
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Iván Darío Ocampo-Ibáñez | + 13875 word(s) | 13875 | 2021-07-26 10:50:51 | | | |
| 2 | Ron Wang | -4319 word(s) | 9556 | 2021-07-27 04:12:13 | | | | |
| 3 | Ron Wang | Meta information modification | 9556 | 2021-07-28 10:06:29 | | | | |
| 4 | Ron Wang | + 104 word(s) | 9660 | 2021-07-28 10:25:39 | | | | |
| 5 | Ron Wang | + 1114 word(s) | 10670 | 2021-07-28 10:28:29 | | |
Video Upload Options
The emergence of bacteria resistant to conventional antibiotics is of great concern in modern medicine because it renders ineffectiveness of the current empirical antibiotic therapies. Infections caused by vancomycin-resistant Staphylococcus aureus (VRSA) and vancomycin-intermediate S. aureus (VISA) strains represent a serious threat to global health due to their considerable morbidity and mortality rates. Therefore, there is an urgent need of research and development of new antimicrobial alternatives against these bacteria. In this context, the use of antimicrobial peptides (AMPs) is considered a promising alternative therapeutic strategy to control resistant strains. Therefore, a wide number of natural, artificial, and synthetic AMPs have been evaluated against VRSA and VISA strains, with great potential for clinical application. In this regard, we aimed to present a comprehensive and systematic review of research findings on AMPs that have shown antibacterial activity against vancomycin-resistant and vancomycin-intermediate resistant strains and clinical isolates of S. aureus, discussing their classification and origin, physicochemical and structural characteristics, and possible action mechanisms. This is the first review that includes all peptides that have shown antibacterial activity against VRSA and VISA strains exclusively.
1. Introduction
The emergence of bacterial resistance (BR) is one of the most critical public health concerns in recent years. The rapid spread of resistant bacteria compromises the efficacy of antibiotic treatments and has serious implications in the practice of modern medicine [1]. Historically, antibiotics have been used to treat bacterial infections. However, some factors, such as their indiscriminate prescription, inappropriate use in the food industry, lack of discovery of new antibiotics, and poor quality of available antibiotics, have accelerated the emergence of BR [2]. Consequently, BR causes high morbidity and mortality rates, significant increase in healthcare costs, and use of antibacterial agents with increased host toxicity [3][4]. In this context, both Gram-negative and Gram-positive bacteria can be resistant to conventional antibiotics, which limits the number of antimicrobial agents that can be effectively used against these bacterial groups [5]. In particular, BR in Gram-positive species presents a worrisome scenario, since several species show multiple drug resistance and cannot be controlled with conventional antibiotics, leading to the use of last-line drugs in higher concentrations, which can have toxic effects on the patients’ health [6]. Due to the concerns associated with BR and its serious impact on global public health, the World Health Organization (WHO) has recently published a list of priority bacteria resistant to antibiotics [7], through which WHO seeks to guide and promote research and development of new alternatives to control resistant bacteria [2]. The list includes different species of Gram-positive bacteria that cause important community and nosocomial infections, including methicillin-resistant S. aureus (MRSA) and vancomycin-resistant S. aureus (VRSA) [7][8]. S. aureus is a bacterium that is frequently isolated in hospital and community settings, causing various skin and soft tissue infections, as well as severe bone and joint infections. It can also cause endocarditis; bacteremia; and, in more severe cases, toxic shock syndrome and death [9].
Initially, MRSA strains emerged after the introduction of methicillin in 1959 and were only associated with hospital settings. However, these strains that have now widely spread around the world are known to be community-associated MRSA strains with wide genetic diversity, easy transmission, and increased virulence [10]. The evolution of MRSA has been mostly framed in hospital settings, where clonal spread occurs easily from one patient to another and sometimes through healthcare personnel [10]. Methicillin resistance is caused due to the acquisition of mobile chromosomal element known as Staphylococcal cassette chromosome mec (SCCmec) by methicillin-susceptible S. aureus (MSSA) strains [11]. Acquisition of this chromosomal fragment generates an expression of a new penicillin-binding protein (PBP2a) having low affinity for beta-lactams [4]. Initially, MRSA isolates had resistance to only one class of antibiotics; however, nowadays they have multi-antibiotic resistance, including resistance to vancomycin. This generates a serious public health issue since vancomycin is the last line of treatment against infections caused due to resistant strains of S. aureus [12][13][14]. In this regard, resistance to vancomycin causes high mortality rates and increases the risk of premature death when compared with infections caused by susceptible strains, as it increases the length of hospital stay [15][16]. Vancomycin resistance by VRSA strains has been associated with the acquisition of van genes (vanA, vanB, vanC, vanD, vanE, vanF), which generate a low affinity for some glycopeptide antibiotics [17]. However, antibiotics such as oritavacin, a semisynthetic glycopeptide [18], and corbomycin and complestatin, which belong to the type V family of glycopeptides [19], have shown activity against MRSA and VRSA strains. These glycopeptides have several mechanisms of action against cell wall of S. aureus , including the inhibition of peptidoglycan synthesis and the inhibition of fatty acid synthesis [18][19]. Despite these alternatives, resistant strains of S. aureus can have a wide and diverse variety of resistance mechanisms, hindering their control with the use of currently available conventional antibiotics for the treatment of the infections caused by them [12]. In view of this situation, it is crucial to search and develop new antimicrobial alternatives to combat resistant S. aureus strains, especially VRSA strains, which cause significant concern in terms of global public health [7].
In this regard, antimicrobial peptides (AMPs) are a promising alternative to conventional antibiotics because of their great potential to combat resistant bacteria [20]. From a pharmacodynamic point of view, AMPs can have a much higher death rate than antibiotics, even against resistant strains [21]. AMPs are naturally produced small molecules that are part of the innate immune system of different organisms as an effective defense against infections caused by bacteria, fungi, viruses, and some protozoa [22]. Although AMPs are widely diverse, they share common characteristics, such as size (generally between 12 and 50 amino acids) and 3D structures [23]. However, they can differ greatly in terms of amino acid content, activity, targets, action mechanisms, origin, and physicochemical properties [22][24]. According to their activity, AMPs can be classified as antibacterial, antifungal, antiviral, or antiparasitic peptides [25]. The most studied type are the antibacterial AMPs, which are diverse, have different physicochemical properties and can have widely diverse structures, which plays a fundamental role in their biological activity [25]. Antibacterial AMPs have a wide range of action mechanisms and can act on different molecular targets within the bacterial cells, for example, by inducing damage to the bacterial membrane or by inhibiting the synthesis of proteins, enzymes, and nucleic acids at the cytoplasmic level, as well as affecting protein folding [25][26][27]. Because of these characteristics, AMPs have a great potential in the control of bacteria susceptible and resistant to conventional antibiotics that are responsible for infections affecting human health. In this regard, several groups of AMPs have shown high efficacy against bacteria and other pathogens, including strains and clinical isolates of VRSA and vancomycin-intermediate S. aureus (VISA) [28][29]. There is a continuous development in the field of research on peptide activity, their possible molecular targets, and their possible action mechanisms against this particular type of bacterial isolate. The purpose of this review is to comprehensively and systematically describe research findings on AMPs that have shown antibacterial activity against VRSA and VISA strains and clinical isolates, discussing their classification, structure, and possible action mechanisms. This is the first review that collects and classifies all peptides that have shown antibacterial activity against VRSA and VISA strains exclusively.
2. Phenotypic and Genotypic Characteristics of VRSA and VISA Strains That Showed Susceptibility to AMPs
Infections caused by S. aureus are treated with conventional antibiotics that are effective against susceptible strains. However, this efficacy is reduced in the case of resistant strains [30]. Nowadays, a wide diversity of strains and clinical isolates of S. aureus have been reported to show resistance to different antibiotics and contain a wide range of genes in their genomes that make them resistant to antibiotics [10][31]. In light of this situation, AMPs appear as promising alternatives to control this type of bacteria. However, the emergence of strains resistant to AMPs has recently been reported, although it is believed that this resistance is much less likely to evolve than the resistance to conventional antibiotics, and it is believed to occur more easily within in vitro systems than in vivo [21][32][33]. Considering this, it is important to identify the phenotypic and genotypic characteristics of the strains that show susceptibility to AMPs in order to provide relevant information to study resistance to AMPs. There is a scarcity of reports that include genotypic and phenotypic characterization of strains and clinical isolates. Table 1 summarizes the profiles for susceptibility and resistance to conventional antibiotics, as well as the resistance genes identified in VRSA and VISA strains that were evaluated against the AMPs included in this review.
