
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Arun Kumar Kondadi | + 1674 word(s) | 1674 | 2021-07-06 12:04:44 | | | |
| 2 | Vicky Zhou | Meta information modification | 1674 | 2021-07-27 03:33:07 | | |
Video Upload Options
Recent studies using fluorescence super-resolution (SR) microscopy techniques showed unexpected fast movement of cristae and CJs, collectively termed as cristae dynamics. Cristae undergo continuous cycles of membrane remodelling often assisted by the dynamics of CJs in a MICOS-dependent manner, which led to the proposal of the ‘Cristae Fission and Fusion’ (CriFF) model. The field of cristae dynamics is still in infancy, future experiments could provide better insights about the consequences of the reduced cristae or CJ dynamics in the knockouts (KOs) of the MICOS subunits and their relevance in many pathologies associated with the MICOS complex.
1. Introduction
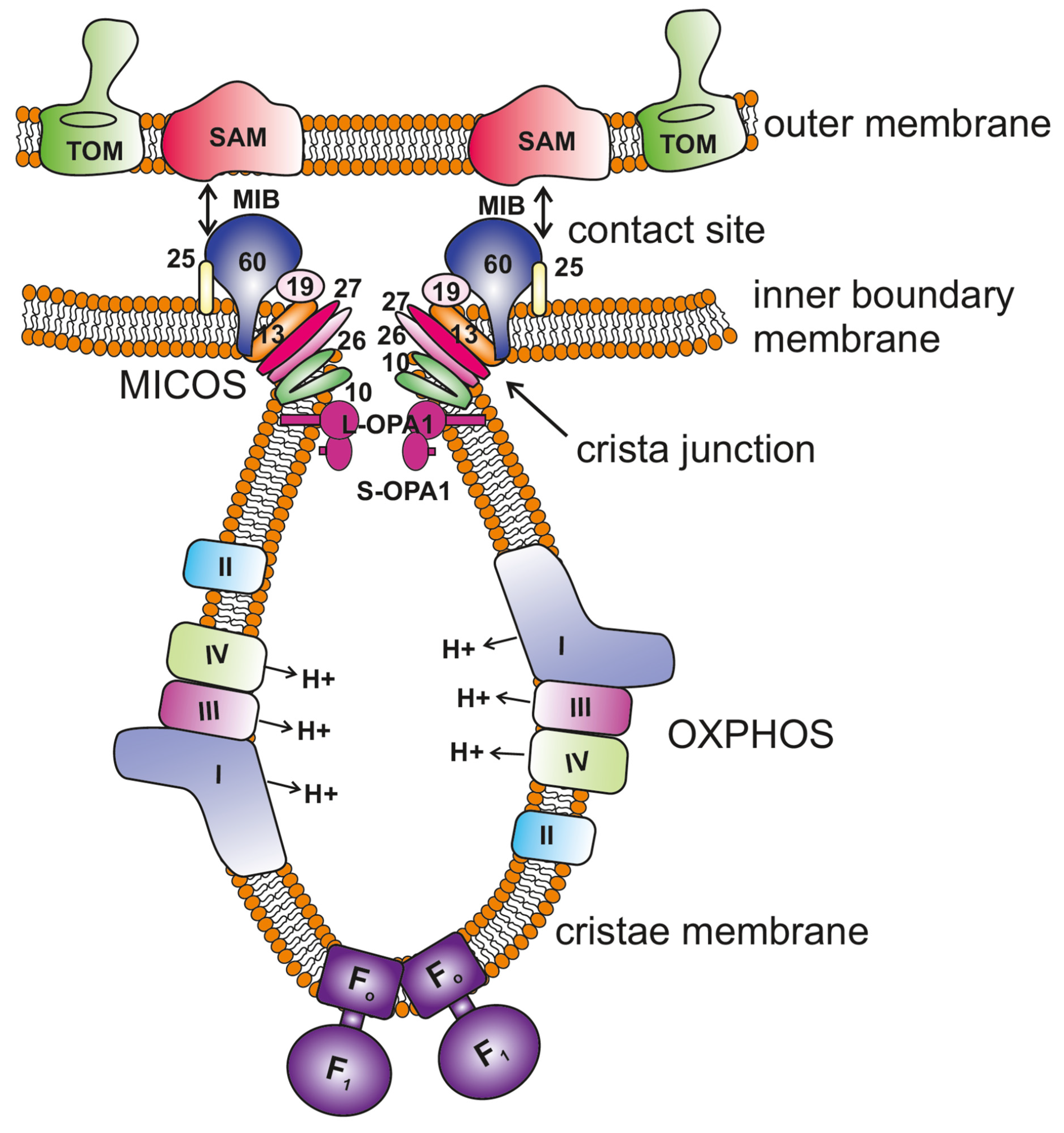 Figure 1. Key regulators of mitochondrial cristae organization. The scheme shows the organization of mitochondrial membranes where the cristae are formed by invagination of the inner membrane towards the matrix. The MICOS (mitochondrial contact site and cristae organizing system) complex resides at the crista junctions (CJs) and is composed of seven subunits, MIC10, MIC13, MIC19, MIC25, MIC26, MIC27 and MIC60 (only the numbers are depicted in the figure for the ease of legibility). MICOS is required to stabilize the CJs and form the contacts between the inner and outer membranes via interaction with the