1000/1000
Hot
Most Recent

Metastasis is the process whereby cancer cells migrate from the primary tumour site to colonise the surrounding or distant tissue or organ. Metastasis is the primary cause of cancer-related mortality and approximately half of all cancer patients present at diagnosis with some form of metastasis. MicroRNAs (miRNAs), a class of small (19-25 nucleotides) non-coding single-strand RNAs, regulate gene expression and play an important role in cancer development and progression including in the metastatic process.
Cancer metastasis, the spread of tumour cells from the primary tumour site, has been reported to account for approximately 67–90% of cancer-related deaths. Approximately half of all cancer patients present with metastasis at the time of diagnosis [1][2][3]. Metastasis is a multistep process, which starts when tumour cells detach from the primary tumour mass, intravasate into lymphatic and circulatory systems to become circulating tumour cells (CTCs), extravasate to leave the circulation, invade and proliferate in a new niche of a distant tissue/organ to form a new tumour [4]. The metastatic process is very inefficient since from the 0.2% of CTCs that survive their time in circulation, only those cells that are the first to reach permissive target organs and are then able to colonize those tissues can initiate metastatic tumour growth [5].
It is well-known that cancer initiation and progression, as well as metastasis, not only depends on tumour cells themselves, but also on the cells of the tumour microenvironment (TME) [6][7][8]. The major components of the TME apart from tumour cells include cancer-associated fibroblasts (CAFs), endothelial cells and immune cells, in addition to other components such as the extracellular matrix [9][10][11]. Hypoxia, cellular oxygen deprivation, is an important factor that drives many aspects of metastasis [12][13], including the initiation of the epithelial–mesenchymal transition (EMT) process that changes the phenotype of tumour cells allowing them to escape from the matrix of the primary tumour [14]. In addition to hypoxia, the interaction between tumour cells and the TME induces a wide range of biological events that are necessary for metastasis including proliferation, immunosuppression and angiogenesis [15][16][17]. Many of these processes are regulated by microRNAs (miRNAs) and are the subject of this review. In addition to the direct control of metastasis by miRNAs, it has been recently shown that they can regulate the metastatic process by acting as mediators of intercellular communication between tumour cells and cells of the TME [18][19].
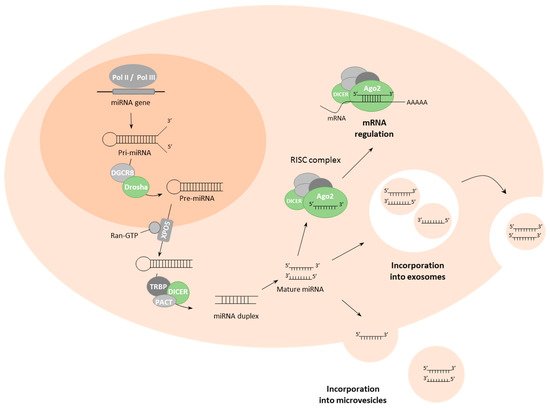
MiRNAs are a class of small (19–25 nucleotides) non-coding single-strand RNAs. Since their initial discovery in Caenorhabditis elegans [20], miRNAs have been demonstrated to play key roles in many, if not all, physiological cellular functions by regulating target genes through primarily negative post-transcriptional regulation of gene expression [21][22][23]. A single miRNA is capable of targeting many genes and, conversely, a single gene can be targeted by many miRNAs leading to a complex regulatory network that encompasses more than 60% of human genes [24]. In addition to their importance under physiological conditions, miRNAs are ubiquitously deregulated in cancer and can act as tumour-promoting miRNAs (oncomiRNAs and metastamiRNAs) targeting messenger RNAs (mRNAs) coding for proteins that act as tumour suppressors or as tumour suppressor miRNAs targeting mRNAs coding for proteins with oncogenic properties [25].
In 2007, two separate reports released in parallel first described the association between miRNAs and metastasis. Ma et al. demonstrated that miR-10b could promote breast cancer metastasis in vitro and in vivo through targeting of the HOXD10 (Homeobox D10) gene [26], whilst Yu et al. demonstrated that let-7 can act as a metastasis suppressor miRNA through targeting of H-RAS and HMGA2 (High Mobility Group AT-Hook 2), leading to a reduction in proliferation, mammosphere formation and metastatic potential, including in breast cancer [27]. Subsequently, many miRNAs have been identified that are associated with metastasis or with associated pathways, such as migration and invasion [28].