| Strain ID * | Interpretive Categories for Conventional Antibiotics | Method | Genotype | Reference | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| PEN 1 | AMX 1 | OXA 1 | ERY 2 | VAN 3 | TET 4 | DAP 5 | LZD 6 | CLI 7 | ||||
| VRSA-1 | R | – | – | – | R | – | – | – | – | MIC (μg/mL) | – | [29] |
| VRSA-2 | R | – | – | – | R | – | – | – | – | MIC (μg/mL) | – | [29] |
| VRSA-3 | – | – | – | – | R | – | – | – | – | – | – | [34] |
| VRSA-4 | R | R | – | R | R | S | – | – | – | Disc diffusion | VanA | [35] |
| VRSA-5 | – | – | – | – | R | – | S | S | R | MIC (μg/mL) | SCCmec II | [36] |
| VRSA-6 | – | – | R | – | R | – | – | – | – | MIC (μM) | – | [37] |
| VRSA-7 | – | – | R | – | R | – | – | – | – | MIC (μM) | – | [37] |
| VRSA-8 | – | – | R | – | R | – | – | – | – | MIC (μM) | – | [37] |
| VRSA-9 | – | – | R | – | R | – | – | – | – | MIC (μM) | – | [37] |
| VRSA-10 | – | – | R | – | R | – | – | – | – | MIC (μM) | – | [37] |
| VRSA-11 | – | – | R | – | R | – | – | – | – | MIC (μM) | – | [37] |
| VRSA-12 | – | – | R | – | R | – | – | – | – | MIC (μM) | – | [37] |
| VRSA-13 | R | – | R | – | R | – | – | – | – | MIC (μg/mL) | – | [38] |
| VRSA-14 | – | – | – | – | R | – | – | – | – | MIC (μg/mL) | – | [39] |
| VRSA-15 | – | – | – | – | R | – | – | – | – | MIC (μM) | – | [40] |
| VRSA-16 | – | – | – | – | R | – | – | – | – | MIC (μM) | – | [40] |
| VRSA-17 | – | – | – | – | R | – | – | – | – | MIC (μM) | – | [40] |
| VRSA-18 | – | – | – | – | R | – | – | – | – | MIC (μg/mL) | – | [41] |
| VRSA-19 | – | – | S | R | R | R | – | – | R | MIC (μg/mL) | – | [42] |
| VRSA-20 | – | – | – | - | R | – | – | – | – | MIC (μg/mL) | – | [43] |
| VRSA-21 | – | – | – | - | R | – | – | – | – | MIC (μg/mL) | – | [43] |
| VRSA-22 | – | – | – | - | R | – | – | – | – | MIC (μg/mL) | – | [43] |
| VRSA-23 | – | – | – | - | R | – | – | – | – | MIC (μg/mL) | – | [44] |
| VRSA-24 | – | – | – | - | R | – | – | – | – | MIC (μg/mL) | – | [45] |
| VRSA-25 | – | – | – | - | R | – | – | – | – | – | – | [28] |
| VRSA-26 | – | – | – | - | R | – | – | – | – | – | – | [28] |
| VRSA-27 | – | – | – | - | R | – | – | – | – | MIC (μg/mL) | MecA | [46] |
| VRSA-28 | – | – | – | - | R | – | – | S | – | – | – | [47] |
| VRSA-29 | – | – | – | - | R | – | – | S | – | – | – | [47] |
| VRSA-30 | – | – | – | - | R | – | – | S | – | – | – | [47] |
| VRSA-31 | – | – | – | - | R | – | – | S | – | – | – | [47] |
| VRSA-32 | – | – | – | - | R | – | – | R | – | – | – | [47] |
| VRSA-33 | – | – | – | - | R | – | – | – | – | – | – | [48] |
| VISA-1 | R | – | – | - | I | – | – | – | – | MIC (μg/mL) | – | [29] |
| VISA-2 | – | – | – | - | I | – | – | – | – | MIC (μg/mL) | – | [34] |
| VISA-3 | – | – | – | - | I | – | R | S | R | MIC (μg/mL) | SCCmec II | [36] |
| VISA-4 | – | – | – | R | I | – | – | S | – | MIC (μM) | – | [37] |
| VISA-5 | – | – | – | R | I | – | – | S | – | MIC (μM) | – | [37] |
| VISA-6 | – | – | – | R | I | – | – | S | – | MIC (μM) | – | [37] |
| VISA-7 | – | – | – | – | I | – | – | – | – | MIC (μg/mL) | – | [41] |
| VISA-8 | – | – | – | – | I | – | – | – | – | Disc diffusion | – | [49] |
| VISA-9 | – | – | – | – | I | – | – | – | – | MIC (μM) | – | [50] |
| VISA-10 | – | – | S | R | I | S | – | – | R | MIC (μg/mL) | – | [42] |
| VISA-11 | – | – | S | R | I | S | – | – | R | MIC (μg/mL) | – | [42] |
| VISA-12 | – | – | – | – | I | – | – | – | – | MIC (μg/mL) | – | [43] |
| VISA-13 | – | – | – | – | I | – | – | – | – | MIC (μg/mL) | – | [43] |
| VISA-14 | – | – | – | – | I | – | – | – | – | MIC (μg/mL) | – | [43] |
| VISA-15 | - | – | – | – | I | – | – | – | – | MIC (μg/mL) | – | [44] |
| VISA-16 | - | – | – | – | I | – | – | – | – | MIC (μg/mL) | GraS | [51] |
| VISA-17 | - | – | – | – | I | – | – | – | – | MIC (μg/mL) | GraS | [51] |
| VISA-18 | - | – | – | – | I | – | – | – | – | – | – | [52] |
| VISA-19 | - | – | – | – | I | – | – | – | – | – | – | [53] |
| VISA-20 | - | – | – | – | I | – | – | – | – | – | – | [53] |
| VISA-21 | - | – | – | – | I | – | – | – | – | – | – | [53] |
| VISA-22 | - | – | – | – | I | – | – | – | – | – | – | [53] |
| VISA-23 | - | – | – | – | I | – | – | – | – | – | – | [53] |
| VISA-24 | - | – | – | – | I | – | – | – | – | – | – | [53] |
| VISA-25 | - | – | – | – | I | – | – | – | – | MIC (μg/mL) | – | [54] |
| VISA-26 | - | – | – | – | I | – | – | – | – | MIC (μg/mL) | – | [54] |
| VISA-27 | - | – | – | – | I | – | – | – | – | – | – | [55] |
| VISA-28 | – | – | – | – | I | – | – | – | – | – | – | [56] |
| VISA-29 | – | – | – | – | I | – | – | – | – | – | – | [57] |
| VISA-30 | – | – | – | – | I | – | – | – | – | – | – | [58] |
| VISA-31 | – | – | – | – | I | – | – | – | – | – | – | [59] |
| VISA-32 | – | – | – | – | I | – | – | – | – | – | – | [60] |
| VISA-33 | – | – | – | – | I | – | – | S | – | – | – | [47] |
Regarding the phenotypic characterization of these strains, the susceptibility and resistance profiles included seven antimicrobial categories, namely, beta-lactams (penicillin, amoxicillin, and oxacillin), macrolides (erythromycin), glycopeptides (vancomycin), tetracyclines (tetracycline), lipopeptides (daptomycin), oxazolidinones (linezolid), and lincosamides (clindamycin) ( Table 1 ). Susceptibility to these antibiotics was evaluated by broth microdilution or disk diffusion, according to the protocols established by the Clinical and Laboratory Standards Institute (CLSI) and the European Committee on Antimicrobial Susceptibility Testing ( Table 1 ). Taking this into account, strains with phenotypic characterization showed resistance to at least one antibiotic from the seven antimicrobial categories mentioned above ( Table 1 ). Thus, a total of 66 strains and clinical isolates with different phenotypic profiles were identified. In this regard, 33 strains were identified as VRSA and 33 as VISA, as they showed resistance and intermediate resistance to vancomycin, respectively ( Table 1 ). Six of these strains can be considered multidrug-resistant, as they showed resistance to at least three different antimicrobial categories, including combined resistance to beta-lactams, macrolides, glycopeptides, tetracyclines, and lincosamides ( Table 1 ). These results are consistent with the numerous reports on the dissemination of VRSA and VISA strains in recent years, especially with the high spread of vancomycin-resistant MRSA strains that represent a serious threat to human health due to the ineffectiveness of conventional antibiotic therapies [34][35][36][37].
Regarding genotypic characterization, there is a dearth of studies that report the presence of resistance genes in the strains tested against AMPs ( Table 1 ). This suggests that most studies do not take into account the presence of resistance-related genetic factors in the strains tested against antimicrobial peptides. A total of two vancomycin-resistant strains had the transposon-like heterogeneous mobile chromosomal element known as SCCmec type II in their genome: one VISA and one VRSA strain; additionally, one VRSA strain had the mecA gene ( Table 1 ). Although these genetic characteristics have been reported in MRSA strains, VRSA and VISA strains with these genetic mechanisms have already been reported [38]. SCCmec type II is related to MRSA clones associated with hospital settings, which could be related to prolonged vancomycin treatment [38][39]. In particular, the mecA gene is responsible for methicillin resistance and can be acquired by susceptible strains through horizontal transfer mediated by SCCmec, which is integrated into the chromosome of strains associated with hospital and community environments [40][41]. SCCmec works as a genetic exchange vehicle for staphylococcal species, in particular as a mechanism of adaptation to environmental conditions including antibiotic selective pressure [42]. The mecA gene codes for the PBP2a protein with a low affinity for beta-lactam antibiotics [43]. PBP2a replaces all other penicillin-binding proteins and provides broad resistance to all beta-lactam antibiotics, which may include penicillins, cephalosporins, and carbapenemics [10]. In addition, it has been documented that the SCCmec element may contain other types of genes resistant to mercury, cadmium, kanamycin, bleomycin, erythromycin, spectinomycin, and fusidic acid, among other categories of antibiotics [44][45]. In this regard, strains containing SCCmec type II showed resistance to vancomycin, daptomycin, and clindamycin ( Table 1 ). On the other hand, the vanA gene was identified in a clinical VRSA isolate. In addition, this strain showed resistance to other antibiotics, such as penicillin, amoxicillin, and erythromycin ( Table 1 ). For some decades now, glycopeptide antibiotics, such as vancomycin, have become an ideal option for treating infections caused by S. aureus strains resistant to multiple antibiotics [17][46][47]. However, strains with resistance and intermediate resistance to vancomycin have been reported during the last few years [46]. VRSA strains are mainly associated with long periods of hospitalization, persistent infections, or failed treatments [48]. Vancomycin interferes with peptidoglycan synthesis by forming non-covalent bonds with d -Ala- d -Ala residues, disrupting bacterial cell wall assembly [49]. Vancomycin resistance is mediated by van genes, which control the substitution of the d -Ala- d -Ala terminus of the peptidoglycan monomer [17][48]. These genes were first found in Enterococcus spp. and were transmitted to other bacterial species, such as S. aureus [17]. To date, 11 van genes are known, which are classified as (1) genes that mediate the substitution of the d -Ala- d -Ala terminus of the peptidoglycan by d -Ala- d -Lactate, such as vanA, vanB, vanD, vanF, vanI, and vanM, which generate high-level resistance, and (2) genes responsible for the substitution of d -Ala- d -Ala by d -Ala- d -Ser, including vanC, vanE, vanG, vanL, and vanN, associated with low-level resistance [48]. Finally, two strains of VISA had the graS gene ( Table 1 ). In particular, VISA strains, despite not having the van genes in their genome, express a resistance phenotype related to a thickening of the cell wall that occurs due to prolonged exposure to vancomycin and results in an increased amount of antibiotic needed for its control [50]. The reasons why VISA strains show intermediate resistance are not known yet, although genetically these strains contain multiple mutations in genes related to cell wall-associated proteins [49]. In this regard, mutations in the graS gene have been found to be associated with the occurrence of VISA isolates [51]. Overexpression of this gene in VISA strains results in an increase in vancomycin MIC and the expression of some genes such as those involved in cell wall synthesis [52]. In addition, the mutant strains with the graS gene show greater sensitization to AMPs [53]. None of the genes reported for VRSA and VISA strains ( Table 1 ) have previously been associated with resistance to AMPs.
Although vancomycin-resistant isolates have not spread as successfully as MRSA, their grade of resistance and their strong clinical impact have increased over time. Therefore, it is necessary to search for new antimicrobials to control VRSA and VISA strains with wide genetic and phenotypic diversity. AMPs are considered promising alternatives for this purpose [38].
3. Classification of AMPs with Antibacterial Activity against VRSA and VISA Strains
AMPs with antimicrobial activity have been found and isolated from different secre- tions, cells, or in many tissues of different organisms, including plants and animals [61]. AMPs have a wide range of comparative advantages over conventional antibiotics due to their ability to interact with bacterial membranes via electrostatic interactions, penetrate cells affecting cellular functions causing bacterial death, and act on a wide spectrum of bacteria and be less likely to induce resistance [61]. Additionally, AMPs can be used in combination with conventional antibiotics, with highly positive synergistic effects that can help fight serious infectious diseases caused even by resistant bacteria [62]. Despite all these advantages, AMPs can have some disadvantages. For example, some naturally occurring peptides can be easily degraded or have strong cytotoxic and hemolytic effects on human cells; hence, it is necessary to optimize and substantially modify sequences in some of their residues to reduce such adverse effects [63]. Thus, peptides with specific or ar- tificial modifications can be developed through bioinformatics strategies or in a laboratory with high production cost, but also with the possibility of reducing their negative effects and improving their clinical application [61][63]. Therefore, there is a wide diversity of antimicrobial peptides, including natural and artificial or modified peptides, which can be produced in a laboratory. Despite this wide diversity, both natural and artificial AMPs that have shown activity against VRSA and VISA strains and clinical isolates can be classified according to (1) organism of origin, (2) structural characteristics, and (3) mechanism of action. A total of 66 AMPs reported in literature showed antibacterial activity against VRSA and VISA strains and clinical isolates (Tables 2–4).