SAM (sorting and assembly machinery) complex. This interaction between MICOS and the SAM complex forms the larger complex called the mitochondrial intermembrane space bridging complex (MIB) that encompasses the intermembrane space. OPA1 is also enriched at the CJs, and interaction between membrane-bound long (L-) forms and soluble short (S-) forms is required to maintain the width of the CJs. F1FO ATP synthase plays an important role in the formation of positive membrane curvature at the tip/rim of cristae. The OXPHOS (oxidative phosphorylation) machinery resides in the cristae membrane.
Figure 1. Key regulators of mitochondrial cristae organization. The scheme shows the organization of mitochondrial membranes where the cristae are formed by invagination of the inner membrane towards the matrix. The MICOS (mitochondrial contact site and cristae organizing system) complex resides at the crista junctions (CJs) and is composed of seven subunits, MIC10, MIC13, MIC19, MIC25, MIC26, MIC27 and MIC60 (only the numbers are depicted in the figure for the ease of legibility). MICOS is required to stabilize the CJs and form the contacts between the inner and outer membranes via interaction with the SAM (sorting and assembly machinery) complex. This interaction between MICOS and the SAM complex forms the larger complex called the mitochondrial intermembrane space bridging complex (MIB) that encompasses the intermembrane space. OPA1 is also enriched at the CJs, and interaction between membrane-bound long (L-) forms and soluble short (S-) forms is