- Mikkat, S.; Koy, C.; Ulbrich, M.; Ringel, B.; Glocker, M.O. Mass spectrometric protein structure characterization reveals cause of migration differences of haptoglobin α chains in two-dimensional gel electrophoresis. Proteomics 2004, 4, 3921–3932.
- Fasano, A. Zonulin and its regulation of intestinal barrier function: The biological door to inflammation, autoimmunity, and cancer. Physiol. Rev. 2011, 91, 151–175.
- Tripathi, A.; Lammers, K.M.; Goldblum, S.; Shea-Donohue, T.; Netzel-Arnett, S.; Buzza, M.S.; Antalis, T.M.; Vogel, S.N.; Zhao, A.; Yang, S.; et al. Identification of human zonulin, a physiological modulator of tight junctions, as prehaptoglobin-2. Proc. Natl. Acad. Sci. USA 2009, 106, 16799–16804.
- Ohlsson, B.; Orho-Melander, M.; Nelsson, P.M. Higher Levels of Serum Zonulin May Rather Be Associated with Increased Risk of Obesity and Hyperlipidemia, Than with Gastrointestinal Symptoms or Disease Manifestations. Int. J. Mol. Sci. 2017, 18, 582.
- Oh, M.K.; Park, H.J.; Lee, J.H.; Bae, H.M.; Kim, I.S. Single chain precursor prohaptoglobin promotes angiogenesis by upregulating expression of vascular endothelial growth factor (VEGF) and VEGF receptor2. FEBS Lett. 2015, 589, 1009–1017.
- Skardelly, M.; Armbruster, F.P.; Meixensberger, J.; Hilbig, H. Expression of zonulin, c-kit, and glial fibrillary acidic protein in human gliomas. Transl. Oncol. 2009, 2, 117–120.
- Díaz-Coránguez, M.; Segovia, J.; López-Ornelas, A.; Puerta-Guardo, H.; Ludert, J.; Chávez, B.; Meraz-Cruz, N.; González-Mariscal, L. Transmigration of Neural Stem Cells across the Blood Brain Barrier Induced by Glioma Cells. PLoS ONE 2013, 8, e60655.