3.1. AMP Classification Based on Their Origin
AMPs act as a defense mechanism against invading cells in animals, plants, fungi, and microorganisms [61]. The immune system of most organisms has primary reaction mechanisms using AMPs to target a specific class of microorganisms through rapid and lethal action mechanisms [64]. Particularly in animals, AMPs trigger different defense mechanisms according to their interactions with the environment in which they live [64]. In this context, it is known that the highest concentrations of these antimicrobial molecules are generally found in epithelial tissues frequently exposed to pathogens or in cells in- volved in host defense [64]. Thus, some body fluids, such as blood, sweat, saliva, plasma, white blood cell secretions, and granule extracts, have been extensively studied for their antimicrobial characteristics [65][66][67]. A wide variety of AMPs from various animal species have been found to show high antimicrobial capacity against bacteria susceptible and resistant to conventional antibiotics [68]. In particular, AMPs that have shown antibacterial activity against VRSA and VISA strains have been isolated from arthropods, amphibians, mammals, and bacteria. Even artificial AMPs have shown activity against these types of resistant strains.
3.1.1. Animal-Derived AMPs
A total of 28 animal-derived AMPs reported in the literature showed antibacterial activity against vancomycin-resistant S. aureus strains. Table 2 lists all animal-derived AMPs that have demonstrated antimicrobial activity against VRSA and VISA strains. There has been extensive progress in the studies on AMPs obtained from invertebrate animals, and in this regard, a large proportion of AMPs with activity against these strains have been isolated and identified in arthropods [69]. A wide variety of natural insect-produced peptides and their analogs with antibacterial, antifungal, and antiviral activity are now known [88]. Insects, which constitute the largest class of animals on earth, accounting for about 50% of all known species, have simple but well-developed immune systems with a wide arsenal of bioactive molecules, including AMPs [70]. Among the best-known families of insect-derived antimicrobial peptides are the melittins and defensins. Melittin is a peptide isolated from the venom of the Apis mellifera bee that has been extensively studied and has shown bactericidal activity against resistant S. aureus strains [71]. This AMP exhibited potent antibacterial activity against VISA clinical isolates, with an MIC of 2 μM and an MBC of 4 μM [50]. Despite its bactericidal effect, melittin has shown strong cytotoxic and hemolytic activity. As a result, a wide variety of peptide analogs have been designed, synthesized, and tested on the basis of melittin, such as the Hec peptide [72]. This AMP showed antibacterial activity against Gram-positive bacteria, including VRSA strains (MIC > 80 μM), and its toxic effect was detected only at very high concentrations [35]. In addition, when Hec was tested in combination with vancomycin, its activity was signifi- cantly enhanced. However, a high toxic effect on epithelial cells was observed [35][73]. The defensin family includes AMPs that effectively combat Gram-positive bacteria and can be found in various insect species, including diptera [69]. Recently, formicin C was identified in the house fly Musca domestica, a type of defensin that was shown to effectively combat wounds infected with resistant strains of S. aureus in an in vivo model of Hermetia illu- cens larvae [46]. Formicin C successfully inhibited the growth of vancomycin-resistant MRSA strains, with MIC of 32 μg/mL [46]. More specifically, this peptide was able to negatively affect the expression of genes with a significant role in the formation of biofilms by resistant strains of S. aureus [46]. On the other hand, the AMP cecropin A identified in the giant silk moth Hyalophora cecropia showed broad-spectrum activity against resistant bacteria, including VISA strains [74]. In in vitro assays, this peptide showed a minimum inhibitory concentration of 64 μg/mL against the growth of VISA strains and caused a low toxic effect in human cells [49]. In addition, when murine models were intravenously infected with VISA strains and treated with this peptide, a 60% reduction in mortality was observed [49]. Finally, several AMPs with antimicrobial activity against Gram-positive bac- teria have been identified in wasp [48][75]. In this respect, agelaia-MPI and protonectin are venom-derived peptides isolated from two species of wasps, Parachartergus fraternus and Agelaia pallipes pallipes, which showed antimicrobial activity against vancomycin-resistant S. aureus strains [48]. Agelaia-MPI is a peptide highly hemolytic that exhibited a potent antibacterial effect against VRSA strains (MIC between 4 and 8 μg/mL) [48]. Despite its moderated hemolytic effect against human erythrocytes, protonectin showed antibacte- rial activity against VRSA strains (MIC = 16 μg/mL) [48]. NeuroVAL and protonectin-F, analogues peptides of agelaia-MPI and protonectin, respectively, were designed to re- duce nonspecific toxicity and improve potency [48]. Despite its reduced toxic effect on eukaryotic cells, NeuroVAL showed higher inhibitory concentration against VRSA strains (MIC > 128 μg/mL) compared to the canonical agelaia-MPI peptide [48]. The antimicrobial activity against VRSA strains and the toxic effect on cancerous and non-cancerous cell lines were very similar between protonectin-F and the canonical protonectin [48].
Arachnids are another group of arthropods that attract great interest because they are a rich source of molecules with promising characteristics for drug therapy. The venom of these animals has shown a cocktail of AMPs with antimicrobial characteristics with potential to combat bacteria that are resistant to conventional antibiotics [76]. In this sense, AMP ctriporin has been identified in the venom of the scorpion Chaerilus tricostatus, which showed inhibitory activity on the growth of resistant Gram-positive bacterial strains, such as VRSA, VISA, methicillin-resistant coagulase-negative Staphylococcus, and penicillin- resistant Staphylococcus epidermis [29]. In vitro and in vivo experiments carried out with this peptide showed an MIC of 10 μg/mL against both VRSA and VISA strains, as well as a significantly positive skin recovery effect in rabbits [29]. The AMP Smp24 isolated from the venom of the North African scorpion Scorpio maurus palmatus has shown broad activity against Gram-negative and Gram-positive bacteria [54]. In particular, this peptide showed an antibacterial effect against VISA strains (MIC between 32 and 64 μg/mL), with a low hemolytic effect against sheep erythrocytes [54]. On the other hand, it has been established that some ectoparasites, which are vectors of animal diseases, have immune systems with arsenals of defense molecules rich in AMPs [77]. Persulcatusin (PI) was found in the midgut of the tick Ixodes persulcatus, a peptide that showed MIC between 2 and 8 μg/mL against VRSA and VISA strains [34]. Similarly, the IR peptide derived from Ixodes ricinus showed activity against VISA (MIC > 32 μg/mL) and VRSA (MIC = 32 μg/mL) strains [78]. HAE and OMBAC peptides identified in the tick species Haemaphysalis longicornis and Ornithodoros moubata, respectively, also showed antibacterial activity against resistant S. aureus strains [34]. The inhibitory concentrations for HAE against VISA and VRSA (MIC > 32 μg/mL) were comparable to those found for OMBAC against these same strains (MIC > 32 μg/mL for VISA and MIC = 8 μg/mL for VRSA) [34].
Vertebrate animals have complex and well-developed defense mechanisms that pro- tect them from invading pathogens. Amphibians have a rich chemical arsenal in their skin, including a great diversity of AMPs [79]. Amphibian skin provides protection from external agents and also performs a variety of functions including respiration, osmoregulation, and thermoregulation [79]. Many amphibian-derived AMPs have demonstrated antimicrobial activity against VRSA and VISA strains and clinical isolates (Table 2). Magainins, including magainin-1 and -2, are a family of AMPs isolated from the skin of the African frog Xeno- pus laevis belonging to the Pipidae family, which have demonstrated antimicrobial activity against fungi, protozoa, and Gram-positive and Gram-negative bacteria [80]. Magainin-2 has been extensively studied and it possesses activity against Gram-positive bacteria and has a low hemolytic effect [98]. Magainin-2 exhibited potent activity against VISA strains, with inhibitory concentrations of 16 μg/mL [49]. A 50% reduction in mortality was ob- served when murine models were intravenously infected with VISA strains and treated with this peptide [49]. Additionally, temporins are a large family of AMPs identified and isolated from frog skin with antibacterial activity against Gram-positive bacteria [81]. In particular, temporin-CPa and temporin-CPb from Lithobates capito, showed moderate activity against VISA strains with MIC of > 25 μM and 12.5 μM, respectively, and low hemolytic effect on human erythrocytes [56]. Temporin-1SPa from Rana septentrionalis showed activity against VISA strains (MIC = 12.5 μM) and moderate hemolytic activity [56]. Temporin-1Oc from Rana ornativentris, temporin-1Ga from Rana grylio, and temporin-1OLa from Rana okaloosae showed potent antimicrobial activity against VISA strain Mu50 (MIC of 1.6 μM, 6.2 μM, and 3.1 μM, respectively), but these AMPs showed a strong hemolysis against human red blood cells, with hemolytic concentrations between 12.5 and 50 μM [56]. Finally, fallaxin isolated from Leptodactylus fallax, is another amphibian-derived AMPs that has shown antimicrobial activity against Gram-negative bacteria exclusively [60]. A total of 65 analog peptides of fallaxin were designed through rational substitution of amino acids in the canonical sequence, and then tested for hemolytic activity and antibacterial activity against Gram-positive bacteria [60]. In this respect, the analogs FL9, FL10, FA12, and FL14 showed the lowest inhibitory concentrations against VISA strains (MIC values of 50 μM); however, they showed the highest hemolytic activity [60].