required to maintain the width of the CJs. F1FO ATP synthase plays an important role in the formation of positive membrane curvature at the tip/rim of cristae. The OXPHOS (oxidative phosphorylation) machinery resides in the cristae membrane.2. Understanding Cristae Architecture Using Recent Technological Advancements
2.1. Application of Fluorescence Super-Resolution Techniques to IM Reveal Novel Insights
2.2. MICOS Complex Regulates Apparent Cristae Fusion and Fission Cycles
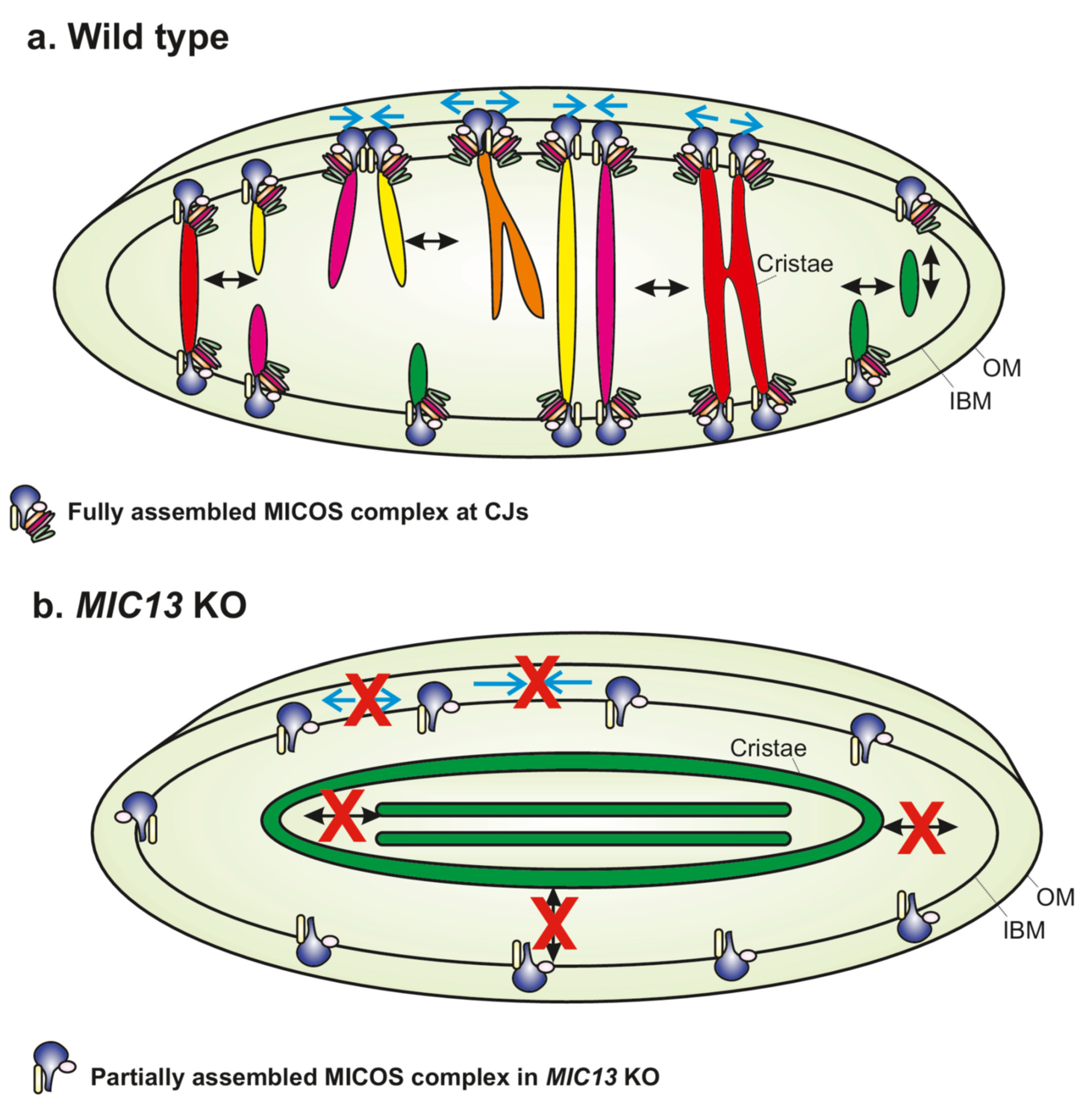
Recent studies using SR techniques showed unexcepted dynamics of cristae [17][19][21][27][28][29]. Both cristae and CJs constantly changed their position within mitochondria, confirming that they are highly dynamic within seconds [17]. Tracking movements of CJs using live-cell STED nanoscopy showed that they repeatedly come together and then move apart in a balanced and reversible manner. Interestingly, two CJs coming together bring with them the adjoining cristae so that the cristae appear as the letter ‘Y’, as visualized using a time-lapse movie of mitochondria expressing MIC13-SNAP, which dually marked cristae and CJs (Figure 2). Deletion of MIC13, impairing, e.g., MICOS assembly, leads to a drastic reduction in merging and splitting events of cristae and CJs, identifying MICOS as the first molecular player that is required for the dynamics of cristae and CJs (Figure 2). Evidence showing exchange of content between cristae during merging events led to the proposal of Cristae Fission and Fusion (‘CriFF’) model [17].