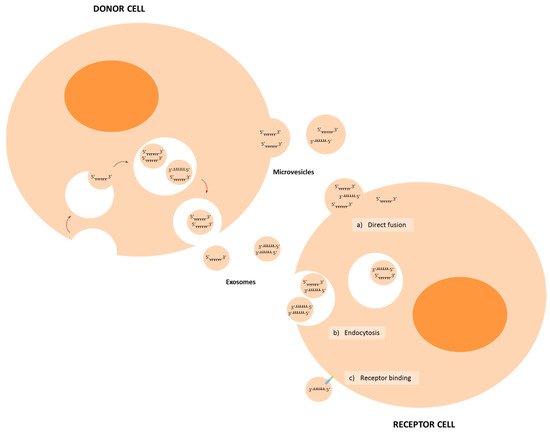
| miRNA | Cancer | Donor Cells | Receptor Cells | Target | Ref. |
|---|---|---|---|---|---|
| miR-9 | Breast | Tumour cells | Fibroblasts | E-cadherin | [51] |
| miR-10b-5p | HCC | Tumour cells | Tumour cells | - | [52] |
| miR-15b-5p | GBM | Tumour cells | Non-tumour brain cells | MAPK/ERK | [53] |
| miR-17-5p | CRC | CAFs | Tumour cells | RUNX3 | [54] |
| miR-19b-3p | ccRCC | CSCs | Tumour cells | PTEN | [55] |
| miR-20a-5p | Breast | Tumour cells | BMMs | SRCIN1 | [56] |
| miR-21 | Lung | Tumour cells | Pre-osteoclasts | PTEN | [57] |
| Lung | Tumour cells | Macrophages | TLR8 | [58] | |
| miR-21-5p | GBM | Tumour cells | Non-tumour brain cells | MAPK/ERK | [53] |
| HCC | Tumour cells | Tumour cells | - | [52] | |
| Breast | Tumour cells | Osteoclasts | PDCD4 | [59] | |
| Breast | CAFs | Tumour cells | - | [60] | |
| Bladder | Tumour cells | Macrophages | PTEN | [61] | |
| Colon | TAMs | Tumour cells | BRG1 | [62] | |
| miR-25-3p | CRC | Tumour cells | Endothelial cells | KLF2/KLF4 | [63] |
| miR-29a | Lung | Tumour cells | Macrophages | TLR8 | [58] |
| miR-30c-5p | GBM | Tumour cells | Non-tumour brain cells | MAPK/ERK | [53] |
| miR-30d-5p | GBM | Tumour cells | Non-tumour brain cells | MAPK/ERK | [53] |
| miR-103a-3p | HCC | Tumour cells | Endothelial cells | VE-Cadherin | [64] |
| miR-105 | Breast | Tumour cells | Endothelial cells | ZO-1 | [65] |
| miR-122 | Breast | Tumour cells | Fibroblasts/astrocytes | PKM2/GLUT1 | [66] |
| miR-130b-3p | Gastric | TAMs | Tumour cells | MLL3/GRHL2 | [67] |
| miR-143-3p | Breast | CAFs | Tumour cells | - | [60] |
| miR-155-5p | Colon | TAMs | Tumour cells | BRG1 | [62] |
| miR-210 | Breast | Tumour cells | Endothelial cells | Ephrin A3 | [68] |
| Breast | Tumour cells | Endothelial cells | Ephrin A3/PTP1B | [69] | |
| miR-210-3p | HCC | Tumour cells | Endothelial cells | SMAD4/STAT6 | [70] |
| miR-211 | Melanoma | Tumour cells | Fibroblasts | IGF2R | [71] |
| miR-214 | Lung | Tumour cells | Treg | PTEN | [72] |
| miR-218-5p | Breast | Tumour cells | Pre-osteoblasts | Col1a1 | [73] |
| miR-223-3p | Breast | TAMs | Tumour cells | Mef2c | [74] |
| miR-301a-3p | Pancreatic | Tumour cells | Macrophages | PTEN | [75] |
| miR-342-3p | OSCC | High-metastatic cells | Low-metastatic cells | - | [76] |
| miR-378e | Breast | CAFs | Tumour cells | - | [60] |
| miR-501-3p | PDAC | TAMs | Tumour cells | TGDBR3 | [77] |
| miR-934 | CRC | Tumour cells | Macrophages | PTEN | [78] |
| miR-939-5p | Breast | Tumour cells | Endothelial cells | VE-cadherin | [79] |
| miR-940 | Prostate | Tumour cells | MSCs | ARHGAP1 FAM134A |
[80] |
| miR-1246 | OSCC | High-metastatic cells | Low-metastatic cells | DENND2D | [76] |
| miR-1247-3p | HCC | High-metastatic cells | Fibroblasts | B4GALT3 | [81] |
Several miRNAs have been described to participate in the communication between tumour cells and cells of the TME, such as fibroblasts, endothelial cells or immune cells, among others; and several of them regulate expression of genes that are involved in the metastasis process (Figure 3).