| Source | AMP Name | Strain ID | MIC Value | Reference | Toxicity/Properties |
|---|---|---|---|---|---|
| Apis mellifera | Melittin | VISA-9 | 2 μM | [50] | High toxicity to erythrocytes and other human cells |
| Mellitin analog | Hec | VRSA-4 | 80 μM | [35] | Moderate toxic effect at high concentrations |
| Musca domestica | Formicin C | VRSA-27 | 32 μg/mL | [46] | Non toxic to the intradermal model of the larva Hermetia illucens |
| Hyalophora cecropia | Cecropin A | VISA-8 | 64 μg/mL | [49] | Low cytotoxic effect on human lung carcinoma |
| Parachartergus fraternus and Agelaia pallipes pallipes | Agelaia-MPI | VRSA-33 | 4–8 μg/mL | [48] | Strong hemolytic effect on human erythrocytes |
| Protonectin | VRSA-33 | 16 μg/mL | [48] | Toxic to cancerous and non-cancerous cell lines, but moderated hemolytic effect against human erythrocytes | |
| Agelaia-MPI analog | NeuroVAL | VRSA-33 | >128 μg/mL | [48] | Non toxic to human erythrocytes, and cancerous and non-cancerous cells lines. |
| Protonectin analog | Protonectin-F | VRSA-33 | 16 μg/mL | [48] | Toxic to cancerous and non-cancerous cell lines, but moderated hemolytic effect against human erythrocytes |
| Chaerilus tricostatus | Ctriporin | VRSA-1 | 10 μg/mL | [29] | Histological results showed recovery of the skin |
| VRSA-2 | 10 μg/mL | [29] | |||
| VISA-1 | 10 μg/mL | [29] | |||
| Scorpio maurus palmatus | Smp24 | VISA-25 | 32 μg/mL | [54] | Toxic to sheep erythrocytes |
| VISA-26 | 64 μg/mL | [54] | |||
| Ixodes persulcatus | Persulcatusin (IP) | VRSA-3 | 2 μg/mL | [34] | Non toxic to fibroblasts, colon epithelial cells, and erythrocytes |
| VISA-2 | 8 μg/mL | [34] | |||
| Ixodes ricinus | IR | VRSA-3 | 32 μg/mL | [34] | – |
| VISA-2 | >32 μg/mL | [34] | – | ||
| Haemaphysalis longicornis | HAE | VRSA-3 | >32 μg/mL | [34] | – |
| VISA-2 | >32 μg/mL | [34] | – | ||
| Ornithodoros moubata | OMBAC | VRSA-3 | 8 μg/mL | [34] | – |
| VISA-2 | >32 μg/mL | [34] | – | ||
| Xenopus laevis | Magainin-2 | VISA-8 | 16 μg/mL | [49] | – |
| Lithobates capito | Temporin-CPa | VISA-28 | >25 μM | [56] | Hemolysis of human erythrocytes at high concentrations |
| Temporin-CPb | VISA-28 | 12.5 μM | [56] | Hemolysis of human erythrocytes at high concentrations | |
| Rana grylio | Temporin-1Ga | VISA-28 | 6.2 μM | [56] | Strong hemolytic effect on human erythrocytes |
| Rana okaloosae | Temporin-1OLa | VISA-28 | 3.1 μM | [56] | Strong hemolytic effect on human erythrocytes |
| Rana septentrionalis | Temporin-1 SPa | VISA-28 | 12.5 μM | [56] | Moderate hemolytic effect on human erythrocytes |
| Rana ornativentris | Temporin-1Oc | VISA-28 | 1.6 μM | [56] | Strong hemolytic effect on human erythrocytes |
| Fallaxin analogs | FL10 | VISA-32 | 50 μM | [60] | High hemolytic effect on human erythrocytes |
| FL9 | VISA-32 | 50 μM | [60] | Moderate hemolytic effect on human erythrocytes | |
| FA12 | VISA-32 | 50 μM | [60] | High hemolytic effect on human erythrocytes | |
| FL14 | VISA-32 | 50 μM | [60] | High hemolytic effect on human erythrocytes | |
| Homo sapiens | LL-37 | VRSA-18 | 64 μg/mL | [41] | Low cytotoxic effect |
| VISA-7 | 64 μg/mL | [41] | |||
| Derived from LL-37 | LL-13 | VRSA-18 | 512 μg/mL | [41] | – |
| VISA-7 | 1024 μg/mL | [41] | – | ||
| Derived from LL-37 | LL-17 | VRSA-18 | 512 μg/mL | [41] | – |
| VISA-7 | 1024 μg/mL | [41] | – |
3.1.2. Bacteria-Derived AMPs
| Source | AMP Name | Strain ID | MIC Value | Reference | Toxicity/Properties |
|---|---|---|---|---|---|
| Lactococcus lactis | Nisin | VISA-19 | 4.1 mg/L | [53] | Hemolytic effect on sheep erythrocytes |
| VISA-20 | 8.3 mg/L | [53] | |||
| VISA-21 | 4.1 mg/L | [53] | |||
| VISA-22 | 8.3 mg/L | [53] | |||
| VISA-23 | 8.3 mg/L | [53] | |||
| VISA-24 | 4.1 mg/L | [53] | |||
| Staphylococcus hominis | Hominicin | VISA-18 | 3.82 μg/mL | [52] | – |
| Streptococcus mutans | Mutacin 1140 | VRSA-23 | 4–8 μg/mL | [44] | – |
| VISA-15 | 4 μg/mL | [44] | – | ||
| Bacillus sp. | Mersacidin | VISA-27 | 35 μg/mL | [55] | – |
| Lactobacillus salivarius | Bactofencin A (analog 5) | VRSA-25 | 4.3 μM | [28] | – |
| VRSA-26 | 100 μM | [28] | – | ||
| Staphylococcus lugdunensis | Lugdunin | VISA-30 | 3 μg/mL | [58] | No lysis of primary human erythrocytes or neutrophils. |
| Bacillus sp. | BCP61 | VRSA-24 | 10 μg/mL | [45] | – |
| Bacillus subtilis subsp. inaquosorum | P138-C | VRSA-14 | 20 μg/mL | [39] | – |
| Bacillus amyloliquefaciens | CSPK14 | VRSA-13 | 64 μg/mL | [38] | – |
| Fusaricidin analogs | LI-F04a analog 5 | VISA-29 | 16 μg/mL | [57] | Hemolysis on human erythrocytes |
| LI-F04a analog 6 | VISA-29 | 16 μg/mL | [57] | Hemolysis on human erythrocytes | |
| LI-F04a analog 8 | VISA-29 | 16 μg/mL | [57] | Hemolysis on human erythrocytes | |
| LI-F04a analog 11 | VISA-29 | 16 μg/mL | [57] | Hemolysis on human erythrocytes |
3.1.3. Artificial AMPs
| AMP Name | Strain ID | MIC Value | Reference | Toxicity/Properties |
|---|---|---|---|---|
| LTX-109 | VRSA-5 | 2–4 μg/mL | [36] | Phase III of a clinical trial |
| VISA-3 | 2–4 μg/mL | [36] | ||
| Omiganan (Indolicidin analog) |
VRSA-19 | 16 μg/mL | [42] | Topical antimicrobial agent in phase III of a clinical trial |
| VISA-10 | 16 μg/mL | [42] | ||
| VISA-11 | 16 μg/mL | [42] | ||
| WR12 | VRSA-6 | 4 μM | [37] | – |
| VRSA-7 | 8 μM | [37] | – | |
| VRSA-8 | 8 μM | [37] | – | |
| VRSA-9 | 4 μM | [37] | – | |
| VRSA-10 | 4 μM | [37] | – | |
| VRSA-11 | 8 μM | [37] | – | |
| VRSA-12 | 4 μM | [37] | – | |
| VISA-4 | 1 μM | [37] | – | |
| VISA-5 | 1 μM | [37] | – | |
| VISA-6 | 1 μM | [37] | – | |
| DIK-8 | VRSA-6 | 8 μM | [37] | – |
| VRSA-7 | 16 μM | [37] | – | |
| VRSA-8 | 16 μM | [37] | – | |
| VRSA-9 | 16 μM | [37] | – | |
| VRSA-10 | 16 μM | [37] | – | |
| VRSA-11 | 16 μM | [37] | – | |
| VRSA-12 | 16 μM | [37] | – | |
| VISA-4 | 8 μM | [37] | – | |
| VISA-5 | 8 μM | [37] | – | |
| VISA-6 | 8 μM | [37] | – | |
| MP196 | VISA-16 | 16 μg/mL | [51] | Light hemolytic effect on cell lines of breast cancer. Acute toxicity in mice cells. |
| VISA-17 | 64 μg/mL | [51] | ||
| P-113 | VRSA-20 | >64 μg/mL | [43] | – |
| VRSA-21 | >64 μg/mL | [43] | – | |
| VRSA-22 | >64 μg/mL | [43] | – | |
| VISA-12 | >64 μg/mL | [43] | – | |
| VISA-13 | >64 μg/mL | [43] | – | |
| VISA-14 | >64 μg/mL | [43] | – | |
| Phe-P-113 | VRSA-20 | >64 μg/mL | [43] | – |
| VRSA-21 | >64 μg/mL | [43] | – | |
| VRSA-22 | >64 μg/mL | [43] | – | |
| VISA-12 | >64 μg/mL | [43] | – | |
| VISA-13 | >64 μg/mL | [43] | – | |
| VISA-14 | >64 μg/mL | [43] | – | |
| Bip-P-113 | VRSA-20 | 16 μg/mL | [43] | – |
| VRSA-21 | 16 μg/mL | [43] | – | |
| VRSA-22 | 8 μg/mL | [43] | – | |
| VISA-12 | 16 μg/mL | [43] | – | |
| VISA-13 | 16 μg/mL | [43] | – | |
| VISA-14 | 8 μg/mL | [43] | – | |
| Dip-P-113 | VRSA-20 | 32 μg/mL | [43] | – |
| VRSA-21 | 32 μg/mL | [43] | – | |
| VRSA-22 | 32 μg/mL | [43] | – | |
| VISA-12 | 16 μg/mL | [43] | – | |
| VISA-13 | 16 μg/mL | [43] | – | |
| VISA-14 | 16 μg/mL | [43] | – | |
| Nal-P-113 | VRSA-20 | 8 μg/mL | [43] | – |
| VRSA-21 | 8 μg/mL | [43] | – | |
| VRSA-22 | 16 μg/mL | [43] | – | |
| VISA-12 | 8 μg/mL | [43] | – | |
| VISA-13 | 8 μg/mL | [43] | – | |
| VISA-14 | 8 μg/mL | [43] | – | |
| Lipopeptide 1 | VRSA-15 | 0.5 μM | [40] | Low toxicity in human embryonic and kidney cells |
| VRSA-16 | 0.7 μM | [40] | ||
| VRSA-17 | 0.9 μM | [40] | ||
| Lipopeptide 2 | VRSA-15 | 2.8 μM | [40] | Low toxicity in human embryonic and kidney cells |
| VRSA-16 | 1.9 μM | [40] | ||
| VRSA-17 | 2.8 μM | [40] | ||
| Lipopeptide 3 | VRSA-15 | >30 μM | [40] | Low toxicity in human embryonic and kidney cells |
| VRSA-16 | >30 μM | [40] | ||
| VRSA-17 | >30 μM | [40] | ||
| Lipopeptide 4 | VRSA-15 | >30 μM | [40] | Low toxicity in human embryonic and kidney cells |
| VRSA-16 | >30 μM | [40] | ||
| VRSA-17 | >30 μM | [40] | ||
| Lipopeptide 5 | VRSA-15 | 0.2 μM | [40] | Low toxicity in human embryonic and kidney cells |
| VRSA-16 | 0.1 μM | [40] | ||
| VRSA-17 | 0.1 μM | [40] | ||
| Lipopeptide 6 | VRSA-15 | 2.8 μM | [40] | Low toxicity in human embryonic and kidney cells |
| VRSA-16 | 1.9 μM | [40] | ||
| VRSA-17 | 1.9 μM | [40] | ||
| C14-KK | VISA-31 | 12.5 μM | [59] | Strong hemolytic effect on human erythrocytes |
| C14-RRR | VISA-31 | 3.1 μM | [59] | Strong hemolytic effect on human erythrocytes |
| C14-LK | VISA-31 | 1.56 μM | [59] | Strong hemolytic effect on human erythrocytes |
| C14-RW | VISA-31 | >12.5 μM | [59] | Strong hemolytic effect on human erythrocytes |
| C14-WR | VISA-31 | 3.1 μM | [59] | Strong hemolytic effect on human erythrocytes |