2.3. Possible Implications of Cristae Dynamics
3. Conclusions
References
- Vogel, F.; Bornhovd, C.; Neupert, W.; Reichert, A.S. Dynamic subcompartmentalization of the mitochondrial inner membrane. J. Cell Biol. 2006, 175, 237–247.
- Wurm, C.A.; Jakobs, S. Differential protein distributions define two sub-compartments of the mitochondrial inner membrane in yeast. FEBS Lett. 2006, 580, 5628–5634.
- Perkins, G.; Renken, C.; Martone, M.E.; Young, S.J.; Ellisman, M.; Frey, T. Electron tomography of neuronal mitochondria: Three-dimensional structure and organization of cristae and membrane contacts. J. Struct. Biol. 1997, 119, 260–272.
- Mannella, C.A. Structural diversity of mitochondria: Functional implications. Ann. N. Y. Acad. Sci. 2008, 1147, 171–179.
- Zick, M.; Rabl, R.; Reichert, A.S. Cristae formation-linking ultrastructure and function of mitochondria. Biochim. Biophys. Acta 2009, 1793, 5–19.
- Hackenbrock, C.R. Ultrastructural bases for metabolically linked mechanical activity in mitochondria. I. Reversible ultrastructural changes with change in metabolic steady state in isolated liver mitochondria. J. Cell Biol. 1966, 30, 269–297.
- Hackenbrock, C.R. Chemical and physical fixation of isolated mitochondria in low-energy and high-energy states. Proc. Natl. Acad. Sci. USA 1968, 61, 598–605.
- Cogliati, S.; Enriquez, J.A.; Scorrano, L. Mitochondrial Cristae: Where Beauty Meets Functionality. Trends Biochem. Sci. 2016, 41, 261–273.
- Baker, N.; Patel, J.; Khacho, M. Linking mitochondrial dynamics, cristae remodeling and supercomplex formation: How mitochondrial structure can regulate bioenergetics. Mitochondrion 2019, 49, 259–268.
- Dlaskova, A.; Spacek, T.; Engstova, H.; Spackova, J.; Schrofel, A.; Holendova, B.; Smolkova, K.; Plecita-Hlavata, L.; Jezek, P. Mitochondrial cristae narrowing upon higher 2-oxoglutarate load. Biochim. Biophys. Acta Bioenerg. 2019, 1860, 659–678.
- Gilkerson, R.W.; Selker, J.M.L.; Capaldi, R.A. The cristal membrane of mitochondria is the principal site of oxidative phosphorylation. FEBS Lett. 2003, 546, 355–358.
- Colina-Tenorio, L.; Horten, P.; Pfanner, N.; Rampelt, H. Shaping the mitochondrial inner membrane in health and disease. J. Intern. Med. 2020, 287, 645–664.
- Rasmussen, N. Mitochondrial structure and the practice of cell biology in the 1950s. J. Hist. Biol. 1995, 28, 381–429.
- Barbot, M.; Meinecke, M. Reconstitutions of mitochondrial inner membrane remodeling. J. Struct. Biol. 2016, 196, 20–28.
- Ikon, N.; Ryan, R.O. Cardiolipin and mitochondrial cristae organization. Biochim. Biophys. Acta Biomembr. 2017, 1859, 1156–1163.
- Kondadi, A.K.; Anand, R.; Reichert, A.S. Functional Interplay between Cristae Biogenesis, Mitochondrial Dynamics and Mitochondrial DNA Integrity. Int. J. Mol. Sci. 2019, 20, 4311.
- Kondadi, A.K.; Anand, R.; Hansch, S.; Urbach, J.; Zobel, T.; Wolf, D.M.; Segawa, M.; Liesa, M.; Shirihai, O.S.; Weidtkamp-Peters, S.; et al. Cristae undergo continuous cycles of membrane remodelling in a MICOS-dependent manner. EMBO Rep. 2020, 21, e49776.