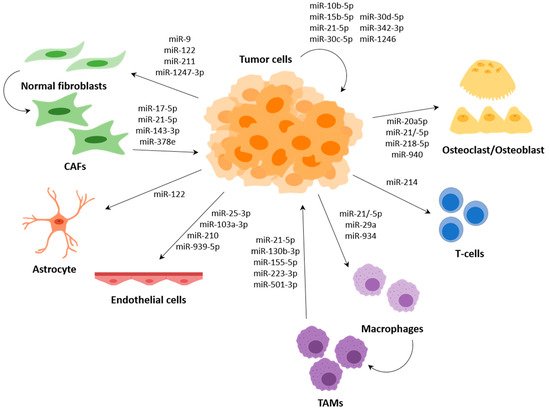
Macrophages are the most abundant infiltrative immune cells present in and around tumours and play a critical role in inflammation [96]. Macrophages are known to polarize, depending on different stimuli, to the M1 phenotype with anti-tumour activity or to the M2 phenotype with pro-tumoral activity. Tumour-associated macrophages (TAMs), which are considered to be M2-like, support different aspects of tumour development, including tumour formation, growth and metastasis [97][98].
The premetastatic inflammatory response generated by TAMs leads to tumour growth and metastasis. MiR-21 and miR-29 secreted by lung tumour cells target TLR8 (toll-like receptor 8) within intracellular endosomes leading to induction of NF-κB (nuclear factor kappa-light-chain-enhancer of activated B cells) and NF-κB-mediated secretion of the proinflammatory cytokines TNF-α (tumour necrosis factor alpha) and IL-6 (interleukin-6) [58]. In bladder cancer, macrophages take up exosomal miR-21-5p from tumour cells leading to promotion of M2 polarization and enhanced migration and invasion of tumour cells [61]. Both CRC cell-derived exosomal miR-934 and hypoxic pancreatic cell-derived exosomal miR-301a-3p were demonstrated to activate PI3K/AKT (phosphatidylinositol 3-kinase/protein kinase B) signalling pathway and enhance metastatic capacity of tumour cells through PTEN targeting [78][75]. PTEN also plays an important role in the regulation of T cells, and it has been demonstrated that EV-associated miR-214 from a range of tumour cells including breast cancer, hepatocellular carcinoma, non-small-cell lung cancer (NSCLC) or pancreatic cancer could transfer to T cells leading to downregulation of PTEN and promoting T-reg (regulatory T cells) expansion and IL-10 (interleukin-10) secretion, which in turn promotes tumour growth and enhanced immune suppression in vivo [72].
MicroRNAs have also been shown to be involved in the recruitment of immune cells to the TME, but rather than through the transfer of miRNAs from tumour cells to immune cells, this occurs through the secretion of attractant molecules. For example, both miR-149 in triple-negative breast cancer (TNBC) and miR-148b in HCC have been demonstrated to target colony-stimulating factor-1 (CSF-1) miRNAs [99][100]. In TNBC, downregulation of miR-149 promoted lung metastasis by enhancing CSF1-dependent recruitment and M2 polarization of macrophages, which also correlated with macrophage infiltration and reduced survival in patient samples [99]. Downregulation of miR-148b in HCC patients correlated positively with recurrence, metastasis and poor prognosis. Moreover, in vitro and in vivo metastatic HCC cells showed decreased levels of miR-148b that correlated with increased CSF1, which promoted HCC growth and metastasis through CSF1/CSF1R (colony-stimulating factor-1 receptor)-mediated TAM infiltration [100]. Similarly, miR-561-5p, which directly target chemokine (C–X3–C motif) ligand 1 (CX3CL1), in metastatic HCC downregulated CX3CL1 leading to low infiltration of CX3CR1 (CX3C chemokine receptor 1)-positive NK cells and resulting in promoted tumorigenesis and metastasis [101].
Moreover, it has been described that TAMs in the TME can release extracellular vesicles with miRNAs, and these can be transferred to tumour cells, generally promoting migration, invasion and metastasis. In CRC, exosomal miR-21-5p and miR-155-5p from macrophages directly target the BRG1 coding gene in tumour cells; this gene is a key factor promoting metastasis [62]. Another example is macrophage-derived EVs in gastric cancer (GC), which contain high levels of miR-130b-3p and promote survival, migration, invasion and angiogenesis in GC cells through the modulation of MLL3 (mixed-lineage leukemia protein 3) and GRHL2 (grainyhead-like protein 2 homolog) [67]. MiR-501-3p has been found to be highly expressed in pancreatic ductal adenocarcinoma (PDAC) tissues and TAM-derived exosomes. Exosomal miR-501-3p promotes cancer cell migration and invasion, as well as tumour formation and metastasis in vivo through regulation of TGFBR3 (transforming growth factor beta receptor 3) [77]. Finally, a study detected miR-223-3p in exosomes released by IL-4 (interleukin-4)-activated macrophages; this miRNA has been shown to transfer to breast tumour cells where it regulates invasion through the Mef2c (myocyte enhancer factor 2C)-β-catenin pathway [74].