| C14-KWI | VISA-31 | 12.5 μM | [59] | Strong hemolytic effect on human erythrocytes |
| C14-LKK | VISA-31 | 3.1 μM | [59] | Strong hemolytic effect on human erythrocytes |
| RRIKA | VISA-33 | 2 μM | [47] | Low hemolytic activity, but show toxicity in mammalian cell lines |
| VRSA-29 | 4 μM | [47] | ||
| VRSA-30 | 4 μM | [47] | ||
| VRSA-31 | 4 μM | [47] | ||
| VRSA-32 | 4 μM | [47] | ||
| VRSA-33 | 4 μM | [47] | ||
| RR | VISA-33 | 16 μM | [47] | Low hemolytic activity, but show toxicity in mammalian cell lines |
| VRSA-29 | 32 μM | [47] | ||
| VRSA-30 | 16 μM | [47] | ||
| VRSA-31 | 16 μM | [47] | ||
| VRSA-32 | 32 μM | [47] | ||
| VRSA-33 | 32 μM | [47] |
3.2. AMPs Classification Based on Their Physicochemical and Structural Properties
| AMP Name | 3D Structure | Sequence | L | C | IP | H | %H | Reference |
|---|---|---|---|---|---|---|---|---|
| Cecropin A |  |
GIGKFLHSAKKFGKAFVGEIMNS | 23 | +6 | 10.6 | 40.19 | 43.48 | [37] |
| Agelaia-MPI |  |
INWLKLGKAIIDAL | 14 | +1 | 9.9 | 45.73 | 64.29 | [48] |
| Protonectin |  |
ILGTILGLLKGL | 12 | +1 | 10.1 | 47.67 | 58.33 | [48] |
| Protonectin-F |  |
IFGTILGFLKGL | 12 | +1 | 10.1 | 50.16 | 58.33 | [48] |
| LL-37 |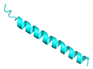  |
LLGDFFRKSKEKIGKEFKRIVQRIKDFLRNLVPRTES | 37 | +6 | 11.1 | 35.14 | 34.62 | [41] |
| LL-13 |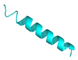  |
IGKEFKRIVQRIKDFLRNLVPRTES | 25 | +4 | 11.4 | 39.37 | 36.00 | [41] |
| LL-17 |  |
LLGDFFRKSKEKIGKEFKRIVQRIKDFLRNLVPRTES | 13 | +4 | 12.2 | 35.69 | 46.15 | [41] |
| Melittin |  |
GIGAVLKVLTTGLPALISWIKRKRQQ | 26 | +5 | 12.5 | 49.39 | 46.15 | [50] |
| Hec |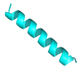  |
FALALKALKKALKKLKKALKKAL | 23 | +9 | 11.4 | 39.47 | 60.87 | [35] |
| Smp24 |  |
IWSFLIKAATKLLPSLFGGG-KKDS | 24 | +4 | 10.6 | 50.39 | 45.83 | [54] |
| Ctriporin |  |
FLWGLIPGAVTSLIAISKK | 19 | +2 | 10.6 | 55.47 | 57.89 | [29] |
| Magainin-2 |  |
GIGKFLHSAKKFGKAFVGEIMNS | 23 | +3 | 10.6 | 40.19 | 43.48 | [49] |
| Temporin-CPa |  |
IPPFIKKVLTTVF | 13 | +2 | 10.6 | 41.02 | 53 | [56] |
| Temporin-CPb |  |
FLPIVGRLISGIL | 13 | +1 | 11.1 | 46.35 | 61 | [56] |
| Temporin-1Ga |  |
SILPTIVSFLSKVF | 14 | +1 | 10.1 | 52.43 | 57 | [56] |
| Temporin-1OLa |  |
FLPFLKSILGKIL | 13 | +2 | 10.6 | 48.08 | 61 | [56] |
| Temporin-1Spa |  |
FLSAITSILGKFF | 13 | +1 | 10.1 | 47.05 | 61 | [56] |
| Temporin-1Oc |  |
FLPLLASLFSRLF | 13 | +1 | 11.1 | 59.16 | 69 | [56] |
| FL9 |  |
GVVDILKGLAKDIAGHLASKVMNKL | 25 | +2 | 10.2 | 41.54 | 52 | [60] |
| FL10 |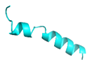  |
GVVDILKGALKDIAGHLASKVMNKL | 25 | +2 | 10.2 | 41.31 | 52 | [60] |
| FA-12 |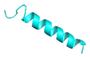  |
GVVDILKGAAKAIAGHLASKVMNKL | 25 | +3 | 10.6 | 37.87 | 56 | [60] |
| FL-14 |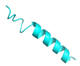  |
GVVDILKGAAKDILGHLASKVMNKL | 25 | +2 | 10.2 | 41.54 | 52 | [60] |
| CSPK-14 |  |
HYDPGDDSGNTG | 12 | −2.9 | 3.6 | 5.66 | 0 | [38] |
| WR12 |  |
RWWRWWRRWWRR | 12 | +6 | 13.2 | 50.42 | 50.00 | [37] |
| RR |  |
WLRRIKAWLRR | 11 | +5 | 13.0 | 33.04 | 54 | [47] |
| RRIKA |  |
WLRRIKAWLRRIKA | 14 | +6 | 13.0 | 39.90 | 57 | [47] |
| Formicin C |  |
ATCDLLSGTGVGHSACAAHCLLRGNRGGYCNGKGVCVCRN | 40 | +3 | 8.3 | 30.58 | 42.50 | [46] |
| IP |  |
GFGCPFNQGACHRHCRSIGRRGGYCAGLFKQTCTCYSR | 38 | +6 | 9.3 | 29.58 | 34.21 | [34] |
| IR |  |
GGYYCPFFQDKCHRHCRSFGRKAGYCGGFLKKTCICV | 37 | +6 | 9.2 | 36.11 | 37.84 | [34] |
| HAE |  |
GCPLNQGACHNHCRSIGRRGGYCAGIIKQTCTCYRK | 36 | +6 | 9.3 | 23.43 | 33.33 | [41] |
| OMBAC |  |
GFGCPFNQYECHAHCSGVPGYKGGYCKGLFKQTCNCY | 37 | +2 | 8.0 | 32.12 | 32.43 | [41] |
3.2.1. α-helix AMPs
| AMP Name | 3D Structure | Sequence | L | C | IP | H | %H | Reference |
|---|---|---|---|---|---|---|---|---|
| Cecropin A |  |
GIGKFLHSAKKFGKAFVGEIMNS | 23 | +6 | 10.6 | 40.19 | 43.48 | [37] |
| Agelaia-MPI |  |
INWLKLGKAIIDAL | 14 | +1 | 9.9 | 45.73 | 64.29 | [48] |
| Protonectin |  |
ILGTILGLLKGL | 12 | +1 | 10.1 | 47.67 | 58.33 | [48] |
| Protonectin-F |  |
IFGTILGFLKGL | 12 | +1 | 10.1 | 50.16 | 58.33 | [48] |
| LL-37 |  |
LLGDFFRKSKEKIGKEFKRIVQRIKDFLRNLVPRTES | 37 | +6 | 11.1 | 35.14 | 34.62 | [41] |
| LL-13 |  |
IGKEFKRIVQRIKDFLRNLVPRTES | 25 | +4 | 11.4 | 39.37 | 36.00 | [41] |
| LL-17 |  |
LLGDFFRKSKEKIGKEFKRIVQRIKDFLRNLVPRTES | 13 | +4 | 12.2 | 35.69 | 46.15 | [41] |
| Melittin |  |
GIGAVLKVLTTGLPALISWIKRKRQQ | 26 | +5 | 12.5 | 49.39 | 46.15 | [50] |
| Hec |  |
FALALKALKKALKKLKKALKKAL | 23 | +9 | 11.4 | 39.47 | 60.87 | [35] |
| Smp24 |  |
IWSFLIKAATKLLPSLFGGG-KKDS | 24 | +4 | 10.6 | 50.39 | 45.83 | [54] |
| Ctriporin |  |
FLWGLIPGAVTSLIAISKK | 19 | +2 | 10.6 | 55.47 | 57.89 | [29] |
| Magainin-2 |  |
GIGKFLHSAKKFGKAFVGEIMNS | 23 | +3 | 10.6 | 40.19 | 43.48 | [49] |
| Temporin-CPa |  |
IPPFIKKVLTTVF | 13 | +2 | 10.6 | 41.02 | 53 | [56] |
| Temporin-CPb |  |
FLPIVGRLISGIL | 13 | +1 | 11.1 | 46.35 | 61 | [56] |
| Temporin-1Ga |  |
SILPTIVSFLSKVF | 14 | +1 | 10.1 | 52.43 | 57 | [56] |
| Temporin-1OLa |  |
FLPFLKSILGKIL | 13 | +2 | 10.6 | 48.08 | 61 | [56] |
| Temporin-1Spa | 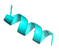 |
FLSAITSILGKFF | 13 | +1 | 10.1 | 47.05 | 61 | [56] |
| Temporin-1Oc |  |
FLPLLASLFSRLF | 13 | +1 | 11.1 | 59.16 | 69 | [56] |
| FL9 |  |
GVVDILKGLAKDIAGHLASKVMNKL | 25 | +2 | 10.2 | 41.54 | 52 | [60] |
| FL10 |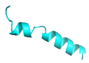  |
GVVDILKGALKDIAGHLASKVMNKL | 25 | +2 | 10.2 | 41.31 | 52 | [60] |
| FA-12 |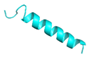  |
GVVDILKGAAKAIAGHLASKVMNKL | 25 | +3 | 10.6 | 37.87 | 56 | [60] |
| FL-14 |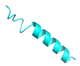  |
GVVDILKGAAKDILGHLASKVMNKL | 25 | +2 | 10.2 | 41.54 | 52 | [60] |
| CSPK-14 |  |
HYDPGDDSGNTG | 12 | −2.9 | 3.6 | 5.66 | 0 | [38] |
| WR12 |  |
RWWRWWRRWWRR | 12 | +6 | 13.2 | 50.42 | 50.00 | [37] |
| RR |  |
WLRRIKAWLRR | 11 | +5 | 13.0 | 33.04 | 54 | [47] |
| RRIKA |  |
WLRRIKAWLRRIKA | 14 | +6 | 13.0 | 39.90 | 57 | [47] |
| Formicin C |  |
ATCDLLSGTGVGHSACAAHCLLRGNRGGYCNGKGVCVCRN | 40 | +3 | 8.3 | 30.58 | 42.50 | [46] |
| IP |  |
GFGCPFNQGACHRHCRSIGRRGGYCAGLFKQTCTCYSR | 38 | +6 | 9.3 | 29.58 | 34.21 | [34] |
| IR |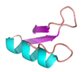  |
GGYYCPFFQDKCHRHCRSFGRKAGYCGGFLKKTCICV | 37 | +6 | 9.2 | 36.11 | 37.84 | [34] |
| HAE |  |
GCPLNQGACHNHCRSIGRRGGYCAGIIKQTCTCYRK | 36 | +6 | 9.3 | 23.43 | 33.33 | [41] |
| OMBAC | 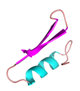 |
GFGCPFNQYECHAHCSGVPGYKGGYCKGLFKQTCNCY | 37 | +2 | 8.0 | 32.12 | 32.43 | [41] |
3.2.2. AMPs Forming β-Pleated Sheet Peptides
3.2.3. Mixed AMPs
3.2.4. AMPs of Atypical Structure: Cyclic, Complex, and with Unusual Amino Acids
| AMP Name | Aminoacid Sequences and Structures | Molecular Weight (KDa) | Reference |
|---|---|---|---|
| Nisin | 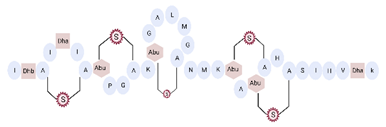 |
3.35 | [53] |
| Hominicin |  |
2.03 | [52] |
| Mutacin 1140 (MU1140) | 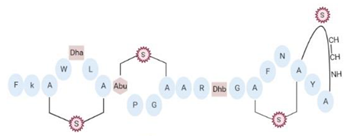 |
2.26 | [44] |
| Mersacidin | 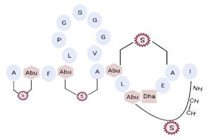 |
1.82 | [55][73] |
| Bactofencin A (analog 5) | 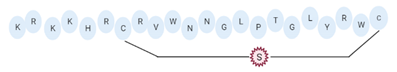 |
2.77 | [28] |
| BCP61 | 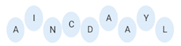 |
9.50 | [45] |
| Lugdunin | 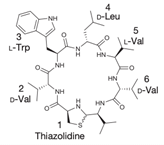 |
0.78 | [58] |
| LI-F04a analog 5 | 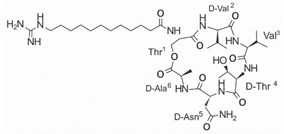 |
– | [58] |
| LI-F04a analog 6 | 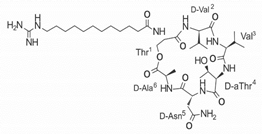 |
– | [58] |
| LI-F04a analog 8 | 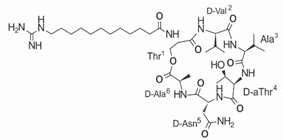 |
– | [58] |
| LI-F04a analog 11 | 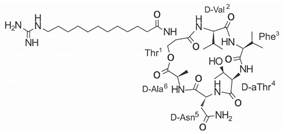 |
– | [58] |
| Peptide Name | Chemical Structure | Molecular Weight (KDa) | Reference |
|---|---|---|---|
| Omiganan | 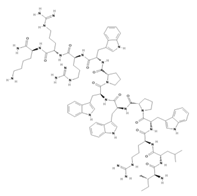 |
1.96 | [42][74] |
| Lipopeptide 1 * | 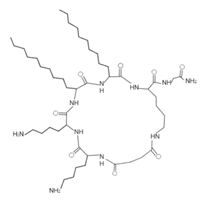 |
– | [40] |
| Lipopeptide 2 * | 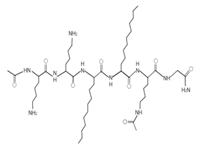 |
– | [40] |
| Lipopeptide 3 * | 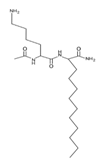 |
– | [40] |
| Lipopeptide 4 * | 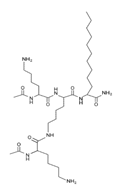 |
– | [40] |
| Lipopeptide 5 * | 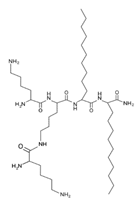 |
– | [40] |
| Lipopeptide 6 * | 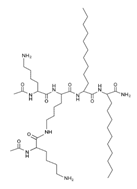 |
– | [40] |
3.3. Mechanisms of Action of AMPs with Antibacterial Activity against VRSA and VISA
3.3.1. AMPs That Permeabilize Bacterial Membranes
3.3.2. AMPs That Interact with DNA
References
- Lima, W.G.; de Brito, J.C.M.; Cardoso, V.N.; Fernandes, S.O.A. In-depth characterization of antibacterial activity of melittin against Staphylococcus aureus and use in a model of non-surgical MRSA-infected skin wounds. Eur. J. Pharm. Sci. 2021, 156, 105592.