- Stephan, T.; Bruser, C.; Deckers, M.; Steyer, A.M.; Balzarotti, F.; Barbot, M.; Behr, T.S.; Heim, G.; Hubner, W.; Ilgen, P.; et al. MICOS assembly controls mitochondrial inner membrane remodeling and crista junction redistribution to mediate cristae formation. EMBO J. 2020, 39, e104105.
- Hu, C.; Shu, L.; Huang, X.; Yu, J.; Li, L.; Gong, L.; Yang, M.; Wu, Z.; Gao, Z.; Zhao, Y.; et al. OPA1 and MICOS Regulate mitochondrial crista dynamics and formation. Cell Death Dis. 2020, 11, 940.
- Wolf, D.M.; Segawa, M.; Kondadi, A.K.; Anand, R.; Bailey, S.T.; Reichert, A.S.; van der Bliek, A.M.; Shackelford, D.B.; Liesa, M.; Shirihai, O.S. Individual cristae within the same mitochondrion display different membrane potentials and are functionally independent. EMBO J. 2019, 38, e101056.
- Kondadi, A.K.; Anand, R.; Reichert, A.S. Cristae Membrane Dynamics—A Paradigm Change. Trends Cell Biol. 2020, 30, 923–936.
- Anand, R.; Kondadi, A.K.; Meisterknecht, J.; Golombek, M.; Nortmann, O.; Riedel, J.; Peifer-Weiss, L.; Brocke-Ahmadinejad, N.; Schlutermann, D.; Stork, B.; et al. MIC26 and MIC27 cooperate to regulate cardiolipin levels and the landscape of OXPHOS complexes. Life Sci. Alliance 2020, 3, e202000711.
- Jans, D.C.; Wurm, C.A.; Riedel, D.; Wenzel, D.; Stagge, F.; Deckers, M.; Rehling, P.; Jakobs, S. STED super-resolution microscopy reveals an array of MINOS clusters along human mitochondria. Proc. Natl. Acad. Sci. USA 2013, 110, 8936–8941.
- Stoldt, S.; Stephan, T.; Jans, D.C.; Bruser, C.; Lange, F.; Keller-Findeisen, J.; Riedel, D.; Hell, S.W.; Jakobs, S. Mic60 exhibits a coordinated clustered distribution along and across yeast and mammalian mitochondria. Proc. Natl. Acad. Sci. USA 2019, 116, 9853–9858.
- Friedman, J.R.; Mourier, A.; Yamada, J.; McCaffery, J.M.; Nunnari, J. MICOS coordinates with respiratory complexes and lipids to establish mitochondrial inner membrane architecture. eLife 2015, 4.
- Pape, J.K.; Stephan, T.; Balzarotti, F.; Buchner, R.; Lange, F.; Riedel, D.; Jakobs, S.; Hell, S.W. Multicolor 3D MINFLUX nanoscopy of mitochondrial MICOS proteins. Proc. Natl. Acad. Sci. USA 2020, 117, 20607–20614.
- Huang, X.; Fan, J.; Li, L.; Liu, H.; Wu, R.; Wu, Y.; Wei, L.; Mao, H.; Lal, A.; Xi, P.; et al. Fast, long-term, super-resolution imaging with Hessian structured illumination microscopy. Nat. Biotechnol. 2018, 36, 451–459.
- Wang, C.; Taki, M.; Sato, Y.; Tamura, Y.; Yaginuma, H.; Okada, Y.; Yamaguchi, S. A photostable fluorescent marker for the superresolution live imaging of the dynamic structure of the mitochondrial cristae. Proc. Natl. Acad. Sci. USA 2019, 116, 15817–15822.
- Stephan, T.; Roesch, A.; Riedel, D.; Jakobs, S. Live-cell STED nanoscopy of mitochondrial cristae. Sci. Rep. 2019, 9, 12419.