- Jelinkova, P.; Splichal, Z.; Jimenez, A.M.J.; Haddad, Y.; Mazumdar, A.; Sur, V.P.; Milosavljevic, V.; Kopel, P.; Buchtelova, H.; Guran, R.; et al. Novel vancomycin–peptide conjugate as potent antibacterial agent against vancomycin-resistant Staphylococcus aureus. Infect Drug Resist. 2018, 11, 1807–1817.
- Wang, B.; Yao, Y.; Wei, P.; Song, C.; Wan, S.; Yang, S.; Zhu, G.M.; Liu, H.M. Housefly Phormicin inhibits Staphylococcus aureus and MRSA by disrupting biofilm formation and altering gene expression In Vitro and In Vivo. Int. J. Biol. Macromol. 2020.
- Cirioni, O.; Silvestri, C.; Ghiselli, R.; Giacometti, A.; Orlando, F.; Mocchegiani, F.; Chiodi, L.; Della Vittoria, A.; Saba, V.; Scalise, G. Experimental study on the efficacy of combination of α-helical antimicrobial peptides and vancomycin against Staphylococcus aureus with intermediate resistance to glycopeptides. Peptides 2006, 27, 2600–2606.
- Muller, J.A.I.; Lawrence, N.; Chan, L.Y.; Harvey, P.J.; Elliott, A.G.; Blaskovich, M.A.T.; Gonçalves, J.C.; Galante, P.; Mortari, M.R.; Toffoli-Kadri, M.C.; et al. Antimicrobial and Anticancer Properties of Synthetic Peptides Derived from the Wasp Parachartergus fraternus. Chem. Biol. Chem. 2021, 22, 1415–1423.
- Fan, Z.; Cao, L.; He, Y.; Hu, J.; Di, Z.; Wu, Y.; Li, W.; Cao, Z. Ctriporin, a new anti-methicillin-resistant Staphylococcus aureus peptide from the venom of the scorpion Chaerilus tricostatus. Antimicrob. Agents Chemother. 2011, 55, 5220–5229.
- Harrison, P.L.; Abdel-Rahman, M.A.; Strong, P.N.; Tawfik, M.M.; Miller, K. Characterisation of three alpha-helical antimicrobial peptides from the venom of Scorpio maurus palmatus. Toxicon 2016, 117, 30–36.
- Miyoshi, N.; Isogai, E.; Hiramatsu, K.; Sasaki, T. Activity of tick antimicrobial peptide from Ixodes persulcatus (persulcatusin) against cell membranes of drug-resistant Staphylococcus aureus. J. Antibiot. (Tokyo) 2017, 70, 142–146.
- Mishra, B.; Wang, X.; Lushnikova, T.; Zang, Y.; Golla, R.; Lakshmaiah, J.; Wang, C.; Mcguire, T.; Wang, G. Antibacterial, Antifungal, Anticancer Activities and Structural Bioinformatics Analysis of Six Naturally Occurring Temporins. Peptides 2018, 1, 9–20.
- Nielsen, S.L.; Frimodt-Møller, N.; Kragelund, B.B.; Hansen, P.R. Structure-activity study of the antibacterial peptide fallaxin. Protein Sci. 2007, 16, 1969–1976.
- Shurko, J.F.; Galega, R.S.; Li, C.; Lee, G.C. Evaluation of LL-37 antimicrobial peptide derivatives alone and in combination with vancomycin against S. aureus. J. Antibiot. 2018, 71, 971–974.
- Piper, C.; Draper, L.A.; Cotter, P.D.; Ross, R.P.; Hill, C. A comparison of the activities of lacticin 3147 and nisin against drug-resistant Staphylococcus aureus and Enterococcus species. J. Antimicrob. Chemother. 2009, 64, 546–551.
- Kim, P.I.; Sohng, J.K.; Sung, C.; Joo, H.S.; Kim, E.M.; Yamaguchi, T.; Park, D.; Kim, B.G. Characterization and structure identification of an antimicrobial peptide, hominicin, produced by Staphylococcus hominis MBBL 2-9. Biochem. Biophys. Res. Commun. 2010, 399, 133–138.
- Ghobrial, O.G.; Derendorf, H.; Hillman, J.D. Pharmacodynamic activity of the lantibiotic MU1140. Int. J. Antimicrob. Agents 2009, 33, 70–74.
- Sass, P.; Jansen, A.; Szekat, C.; Sass, V.; Sahl, H.G.; Bierbaum, G. The lantibiotic mersacidin is a strong inducer of the cell wall stress response of Staphylococcus aureus. BMC Microbiol. 2008, 8, 1–11.
- Bédard, F.; Fliss, I.; Biron, E. Structure-Activity Relationships of the Bacteriocin Bactofencin A and Its Interaction with the Bacterial Membrane. ACS Infect Dis. 2019, 5, 199–207.
- Zipperer, A.; Konnerth, M.C.; Laux, C.; Berscheid, A.; Janek, D.; Weidenmaier, C.; Burian, M.; Schilling, N.A.; Slavetinsky, C.; Marschal, M.; et al. Human commensals producing a novel antibiotic impair pathogen colonization. Nature 2016, 535, 511–516.
- Choi, Y.H.; Cho, S.S.; Simkhada, J.R.; Yoo, J.C. A novel thermotolerant and acidotolerant peptide produced by a Bacillus strain newly isolated from a fermented food (kimchi) shows activity against multidrug-resistant bacteria. Int. J. Antimicrob. Agents 2012, 40, 80–83.
- Regmi, S.; Choi, Y.S.; Choi, Y.H.; Kim, Y.K.; Cho, S.S.; Yoo, J.C.; Suh, J.W. Antimicrobial peptide from Bacillus subtilis CSB138: Characterization, killing kinetics, and synergistic potency. Int. Microbiol. 2017, 20, 45–53.
- Regmi, S.; Choi, Y.H.; Choi, Y.S.; Kim, M.R.; Yoo, J.C. Antimicrobial peptide isolated from Bacillus amyloliquefaciens K14 revitalizes its use in combinatorial drug therapy. Folia Microbiol. (Praha) 2017, 62, 127–138.
- Bionda, N.; Stawikowski, M.; Stawikowski, R.; Cudic, M.; Lopéz, F.; Treitl, D.; Medina, J.; Cudic, P. Effects of cyclic lipodepsipeptide structural modulation on stability, antibacterial activity and human cell toxicity. Chem. Med. Chem. 2012, 7, 871–882.
- Saravolatz, L.D.; Pawlak, J.; Johnson, L.; Bonilla, H.; Saravolatz, L.D.; Fakih, M.G.; Fugelli, A.; Olsen, W.M. In Vitro activities of LTX-109, a synthetic antimicrobial peptide, against methicillin-resistant, vancomycin-intermediate, vancomycin-resistant, daptomycin-nonsusceptible, and linezolid-nonsusceptible Staphylococcus aureus. Antimicrob. Agents Chemother. 2012, 56, 4478–4482.
- Fritsche, T.R.; Rhomberg, P.R.; Sader, H.S.; Jones, R.N. In Vitro activity of omiganan pentahydrochloride tested against vancomycin-tolerant, -intermediate, and -resistant Staphylococcus aureus. Diagn. Microbiol. Infect Dis. 2008, 60, 399–403.
- Mohamed, M.F.; Abdelkhalek, A.; Seleem, M.N. Evaluation of short synthetic antimicrobial peptides for treatment of drug-resistant and intracellular Staphylococcus aureus. Sci. Rep. 2016, 6, 1–14.
- Wenzel, M.; Prochnow, P.; Mowbray, C.; Vuong, C.; Höxtermann, S.; Stepanek, J.J.; Albada, H.B.; Hall, J.; Metzler-Nolte, N.; Bandow, J.E. Towards profiles of resistance development and toxicity for the small cationic hexapeptide RWRWRW-NH 2. Front. Cell Dev. Biol. 2016, 4, 1–9.
- Wu, C.; Hsueh, J.; Yip, B.; Chih, Y.; Peng, K.; Cheng, J. Antimicrobial Peptides Display Strong Synergy with Vancomycin Against Vancomycin-Resistant. Int. J. Mol. Sci. 2020, 21, 4578.
- Azmi, F.; Elliott, A.G.; Marasini, N.; Ramu, S.; Ziora, Z.; Kavanagh, A.M.; Blaskovich, M.A.T.; Cooper, M.A.; Skwarczynski, M.; Toth, I. Short cationic lipopeptides as effective antibacterial agents: Design, physicochemical properties and biological evaluation. Bioorganic Med. Chem. 2016, 24, 2235–2241.
- Mishra, B.; Lushnikova, T.; Wang, G. Small lipopeptides possess anti-biofilm capability comparable to daptomycin and vancomycin. RSC Adv. 2015, 5, 59758–59769.
- Mohamed, M.F.; Hamed, M.I.; Panitch, A.; Seleem, M.N. Targeting methicillin-resistant Staphylococcus aureus with short salt-resistant synthetic peptides. Antimicrob. Agents Chemother. 2014, 58, 4113–4122.
- Saravolatz, L.D.; Pawlak, J.; Johnson, L.; Bonilla, H.; Saravolatz, L.D.; Fakih, M.G.; Fugelli, A.; Olsen, W.M. In Vitro activities of LTX-109, a synthetic antimicrobial peptide, against methicillin-resistant, vancomycin-intermediate, vancomycin-resistant, daptomycin-nonsusceptible, and linezolid-nonsusceptible Staphylococcus aureus. Antimicrob. Agents Chemother. 2012, 56, 4478–4482.
- Rubinchik, E.; Dugourd, D.; Algara, T.; Pasetka, C.; Friedland, H.D. Antimicrobial and antifungal activities of a novel cationic antimicrobial peptide, omiganan, in experimental skin colonisation models. Int. J. Antimicrob. Agents 2009, 34, 457–461.
- Fritsche, T.R.; Rhomberg, P.R.; Sader, H.S.; Jones, R.N. In Vitro activity of omiganan pentahydrochloride tested against vancomycin-tolerant, -intermediate, and -resistant Staphylococcus aureus. Diagn. Microbiol. Infect Dis. 2008, 60, 399–403.
- Sader, H.S.; Fedler, K.A.; Rennie, R.P.; Stevens, S.; Jones, R.N. Omiganan pentahydrochloride (MBI 226), a topical 12-amino-acid cationic peptide: Spectrum of antimicrobial activity and measurements of bactericidal activity. Antimicrob. Agents Chemother. 2004, 48, 3112–3118.
- Mohamed, M.F.; Abdelkhalek, A.; Seleem, M.N. Evaluation of short synthetic antimicrobial peptides for treatment of drug-resistant and intracellular Staphylococcus aureus. Sci. Rep. 2016, 6, 1–14.
- Wenzel, M.; Prochnow, P.; Mowbray, C.; Vuong, C.; Höxtermann, S.; Stepanek, J.J.; Albada, H.B.; Hall, J.; Metzler-Nolte, N.; Bandow, J.E. Towards profiles of resistance development and toxicity for the small cationic hexapeptide RWRWRW-NH 2. Front. Cell Dev. Biol. 2016, 4, 1–9.
- Lohan, S.; Monga, J.; Cameotra, S.S.; Bisht, G.S. In Vitro and In Vivo antibacterial evaluation and mechanistic study of ornithine based small cationic lipopeptides against antibiotic resistant clinical isolates. Eur. J. Med. Chem. 2014, 88, 19–27.
- Wu, C.; Hsueh, J.; Yip, B.; Chih, Y.; Peng, K.; Cheng, J. Antimicrobial Peptides Display Strong Synergy with Vancomycin Against Vancomycin-Resistant. Int. J. Mol. Sci. 2020, 21, 4578.
- Azmi, F.; Elliott, A.G.; Marasini, N.; Ramu, S.; Ziora, Z.; Kavanagh, A.M.; Blaskovich, M.A.T.; Cooper, M.A.; Skwarczynski, M.; Toth, I. Short cationic lipopeptides as effective antibacterial agents: Design, physicochemical properties and biological evaluation. Bioorganic Med. Chem. 2016, 24, 2235–2241.
- Mishra, B.; Lushnikova, T.; Wang, G. Small lipopeptides possess anti-biofilm capability comparable to daptomycin and vancomycin. RSC Adv. 2015, 5, 59758–59769.
- Mohamed, M.F.; Hamed, M.I.; Panitch, A.; Seleem, M.N. Targeting methicillin-resistant Staphylococcus aureus with short salt-resistant synthetic peptides. Antimicrob. Agents Chemother. 2014, 58, 4113–4122.
- BARBER, M. Methicillin-resistant staphylococci. J. Clin. Pathol. 1961, 14, 385–393.
- Hiramatsu, K.; Katayama, Y.; Matsuo, M.; Sasaki, T.; Morimoto, Y.; Sekiguchi, A.; Baba, T. Multi-drug-resistant Staphylococcus aureus and future chemotherapy. J. Infect Chemother 2014, 20, 593–601.
- Chambers, H.F. Methicillin resistance in staphylococci: Molecular and biochemical basis and clinical implications. Clin. Microbiol. Rev. 1997, 10, 781–791.
- Ito, T.; Katayama, Y.; Asada, K.; Mori, N.; Tsutsumimoto, K.; Tiensasitorn, C.; Hiramatsu, K. Structural comparison of three types of staphylococcal cassette chromosome mec integrated in the chromosome in methicillin-resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 2001, 45, 1323–1336.
- Miragaia, M. Factors contributing to the evolution of Meca-mediated β-lactam resistance in staphylococci: Update and new insights from whole genome sequencing (WGS). Front. Microbiol. 2018, 9, 2723.
- Mella, S.S. Staphylococcus aureus resistente a vancomicina. Rev. Chil. Infectol. 2002, 19, 575–587.
- CLSI. Performance Standards for Antimicrobial Susceptibility Testing; Twenty-Seventh Informational Supplement, CLSI Document M100-S27; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2017; ISBN 1562387855.
- Cong, Y.; Yang, S.; Rao, X. Vancomycin resistant Staphylococcus aureus infections: A review of case updating and clinical features. J. Adv. Res. 2020, 21, 169–176.
- McGuinness, W.A.; Malachowa, N.; DeLeo, F.R. Vancomycin resistance in Staphylococcus aureus. Yale J. Biol. Med. 2017, 90, 269–281.
- Loomba, P.; Taneja, J.; Mishra, B. Methicillin and vancomycin resistant S. aureus in hospitalized patients. J. Glob. Infect Dis. 2010, 2, 275.
- Wenzel, M.; Prochnow, P.; Mowbray, C.; Vuong, C.; Höxtermann, S.; Stepanek, J.J.; Albada, H.B.; Hall, J.; Metzler-Nolte, N.; Bandow, J.E. Towards profiles of resistance development and toxicity for the small cationic hexapeptide RWRWRW-NH 2. Front. Cell Dev. Biol. 2016, 4, 1–9.
- Howden, B.P.; Davies, J.K.; Johnson, P.D.R.; Stinear, T.P.; Grayson, M.L. Reduced vancomycin susceptibility in Staphylococcus aureus, including vancomycin-intermediate and heterogeneous vancomycin-intermediate strains: Resistance mechanisms, laboratory detection, and clinical implications. Clin. Microbiol. Rev. 2010, 23, 99–139.
- Chaili, S.; Cheung, A.L.; Bayer, A.S.; Xiong, Y.Q.; Waring, A.J.; Memmi, G.; Donegan, N.; Yang, S.J.; Yeaman, M.R. The GraS sensor in Staphylococcus aureus mediates resistance to host defense peptides differing in mechanisms of action. Infect Immun. 2016, 84, 459–466.
- Wang, J.; Dou, X.; Song, J.; Lyu, Y.; Zhu, X.; Xu, L.; Li, W.; Shan, A. Antimicrobial peptides: Promising alternatives in the post feeding antibiotic era. Med. Res. Rev. 2018, 39, 831–859.
- Zelezetsky, I.; Tossi, A. Alpha-helical antimicrobial peptides-Using a sequence template to guide structure-activity relationship studies. Biochim. Biophys. Acta Biomembr. 2006, 1758, 1436–1449.
- Mojsoska, B.; Jenssen, H. Peptides and peptidomimetics for antimicrobial drug design. Pharmaceuticals 2015, 8, 366–415.
- Wu, Q.; Patočka, J.; Kuča, K. Insect Antimicrobial Peptides, a Mini Review. Toxins 2018, 10, 461.
- Sancho-Vaello, E.; Gil-Carton, D.; François, P.; Bonetti, E.J.; Kreir, M.; Pothula, K.R.; Kleinekathöfer, U.; Zeth, K. The structure of the antimicrobial human cathelicidin LL-37 shows oligomerization and channel formation in the presence of membrane mimics. Sci. Rep. 2020, 10, 1–16.
- Muller, J.A.I.; Lawrence, N.; Chan, L.Y.; Harvey, P.J.; Elliott, A.G.; Blaskovich, M.A.T.; Gonçalves, J.C.; Galante, P.; Mortari, M.R.; Toffoli-Kadri, M.C.; et al. Antimicrobial and Anticancer Properties of Synthetic Peptides Derived from the Wasp Parachartergus fraternus. Chem. Biol. Chem. 2021, 22, 1415–1423.
- Seil, M.; Nagant, C.; Dehaye, J.P.; Vandenbranden, M.; Lensink, M.F. Spotlight on human LL-37, an immunomodulatory peptide with promising cell-penetrating properties. Pharmaceuticals 2010, 3, 3435–3460.
- Raghuraman, H.; Chattopadhyay, A. Melittin: A membrane-active peptide with diverse functions. Biosci. Rep. 2007, 27, 189–223.
- Ceremuga, M.; Stela, M.; Janik, E.; Gorniak, L.; Synowiec, E.; Sliwinski, T.; Sitarek, P.; Saluk-Bijak, J.; Bijak, M. Melittin—a natural peptide from bee venom which induces apoptosis in human leukaemia cells. Biomolecules 2020, 10, 247.
- Jelinkova, P.; Splichal, Z.; Jimenez, A.M.J.; Haddad, Y.; Mazumdar, A.; Sur, V.P.; Milosavljevic, V.; Kopel, P.; Buchtelova, H.; Guran, R.; et al. Novel vancomycin–peptide conjugate as potent antibacterial agent against vancomycin-resistant Staphylococcus aureus. Infect Drug Resist. 2018, 11, 1807–1817.
- Harrison, P.L.; Abdel-Rahman, M.A.; Strong, P.N.; Tawfik, M.M.; Miller, K. Characterisation of three alpha-helical antimicrobial peptides from the venom of Scorpio maurus palmatus. Toxicon 2016, 117, 30–36.
- Matsuzaki, K.; Sugishita, K.I.; Harada, M.; Fujii, N.; Miyajima, K. Interactions of an antimicrobial peptide, magainin 2, with outer and inner membranes of Gram-negative bacteria. Biochim. Biophys. Acta Biomembr. 1997, 1327, 119–130.
- Mishra, B.; Wang, X.; Lushnikova, T.; Zang, Y.; Golla, R.; Lakshmaiah, J.; Wang, C.; Mcguire, T.; Wang, G. Antibacterial, Antifungal, Anticancer Activities and Structural Bioinformatics Analysis of Six Naturally Occurring Temporins. Peptides 2018, 1, 9–20.
- Nielsen, S.L.; Frimodt-Møller, N.; Kragelund, B.B.; Hansen, P.R. Structure-activity study of the antibacterial peptide fallaxin. Protein Sci. 2007, 16, 1969–1976.
- Regmi, S.; Choi, Y.H.; Choi, Y.S.; Kim, M.R.; Yoo, J.C. Antimicrobial peptide isolated from Bacillus amyloliquefaciens K14 revitalizes its use in combinatorial drug therapy. Folia Microbiol. (Praha) 2017, 62, 127–138.
- Ennahar, S.; Sashihara, T.; Sonomoto, K.; Ishizaki, A. Class IIa bacteriocins: Biosynthesis, structure and activity. FEMS Microbiol. Rev. 2000, 24, 85–106.
- Müller, A.; Wenzel, M.; Strahl, H.; Grein, F.; Saaki, T.N.V.; Kohl, B.; Siersma, T.; Bandow, J.E.; Sahl, H.G.; Schneider, T.; et al. Daptomycin inhibits cell envelope synthesis by interfering with fluid membrane microdomains. Proc. Natl. Acad. Sci. USA 2016, 113, E7077–E7086.
- Mohamed, M.F.; Abdelkhalek, A.; Seleem, M.N. Evaluation of short synthetic antimicrobial peptides for treatment of drug-resistant and intracellular Staphylococcus aureus. Sci. Rep. 2016, 6, 1–14.
- Mohamed, M.F.; Hamed, M.I.; Panitch, A.; Seleem, M.N. Targeting methicillin-resistant Staphylococcus aureus with short salt-resistant synthetic peptides. Antimicrob. Agents Chemother. 2014, 58, 4113–4122.
- Kumar, P.; Kizhakkedathu, J.N.; Straus, S.K. Antimicrobial peptides: Diversity, mechanism of action and strategies to improve the activity and biocompatibility In Vivo. Biomolecules 2018, 8, 4.
- Zhong, G.; Cheng, J.; Liang, Z.C.; Xu, L.; Lou, W.; Bao, C.; Ong, Z.Y.; Dong, H.; Yang, Y.Y.; Fan, W. Short Synthetic β-Sheet Antimicrobial Peptides for the Treatment of Multidrug-Resistant Pseudomonas aeruginosa Burn Wound Infections. Adv. Healthc. Mater. 2017, 6.
- Jin, Y.; Hammer, J.; Pate, M.; Zhang, Y.; Zhu, F.; Zmuda, E.; Blazyk, J. Antimicrobial activities and structures of two linear cationic peptide families with various amphipathic β-sheet and α-helical potentials. Antimicrob. Agents Chemother. 2005, 49, 4957–4964.
- Rausch, J.M.; Marks, J.R.; Wimley, W.C. Rational combinatorial design of pore-forming β-sheet peptides. Proc. Natl. Acad. Sci. USA 2005, 102, 10511–10515.
- Santos-Silva, C.A.; Zupin, L.; Oliveira-Lima, M.; Vilela, L.M.; Bezerra-Neto, J.P.; Ferreira-Neto, J.R.; Ferreira, J.D.; Oliveira-Silva, R.L.; Pires, C.D.; Aburjaile, F.F.; et al. Plant Antimicrobial Peptides: State of the Art, In Silico Prediction and Perspectives in the Omics Era. Bioinform. Biol. Insights 2020, 14.
- Greco, S.; Gerdol, M.; Edomi, P.; Pallavicini, A. Molecular Diversity of Mytilin-Like Defense Peptides (Mollusca, Bivalvia). Antibiotics 2020, 9, 37.
- Dias, R.D.O.; Franco, O.L. Cysteine-stabilized αβ defensins: From a common fold to antibacterial activity. Peptides 2015, 72, 64–72.
- Wang, B.; Yao, Y.; Wei, P.; Song, C.; Wan, S.; Yang, S.; Zhu, G.M.; Liu, H.M. Housefly Phormicin inhibits Staphylococcus aureus and MRSA by disrupting biofilm formation and altering gene expression In Vitro and In Vivo. Int. J. Biol. Macromol. 2020.
- Simons, A.; Alhanout, K.; Duval, R.E. Bacteriocins, antimicrobial peptides from bacterial origin: Overview of their biology and their impact against multidrug-resistant bacteria. Microorganisms 2020, 8, 639.
- Lee, D.W.; Kim, B.S. Antimicrobial cyclic peptides for plant disease control. Plant. Pathol. J. 2015, 31, 1–11.
- Jeong Hee, J.; Ha Chul, S. Crystal Structure of NisI in a Lipid-Free Form, the Nisin Immunity Protein, from Lactococcus lactis. Antimicrob. Agents Chemother. 2018, 62, 1–12.
- Williams, G.C.; Delves-Broughton, J. NISIN. In Encyclopedia of Food Sciences and Nutrition, 2nd ed.; Caballero, B., Ed.; Academic Press: Oxford, UK, 2003; pp. 4128–4135. ISBN 978-0-12-227055-0.
- Sass, P.; Jansen, A.; Szekat, C.; Sass, V.; Sahl, H.G.; Bierbaum, G. The lantibiotic mersacidin is a strong inducer of the cell wall stress response of Staphylococcus aureus. BMC Microbiol. 2008, 8, 1–11.
- Smith, L.; Zachariah, C.; Thirumoorthy, R.; Rocca, J.; Novák, J.; Hillman, J.D.; Edison, A.S. Structure and dynamics of the lantibiotic mutacin 1140. Biochemistry 2003, 42, 10372–10384.
- Kim, P.I.; Sohng, J.K.; Sung, C.; Joo, H.S.; Kim, E.M.; Yamaguchi, T.; Park, D.; Kim, B.G. Characterization and structure identification of an antimicrobial peptide, hominicin, produced by Staphylococcus hominis MBBL 2-9. Biochem. Biophys. Res. Commun. 2010, 399, 133–138.
- O’Shea, E.F.; O’Connor, P.M.; O’Sullivan, O.; Cotter, P.D.; Ross, R.P.; Hill, C. Bactofencin A, a new type of cationic bacteriocin with Unusual Immunity. MBio 2013, 4, 1–9.
- Choi, Y.H.; Cho, S.S.; Simkhada, J.R.; Yoo, J.C. A novel thermotolerant and acidotolerant peptide produced by a Bacillus strain newly isolated from a fermented food (kimchi) shows activity against multidrug-resistant bacteria. Int. J. Antimicrob. Agents 2012, 40, 80–83.
- Zipperer, A.; Konnerth, M.C.; Laux, C.; Berscheid, A.; Janek, D.; Weidenmaier, C.; Burian, M.; Schilling, N.A.; Slavetinsky, C.; Marschal, M.; et al. Human commensals producing a novel antibiotic impair pathogen colonization. Nature 2016, 535, 511–516.
- Bionda, N.; Stawikowski, M.; Stawikowski, R.; Cudic, M.; Lopéz, F.; Treitl, D.; Medina, J.; Cudic, P. Effects of cyclic lipodepsipeptide structural modulation on stability, antibacterial activity and human cell toxicity. Chem. Med. Chem. 2012, 7, 871–882.
- Lei, J.; Sun, L.C.; Huang, S.; Zhu, C.; Li, P.; He, J.; Mackey, V.; Coy, D.H.; He, Q.Y. The antimicrobial peptides and their potential clinical applications. Am. J. Transl. Res. 2019, 11, 3919–3931.
- Fritsche, T.R.; Rhomberg, P.R.; Sader, H.S.; Jones, R.N. In Vitro activity of omiganan pentahydrochloride tested against vancomycin-tolerant, -intermediate, and -resistant Staphylococcus aureus. Diagn. Microbiol. Infect Dis. 2008, 60, 399–403.
- Rubinchik, E.; Dugourd, D. Omiganan Pentahydrochloride: A Novel, Broad-Spectrum Antimicrobial Peptide for Topical Use; Wiley-VCH Verlag GmbH & Co, Boschstr.: Weinheim, Germany, 2011; pp. 157–169. ISBN 9783527328918.
- Yu, H.-Y.; Tu, C.-H.; Yip, B.-S.; Chen, H.-L.; Cheng, H.-T.; Huang, K.-C.; Lo, H.-J.; Cheng, J.-W. Easy strategy to increase salt resistance of antimicrobial peptides. Antimicrob. Agents Chemother. 2011, 55, 4918–4921.
- Wu, C.; Hsueh, J.; Yip, B.; Chih, Y.; Peng, K.; Cheng, J. Antimicrobial Peptides Display Strong Synergy with Vancomycin Against Vancomycin-Resistant. Int. J. Mol. Sci. 2020, 21, 4578.
- Azmi, F.; Elliott, A.G.; Marasini, N.; Ramu, S.; Ziora, Z.; Kavanagh, A.M.; Blaskovich, M.A.T.; Cooper, M.A.; Skwarczynski, M.; Toth, I. Short cationic lipopeptides as effective antibacterial agents: Design, physicochemical properties and biological evaluation. Bioorganic Med. Chem. 2016, 24, 2235–2241.
- Datta, S.; Roy, A. Antimicrobial Peptides as Potential Therapeutic Agents: A Review. Int. J. Pept. Res. Ther. 2021, 27, 555–577.
- Martinez de Tejada, G.; Sanchez-Gomez, S.; Razquin-Olazaran, I.; Kowalski, I.; Kaconis, Y.; Heinbockel, L.; Andra, J.; Schurholz, T.; Hornef, M.; Dupont, A.; et al. Bacterial Cell Wall Compounds as Promising Targets of Antimicrobial Agents I. Antimicrobial Peptides and Lipopolyamines. Curr. Drug Targets 2012, 13, 1121–1130.
- Malanovic, N.; Lohner, K. Antimicrobial Peptides Targeting Gram-Positive Bacteria. Pharmaceuticals 2016, 9, 59.
- Ren, H.; Gray, W.M. Amino Acid Composition Determines Peptide Activity Spectrum and Hot-Spot-Based Design of Merecidin. Physiol. Behav. 2019, 176, 139–148.
- Yin, L.M.; Edwards, M.A.; Li, J.; Yip, C.M.; Deber, C.M. Roles of hydrophobicity and charge distribution of cationic antimicrobial peptides in peptide-membrane interactions. J. Biol. Chem. 2012, 287, 7738–7745.
- Sohlenkamp, C.; Geiger, O. Bacterial membrane lipids: Diversity in structures and pathways. FEMS Microbiol. Rev. 2015, 40, 133–159.
- Van Den Bogaart, G.; Guzmán, J.V.; Mika, J.T.; Poolman, B. On the mechanism of pore formation by melittin. J. Biol. Chem. 2008, 283, 33854–33857.
- Matsuzaki, K.; Sugishita, K.; Ishibe, N.; Ueha, M.; Nakata, S.; Miyajima, K.; Epand, R.M. Relationship of Membrane Curvature to the Formation of Pores by Magainin 2. Biochemistry 1998, 37, 11856–11863.
- Miyoshi, N.; Isogai, E.; Hiramatsu, K.; Sasaki, T. Activity of tick antimicrobial peptide from Ixodes persulcatus (persulcatusin) against cell membranes of drug-resistant Staphylococcus aureus. J. Antibiot. (Tokyo) 2017, 70, 142–146.
- Ghobrial, O.G.; Derendorf, H.; Hillman, J.D. Pharmacodynamic activity of the lantibiotic MU1140. Int. J. Antimicrob. Agents 2009, 33, 70–74.
- Subbalakshmi, C.; Sitaram, N. Mechanism of antimicrobial action of indolicidin. FEMS Microbiol. Lett. 1998, 160, 91–96.




